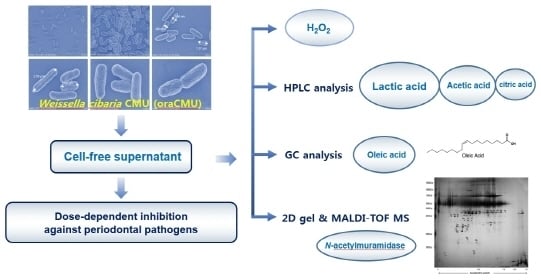
Characterization of Antibacterial Cell-Free Supernatant from Oral Care Probiotic Weissella cibaria, CMU
Abstract
1. Introduction
2. Results
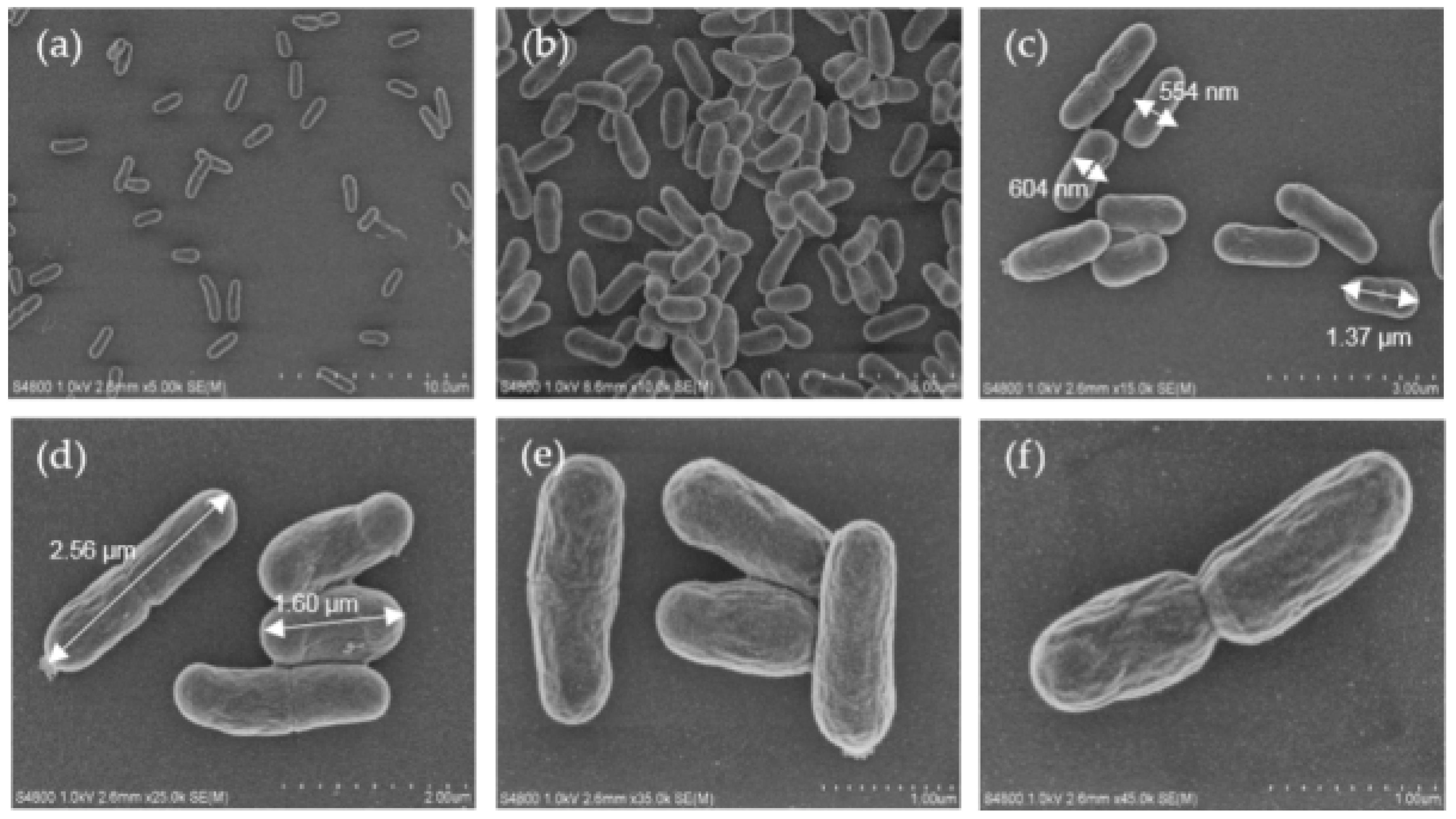
2.1. Cell Morphology of oraCMU
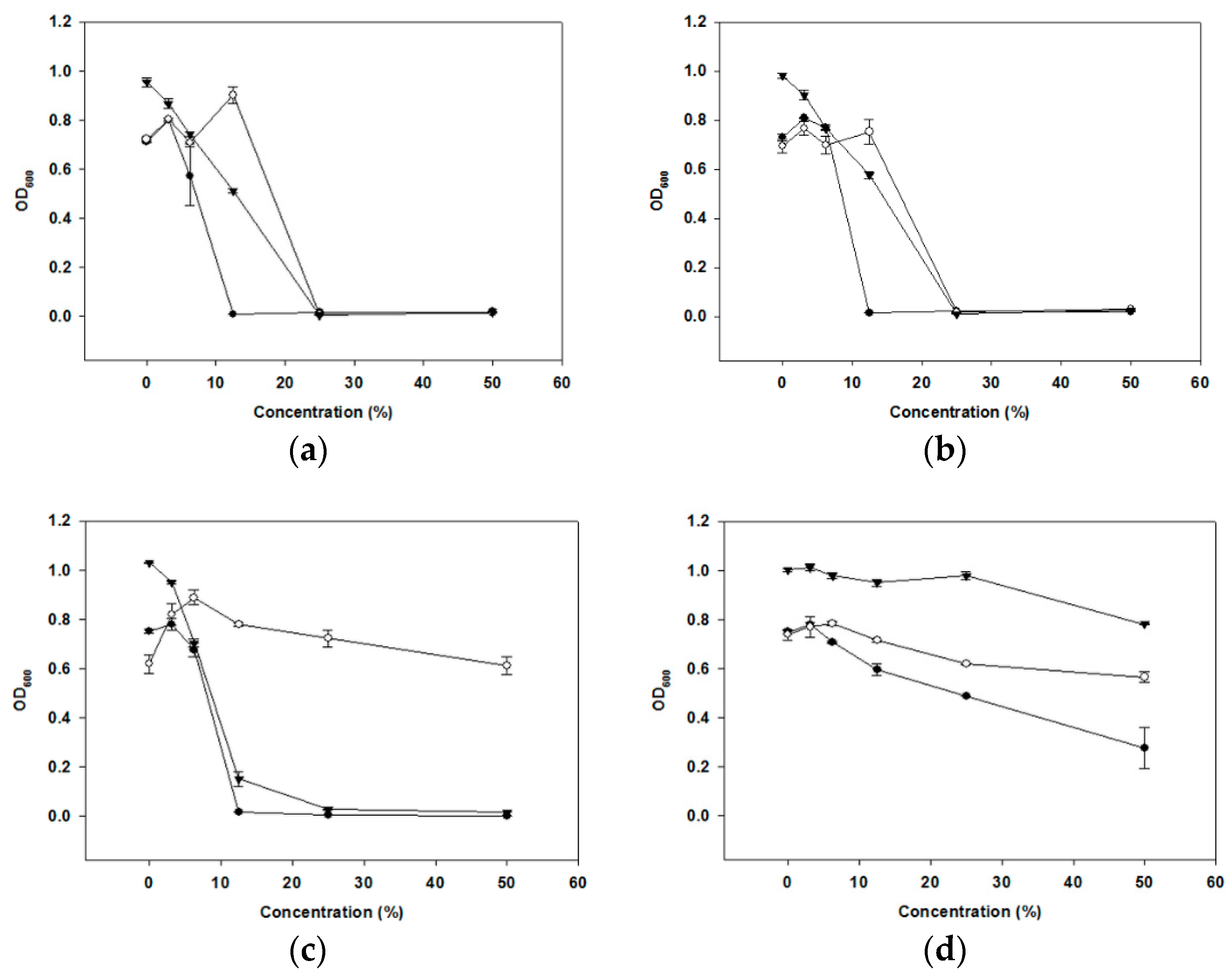
2.2. Characterization of Antibacterial Substances
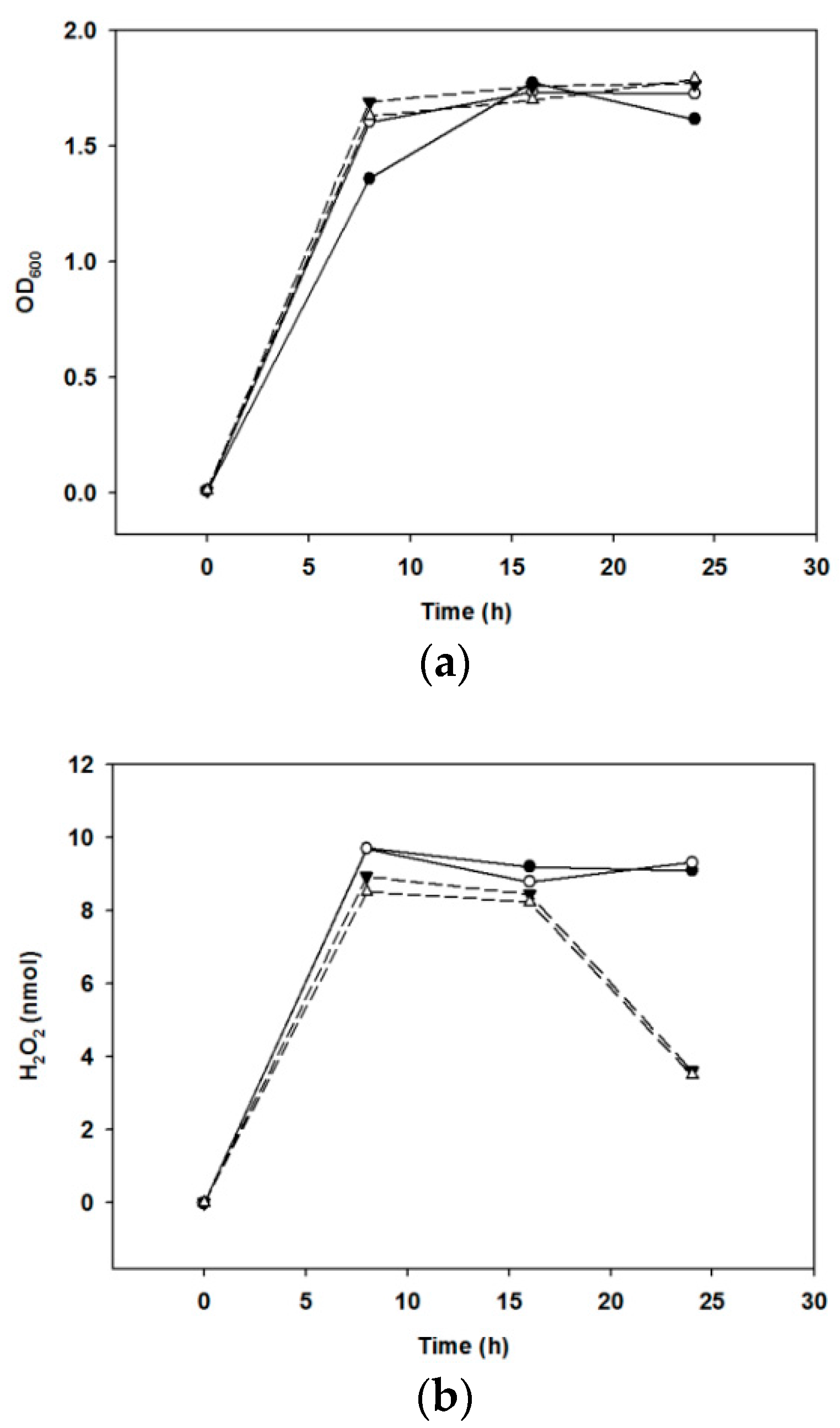
2.3. Kinetics of H2O2 Production
2.4. Determination of Organic and Fatty Acid Production
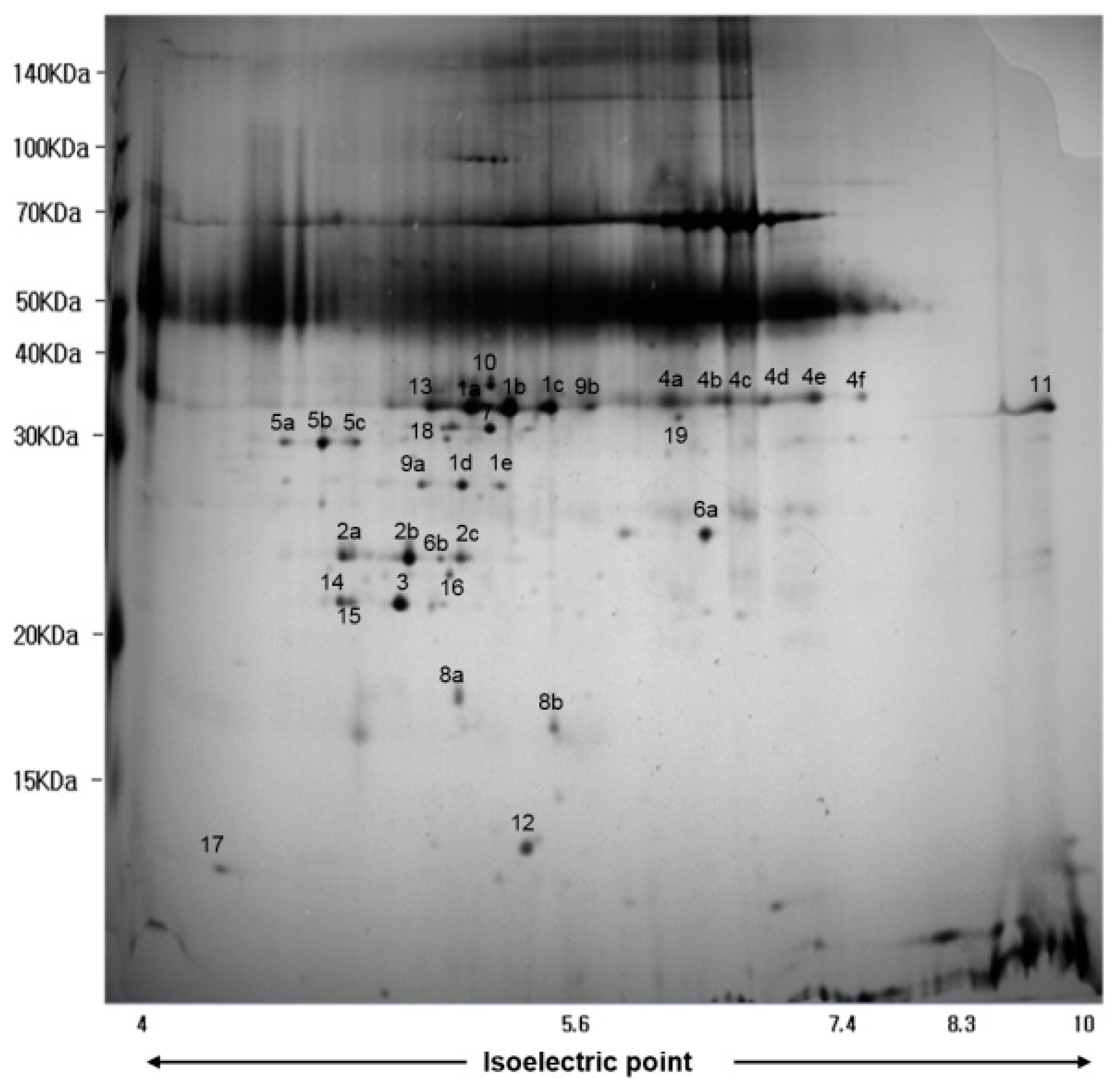
2.5. Secretome Analysis
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Growth Conditions
4.2. Growth Inhibition of Periodontopathogenic Bacteria
4.3. SEM Analysis
4.4. H2O2 Determination
4.5. Analysis of Organic Acids by HPLC
4.6. Analysis of Fatty Acids by GC
4.7. Analysis of Secretory Proteins by MALDI-TOF MS
Author Contributions
Funding
Conflicts of Interest
References
- Samot, J.; Badet, C. Antibacterial activity of probiotic candidates for oral health. Anaerobe 2013, 19, 34–38. [Google Scholar] [CrossRef] [PubMed]
- Terai, T.; Okumura, T.; Imai, S.; Nakao, M.; Yamaji, K.; Ito, M.; Nagata, T.; Kaneko, K.; Miyazaki, K.; Okada, A.; et al. Screening of probiotic candidates in human oral bacteria for the prevention of dental disease. PLoS ONE 2015, 8, e0128657. [Google Scholar] [CrossRef] [PubMed]
- López-López, A.; Camelo-Castillo, A.; Ferrer, M.D.; Simon-Soro, Á.; Mira, A. Health-associated niche inhabitants as oral probiotics: The Case of Streptococcus dentisani. Front. Microbiol. 2017, 10, 379. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.J.; Kang, M.S.; Yi, S.H.; Hong, J.Y.; Hong, S.P. Comparative study on the characteristics of Weissella cibaria CMU and probiotic strains for oral care. Molecules 2016, 20, 1752. [Google Scholar] [CrossRef] [PubMed]
