MicroRNAs as Biomarkers for Liver Disease and Hepatocellular Carcinoma
Abstract
:1. Introduction
2. MicroRNAs
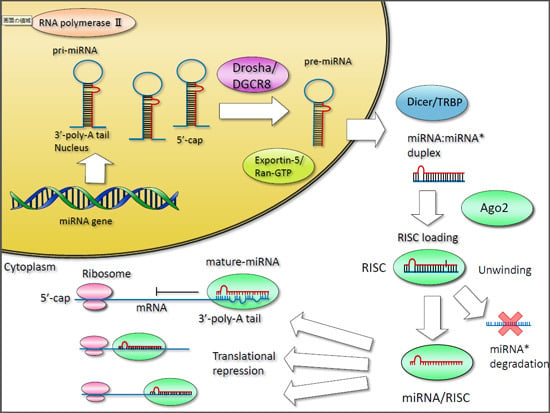
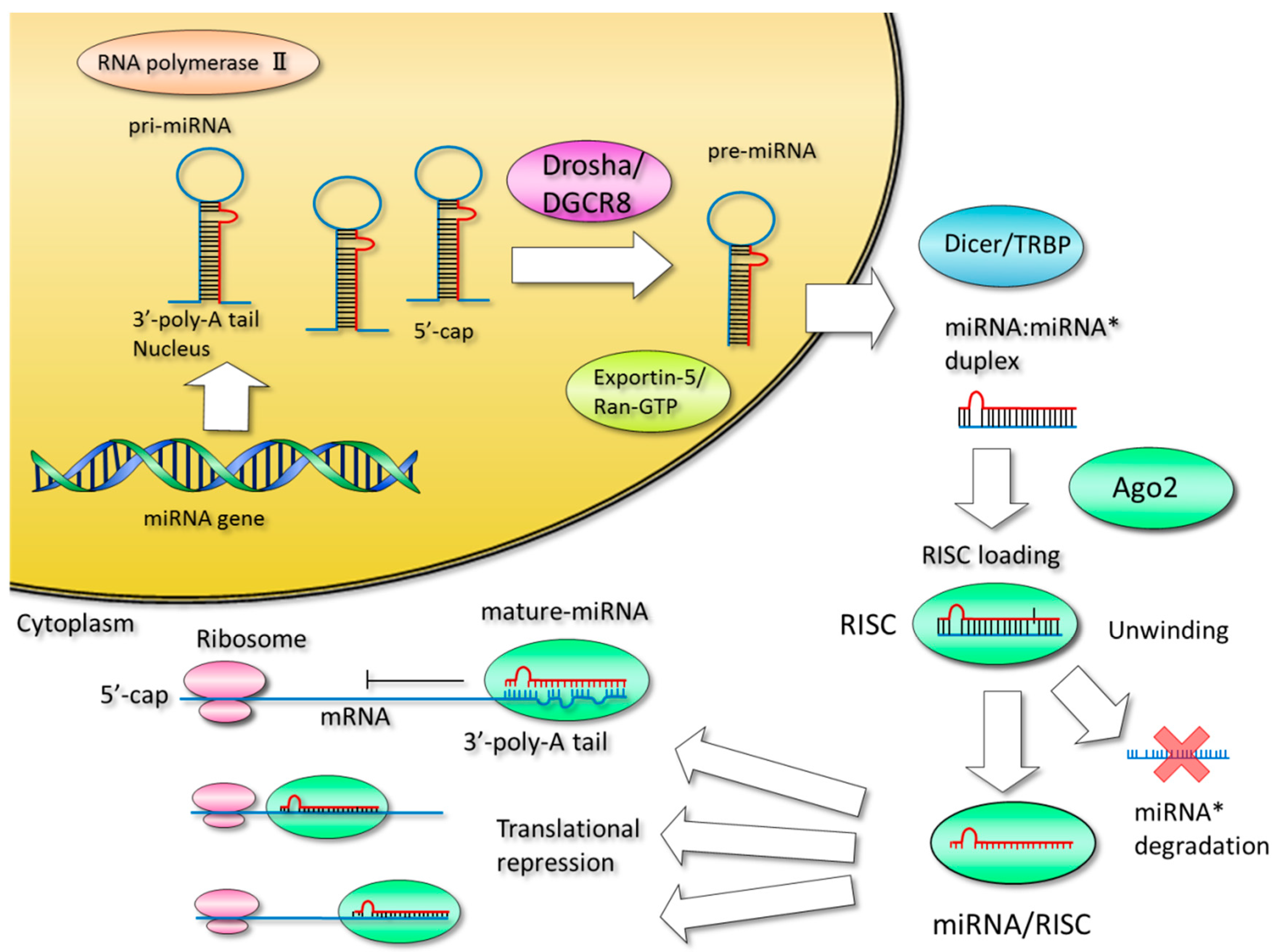
3. Production of MicroRNAs
4. Serum MicroRNAs
5. Exosomes
6. Measurement of Serum MicroRNAs
7. Baseline MicroRNA Expression in the Liver
8. The Role of MicroRNAs in Liver Injury and Disease
8.1. Autoimmune Liver Diseases
8.2. Drug-Induced Liver Injury
8.3. Inflammatory Liver Damage
8.4. Alcoholic Liver Disease
8.5. Non-Alcoholic Fatty Liver Disease
8.6. Fibrosis
9. The Role of MicroRNAs in Viral Hepatitis
9.1. Hepatitis B Virus
9.2. Hepatitis C Virus
10. Hepatocellular Carcinoma
11. Mechanisms of MicroRNA Dysregulation
12. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin. 2011, 61, 69–90. [Google Scholar] [CrossRef] [PubMed]
- Fattovich, G.; Stroffolini, T.; Zagni, I.; Donato, F. Hepatocellular carcinoma in cirrhosis: Incidence and risk factors. Gastroenterology 2004, 127, S35–S50. [Google Scholar] [CrossRef] [PubMed]
- Parkin, D.M.; Bray, F.; Ferlay, J.; Pisani, P. Global cancer statistics, 2002. CA Cancer J. Clin. 2005, 55, 74–108. [Google Scholar] [CrossRef] [PubMed]
- Aleman, S.; Rahbin, N.; Weiland, O.; Davidsdottir, L.; Hedenstierna, M.; Rose, N.; Verbaan, H.; Stal, P.; Carlsson, T.; Norrgren, H.; et al. A risk for hepatocellular carcinoma persists long-term after sustained virologic response in patients with hepatitis C-associated liver cirrhosis. Clin. Infect. Dis. 2013, 57, 230–236. [Google Scholar] [CrossRef] [PubMed]
- Turchinovich, A.; Weiz, L.; Langheinz, A.; Burwinkel, B. Characterization of extracellular circulating microrna. Nucleic Acids Res. 2011, 39, 7223–7233. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Anwar, S.L.; Lehmann, U. Micrornas: Emerging novel clinical biomarkers for hepatocellular carcinomas. J. Clin. Med. 2015, 4, 1631–1650. [Google Scholar] [CrossRef] [PubMed]
- Kerr, T.A.; Korenblat, K.M.; Davidson, N.O. MicroRNAs and liver disease. Transl. Res. 2011, 157, 241–252. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yang, R.; Fan, X.; Gu, T.; Zhao, Z.; Chang, D.; Wang, W.; Wang, C. MicroRNA array analysis of microRNAs related to systemic scleroderma. Rheumatol. Int. 2012, 32, 307–313. [Google Scholar] [CrossRef] [PubMed]
- Te, J.L.; Dozmorov, I.M.; Guthridge, J.M.; Nguyen, K.L.; Cavett, J.W.; Kelly, J.A.; Bruner, G.R.; Harley, J.B.; Ojwang, J.O. Identification of unique microRNA signature associated with lupus nephritis. PLoS ONE 2010, 5, e10344. [Google Scholar] [CrossRef] [PubMed]
- Croce, C.M.; Calin, G.A. MiRNAs, cancer, and stem cell division. Cell 2005, 122, 6–7. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Hagedorn, C.H.; Cullen, B.R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 2004, 10, 1957–1966. [Google Scholar] [CrossRef] [PubMed]
- Lawrie, C.H.; Gal, S.; Dunlop, H.M.; Pushkaran, B.; Liggins, A.P.; Pulford, K.; Banham, A.H.; Pezzella, F.; Boultwood, J.; Wainscoat, J.S.; et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br. J. Haematol. 2008, 141, 672–675. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.M.; Zhang, C.; Burchard, J.; Fan, S.T.; Wong, K.F.; Dai, H.; Poon, R.T.; Luk, J.M. Global regulation on microRNA in hepatitis B virus-associated hepatocellular carcinoma. Omics 2011, 15, 187–191. [Google Scholar] [CrossRef] [PubMed]
- Bala, S.; Marcos, M.; Szabo, G. Emerging role of microRNAs in liver diseases. World J. Gastroenterol. 2009, 15, 5633–5640. [Google Scholar] [CrossRef] [PubMed]
- Ji, F.; Yang, B.; Peng, X.; Ding, H.; You, H.; Tien, P. Circulating microRNAs in hepatitis B virus-infected patients. J. Viral Hepat. 2011, 18, e242–e251. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Ura, S.; Honda, M.; Yamashita, T.; Ueda, T.; Takatori, H.; Nishino, R.; Sunakozaka, H.; Sakai, Y.; Horimoto, K.; Kaneko, S. Differential microRNA expression between hepatitis B and hepatitis C leading disease progression to hepatocellular carcinoma. Hepatology 2009, 49, 1098–1112. [Google Scholar] [CrossRef] [PubMed]
