Do Entropic Approaches Improve Understanding of Biology?
A topical collection in Entropy (ISSN 1099-4300). This collection belongs to the section "Entropy and Biology".
Submission Status:
Closed (29 January 2026)
|
Viewed by 29676
Share This Topical Collection
Editors
 Prof. Dr. William B. Sherwin
Prof. Dr. William B. Sherwin
 Prof. Dr. William B. Sherwin
Prof. Dr. William B. Sherwin
E-Mail
Website
Collection Editor
Evolution & Ecology Research Centre, School of Biological Earth and Environmental Science, UNSW Sydney, Sydney NSW 2052, Australia
Interests: molecular ecology; evolutionary genetics; biodiversity; genomics, transciptomics; mathematics of forecasting and measuring biodiversity; conservation genetics and demography; endangered harvested and invasive species; evolutionary ecology of parentage, relatedness and group formation, especially in dolphins
Special Issues, Collections and Topics in MDPI journals
 Dr. Robert Niven
Dr. Robert Niven
 Dr. Robert Niven
Dr. Robert Niven
E-Mail
Website
Collection Editor
School of Engineering and Information Technology, The University of New South Wales, Canberra, ACT 2600, Australia
Interests: Bayesian and maximum entropy methods for the analysis of engineering and scientific systems; theoretical foundations of Bayesian inference; Bayesian estimation and plausible reasoning; entropy-based inference and extremum methods; Bayesian risk assessment; heuristics and methods for the selection of prior probabilities; probabilistic transport and evolution equations and operators
Special Issues, Collections and Topics in MDPI journals
Topical Collection Information
Dear Colleagues,
Biology and medicine, from molecules to landscapes, are ideally suited to entropy or information approaches, because biological systems are highly variable, with stochastic processes of Innovation, Transmission, Adaptation and Movement. We are now seeing many papers being published for all biological levels from biomolecules to landscapes. However, these papers often do not assess whether the entropic approach actually gives a better performance than other forecasts or measures. Therefore this Topical Issue of ‘Entropy’ encourages the submission of manuscripts that do such comparisons, using either or both simulations and biological data.
We encourage authors to make their contribution accessible to a wide range of science graduates, without compromising scientific content or flow, for example via a table of symbols and jargon, containing definitions understandable to most science graduates. We also encourage the addition of a supplementary short (e.g., three-minute) video, which explains in plain language the general significance of the major finding(s).
Prof. William B. Sherwin
Prof. Dr. Robert Niven
Guest Editors
Manuscript Submission Information
Manuscripts should be submitted online at www.mdpi.com by registering and logging in to this website. Once you are registered, click here to go to the submission form. All submissions that pass pre-check are peer-reviewed. Accepted papers will be published continuously in the journal (as soon as accepted) and will be listed together on the collection website. Research articles, review articles as well as short communications are invited. For planned papers, a title and short abstract (about 250 words) can be sent to the Editorial Office for assessment.
Submitted manuscripts should not have been published previously, nor be under consideration for publication elsewhere (except conference proceedings papers). All manuscripts are thoroughly refereed through a single-blind peer-review process. A guide for authors and other relevant information for submission of manuscripts is available on the Instructions for Authors page. Entropy is an international peer-reviewed open access monthly journal published by MDPI.
Please visit the Instructions for Authors page before submitting a manuscript.
The Article Processing Charge (APC) for publication in this open access journal is 2600 CHF (Swiss Francs).
Submitted papers should be well formatted and use good English. Authors may use MDPI's
English editing service prior to publication or during author revisions.
Published Papers (5 papers)
Open AccessArticle
Potential Benefits and Challenges of Quantifying Pseudoreplication in Genomic Data with Entropy Statistics
by
Eric J. Ward and Robin S. Waples
Viewed by 1544
Abstract
Generating vast arrays of genetic markers for evolutionary ecology studies has become routine and cost-effective. However, analyzing data from large numbers of loci associated with a small number of finite chromosomes introduces a challenge: loci on the same chromosome do not assort independently,
[...] Read more.