- Flichy-Fernández, A.J.; Ata-Ali, J.; Alegre-Domingo, T.; Candel-Martí, E.; Ata-Ali, F.; Palacio, J.R.; Peñarrocha-Diago, M. The effect of orally administered probiotic Lactobacillus reuteri-containing tablets in peri-implant mucositis: A double-blind randomized controlled trial. J. Period. Res. 2015, 50, 775–785. [Google Scholar] [CrossRef] [PubMed]
- Nishihara, T.; Suzuki, N.; Yoneda, M.; Hirofuji, T. Effects of Lactobacillus salivarius-containing tablets on caries risk factors: A randomized open-label clinical trial. BMC Oral Health 2014, 14, 110. [Google Scholar] [CrossRef] [PubMed]
- Masdea, L.; Kulik, E.M.; Hauser-Gerspach, I.; Ramseier, A.M.; Filippi, A.; Waltimo, T. Antimicrobial activity of Streptococcus salivarius K12 on bacteria involved in oral malodour. Arch. Oral Biol. 2012, 57, 1041–1047. [Google Scholar] [CrossRef] [PubMed]
- Maekawa, T.; Hajishengallis, G. Topical treatment with probiotic Lactobacillus brevis CD2 inhibits experimental periodontal inflammation and bone loss. J. Period. Res. 2014, 49, 785–791. [Google Scholar] [CrossRef] [PubMed]
- Food and Agriculture Organization of the United Nations (FAO); World Health Organization (WHO). Health and Nutritional Properties of Probiotics in Food including Powder Milk with Live Lactic Acid Bacteria; Food and Agriculture Organization of the United Nations (FAO): Rome, Italy; World Health Organization (WHO): Geneva, Switzerland, 2001. [Google Scholar]
- Kanmani, P.; Satish Kumar, R.; Yuvaraj, N.; Paari, K.A.; Pattukumar, V.; Arul, V. Probiotics and its functionally valuable products-a review. Crit. Rev. Food Sci. Nutr. 2013, 53, 641–658. [Google Scholar] [CrossRef] [PubMed]
- Popova, M.; Molimard, P.; Courau, S.; Crociani, J.; Dufour, C.; Le Vacon, F.; Carton, T. Beneficial effects of probiotics in upper respiratory tract infections and their mechanical actions to antagonize pathogens. J. Appl. Microbiol. 2012, 113, 1305–1318. [Google Scholar] [CrossRef] [PubMed]
