Multiple Evolutionary Selections Involved in Synonymous Codon Usages in the Streptococcus agalactiae Genome
Abstract
:1. Introduction
2. Results
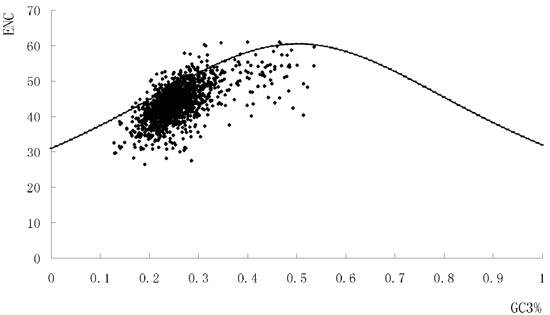
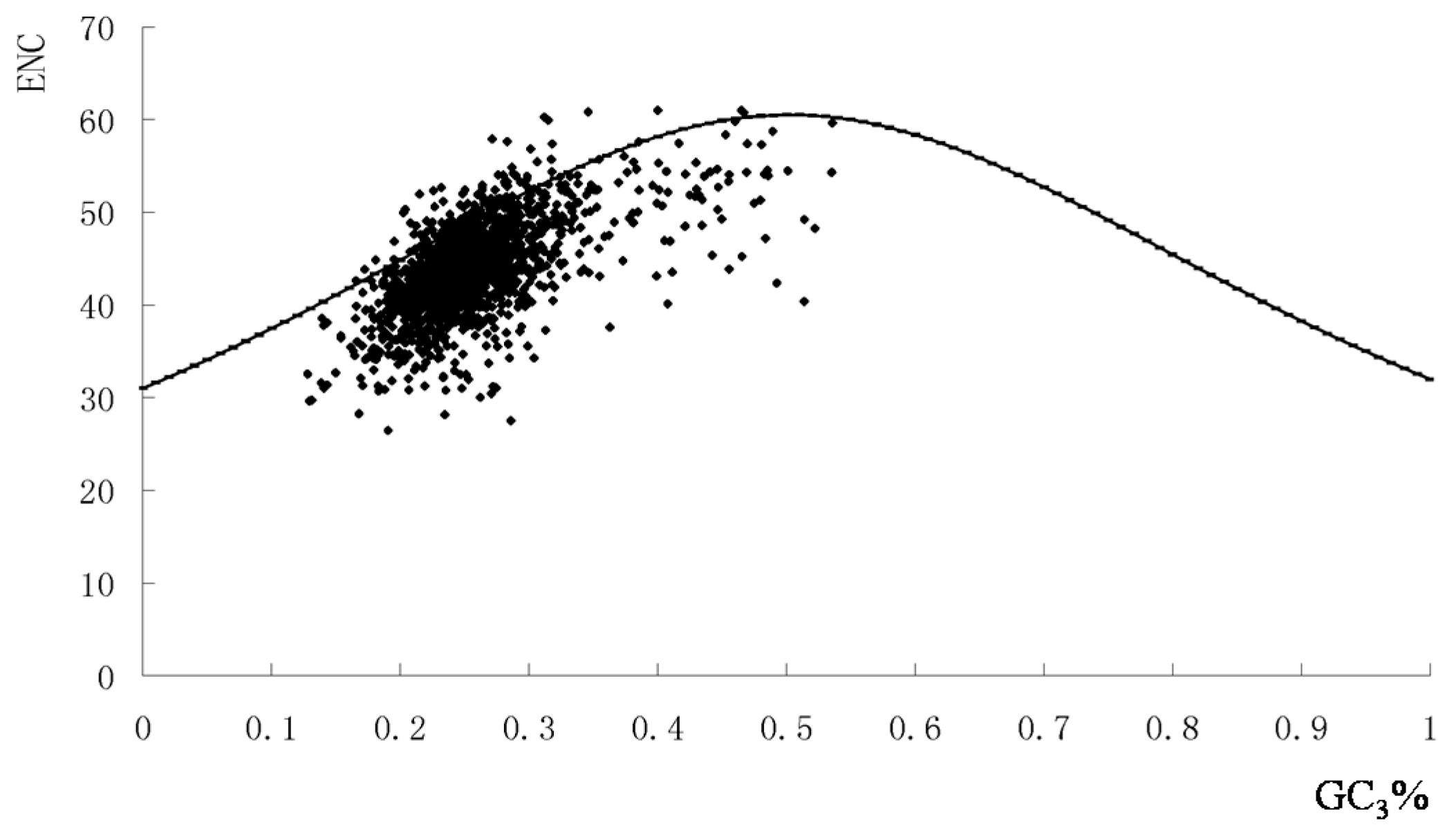
2.1. The Synonymous Codon Usage Pattern of S. agalactiae Genomes
2.2. The Overall Codon Usage Pattern of S. agalactiae
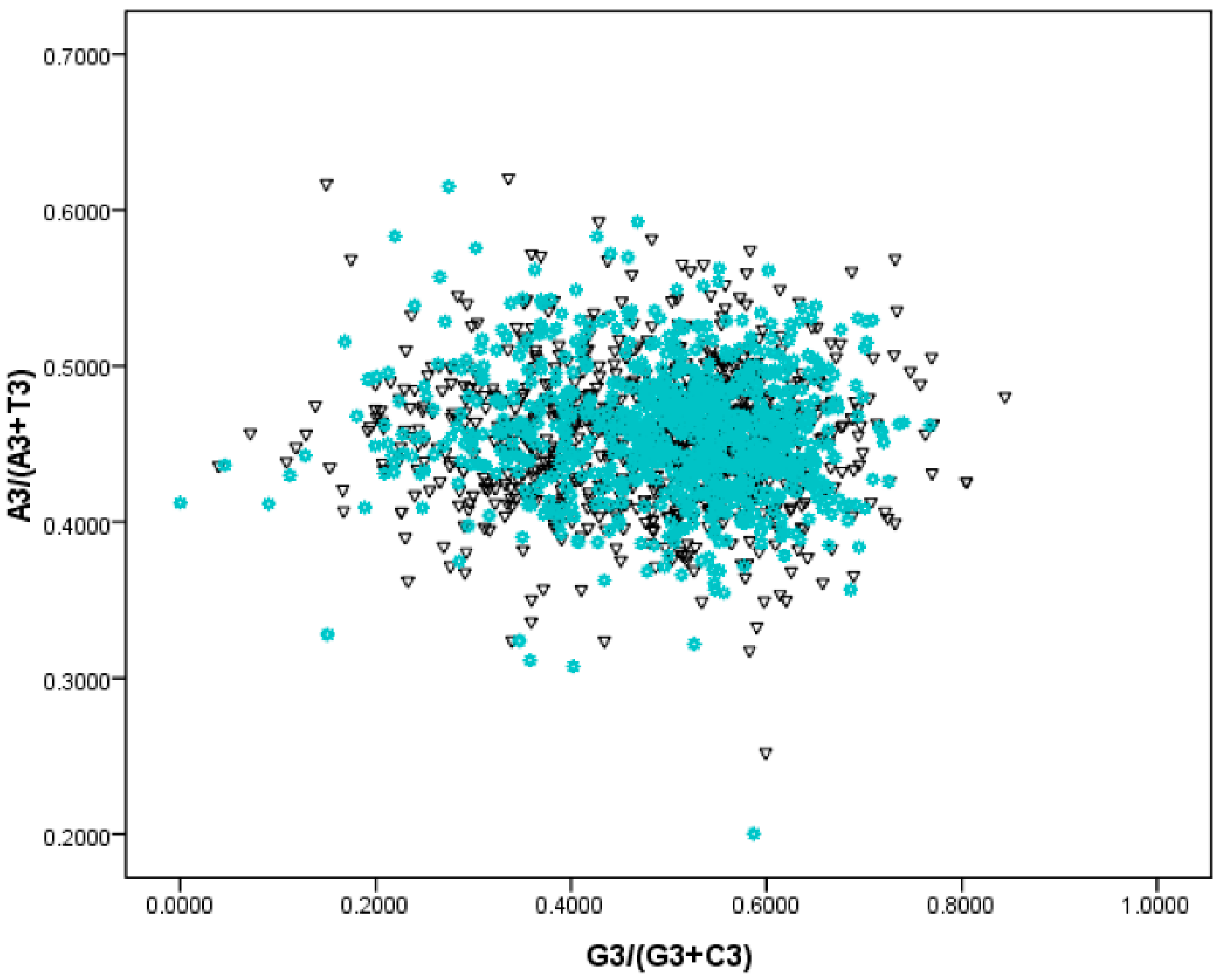
2.3. Using PR—2 Plots to Differentiate Replication and Transcription Associated Biases
2.4. Genetic Diversities Based on Synonymous Codon Usage by PCA
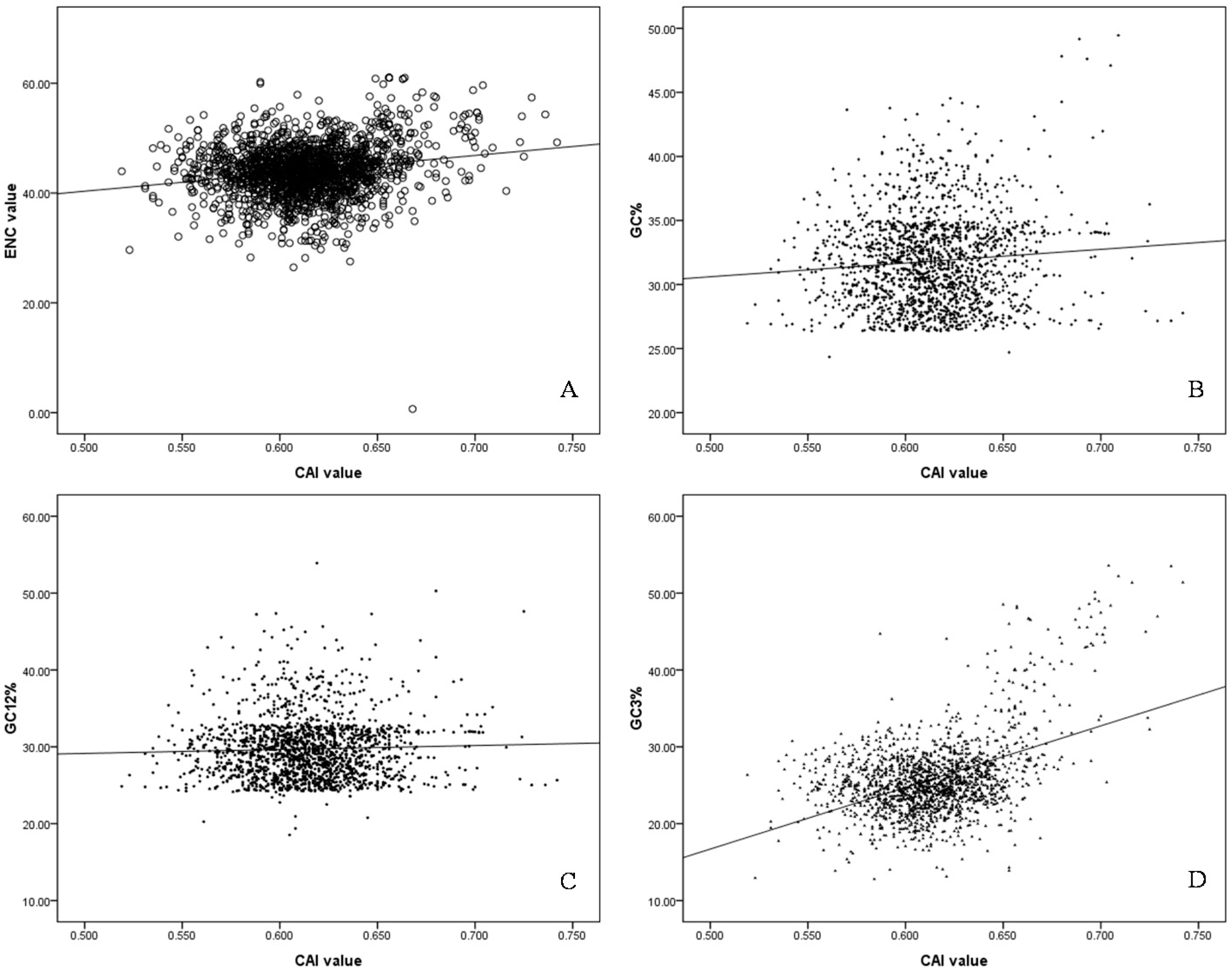
2.5. Codon Adaptation Index (CAI) and Identification of Translationally Optimal Codons
2.6. Effects of the Overall Codon Usage of Hosts on that of S. agalactiae
3. Discussion
4. Materials and Methods
4.1. Sequence Data
4.2. Codon Usage Pattern of Functional Genes of S. agalactiae
4.3. Analysis of the Genetic Diversities of the S. agalactiae ORFs by Principle Component Analysis (PCA)
4.4. Nucleotide Composition Statistics for Genes in S. agalactiae
4.5. Replication and Transcription Associated Biases Differentiation
4.6. Preferred Gene Analysis by Codon Adaptation Index (CAI)
4.7. Computation of tRNA Adaptation Index (tAI) and Frequency of Optimal Codons (Fop)
4.8. Estimating Effects of the Overall Codon Usage of Human and Tilapia on that of S. agalactiae
4.9. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kong, W.; Kong, J.; Hu, S.; Lu, W.; Wang, K.; Ji, M. Enhanced expression of PCV2 capsid protein in Escherichia coli and Lactococcus lactis by codon optimization. World J. Microbiol. Biotechnol. 2011, 27, 651–657. [Google Scholar] [CrossRef]