Generating vast arrays of genetic markers for evolutionary ecology studies has become routine and cost-effective. However, analyzing data from large numbers of loci associated with a small number of finite chromosomes introduces a challenge: loci on the same chromosome do not assort independently, leading to pseudoreplication. Previous studies have demonstrated that pseudoreplication can substantially reduce precision of genetic analyses (and make confidence intervals wider), such as
FST and linkage disequilibrium (LD) measures between pairs of loci. In LD analyses, another type of dependency (overlapping pairs of the same loci) also creates pseudoreplication. Building on previous work, we explore the potential of entropy metrics to improve the status quo, particularly total correlation (TC), to assess pseudoreplication in LD studies. Our simulations, performed on a monoecious population with a range of effective population sizes (
Ne) and numbers of loci, attempted to isolate the overlapping-pairs-of-loci effect by considering unlinked loci and using entropy to quantify inter-locus relationships. We hypothesized a positive correlation between TC and the number of loci (L), and a negative correlation between TC and
Ne. Results from our statistical models predicting TC demonstrate a strong effect of the number of loci, and muted effects of
Ne and other predictors, adding support to the use of entropy-based metrics as a tool for estimating the statistical information of complex genetic datasets. Our results also highlight a challenge regarding scalability; computational limitations arise as the number of loci grows, making our current approach limited to smaller datasets. Despite these challenges, this work further refines our understanding of entropy measures, and offers insights into the complex dynamics of genetic information in evolutionary ecology research.
Full article
►▼
Show Figures
Open AccessArticle
A Comparison of Entropic Diversity and Variance in the Study of Population Structure
by
Eric F. Karlin
Cited by 4 | Viewed by 2084
Abstract
AMOVA is a widely used approach that focuses on variance within and among strata to study the hierarchical genetic structure of populations. The recently developed Shannon Informational Diversity Translation Analysis (SIDTA) instead tackles exploration of hierarchical genetic structure using entropic allelic diversity. A
[...] Read more.
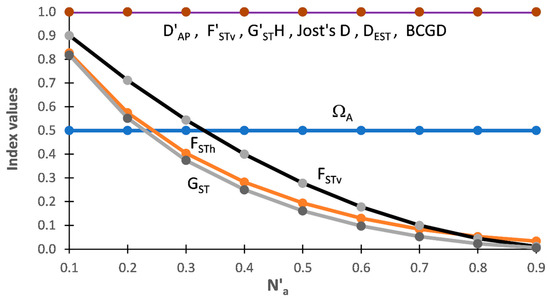
AMOVA is a widely used approach that focuses on variance within and among strata to study the hierarchical genetic structure of populations. The recently developed Shannon Informational Diversity Translation Analysis (SIDTA) instead tackles exploration of hierarchical genetic structure using entropic allelic diversity. A mix of artificial and natural population data sets (including allopolyploids) is used to compare the performance of SIDTA (a ‘q = 1’ diversity measure) vs. AMOVA (a ‘q = 2’ measure) under different conditions. An additive allelic differentiation index based on entropic allelic diversity measuring the mean difference among populations (Ω
AP) was developed to facilitate the comparison of SIDTA with AMOVA. These analyses show that the genetic population structure seen by AMOVA is notably different in many ways from that provided by SIDTA, and the extent of this difference is greatly affected by the stability of the markers employed. Negative among group values are lacking with SIDTA but occur with AMOVA, especially with allopolyploids. To provide more focus on measuring allelic differentiation among populations, additional measures were also tested including Bray–Curtis Genetic Differentiation (BCGD) and several expected heterozygosity-based indices (e.g., G
ST, G″
ST, Jost’s D, and D
EST). Corrections, such as almost unbiased estimators, that were designed to work with heterozygosity-based fixation indices (e.g., F
ST, G
ST) are problematic when applied to differentiation indices (eg., D
EST, G″
ST, G′
STH).
Full article
►▼
Show Figures
Open AccessFeature PaperArticle
The Eco-Evo Mandala: Simplifying Bacterioplankton Complexity into Ecohealth Signatures
by
Elroy Galbraith and Matteo Convertino
Cited by 8 | Viewed by 3789
Abstract
The microbiome emits informative signals of biological organization and environmental pressure that aid ecosystem monitoring and prediction. Are the many signals reducible to a habitat-specific portfolio that characterizes ecosystem health? Does an optimally structured microbiome imply a resilient microbiome? To answer these questions,
[...] Read more.
