Global Sensitivity Analysis and Uncertainty Quantification for Simulated Atrial Electrocardiograms
Abstract
1. Introduction
2. Materials and Methods
2.1. The Atrial Model
2.2. Global Sensitivity Analysis, Uncertainty Quantification and Sobol Indices
2.3. Polynomial Chaos Expansion
2.4. Data Description
3. Results
3.1. Convergence of the Surrogate Model
3.1.1. Error Estimation
3.1.2. Normalization of the Surrogate Error
3.1.3. Surrogate Error Convergence
3.1.4. Convergence with Polynomial Order
3.1.5. Convergence with Sample Size
3.2. Sensitivity Analysis
3.2.1. Sobol Indices along the Signal
3.2.2. Time-Integrated Sobol Indices and Dataset Interpretation
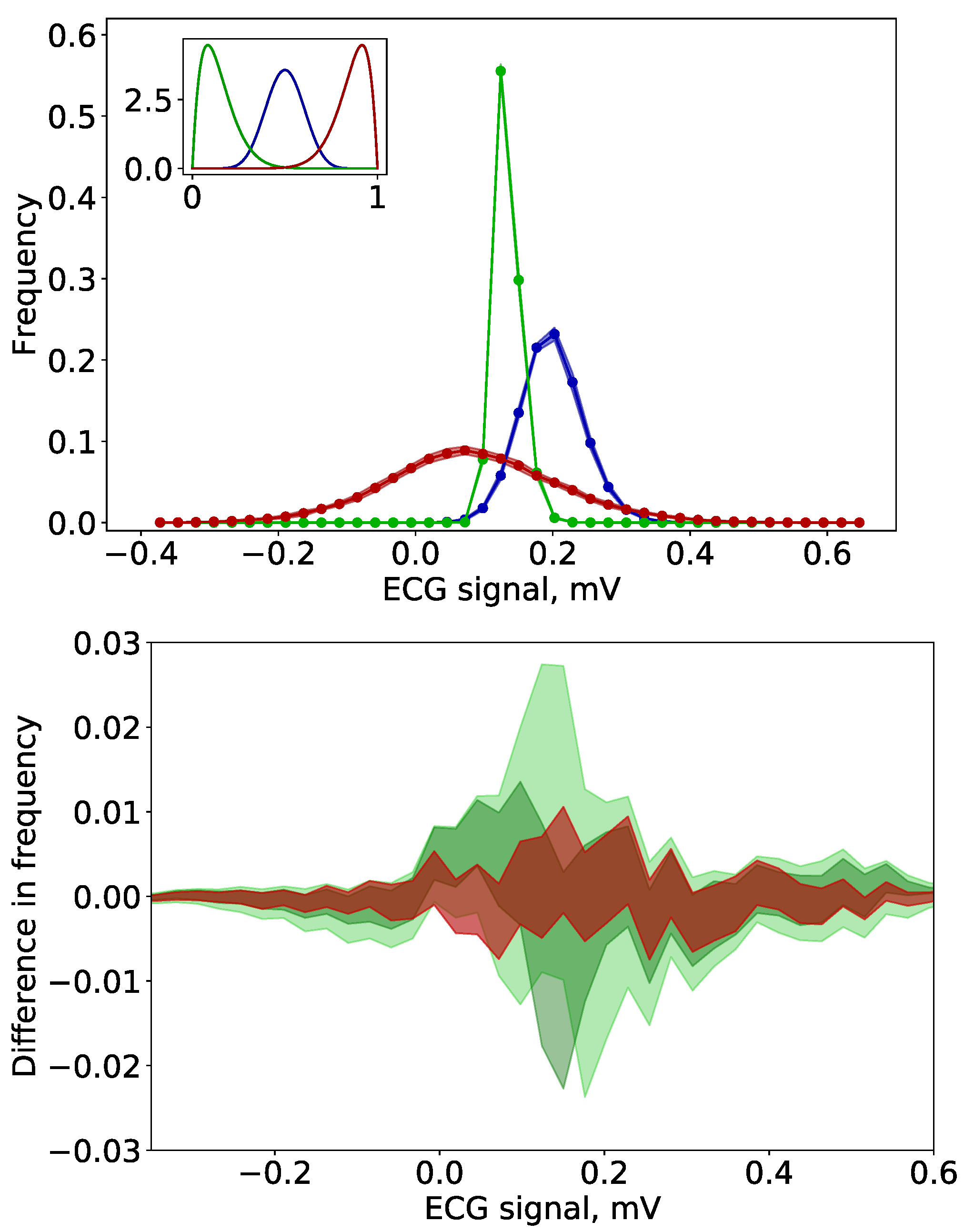
3.3. Uncertainty Quantification
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| PCE | Polynomial Chaos Expansion |
| SA | Sensitivity Analysis |
| UQ | Uncertainty Quantification |
| ECG | Electrocardiogram |
| SI | Sobol index |
| DS | dataset |
| GTD | ground truth data |
Appendix A. Mean Signals


Appendix B. Usage, Limitations and Further Directions
References
- Karma, A. Physics of cardiac arrhythmogenesis. Annu. Rev. Condens. Matter Phys. 2013, 4, 313–337. [Google Scholar] [CrossRef]
- Qu, Z.; Hu, G.; Garfinkel, A.; Weiss, J.N. Nonlinear and stochastic dynamics in the heart. Phys. Rep. 2014, 543, 61–162. [Google Scholar] [CrossRef]
- Alonso, S.; Bär, M.; Echebarria, B. Nonlinear physics of electrical wave propagation in the heart: A review. Rep. Prog. Phys. 2016, 79, 096601. [Google Scholar] [CrossRef] [PubMed]
- Rappel, W.J. The physics of heart rhythm disorders. Phys. Rep. 2022, 978, 1–45. [Google Scholar] [CrossRef]
- Loewe, A.; Wilhelms, M.; Fischer, F.; Scholz, E.P.; Dössel, O.; Seemann, G. Arrhythmic potency of human ether-a-go-go-related gene mutations L532P and N588K in a computational model of human atrial myocytes. Europace 2014, 16, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Clayton, R.; Bernus, O.; Cherry, E.; Dierckx, H.; Fenton, F.H.; Mirabella, L.; Panfilov, A.V.; Sachse, F.B.; Seemann, G.; Zhang, H. Models of cardiac tissue electrophysiology: Progress, challenges and open questions. Prog. Biophys. Mol. Biol. 2011, 104, 22–48. [Google Scholar] [CrossRef] [PubMed]
- Bragard, J.R.; Camara, O.; Echebarria, B.; Giorda, L.G.; Pueyo, E.; Saiz, J.; Sebastián, R.; Soudah, E.; Vázquez, M. Cardiac computational modelling. Rev. Esp. Cardiol. (Engl. Ed.) 2021, 74, 65–71. [Google Scholar] [CrossRef]
- Dössel, O.; Luongo, G.; Nagel, C.; Loewe, A. Computer modeling of the heart for ECG interpretation—A review. Hearts 2021, 2, 350–368. [Google Scholar] [CrossRef]
- Corral-Acero, J.; Margara, F.; Marciniak, M.; Rodero, C.; Loncaric, F.; Feng, Y.; Gilbert, A.; Fernandes, J.F.; Bukhari, H.A.; Wajdan, A.; et al. The ‘Digital Twin’ to enable the vision of precision cardiology. Eur. Heart J. 2020, 41, 4556–4564. [Google Scholar] [CrossRef]