- Bulmer, M. Codon usage and intragenic position. J. Theor. Biol. 1988, 133, 67–71. [Google Scholar] [CrossRef]
- Sharp, P.M.; Cowe, E.; Higgin, D.G.; Shields, D.C.; Wolfe, K.H.; Wright, F. Codon usage patterns in Escherichia coli, Bacillus subtilis, Saccharomyces cerevisiae, Schizosaccharomyces pombe, Drosophila melanogaster and Homo sapiens; a review of the considerable within-species diversity. Nucleic Acids. Res. 1988, 16, 8207–8211. [Google Scholar] [CrossRef] [PubMed]
- Karlin, S.; Mrázek, J. What drives codon choices in human genes? J. Mol. Biol. 1996, 262, 459–472. [Google Scholar] [CrossRef] [PubMed]
- Kanaya, S.; Kinouchi, M.; Abe, T.; Kudo, Y.; Yamada, Y.; Nishi, T.; Mori, H.; Ikemura, T. Analysis of codon usage diversity of bacterial genes with a self-organizing map (SOM): Characterization of horizontally transferred genes with emphasis on the E. coli O157 genome. Gene 2001, 276, 89–99. [Google Scholar] [CrossRef]
- Yadav, M.K.; Swati, D. Comparative genome analysis of six malarial parasites using codon usage bias based tools. Bioinformation 2012, 8, 1230–1239. [Google Scholar] [CrossRef] [PubMed]
- Doolittle, W.F. You are what you eat: A gene trandfer ratchet could account for bacterial genes in eukaryotic nuclear genomes. Trends Genet. 1998, 14, 307–311. [Google Scholar] [CrossRef]