The microbiome emits informative signals of biological organization and environmental pressure that aid ecosystem monitoring and prediction. Are the many signals reducible to a habitat-specific portfolio that characterizes ecosystem health? Does an optimally structured microbiome imply a resilient microbiome? To answer these questions, we applied our novel Eco-Evo Mandala to bacterioplankton data from four habitats within the Great Barrier Reef, to explore how patterns in community structure, function and genetics signal habitat-specific organization and departures from theoretical optimality. The Mandala revealed communities departing from optimality in habitat-specific ways, mostly along structural and functional traits related to bacterioplankton abundance and interaction distributions (reflected by
and
as power law and exponential distribution parameters), which are not linearly associated with each other. River and reef communities were similar in their relatively low abundance and interaction disorganization (low
and
) due to their protective structured habitats. On the contrary, lagoon and estuarine inshore reefs appeared the most disorganized due to the ocean temperature and biogeochemical stress. Phylogenetic distances (
D) were minimally informative in characterizing bacterioplankton organization. However, dominant populations, such as Proteobacteria, Bacteroidetes, and Cyanobacteria, were largely responsible for community patterns, being generalists with a large functional gene repertoire (high
D) that increases resilience. The relative balance of these populations was found to be habitat-specific and likely related to systemic environmental stress. The position on the Mandala along the three fundamental traits, as well as fluctuations in this ecological state, conveys information about the microbiome’s health (and likely ecosystem health considering bacteria-based multitrophic dependencies) as divergence from the expected relative optimality. The Eco-Evo Mandala emphasizes how habitat and the microbiome’s interaction network topology are first- and second-order factors for ecosystem health evaluation over taxonomic species richness. Unhealthy microbiome communities and unbalanced microbes are identified not by macroecological indicators but by mapping their impact on the collective proportion and distribution of interactions, which regulates the microbiome’s ecosystem function.
Full article
►▼
Show Figures
Open AccessArticle
Bridging Offline Functional Model Carrying Aging-Specific Growth Rate Information and Recombinant Protein Expression: Entropic Extension of Akaike Information Criterion
by
Renaldas Urniezius, Benas Kemesis and Rimvydas Simutis
Cited by 8 | Viewed by 3280
Abstract
This study presents a mathematical model of recombinant protein expression, including its development, selection, and fitting results based on seventy fed-batch cultivation experiments from two independent biopharmaceutical sites. To resolve the overfitting feature of the Akaike information criterion, we proposed an entropic extension,
[...] Read more.
This study presents a mathematical model of recombinant protein expression, including its development, selection, and fitting results based on seventy fed-batch cultivation experiments from two independent biopharmaceutical sites. To resolve the overfitting feature of the Akaike information criterion, we proposed an entropic extension, which behaves asymptotically like the classical criteria. Estimation of recombinant protein concentration was performed with pseudo-global optimization processes while processing offline recombinant protein concentration samples. We show that functional models including the average age of the cells and the specific growth at induction or the start of product biosynthesis are the best descriptors for datasets. We also proposed introducing a tuning coefficient that would force the modified Akaike information criterion to avoid overfitting when the designer requires fewer model parameters. We expect that a lower number of coefficients would allow the efficient maximization of target microbial products in the upstream section of contract development and manufacturing organization services in the future. Experimental model fitting was accomplished simultaneously for 46 experiments at the first site and 24 fed-batch experiments at the second site. Both locations contained 196 and 131 protein samples, thus giving a total of 327 target product concentration samples derived from the bioreactor medium.
Full article
Open AccessReview
Entropy and the Brain: An Overview
by
Soheil Keshmiri
Cited by 136 | Viewed by 16928
Abstract
Entropy is a powerful tool for quantification of the brain function and its information processing capacity. This is evident in its broad domain of applications that range from functional interactivity between the brain regions to quantification of the state of consciousness. A number
[...] Read more.
Entropy is a powerful tool for quantification of the brain function and its information processing capacity. This is evident in its broad domain of applications that range from functional interactivity between the brain regions to quantification of the state of consciousness. A number of previous reviews summarized the use of entropic measures in neuroscience. However, these studies either focused on the overall use of nonlinear analytical methodologies for quantification of the brain activity or their contents pertained to a particular area of neuroscientific research. The present study aims at complementing these previous reviews in two ways. First, by covering the literature that specifically makes use of entropy for studying the brain function. Second, by highlighting the three fields of research in which the use of entropy has yielded highly promising results: the (altered) state of consciousness, the ageing brain, and the quantification of the brain networks’ information processing. In so doing, the present overview identifies that the use of entropic measures for the study of consciousness and its (altered) states led the field to substantially advance the previous findings. Moreover, it realizes that the use of these measures for the study of the ageing brain resulted in significant insights on various ways that the process of ageing may affect the dynamics and information processing capacity of the brain. It further reveals that their utilization for analysis of the brain regional interactivity formed a bridge between the previous two research areas, thereby providing further evidence in support of their results. It concludes by highlighting some potential considerations that may help future research to refine the use of entropic measures for the study of brain complexity and its function. The present study helps realize that (despite their seemingly differing lines of inquiry) the study of consciousness, the ageing brain, and the brain networks’ information processing are highly interrelated. Specifically, it identifies that the complexity, as quantified by entropy, is a fundamental property of conscious experience, which also plays a vital role in the brain’s capacity for adaptation and therefore whose loss by ageing constitutes a basis for diseases and disorders. Interestingly, these two perspectives neatly come together through the association of entropy and the brain capacity for information processing.
Full article
►▼
Show Figures