- Heller, F.; Duchmann, R. Intestinal flora and mucosal immune responses. Int. J. Med. Microbiol. 2003, 293, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.S.; Yeu, J.E.; Oh, J.S.; Shin, B.A.; Kim, J.H. Complete Genome Sequences of Weissella cibaria Strains CMU, CMS1, CMS2, and CMS3 isolated from infant saliva in South Korea. Genome Announc. 2017, 5, e01103–e01117. [Google Scholar] [CrossRef] [PubMed]
- Kang, B.K.; Cho, M.S.; Park, D.S. Red pepper powder is a crucial factor that influences the ontogeny of Weissella cibaria during kimchi fermentation. Sci. Rep. 2016, 6, 28232. [Google Scholar] [CrossRef] [PubMed]
- Sengun, I.Y.; Nielsen, D.S.; Karapinar, M.; Jakobsen, M. Identification of lactic acid bacteria isolated from Tarhana, a traditional Turkish fermented food. Int. J. Food Microbiol. 2009, 135, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Bounaix, M.S.; Robert, H.; Gabriel, V.; Morel, S.; Remaud-Siméon, M.; Gabriel, B.; Fontagné-Faucher, C. Characterization of dextran-producing Weissella strains isolated from sourdoughs and evidence of constitutive dextransucrase expression. FEMS Microbiol. Lett. 2010, 311, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.S.; Kim, B.G.; Chung, J.; Lee, H.C.; Oh, J.S. Inhibitory effect of Weissella cibaria isolates on the production of volatile sulphur compounds. J. Clin. Periodontol. 2006, 33, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.S.; Chung, J.; Kim, S.M.; Yang, K.H.; Oh, J.S. Effect of Weissella cibaria isolates on the formation of Streptococcus mutans biofilm. Caries Res. 2006, 40, 418–425. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.S.; Na, H.S.; Oh, J.S. Coaggregation ability of Weissella cibaria isolates with Fusobacterium nucleatum and their adhesiveness to epithelial cells. FEMS Microbiol. Lett. 2005, 253, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.S.; Lim, H.S.; Kim, S.M.; Lee, H.C.; Oh, J.S. Effect of Weissella cibaria on Fusobacterium nucleatum-induced Interleukin-6 and Interleukin-8 Production in KB Cells. J. Bacteriol. Virol. 2011, 41, 9–18. [Google Scholar] [CrossRef]