- Ochman, H.; Lawrence, J.G.; Groisman, E.A. Lateral gene transfer and the nature of bacterial innovation. Nature 2000, 405, 299–304. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Huang, J.; Liang, C.; Liu, S.; Xie, J.; Liu, C. Is there a close relationship between synonymous codon bias and codon-anticodon binding strength in human genes? Chin. Sci. Bull. 2001, 46, 1015–1019. [Google Scholar] [CrossRef]
- Woese, C.R. On the evolution of cells. Proc. Natl. Acad. Sci. USA 2002, 99, 8742–8747. [Google Scholar] [CrossRef] [PubMed]
- Romero, H.; Zavala, A.; Musto, H.; Bernardi, G. The influence of translational selection on codon usage in fishes from the family Cyprinidae. Gene 2003, 317, 141–147. [Google Scholar] [CrossRef]
- Guo, F.B.; Yuan, J.B. Codon usages of genes on chromosome, and surprisingly, genes in plasmid are primarily affected by strandspecific mutational biases in Lawsonia intracellularis. DNA Res. 2009, 16, 91–104. [Google Scholar] [CrossRef] [PubMed]
- Evans, J.J.; Klesius, P.H.; Gilbert, P.M. Characterization of β-haemolytic Group B Streptococcus agalactiae in cultured seabream, Sparus auratus L., and wild mullet, Liza klunzingeri (Day), in Kuwait. J. Fish Dis. 2002, 25, 505–551. [Google Scholar] [CrossRef]
- Evans, J.J.; Pasnik, D.J.; Klesius, P.H.; Ai-Ablani, S. First report of Streptococcus agalactiae and Lactococcus garvieae from a wild bottlenose dolphin (Tursiops truncatus). J. Wildl. Dis. 2006, 42, 561–569. [Google Scholar] [CrossRef] [PubMed]