- Hayes, C.N.; Akamatsu, S.; Tsuge, M.; Miki, D.; Akiyama, R.; Abe, H.; Ochi, H.; Hiraga, N.; Imamura, M.; Takahashi, S.; et al. Hepatitis B virus-specific miRNAs and argonaute2 play a role in the viral life cycle. PLoS ONE 2012, 7, e47490. [Google Scholar] [CrossRef] [PubMed]
- Shwetha, S.; Gouthamchandra, K.; Chandra, M.; Ravishankar, B.; Khaja, M.N.; Das, S. Circulating miRNA profile in HCV infected serum: Novel insight into pathogenesis. Sci. Rep. 2013, 3, 1555. [Google Scholar] [CrossRef] [PubMed]
- Novellino, L.; Rossi, R.L.; Bonino, F.; Cavallone, D.; Abrignani, S.; Pagani, M.; Brunetto, M.R. Circulating hepatitis b surface antigen particles carry hepatocellular microRNAs. PLoS ONE 2012, 7, e31952. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.M.; Yao, T.J.; Wang, W.; Wong, K.F.; Lee, N.P.; Fan, S.T.; Poon, R.T.; Gao, C.; Luk, J.M. Circulating miR-15b and miR-130b in serum as potential markers for detecting hepatocellular carcinoma: A retrospective cohort study. BMJ Open 2012, 2, e000825. [Google Scholar] [CrossRef] [PubMed]
- Cortez, M.A.; Bueso-Ramos, C.; Ferdin, J.; Lopez-Berestein, G.; Sood, A.K.; Calin, G.A. MicroRNAs in body fluids—The mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 2011, 8, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [PubMed]
- Gallo, A.; Tandon, M.; Alevizos, I.; Illei, G.G. The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 2012, 7, e30679. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Yuan, T.; Tschannen, M.; Sun, Z.; Jacob, H.; Du, M.; Liang, M.; Dittmar, R.L.; Liu, Y.; Kohli, M.; et al. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genom. 2013, 14, 319. [Google Scholar] [CrossRef] [PubMed]
- Conde-Vancells, J.; Rodriguez-Suarez, E.; Embade, N.; Gil, D.; Matthiesen, R.; Valle, M.; Elortza, F.; Lu, S.C.; Mato, J.M.; Falcon-Perez, J.M. Characterization and comprehensive proteome profiling of exosomes secreted by hepatocytes. J. Proteome Res. 2008, 7, 5157–5166. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, K.; Liu, Y.; Xu, Y.; Zhang, F.; Yang, H.; Liu, J.; Pan, T.; Chen, J.; Wu, M.; et al. Exosomes mediate the cell-to-cell transmission of IFN-α-induced antiviral activity. Nat. Immunol. 2013, 14, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Taylor, C.J.; Satoor, S.N.; Ranjan, A.K.; Pereira e Cotta, M.V.; Joglekar, M.V. A protocol for measurement of noncoding RNA in human serum. Exp. Diabetes Res. 2012, 2012, 168368. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, S.K.; Pachunka, J.M.; Mott, J.L. Role of microRNAs in alcohol-induced multi-organ injury. Biomolecules 2015, 5, 3309–3338. [Google Scholar] [CrossRef] [PubMed]
- Marabita, F.; de Candia, P.; Torri, A.; Tegner, J.; Abrignani, S.; Rossi, R.L. Normalization of circulating microRNA expression data obtained by quantitative real-time RT-PCR. Brief. Bioinform. 2015. [Google Scholar] [CrossRef] [PubMed]
- Benz, F.; Roderburg, C.; Vargas Cardenas, D.; Vucur, M.; Gautheron, J.; Koch, A.; Zimmermann, H.; Janssen, J.; Nieuwenhuijsen, L.; Luedde, M.; et al. U6 is unsuitable for normalization of serum miRNA levels in patients with sepsis or liver fibrosis. Exp. Mol. Med. 2013, 45, e42. [Google Scholar] [CrossRef] [PubMed]
- McDonald, J.S.; Milosevic, D.; Reddi, H.V.; Grebe, S.K.; Algeciras-Schimnich, A. Analysis of circulating microRNA: Preanalytical and analytical challenges. Clin. Chem. 2011, 57, 833–840. [Google Scholar] [CrossRef] [PubMed]
- Li, L.Z.; Zhang, C.Z.; Liu, L.L.; Yi, C.; Lu, S.X.; Zhou, X.; Zhang, Z.J.; Peng, Y.H.; Yang, Y.Z.; Yun, J.P. MiR-720 inhibits tumor invasion and migration in breast cancer by targeting twist1. Carcinogenesis 2014, 35, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Schopman, N.C.; Heynen, S.; Haasnoot, J.; Berkhout, B. A miRNA-tRNA mix-up: TRNA origin of proposed miRNA. RNA Biol. 2010, 7, 573–576. [Google Scholar] [CrossRef] [PubMed]