- Tingirikari, J.M.; Kothari, D.; Shukla, R.; Goyal, A. Structural and biocompatibility properties of dextran from Weissella cibaria JAG8 as food additive. Int. J. Food Sci. Nutr. 2014, 65, 686–691. [Google Scholar] [CrossRef] [PubMed]
- Björkroth, K.J.; Schillinger, U.; Geisen, R.; Weiss, N.; Hoste, B.; Holzapfel, W.H.; Korkeala, H.J.; Vandamme, P. Taxonomic study of Weissella confusa and description of Weissella cibaria sp. nov., detected in food and clinical samples. Int. J. Syst. Evol. Microbiol. 2002, 52, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Kreth, J. The Role of hydrogen peroxide in environmental adaptation of oral microbial communities. Oxid. Med. Cell Longev. 2012, 2012, 717843. [Google Scholar] [CrossRef] [PubMed]
- Thomas, E.L. Myeloperoxidase-hydrogen peroxide-chloride antimicrobial system: Effect of exogenous amines on antibacterial action against Escherichia coli. Infect. Immun. 1979, 25, 110–116. [Google Scholar] [PubMed]
- Hung, J.; Cooper, D.; Turner, M.S.; Walsh, T.; Giffard, P.M. Cystine uptake prevents production of hydrogen peroxide by Lactobacillus fermentum BR11. FEMS Microbiol. Lett. 2003, 227, 93–99. [Google Scholar] [CrossRef]
- Konings, W.N.; Poolman, B.; Driessen, A.J. Bioenergetics and solute transport in lactococci. CRC Crit. Rev. Microbiol. 1989, 16, 419–476. [Google Scholar] [CrossRef] [PubMed]
- Speert, D.P.; Wannamaker, L.W.; Gray, E.D.; Clawson, C.C. Bactericidal effect of oleic acid on group A streptococci: Mechanism of action. Infect. Immun. 1979, 26, 1202–1210. [Google Scholar] [PubMed]
- Wong, C.B.; Khoo, B.Y.; Sasidharan, S.; Piyawattanametha, W.; Kim, S.H.; Khemthongcharoen, N.; Ang, M.Y.; Chuah, L.O.; Liong, M.T. Inhibition of Staphylococcus aureus by crude and fractionated extract from lactic acid bacteria. Benef. Microbes 2015, 6, 129–139. [Google Scholar] [CrossRef] [PubMed]
- Nuñez de Kairuz, M.S.; Olazabal, M.E.; Oliver, G.; Pesce de Ruiz Holgado, A.A.; Massa, E.; Farías, R.N. Fatty acid dependent hydrogen peroxide production in Lactobacillus. Biochem. Biophys. Res. Commun. 1988, 152, 113–121. [Google Scholar] [CrossRef]