- Gillette, K.; Gsell, M.A.; Prassl, A.J.; Karabelas, E.; Reiter, U.; Reiter, G.; Grandits, T.; Payer, C.; Štern, D.; Urschler, M.; et al. A framework for the generation of digital twins of cardiac electrophysiology from clinical 12-leads ECGs. Med. Image Anal. 2021, 71, 102080. [Google Scholar] [CrossRef]
- Azzolin, L.; Eichenlaub, M.; Nagel, C.; Nairn, D.; Sánchez, J.; Unger, L.; Dössel, O.; Jadidi, A.; Loewe, A. AugmentA: Patient-specific Augmented Atrial model Generation Tool. medRxiv 2022. [Google Scholar] [CrossRef]
- Azzolin, L.; Eichenlaub, M.; Nagel, C.; Nairn, D.; Sanchez, J.; Unger, L.; Dössel, O.; Jadidi, A.; Loewe, A. Personalized ablation vs. conventional ablation strategies to terminate atrial fibrillation and prevent recurrence. EP Eur. 2022, 1–12. [Google Scholar] [CrossRef]
- Niederer, S.; Aboelkassem, Y.; Cantwell, C.D.; Corrado, C.; Coveney, S.; Cherry, E.M.; Delhaas, T.; Fenton, F.H.; Panfilov, A.; Pathmanathan, P.; et al. Creation and application of virtual patient cohorts of heart models. Philos. Trans. R. Soc. A 2020, 378, 20190558. [Google Scholar] [CrossRef]
- Nagel, C.; Luongo, G.; Azzolin, L.; Schuler, S.; Dössel, O.; Loewe, A. Non-Invasive and Quantitative Estimation of Left Atrial Fibrosis Based on P Waves of the 12-Lead ECG-A Large-Scale Computational Study Covering Anatomical Variability. J. Clin. Med. 2021, 10, 1797. [Google Scholar] [CrossRef]
- Wagner, P.; Strodthoff, N.; Bousseljot, R.D.; Kreiseler, D.; Lunze, F.I.; Samek, W.; Schaeffter, T. PTB-XL, a large publicly available electrocardiography dataset. Sci. Data 2020, 7, 154. [Google Scholar] [CrossRef]
- Zheng, J.; Fu, G.; Anderson, K.; Chu, H.; Rakovski, C. A 12-Lead ECG database to identify origins of idiopathic ventricular arrhythmia containing 334 patients. Sci. Data 2020, 7, 98. [Google Scholar] [CrossRef]
- Strodthoff, N.; Wagner, P.; Schaeffter, T.; Samek, W. Deep learning for ECG analysis: Benchmarks and insights from PTB-XL. IEEE J. Biomed. Health Inform. 2020, 25, 1519–1528. [Google Scholar] [CrossRef]
- Venton, J.; Harris, P.M.; Sundar, A.; Smith, N.A.; Aston, P.J. Robustness of convolutional neural networks to physiological electrocardiogram noise. Philos. Trans. R. Soc. A 2021, 379, 20200262. [Google Scholar] [CrossRef]
- Mehari, T.; Strodthoff, N. Self-supervised representation learning from 12-lead ECG data. Comput. Biol. Med. 2022, 141, 105114. [Google Scholar] [CrossRef]
- Mirams, G.; Pathmanathan, P.; Gray, R.; Challenor, P.; Clayton, R. White Paper: Uncertainty and variability in computational and mathematical models of cardiac physiology. J. Physiol. 2016, 594, 6833–6847. [Google Scholar] [CrossRef]
- Pathmanathan, P.; Gray, R. Ensuring reliability of safety-critical clinical applications of computational cardiac models. Front. Physiol. 2013, 4, 358. [Google Scholar] [CrossRef] [PubMed]
- Eck, V.G.; Donders, W.P.; Sturdy, J.; Feinberg, J.; Delhaas, T.; Hellevik, L.R.; Huberts, W. A guide to uncertainty quantification and sensitivity analysis for cardiovascular applications. Int. J. Numer. Methods Biomed. Eng. 2016, 32, e02755. [Google Scholar] [CrossRef] [PubMed]