- Langenberger, D.; Bartschat, S.; Hertel, J.; Hoffmann, S.; Tafer, H.; Stadler, P. MicroRNA or not microRNA? In Advances in Bioinformatics and Computational Biology; Norberto de Souza, O., Telles, G., Palakal, M., Eds.; Springer: Berlin Heidelberg, Germany, 2011; Volume 6832, pp. 1–9. [Google Scholar]
- Dhahbi, J.M.; Spindler, S.R.; Atamna, H.; Yamakawa, A.; Boffelli, D.; Mote, P.; Martin, D.I. 5′ tRNA halves are present as abundant complexes in serum, concentrated in blood cells, and modulated by aging and calorie restriction. BMC Genom. 2013, 14, 298. [Google Scholar] [CrossRef] [PubMed]
- Anderson, P.; Ivanov, P. TRNA fragments in human health and disease. FEBS Lett. 2014, 588, 4297–4304. [Google Scholar] [CrossRef] [PubMed]
- Selitsky, S.R.; Baran-Gale, J.; Honda, M.; Yamane, D.; Masaki, T.; Fannin, E.E.; Guerra, B.; Shirasaki, T.; Shimakami, T.; Kaneko, S.; et al. Small tRNA-derived RNAs are increased and more abundant than microRNAs in chronic hepatitis B and C. Sci. Rep. 2015, 5, 7675. [Google Scholar] [CrossRef] [PubMed]
- Law, P.T.; Qin, H.; Ching, A.K.; Lai, K.P.; Co, N.N.; He, M.; Lung, R.W.; Chan, A.W.; Chan, T.F.; Wong, N. Deep sequencing of small RNA transcriptome reveals novel non-coding RNAs in hepatocellular carcinoma. J. Hepatol. 2013, 58, 1165–1173. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Lin, L.; Zhou, W.; Wang, Z.; Ding, G.; Dong, Q.; Qin, L.; Wu, X.; Zheng, Y.; Yang, Y.; et al. Identification of miRNomes in human liver and hepatocellular carcinoma reveals miR-199a/b-3p as therapeutic target for hepatocellular carcinoma. Cancer Cell 2011, 19, 232–243. [Google Scholar] [CrossRef] [PubMed]
- Gamazon, E.R.; Innocenti, F.; Wei, R.; Wang, L.; Zhang, M.; Mirkov, S.; Ramirez, J.; Huang, R.S.; Cox, N.J.; Ratain, M.J.; et al. A genome-wide integrative study of microRNAs in human liver. BMC Genom. 2013, 14, 395. [Google Scholar] [CrossRef] [PubMed]
- Jopling, C. Liver-specific microRNA-122: Biogenesis and function. RNA Biol. 2012, 9, 137–142. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Eghbali, M. Influence of sex differences on microRNA gene regulation in disease. Biol. Sex Differ. 2014, 5, 3. [Google Scholar] [CrossRef] [PubMed]
- Roderburg, C.; Urban, G.W.; Bettermann, K.; Vucur, M.; Zimmermann, H.; Schmidt, S.; Janssen, J.; Koppe, C.; Knolle, P.; Castoldi, M.; et al. Micro-RNA profiling reveals a role for miR-29 in human and murine liver fibrosis. Hepatology 2011, 53, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Migita, K.; Komori, A.; Kozuru, H.; Jiuchi, Y.; Nakamura, M.; Yasunami, M.; Furukawa, H.; Abiru, S.; Yamasaki, K.; Nagaoka, S.; et al. Circulating microRNA profiles in patients with type-1 autoimmune hepatitis. PLoS ONE 2015, 10, e0136908. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Ge, G.; Pan, T.; Wen, D.; Gan, J. A pilot study of serum microRNAs panel as potential biomarkers for diagnosis of nonalcoholic fatty liver disease. PLoS ONE 2014, 9, e105192. [Google Scholar] [CrossRef] [PubMed]
- Li, L.M.; Wang, D.; Zen, K. MicroRNAs in drug-induced liver injury. J. Clin. Transl. Hepatol. 2014, 2, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Bala, S.; Petrasek, J.; Mundkur, S.; Catalano, D.; Levin, I.; Ward, J.; Alao, H.; Kodys, K.; Szabo, G. Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology 2012, 56, 1946–1957. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zhang, S.; Marzolf, B.; Troisch, P.; Brightman, A.; Hu, Z.; Hood, L.E.; Galas, D.J. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc. Natl. Acad. Sci. USA 2009, 106, 4402–4407. [Google Scholar] [CrossRef] [PubMed]
- Fukushima, T.; Hamada, Y.; Yamada, H.; Horii, I. Changes of micro-RNA expression in rat liver treated by acetaminophen or carbon tetrachloride—Regulating role of micro-RNA for RNA expression. J. Toxicol. Sci. 2007, 32, 401–409. [Google Scholar] [CrossRef] [PubMed]