- Corcoran, B.M.; Stanton, C.; Fitzgerald, G.F.; Ross, R.P. Growth of probiotic lactobacilli in the presence of oleic acid enhances subsequent survival in gastric juice. Microbiology 2007, 153, 291–299. [Google Scholar] [CrossRef] [PubMed]
- Tristan, C.; Shahani, N.; Sedlak, T.W.; Sawa, A. The diverse functions of GAPDH: Views from different subcellular compartments. Cell. Signal. 2011, 23, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Hurmalainen, V.; Edelman, S.; Antikainen, J.; Baumann, M.; Lähteenmäki, K.; Korhonen, T.K. Extracellular proteins of Lactobacillus crispatus enhance activation of human plasminogen. Microbiology 2007, 153, 1112–1122. [Google Scholar] [CrossRef] [PubMed]
- Striker, R.; Kline, M.E.; Haak, R.A.; Rest, R.F.; Rosenthal, R.S. Degradation of gonococcal peptidoglycan by granule extract from human neutrophils: Demonstration of N-acetylglucosaminidase activity that utilizes peptidoglycan substrates. Infect. Immun. 1987, 55, 2579–2584. [Google Scholar] [PubMed]
- Kretschmar, S.; Yin, L.; Roberts, F.; London, R.; Flemmig, T.T.; Arushanov, D.; Kaiyala, K.; Chung, W.O. Protease inhibitor levels in periodontal health and disease. J. Period. Res. 2012, 47, 228–235. [Google Scholar] [CrossRef] [PubMed]
- Laugisch, O.; Schacht, M.; Guentsch, A.; Kantyka, T.; Sroka, A.; Stennicke, H.R.; Pfister, W.; Sculean, A.; Potempa, J.; Eick, S. Periodontal pathogens affect the level of protease inhibitors in gingival crevicular fluid. Mol. Oral Microbiol. 2012, 27, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Remold-O’Donnell, E. The ovalbumin family of serpin proteins. FEBS Lett. 1993, 315, 105–108. [Google Scholar] [CrossRef]