- Quicken, S.; Donders, W.P.; van Disseldorp, E.M.; Gashi, K.; Mees, B.M.; van de Vosse, F.N.; Lopata, R.G.; Delhaas, T.; Huberts, W. Application of an adaptive polynomial chaos expansion on computationally expensive three-dimensional cardiovascular models for uncertainty quantification and sensitivity analysis. J. Biomech. Eng. 2016, 138, 121010. [Google Scholar] [CrossRef] [PubMed]
- Pathmanathan, P.; Cordeiro, J.M.; Gray, R.A. Comprehensive Uncertainty Quantification and Sensitivity Analysis for Cardiac Action Potential Models. Front. Physiol. 2019, 10, 721. [Google Scholar] [CrossRef] [PubMed]
- Montes, G.; Oliveira, J.; Alvarez-Lacalle, E.; Alonso, S.; Martins, B.; Weber, R. Combining polynomial chaos expansions and genetic algorithm for the coupling of electrophysiological models. In Lecture Notes in Computer Science: Bioinformatics and Biomedical Engineering 11538; Springer: Cham, Switzerland, 2019; pp. 116–129. [Google Scholar] [CrossRef]
- Costabal, F.S.; Matsuno, K.; Yao, J.; Perdikaris, P.; Kuhl, E. Machine learning in drug development: Characterizing the effect of 30 drugs on the QT interval using Gaussian process regression, sensitivity analysis, and uncertainty quantification. Comput. Methods Appl. Mech. Eng. 2019, 348, 313–333. [Google Scholar] [CrossRef]
- Hoffman, M.J.; Cherry, E.M. Sensitivity of a data-assimilation system for reconstructing three-dimensional cardiac electrical dynamics. Philos. Trans. R. Soc. A 2020, 378, 20190388. [Google Scholar] [CrossRef]
- Rupp, L.C.; Liu, Z.; Bergquist, J.A.; Rampersad, S.; White, D.; Tate, J.D.; Brooks, D.H.; Narayan, A.; MacLeod, R.S. Using uncertainSCI to quantify uncertainty in cardiac simulations. In Proceedings of the 2020 Computing in Cardiology, Rimini, Italy, 13–16 September 2020; pp. 1–4. [Google Scholar]
- Campos, J.; Sundnes, J.; Dos Santos, R.; Rocha, B. Uncertainty quantification and sensitivity analysis of left ventricular function during the full cardiac cycle. Philos. Trans. R. Soc. A 2020, 378, 20190381. [Google Scholar] [CrossRef]
- Lei, C.L.; Ghosh, S.; Whittaker, D.G.; Aboelkassem, Y.; Beattie, K.A.; Cantwell, C.D.; Delhaas, T.; Houston, C.; Novaes, G.M.; Panfilov, A.V.; et al. Considering discrepancy when calibrating a mechanistic electrophysiology model. Philos. Trans. R. Soc. A 2020, 378, 20190349. [Google Scholar] [CrossRef]
- Clayton, R.H.; Aboelkassem, Y.; Cantwell, C.D.; Corrado, C.; Delhaas, T.; Huberts, W.; Lei, C.L.; Ni, H.; Panfilov, A.V.; Roney, C.; et al. An audit of uncertainty in multi-scale cardiac electrophysiology models. Philos. Trans. R. Soc. A 2020, 378, 20190335. [Google Scholar] [CrossRef]
- Tate, J.D.; Good, W.W.; Zemzemi, N.; Boonstra, M.; Dam, P.v.; Brooks, D.H.; Narayan, A.; MacLeod, R.S. Uncertainty quantification of the effects of segmentation variability in ecgi. In Proceedings of the International Conference on Functional Imaging and Modeling of the Heart, Stanford, CA, USA, 21–25 June 2021; pp. 515–522. [Google Scholar]