- Mian, G.F.; Godoy, D.T.; Leal, C.A.; Yuhara, T.Y.; Costa, G.M.; Figueiredo, H.C. Aspects of the natural history and virulence of S.agalactiae infection in Nile tilapia. Vet. Microbiol. 2009, 136, 180–183. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Wang, K.Y.; Huang, X.L.; Chen, D.F.; Li, C.W.; Ren, S.Y.; Liao, Y.T.; Zhou, Z.Y.; Liu, Q.F.; Du, Z.J.; et al. Streptococcus agalactiae, an emerging pathogen for cultured ya-fish, Schizothorax prenanti, in China. Transbound. Emerg. Dis. 2012, 59, 369–375. [Google Scholar] [CrossRef] [PubMed]
- Delannoy, C.M.; Crumlish, M.; Fontaine, M.C.; Pollock, J.; Foster, G.; Dagleish, M.P.; Turnbull, J.F.; Zadoks, R.N. Human Streptococcus agalactiae strains in aquatic mammals and fish. BMC Microbiol. 2013, 13, 41. [Google Scholar] [CrossRef] [PubMed]
- Zamri-Saad, M.; Amal, M.N.; Siti-Zahrah, A. Pathological changes in red tilapias (Oreochromis spp.) naturally infected by Streptococcus agalactiae. J. Comp. Pathol. 2010, 143, 227–229. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, S.; Omar, N.; Yusoff, S.M.; Obukwho, E.B.; Nwunuji, T.P.; Hanan, L.; Samad, J. Clinicopathological features and immunohistochemical detection of antigens in acute experimental Streptococcus agalactiae infection in red tilapia (Oreochromis spp.). Springerplus 2013, 2, 286. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.W.; Liu, L.; Huang, P.R.; Fang, W.; Luo, Z.P.; Peng, H.L.; Wang, Y.X.; Li, A.X. Chronic streptococcosis in Nile tilapia, Oreochromisniloticus (L.), caused by Streptococcus agalactiae. J. Fish Dis. 2014, 37, 757–763. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.P.; Liu, Z.X.; Hao, L.; Ma, J.Y.; Liang, Z.L.; Li, Y.G.; Ke, H. Analysing codon usage bias of cyprinid herpesvirus 3 and adaptation of this virus to the hosts. J. Fish Dis. 2015, 38, 665–673. [Google Scholar]