- Remold-O’Donnell, E.; Chin, J.; Alberts, M. Sequence and molecular characterization of human monocyte/neutrophil elastase inhibitor. Proc. Natl. Acad. Sci. USA 1992, 89, 5635–5639. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, B.; Schmitter, J.M.; Urdaci, M.C. Identification of novel proteins secreted by Lactobacillus rhamnosus GG grown in de Mann-Rogosa-Sharpe broth. Lett. Appl. Microbiol. 2009, 48, 618–622. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.S.; Oh, J.S.; Lee, H.C.; Lim, H.S.; Lee, S.W.; Yang, K.H.; Choi, N.K.; Kim, S.M. Inhibitory effect of Lactobacillus reuteri on periodontopathic and cariogenic bacteria. J. Microbiol. 2011, 49, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Sorrentino, E.; Reale, A.; Tremonte, P.; Maiuro, L.; Succi, M.; Tipaldi, L.; Di Renzo, T.; Pannella, G.; Coppola, R. Lactobacillus plantarum 29 inhibits Penicillium spp. involved in the spoilage of black truffles (Tuber aestivum). J. Food Sci. 2013, 78, M1188–M1194. [Google Scholar] [PubMed]
- Yeo, S.K.; Liong, M.T. Effect of prebiotics on viability and growth characteristics of probiotics in soymilk. J. Sci. Food Agric. 2010, 90, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Oakley, B.R.; Kirsch, D.R.; Morris, N.R. A simplified ultrasensitive silver stain for detecting proteins in polyacrylamide gels. Anal. Biochem. 1980, 105, 361–363. [Google Scholar] [CrossRef]
- Fernandez, J.; Gharahdaghi, F.; Mische, S.M. Routine identification of proteins from sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) gels or polyvinyl difluoride membranes using matrix assisted laser desorption/ionization-time of flight-mass spectrometry (MALDI-TOF-MS). Electrophoresis 1998, 19, 1036–1045. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Not available. |

| Acids | mg/mL | Acids | mg/mL | Acids | mg/mL |
|---|---|---|---|---|---|
| Oxalic acid | ND | Pentadecanoic acid (C15:0) | ND | cis-11-Eicosenoic acid (C20:1) | ND |
| Citric acid | 0.88 a | cis-10-Peptadecenoic acid (C15:1) | ND | cis-11,14-Eicosadienoic acid (C20:2) | ND |
| Succinic acid | ND b | Palmitic acid (C16:0) | 0.00 | cis-11,14,17-Eicosatrienoic acid (C20:3n-3) | ND |