- Starkey Lewis, P.J.; Dear, J.; Platt, V.; Simpson, K.J.; Craig, D.G.; Antoine, D.J.; French, N.S.; Dhaun, N.; Webb, D.J.; Costello, E.M.; et al. Circulating microRNAs as potential markers of human drug-induced liver injury. Hepatology 2011, 54, 1767–1776. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Ding, J.; Ning, J.; Yi, F.; Chen, J.; Zhao, D.; Zheng, J.; Liang, Z.; Hu, Z.; Du, Q. Circulating microRNA-122 as a potential biomarker for liver injury. Mol. Med. Rep. 2012, 5, 1428–1432. [Google Scholar] [CrossRef] [PubMed]
- Dolganiuc, A.; Petrasek, J.; Kodys, K.; Catalano, D.; Mandrekar, P.; Velayudham, A.; Szabo, G. MicroRNA expression profile in lieber-decarli diet-induced alcoholic and methionine choline deficient diet-induced nonalcoholic steatohepatitis models in mice. Alcohol. Clin. Exp. Res. 2009, 33, 1704–1710. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.P.; Jin, X.; Xiang, Z.; Chen, S.H.; Li, Y.M. Circulating microRNAs as potential biomarkers for alcoholic steatohepatitis. Liver Int. 2013, 33, 1257–1265. [Google Scholar] [CrossRef] [PubMed]
- Baffy, G. MicroRNAs in nonalcoholic fatty liver disease. J. Clin. Med. 2015, 4, 1977–1988. [Google Scholar] [CrossRef] [PubMed]
- Zarfeshani, A.; Ngo, S.; Sheppard, A.M. MicroRNA expression relating to dietary-induced liver steatosis and nash. J. Clin. Med. 2015, 4, 1938–1950. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Ye, Y.F.; Chen, S.H.; Yu, C.H.; Liu, J.; Li, Y.M. MicroRNA expression pattern in different stages of nonalcoholic fatty liver disease. Dig. Liver Dis. 2009, 41, 289–297. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Chen, X.; Zhang, H.; Liang, X.; Xiang, Y.; Yu, C.; Zen, K.; Li, Y.; Zhang, C.Y. Differential expression of microRNAs in mouse liver under aberrant energy metabolic status. J. Lipid Res. 2009, 50, 1756–1765. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.Y.; Xu, X.Q.; Ji, C.B.; Shi, C.M.; Guo, X.R.; Fu, J.F. Aberrant hepatic microRNA expression in nonalcoholic fatty liver disease. Cell. Physiol. Biochem. 2014, 34, 1983–1997. [Google Scholar] [CrossRef] [PubMed]
- Cheung, O.; Puri, P.; Eicken, C.; Contos, M.J.; Mirshahi, F.; Maher, J.W.; Kellum, J.M.; Min, H.; Luketic, V.A.; Sanyal, A.J. Nonalcoholic steatohepatitis is associated with altered hepatic microRNA expression. Hepatology 2008, 48, 1810–1820. [Google Scholar] [CrossRef] [PubMed]
- Cermelli, S.; Ruggieri, A.; Marrero, J.A.; Ioannou, G.N.; Beretta, L. Circulating microRNAs in patients with chronic hepatitis c and non-alcoholic fatty liver disease. PLoS ONE 2011, 6, e23937. [Google Scholar] [CrossRef] [PubMed]
- Pirola, C.J.; Fernandez Gianotti, T.; Castano, G.O.; Mallardi, P.; San Martino, J.; Mora Gonzalez Lopez Ledesma, M.; Flichman, D.; Mirshahi, F.; Sanyal, A.J.; Sookoian, S. Circulating microRNA signature in non-alcoholic fatty liver disease: From serum non-coding RNAs to liver histology and disease pathogenesis. Gut 2015, 64, 800–812. [Google Scholar] [CrossRef] [PubMed]
- Teng, K.Y.; Ghoshal, K. Role of noncoding RNAs as biomarker and therapeutic targets for liver fibrosis. Gene Expr. 2015, 16, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Friedman, S.L.; Roll, F.J.; Boyles, J.; Bissell, D.M. Hepatic lipocytes: The principal collagen-producing cells of normal rat liver. Proc. Natl. Acad. Sci. USA 1985, 82, 8681–8685. [Google Scholar] [CrossRef] [PubMed]
- Bataller, R.; Brenner, D.A. Liver fibrosis. J. Clin. Investig. 2005, 115, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wu, L.; Wang, Y.; Zhang, M.; Li, L.; Zhu, D.; Li, X.; Gu, H.; Zhang, C.Y.; Zen, K. Protective role of estrogen-induced miRNA-29 expression in carbon tetrachloride-induced mouse liver injury. J. Biol. Chem. 2012, 287, 14851–14862. [Google Scholar] [CrossRef] [PubMed]