- Ikemura, T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes: A proposal for a synonymous codon choice that is optimal for the E. coli translational system. J. Mol. Biol. 1981, 151, 389–409. [Google Scholar] [CrossRef]
- Duret, L. tRNA gene number and codon usage in the C. elegans genome are co-adapted for optimal translation of highly expressed genes. Trends Genet. 2000, 16, 287–289. [Google Scholar] [CrossRef]
- Roy, A.; Mukhopadhyay, S.; Sarkar, I.; Sen, A. Comparative investigation of the various determinants that influence the codon and amino acid usage patterns in the genus Bifidobacterium. World J. Microbiol. Biotechnol. 2015, 31, 959–981. [Google Scholar] [CrossRef] [PubMed]
- Moriyama, E.N.; Powell, J.R. Codon usage bias and tRNA abundance in Drosophila. J. Mol. Evol. 1997, 45, 514–523. [Google Scholar] [CrossRef] [PubMed]
- Behura, S.K.; Severson, D.W. Coadaptation of isoacceptor tRNA genes and codon usage for translation efficienty in Aedes aegypti and Anopheles gambiae. Insect Mol. Biol. 2011, 20, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Sanjukta, R.; Faroogi, M.S.; Sharma, N.; Rai, A.; Mishra, D.C.; Singh, D.P. Trends in the codon usage patterns of Chromohalobacter salexigens genes. Bioinformation 2012, 8, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- GenBank database. Available online: http://www.ncbi.nlm.nih.gov/GenBank (accessed on 19 February 2016).