| Lactic acid | 6.42 | Palmitoleic acid (C16:1) | ND | cis-8,11,14-Eicosatrienoic acid (C20:3n-6) | ND |
| Acetic acid | 2.78 | Heptadecanoic acid (C17:0) | ND | Arachidonic acid (C20:4n-6) | ND |
| Butyric acid (C4:0) | ND | cis-10-Heptadecenoic acid (C17:1) | ND | cis-5,8,11,14,17-Eicosapentaenoic acid (C20:5n-3) | ND |
| Caproic acid (C6:0) | ND | Stearic acid (C18:0) | ND | Heneicosanoic acid (C21:0) | ND |
| Caprylic acid (C8:0) | ND | Elaidic acid (C18:1n-9, trans) | ND | Behenic acid (C22:0) | ND |
| Capric acid (C10:0) | ND | Oleic acid (C18:1n-9, cis) | 0.2 | Erucic acid (C22:1n-9) | ND |
| Undecanoic acid (C11:0) | ND | Linoleiaidic acid (C18:2n-6, trans) | ND | cis-13,16-Docosadienoic acid (C22:2) | ND |
| Lauric acid (C12:0) | ND | Linoleic acid (C18:2n-6, cis) | ND | cis-4,7,10,13,16,19-Docosahexaenoic acid (C22:6n-3) | ND |
| Tridecanoic acid (C13:0) | ND | α-Linolenic acid (C18:3n-3) | ND | Tricosanoic acid (C23:0) | ND |
| Myristic acid (C14:0) | ND | γ-Linolenic acid (C18:3n-6) | ND | Lignoceric acid (C24:0) | ND |
| Myristoleic acid (C14:1) | ND | Arachidic acid (C20:0) | ND | Nervonic acid (C24:1) | ND |
| Spot No. a | Protein Description b | Accession Number c | MW (kDa) | PI | Protein Score CI (%) d |
|---|---|---|---|---|---|
| 1a–e | Type 1 glyceraldehyde-3-phosphate dehydrogenase | WP_010369259 | 35.756 | 5.03 | 100.00 |
| 2a–c | N-acetylmuramidase | WP_010373609 | 29.343 | 5.40 | 100.00 |
| 3 | TIGR00266 family protein | WP_043941068 | 24.976 | 4.91 | 99.19 |
| 4a–f | Leukocyte elastase inhibitor | XP_003482167 | 42.658 | 6.13 | 100.00 |
| 5a–c | l-lactate dehydrogenase | WP_010372268 | 33.942 | 4.75 | 100.00 |
| 6a,b | Hypothetical protein | WP_043711225 | 23.526 | 5.60 | 99.99 |
| WP_063083480 | 23.318 | 5.04 | 100.00 | ||
| 7 | Aldo/keto reductase | WP_063083131 | 36.627 | 5.08 | 100.00 |
| 8a,b | Single-stranded DNA-binding protein | WP_056973096 | 17.991 | 4.99 | 100.00 |
| WP_043709699 | 15.326 | 5.17 | 99.99 | ||