- Peng, X.; Li, Y.; Walters, K.A.; Rosenzweig, E.R.; Lederer, S.L.; Aicher, L.D.; Proll, S.; Katze, M.G. Computational identification of hepatitis C virus associated microRNA-mRNA regulatory modules in human livers. BMC Genom. 2009, 10, 373. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.L.; Li, Y.X.; Zheng, S.Q.; Liu, M.; Li, X.; Tang, H. Suppression of hepatitis b virus replication by microRNA-199a-3p and microRNA-210. Antivir. Res. 2010, 88, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Potenza, N.; Papa, U.; Mosca, N.; Zerbini, F.; Nobile, V.; Russo, A. Human microRNA hsa-miR-125a-5p interferes with expression of hepatitis B virus surface antigen. Nucleic Acids Res. 2011, 39, 5157–5163. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.L.; Jin, W.B.; Li, J.H.; Guo, A.G. Targets for human encoded microRNAs in hbv genes. Virus Genes 2011, 42, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Kohno, T.; Tsuge, M.; Murakami, E.; Hiraga, N.; Abe, H.; Miki, D.; Imamura, M.; Ochi, H.; Hayes, C.N.; Chayama, K. Human microRNA hsa-miR-1231 suppresses hepatitis B virus replication by targeting core mRNA. J. Viral Hepat. 2014, 21, e89–e97. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Shen, A.; Rider, P.J.; Yu, Y.; Wu, K.; Mu, Y.; Hao, Q.; Liu, Y.; Gong, H.; Zhu, Y.; et al. A liver-specific microRNA binds to a highly conserved RNA sequence of hepatitis B virus and negatively regulates viral gene expression and replication. FASEB J. 2011, 25, 4511–4521. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Qiu, L.; Yan, X.; Jin, W.; Wang, Y.; Chen, L.; Wu, E.; Ye, X.; Gao, G.F.; Wang, F.; et al. Loss of miR-122 expression in patients with hepatitis B enhances hepatitis B virus replication through cyclin G1 modulated P53 activity. Hepatology 2012, 55, 730–741. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Xu, Y.; Hao, J.; Wang, S.; Li, C.; Meng, S. MiR-122 in hepatic function and liver diseases. Protein Cell 2012, 3, 364–371. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Nicolas, E.; Marks, D.; Sander, C.; Lerro, A.; Buendia, M.A.; Xu, C.; Mason, W.S.; Moloshok, T.; Bort, R.; et al. MiR-122, a mammalian liver-specific microRNA, is processed from hcr mRNA and may downregulate the high affinity cationic amino acid transporter CAT-1. RNA Biol. 2004, 1, 106–113. [Google Scholar] [CrossRef] [PubMed]
- Qiu, L.; Fan, H.; Jin, W.; Zhao, B.; Wang, Y.; Ju, Y.; Chen, L.; Chen, Y.; Duan, Z.; Meng, S. MiR-122-induced down-regulation of HO-1 negatively affects miR-122-mediated suppression of HBV. Biochem. Biophys. Res. Commun. 2010, 398, 771–777. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Liu, X.; Lin, L.; Hou, J.; Li, N.; Wang, C.; Wang, P.; Zhang, Q.; Zhang, P.; Zhou, W.; et al. MicroRNA-99a inhibits hepatocellular carcinoma growth and correlates with prognosis of patients with hepatocellular carcinoma. J. Biol. Chem. 2011, 286, 36677–36685. [Google Scholar] [CrossRef] [PubMed]
- Takata, A.; Otsuka, M.; Kojima, K.; Yoshikawa, T.; Kishikawa, T.; Yoshida, H.; Koike, K. MicroRNA-22 and microRNA-140 suppress NF-κB activity by regulating the expression of NF-κB coactivators. Biochem. Biophys. Res. Commun. 2011, 411, 826–831. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Takeshita, F.; Hino, Y.; Fukunaga, S.; Kudo, Y.; Tamaki, A.; Matsunaga, J.; Takahashi, R.U.; Takata, T.; Shimamoto, A.; et al. MiR-22 represses cancer progression by inducing cellular senescence. J. Cell Biol. 2011, 193, 409–424. [Google Scholar] [CrossRef]
- Shi, C.; Xu, X. MicroRNA-22 is down-regulated in hepatitis B virus-related hepatocellular carcinoma. Biomed. Pharmacother. 2013, 67, 375–380. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Wang, X.; Ding, X.; Li, Y.; Zhang, X.; Xie, P.; Yang, J.; Wang, S. MicroRNA-141 represses HBV replication by targeting PPARA. PLoS ONE 2012, 7, e34165. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, E.; Ma, Z.; Pei, R.; Jiang, M.; Schlaak, J.F.; Roggendorf, M.; Lu, M. Modulation of hepatitis b virus replication and hepatocyte differentiation by microRNA-1. Hepatology 2011, 53, 1476–1485. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Tang, S.; Xia, L.; Du, R.; Xie, H.; Song, J.; Fan, R.; Bi, Q.; Chen, Z.; Yang, G.; et al. MicroRNA-501 promotes hbv replication by targeting HBXIP. Biochem. Biophys. Res. Commun. 2013, 430, 1228–1233. [Google Scholar] [CrossRef] [PubMed]