- Codon usage database. Available online: http://www.kazusa.or.jp/codon/ (accessed on 19 February 2016).
- Wright, F. The “effective number of codons” used in a gene. Gene 1990, 87, 23–29. [Google Scholar] [CrossRef]
- Fuglsang, A. The “effective number of codons” revisited. Biochem. Biophys. Res. Commun. 2004, 317, 957–964. [Google Scholar] [CrossRef] [PubMed]
- Lobry, J.R. Asymmetric substitution patterns in the two DNA strands of bacteria. Mol. Biol. Evol. 1996, 13, 660–665. [Google Scholar] [CrossRef] [PubMed]
- McLean, M.J.; Wolfe, K.H.; Devine, K.M. Base composition skews, replication orientation, and gene orientation in 12 prokaryote genomes. J. Mol. Evol. 1998, 47, 691–696. [Google Scholar] [CrossRef] [PubMed]
- Lobry, J.R.; Sueoka, N. Asymmetric directional mutation pressures in bacteria. Genome Biol. 2002, 3, RESEARCH0058. [Google Scholar] [CrossRef] [PubMed]
- Sharp, P.M.; Li, W.H. The rate of synonymous substitution in enterbacterial genes is inversely related to codon usage bias. Mol. Biol. Evol. 1987, 4, 222–230. [Google Scholar] [PubMed]
- Sen, A.; Sur, S.; Bothra, A.K.; Benson, D.R.; Normand, P.; Tisa, L.S. The implication of life style on codon usage patterns and predicted highly expressed genes for three Frankia genomes. Anton Van Leeuwenhoek 2008, 93, 335–346. [Google Scholar] [CrossRef] [PubMed]
- Lü, H.; Zhao, W.M.; Zheng, Y.; Wang, H.; Qi, M.; Yu, X.P. Analysis of synonymous codon usage bias in Chlamydia. Acta Biochim. Biophys. Sin. 2005, 37, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Dos Reis, M.; Savva, R.; Wernisch, L. Solving the riddle of codon usage preferences: A test for translational selection. Nucleic Acids Res. 2004, 32, 5036–5044. [Google Scholar] [CrossRef] [PubMed]
- Dos Reis, M.; Wernisch, L.; Sawa, R. Unexpected correlations between gene expression and codon usage bias from microarray data for the whole Escherichia coli K-12 genome. Nucletic Acids Res. 2003, 31, 6976–6985. [Google Scholar] [CrossRef]
- Peden, J. Analysis of Codon Usage. Ph.D. Thesis, University of Nottingham, Department of Genetics, Nottingham, UK, 1997. [Google Scholar]
- Zhou, J.H.; Ding, Y.Z.; He, Y.; Chu, Y.F.; Zhao, P.; Ma, L.Y.; Wang, X.J.; Li, X.R.; Liu, Y.S. The effect of multiple evolutionary selections on synonymous codon usage of genes in the Mycoplasma bovis genome. PLoS ONE 2014, 9, e108949. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Liu, Y.S.; Zhou, J.H.; Chen, H.T.; Ma, L.N.; Ding, Y.Z.; Liu, W.Q.; Gu, Y.X.; Zhang, J. Analysis of codon usage in Newcastle disease virus. Virus Genes 2011, 42, 45–25. [Google Scholar] [CrossRef] [PubMed]

| Amino Acid | Codon | RSCU a |
|---|---|---|