| 9a,b | NAD-dependent glyceraldehyde-3-phosphate dehydrogenase | KKU03010 | 41.843 | 5.54 | 100.00 |
| 10 | N-acetylglucosamin-6-phosphate deacetylase | WP_0432710743 | 42.715 | 5.17 | 100.00 |
| 11 | BMP family ABC transporter substrate-binding protein | WP_060655478 | 39.514 | 9.18 | 100.00 |
| 12 | Nucleoside-diphosphate kinase | WP_043710743 | 42.715 | 5.17 | 100.00 |
| 13 | Glyceraldehyde 3-phosphate reductase | WP_010370341 | 36.629 | 5.00 | 100.00 |
| 14 | 16S rRNA (cytosine(1402)-N(4))-methyltransferase RsmH | WP_004044730 | 35.71 | 8.73 | 99.64 |
| 15 | HTH-type transcriptional activator RhaR | WP_008787014 | 33.031 | 6.59 | 99.94 |
| 16 | 2,3-diphosphoglycerate-dependent phosphoglycerate mutase | WP_010371458 | 26.717 | 4.94 | 100.00 |
| 17 | Phosphocarrier protein HPr | WP_010370017 | 9.191 | 4.57 | 99.19 |
| 18 | PTS mannose transporter subunit EIIAB | WP_043709678 | 35.417 | 4.97 | 100.00 |
| 19 | Zinc-dependent alcohol dehydrogenase | XP_003482167 | 42.658 | 6.13 | 99.99 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lim, H.-S.; Yeu, J.-E.; Hong, S.-P.; Kang, M.-S. Characterization of Antibacterial Cell-Free Supernatant from Oral Care Probiotic Weissella cibaria, CMU. Molecules 2018, 23, 1984. https://doi.org/10.3390/molecules23081984
Lim H-S, Yeu J-E, Hong S-P, Kang M-S. Characterization of Antibacterial Cell-Free Supernatant from Oral Care Probiotic Weissella cibaria, CMU. Molecules. 2018; 23(8):1984. https://doi.org/10.3390/molecules23081984
Chicago/Turabian StyleLim, Hae-Soon, Ji-Eun Yeu, Sang-Phil Hong, and Mi-Sun Kang. 2018. "Characterization of Antibacterial Cell-Free Supernatant from Oral Care Probiotic Weissella cibaria, CMU" Molecules 23, no. 8: 1984. https://doi.org/10.3390/molecules23081984
APA StyleLim, H.-S., Yeu, J.-E., Hong, S.-P., & Kang, M.-S. (2018). Characterization of Antibacterial Cell-Free Supernatant from Oral Care Probiotic Weissella cibaria, CMU. Molecules, 23(8), 1984. https://doi.org/10.3390/molecules23081984