- Jin, W.B.; Wu, F.L.; Kong, D.; Guo, A.G. Hbv-encoded microRNA candidate and its target. Comput. Biol. Chem. 2007, 31, 124–126. [Google Scholar] [CrossRef] [PubMed]
- Li, L.M.; Hu, Z.B.; Zhou, Z.X.; Chen, X.; Liu, F.Y.; Zhang, J.F.; Shen, H.B.; Zhang, C.Y.; Zen, K. Serum microRNA profiles serve as novel biomarkers for HBV infection and diagnosis of HBV-positive hepatocarcinoma. Cancer Res. 2010, 70, 9798–9807. [Google Scholar] [CrossRef] [PubMed]
- Arataki, K.; Hayes, C.N.; Akamatsu, S.; Akiyama, R.; Abe, H.; Tsuge, M.; Miki, D.; Ochi, H.; Hiraga, N.; Imamura, M.; et al. Circulating microRNA-22 correlates with microRNA-122 and represents viral replication and liver injury in patients with chronic hepatitis B. J. Med. Virol. 2013, 85, 789–798. [Google Scholar] [CrossRef] [PubMed]
- Akamatsu, S.; Hayes, C.N.; Tsuge, M.; Miki, D.; Akiyama, R.; Abe, H.; Ochi, H.; Hiraga, N.; Imamura, M.; Takahashi, S.; et al. Differences in serum microRNA profiles in hepatitis B and C virus infection. J. Infect. 2015, 70, 273–287. [Google Scholar] [CrossRef] [PubMed]
- Giray, B.G.; Emekdas, G.; Tezcan, S.; Ulger, M.; Serin, M.S.; Sezgin, O.; Altintas, E.; Tiftik, E.N. Profiles of serum microRNAs; miR-125b-5p and miR223–3p serve as novel biomarkers for HBV-positive hepatocellular carcinoma. Mol. Biol. Rep. 2014, 41, 4513–4519. [Google Scholar] [CrossRef] [PubMed]
- Ren, M.; Qin, D.; Li, K.; Qu, J.; Wang, L.; Wang, Z.; Huang, A.; Tang, H. Correlation between hepatitis B virus protein and microRNA processor drosha in cells expressing HBV. Antivir. Res. 2012, 94, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Steuerwald, N.M.; Parsons, J.C.; Bennett, K.; Bates, T.C.; Bonkovsky, H.L. Parallel microRNA and mRNA expression profiling of (genotype 1b) human hepatoma cells expressing hepatitis C virus. Liver Int. 2010, 30, 1490–1504. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, I.M.; Cheng, G.; Wieland, S.; Volinia, S.; Croce, C.M.; Chisari, F.V.; David, M. Interferon modulation of cellular microRNAs as an antiviral mechanism. Nature 2007, 449, 919–922. [Google Scholar] [CrossRef] [PubMed]
- Jopling, C.L.; Yi, M.; Lancaster, A.M.; Lemon, S.M.; Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific microRNA. Science 2005, 309, 1577–1581. [Google Scholar] [CrossRef] [PubMed]
- Lanford, R.E.; Hildebrandt-Eriksen, E.S.; Petri, A.; Persson, R.; Lindow, M.; Munk, M.E.; Kauppinen, S.; Orum, H. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis C virus infection. Science 2010, 327, 198–201. [Google Scholar] [CrossRef] [PubMed]
- Banaudha, K.; Kaliszewski, M.; Korolnek, T.; Florea, L.; Yeung, M.L.; Jeang, K.T.; Kumar, A. MicroRNA silencing of tumor suppressor DLC-1 promotes efficient hepatitis C virus replication in primary human hepatocytes. Hepatology 2011, 53, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Ishida, H.; Tatsumi, T.; Hosui, A.; Nawa, T.; Kodama, T.; Shimizu, S.; Hikita, H.; Hiramatsu, N.; Kanto, T.; Hayashi, N.; et al. Alterations in microRNA expression profile in HCV-infected hepatoma cells: Involvement of miR-491 in regulation of HCV replication via the PI3 kinase/akt pathway. Biochem. Biophys. Res. Commun. 2011, 412, 92–97. [Google Scholar] [CrossRef] [PubMed]
- Murakami, Y.; Aly, H.H.; Tajima, A.; Inoue, I.; Shimotohno, K. Regulation of the hepatitis C virus genome replication by miR-199a. J. Hepatol. 2009, 50, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Tian, Q.; Zheng, J.; Bonkovsky, H.L. MicroRNA-196 represses Bach1 protein and hepatitis C virus gene expression in human hepatoma cells expressing hepatitis C viral proteins. Hepatology 2010, 51, 1494–1504. [Google Scholar] [CrossRef] [PubMed]