| Ala | GCA | 1.39 |
| GCC | 0.44 | |
| GCG | 0.37 | |
| GCU | 1.80 | |
| Arg | AGA | 1.55 |
| AGG | 0.56 | |
| CGA | 0.72 | |
| CGC | 0.67 | |
| CGG | 0.18 | |
| CGU | 2.32 | |
| Asn | AAC | 0.50 |
| AAU | 1.50 | |
| Asp | GAC | 0.43 |
| GAU | 1.57 | |
| Cys | UGC | 0.41 |
| UGU | 1.47 | |
| Gln | CAA | 1.55 |
| CAG | 0.44 | |
| Glu | GAA | 1.48 |
| GAG | 0.52 | |
| Gly | GGA | 1.28 |
| GGC | 0.46 | |
| GGG | 0.47 | |
| GGU | 1.79 | |
| His | CAC | 0.53 |
| CAU | 1.46 | |
| Ile | AUA | 0.55 |
| AUC | 0.61 | |
| AUU | 1.84 | |
| Leu | CUA | 0.73 |
| CUC | 0.35 | |
| CUG | 0.25 | |
| CUU | 1.22 | |
| UUA | 2.45 | |
| UUG | 0.98 | |
| Lys | AAA | 1.52 |
| AAG | 0.48 | |
| Phe | UUC | 0.45 |
| UUU | 1.56 | |
| Pro | CCA | 1.80 |
| CCC | 0.32 | |
| CCG | 0.32 | |
| CCU | 1.55 | |
| Ser | AGC | 0.57 |
| AGU | 1.43 | |
| UCA | 1.76 | |
| UCC | 0.32 | |
| UCG | 0.32 | |
| UCU | 1.61 | |
| Thr | ACA | 1.59 |
| ACC | 0.48 | |
| ACG | 0.41 | |
| ACU | 1.52 | |
| Tyr | UAC | 0.51 |
| UAU | 1.48 | |
| Val | GUA | 1.12 |
| GUC | 0.51 | |
| GUG | 0.44 | |
| GUU | 1.92 |
| Skew | Axis 1 | Axis 2 |
|---|---|---|
| GC skew | r = 0.022, p > 0.05 | r = 0.006, p > 0.05 |
| AT skew | r = −0.015, p > 0.05 | r = 0.01, p > 0.05 |
| GC12 skew | r = 0.003, p > 0.05 | r = 0.13, p > 0.05 |
| AT12 skew | r = 0.016, p > 0.05 | r = 0.019, p > 0.05 |
| GC3 skew | r = 0.067 *, p < 0.05 | r = 0.056 *, p < 0.05 |
| AT3 skew | r = 0.628 **, p < 0.01 | r = 0.555 *, p < 0.01 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, Y.-P.; Ke, H.; Liang, Z.-L.; Liu, Z.-X.; Hao, L.; Ma, J.-Y.; Li, Y.-G. Multiple Evolutionary Selections Involved in Synonymous Codon Usages in the Streptococcus agalactiae Genome. Int. J. Mol. Sci. 2016, 17, 277. https://doi.org/10.3390/ijms17030277
Ma Y-P, Ke H, Liang Z-L, Liu Z-X, Hao L, Ma J-Y, Li Y-G. Multiple Evolutionary Selections Involved in Synonymous Codon Usages in the Streptococcus agalactiae Genome. International Journal of Molecular Sciences. 2016; 17(3):277. https://doi.org/10.3390/ijms17030277
Chicago/Turabian StyleMa, Yan-Ping, Hao Ke, Zhi-Ling Liang, Zhen-Xing Liu, Le Hao, Jiang-Yao Ma, and Yu-Gu Li. 2016. "Multiple Evolutionary Selections Involved in Synonymous Codon Usages in the Streptococcus agalactiae Genome" International Journal of Molecular Sciences 17, no. 3: 277. https://doi.org/10.3390/ijms17030277
APA StyleMa, Y.-P., Ke, H., Liang, Z.-L., Liu, Z.-X., Hao, L., Ma, J.-Y., & Li, Y.-G. (2016). Multiple Evolutionary Selections Involved in Synonymous Codon Usages in the Streptococcus agalactiae Genome. International Journal of Molecular Sciences, 17(3), 277. https://doi.org/10.3390/ijms17030277