- Bandyopadhyay, S.; Friedman, R.C.; Marquez, R.T.; Keck, K.; Kong, B.; Icardi, M.S.; Brown, K.E.; Burge, C.B.; Schmidt, W.N.; Wang, Y.; et al. Hepatitis C virus infection and hepatic stellate cell activation downregulate miR-29: MiR-29 overexpression reduces hepatitis C viral abundance in culture. J. Infect. Dis. 2011, 203, 1753–1762. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.C.; Yeh, Y.J.; Tseng, C.P.; Hsu, S.D.; Chang, Y.L.; Sakamoto, N.; Huang, H.D. Let-7b is a novel regulator of hepatitis C virus replication. Cell. Mol. Life Sci. 2012, 69, 2621–2633. [Google Scholar] [CrossRef] [PubMed]
- Bhanja Chowdhury, J.; Shrivastava, S.; Steele, R.; Di Bisceglie, A.M.; Ray, R.; Ray, R.B. Hepatitis C virus infection modulates expression of interferon stimulatory gene IFITM1 by upregulating miR-130a. J. Virol. 2012, 86, 10221–10225. [Google Scholar] [CrossRef] [PubMed]
- Shirasaki, T.; Honda, M.; Shimakami, T.; Horii, R.; Yamashita, T.; Sakai, Y.; Sakai, A.; Okada, H.; Watanabe, R.; Murakami, S.; et al. MicroRNA-27a regulates lipid metabolism and inhibits hepatitis C virus replication in human hepatoma cells. J. Virol. 2013, 87, 5270–5286. [Google Scholar] [CrossRef] [PubMed]
- You, J.S.; Jones, P.A. Cancer genetics and epigenetics: Two sides of the same coin? Cancer Cell 2012, 22, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Ashworth, A.; Lord, C.J.; Reis-Filho, J.S. Genetic interactions in cancer progression and treatment. Cell 2011, 145, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Pogribny, I.P.; Rusyn, I. Role of epigenetic aberrations in the development and progression of human hepatocellular carcinoma. Cancer Lett. 2014, 342, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Bertino, G.; Ardiri, A.M.; Calvagno, G.S.; Bertino, N.; Boemi, P.M. Prognostic and diagnostic value of des-γ-carboxy prothrombin in liver cancer. Drug News Perspect. 2010, 23, 498–508. [Google Scholar] [PubMed]
- Durazo, F.A.; Blatt, L.M.; Corey, W.G.; Lin, J.H.; Han, S.; Saab, S.; Busuttil, R.W.; Tong, M.J. Des-γ-carboxyprothrombin, α-fetoprotein and AFP-L3 in patients with chronic hepatitis, cirrhosis and hepatocellular carcinoma. J. Gastroenterol. Hepatol. 2008, 23, 1541–1548. [Google Scholar] [CrossRef] [PubMed]
- Borel, F.; Konstantinova, P.; Jansen, P.L. Diagnostic and therapeutic potential of miRNA signatures in patients with hepatocellular carcinoma. J. Hepatol. 2012, 56, 1371–1383. [Google Scholar] [CrossRef] [PubMed]
- Ladeiro, Y.; Couchy, G.; Balabaud, C.; Bioulac-Sage, P.; Pelletier, L.; Rebouissou, S.; Zucman-Rossi, J. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology 2008, 47, 1955–1963. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; He, X. The role of microRNAs in liver cancer progression. Br. J. Cancer 2011, 104, 235–240. [Google Scholar] [CrossRef] [PubMed]
- Qi, P.; Cheng, S.Q.; Wang, H.; Li, N.; Chen, Y.F.; Gao, C.F. Serum microRNAs as biomarkers for hepatocellular carcinoma in chinese patients with chronic hepatitis B virus infection. PLoS ONE 2011, 6, e28486. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Lee, A.T.; Ma, J.Z.; Wang, J.; Ren, J.; Yang, Y.; Tantoso, E.; Li, K.B.; Ooi, L.L.; Tan, P.; et al. Profiling microRNA expression in hepatocellular carcinoma reveals microRNA-224 up-regulation and apoptosis inhibitor-5 as a microRNA-224-specific target. J. Biol. Chem. 2008, 283, 13205–13215. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, L.P.; Meng, Z.Q. Serum miR-224 reflects stage of hepatocellular carcinoma and predicts survival. Biomed. Res. Int. 2015, 2015, 731781. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.; Ghosh, A.; Datta, S.; Dasgupta, D.; Das, S.; Ray, S.; Gupta, S.; Datta, S.; Chowdhury, A.; Chatterjee, R.; et al. Hepatic miR-126 is a potential plasma biomarker for detection of hepatitis B virus infected hepatocellular carcinoma. Int. J. Cancer 2016. [Google Scholar] [CrossRef] [PubMed]
- Barshack, I.; Meiri, E.; Rosenwald, S.; Lebanony, D.; Bronfeld, M.; Aviel-Ronen, S.; Rosenblatt, K.; Polak-Charcon, S.; Leizerman, I.; Ezagouri, M.; et al. Differential diagnosis of hepatocellular carcinoma from metastatic tumors in the liver using microRNA expression. Int. J. Biochem. Cell Biol. 2010, 42, 1355–1362. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.J.; Chong, Y.; Guo, Z.W.; Xie, C.; Yang, X.J.; Zhang, Q.; Li, S.P.; Xiong, Y.; Yuan, Y.; Min, J.; et al. A serum microRNA classifier for early detection of hepatocellular carcinoma: A multicentre, retrospective, longitudinal biomarker identification study with a nested case-control study. Lancet Oncol. 2015, 16, 804–815. [Google Scholar] [CrossRef]
- Ji, J.; Shi, J.; Budhu, A.; Yu, Z.; Forgues, M.; Roessler, S.; Ambs, S.; Chen, Y.; Meltzer, P.S.; Croce, C.M.; et al. MicroRNA expression, survival, and response to interferon in liver cancer. N. Engl. J. Med. 2009, 361, 1437–1447. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.T.; Dong, Q.Z.; Sheng, Y.Y.; Wei, J.W.; Wang, G.; Zhou, H.J.; Ren, N.; Jia, H.L.; Ye, Q.H.; Qin, L.X. MicroRNA-29a-5p is a novel predictor for early recurrence of hepatitis b virus-related hepatocellular carcinoma after surgical resection. PLoS ONE 2012, 7, e52393. [Google Scholar] [CrossRef] [PubMed]
- Kourtidis, A.; Ngok, S.P.; Pulimeno, P.; Feathers, R.W.; Carpio, L.R.; Baker, T.R.; Carr, J.M.; Yan, I.K.; Borges, S.; Perez, E.A.; et al. Distinct E-cadherin-based complexes regulate cell behaviour through miRNA processing or Src and p120 catenin activity. Nat. Cell Biol. 2015, 17, 1145–1157. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M.; et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef] [PubMed]
- Anwar, S.L.; Albat, C.; Krech, T.; Hasemeier, B.; Schipper, E.; Schweitzer, N.; Vogel, A.; Kreipe, H.; Lehmann, U. Concordant hypermethylation of intergenic microRNA genes in human hepatocellular carcinoma as new diagnostic and prognostic marker. Int. J. Cancer 2013, 133, 660–670. [Google Scholar] [CrossRef] [PubMed]
- Anwar, S.L.; Lehmann, U. DNA methylation, microRNAs, and their crosstalk as potential biomarkers in hepatocellular carcinoma. World J. Gastroenterol. 2014, 20, 7894–7913. [Google Scholar] [CrossRef] [PubMed]
- Buurman, R.; Gurlevik, E.; Schaffer, V.; Eilers, M.; Sandbothe, M.; Kreipe, H.; Wilkens, L.; Schlegelberger, B.; Kuhnel, F.; Skawran, B. Histone deacetylases activate hepatocyte growth factor signaling by repressing microRNA-449 in hepatocellular carcinoma cells. Gastroenterology 2012, 143, 811–820. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Cui, Y.; Wang, W.; Gu, J.; Guo, S.; Ma, K.; Luo, X. Hypomethylation of the hsa-miR-191 locus causes high expression of hsa-miR-191 and promotes the epithelial-to-mesenchymal transition in hepatocellular carcinoma. Neoplasia 2011, 13, 841–853. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hayes, C.N.; Chayama, K. MicroRNAs as Biomarkers for Liver Disease and Hepatocellular Carcinoma. Int. J. Mol. Sci. 2016, 17, 280. https://doi.org/10.3390/ijms17030280
Hayes CN, Chayama K. MicroRNAs as Biomarkers for Liver Disease and Hepatocellular Carcinoma. International Journal of Molecular Sciences. 2016; 17(3):280. https://doi.org/10.3390/ijms17030280
Chicago/Turabian StyleHayes, C. Nelson, and Kazuaki Chayama. 2016. "MicroRNAs as Biomarkers for Liver Disease and Hepatocellular Carcinoma" International Journal of Molecular Sciences 17, no. 3: 280. https://doi.org/10.3390/ijms17030280
APA StyleHayes, C. N., & Chayama, K. (2016). MicroRNAs as Biomarkers for Liver Disease and Hepatocellular Carcinoma. International Journal of Molecular Sciences, 17(3), 280. https://doi.org/10.3390/ijms17030280