- Narayan, A.; Liu, Z.; Bergquist, J.; Charlebois, C.; Rampersad, S.; Rupp, L.; Brooks, D.; White, D.; Tate, J.; MacLeod, R.S. Uncertainsci: Uncertainty Quantification for Computational Models in Biomedicine and Bioengineering. Comput. Biol. Med. 2022, 152, 106407. [Google Scholar] [CrossRef]
- Steinman, D.A.; Migliavacca, F. Special issue on verification, validation, and uncertainty quantification of cardiovascular models: Towards effective vvuq for translating cardiovascular modelling to clinical utility. Cardiovasc. Eng. Technol. 2018, 9, 539–543. [Google Scholar] [CrossRef]
- Mirams, G.R.; Niederer, S.A.; Clayton, R.H. The fickle heart: Uncertainty quantification in cardiac and cardiovascular modelling and simulation. Philos. Trans. R. Soc. A 2020, 378, 20200119. [Google Scholar] [CrossRef]
- Nagel, C.; Espinosa, C.B.; Gillette, K.; Gsell, M.A.; Sánchez, J.; Plank, G.; Dössel, O.; Loewe, A. Comparison of Propagation Models and Forward Calculation Methods on Cellular, Tissue and Organ Scale Atrial Electrophysiology. IEEE Trans. Biomed. Eng. 2022, 1–12. [Google Scholar] [CrossRef]
- Hoffman, B.F.; Rosen, M.R. Cellular mechanisms for cardiac arrhythmias. Circ. Res. 1981, 49, 1–15. [Google Scholar] [CrossRef]
- Keating, M.T.; Sanguinetti, M.C. Molecular and cellular mechanisms of cardiac arrhythmias. Cell 2001, 104, 569–580. [Google Scholar] [CrossRef]
- Tse, G. Mechanisms of cardiac arrhythmias. J. Arrhythmia 2016, 32, 75–81. [Google Scholar] [CrossRef]
- Luongo, G.; Vacanti, G.; Nitzke, V.; Nairn, D.; Nagel, C.; Kabiri, D.; Almeida, T.P.; Soriano, D.C.; Rivolta, M.W.; Ng, G.A.; et al. Hybrid machine learning to localize atrial flutter substrates using the surface 12-lead electrocardiogram. EP Eur. 2022, 24, 1186–1194. [Google Scholar] [CrossRef]
- BIPM; IFCC; ISO. IUPAP and OIML 2008, Supplement 1 to the ‘Guide to the Expression of Uncertainty in Measurement’—Propagation of Distributions Using a Monte Carlo Method JCGM 101: 2008; JCGM: Geneva, Switzerland, 2008. [Google Scholar]
- Rasmussen, K.; Kondrup, J.B.; Allard, A.; Demeyer, S.; Fischer, N.; Barton, E.; Partridge, D.; Wright, L.; Bär, M.; Fiebach, H.; et al. Novel Mathematical and Statistical Approaches to Uncertainty Evaluation: Best Practice Guide to Uncertainty Evaluation for Computationally Expensive Models; Euramet: Brunswick, Germany, 2015. [Google Scholar]
- Xiu, D. Numerical methods for stochastic computations. In Numerical Methods for Stochastic Computations; Princeton University Press: Princeton, NJ, USA, 2010. [Google Scholar]
- Smith, R.C. Uncertainty Quantification: Theory, Implementation, and Applications; Siam: Philadelphia, PA, USA, 2013; Volume 12. [Google Scholar]
- Ghanem, R.; Higdon, D.; Owhadi, H. Handbook of Uncertainty Quantification; Springer: Berlin/Heidelberg, Germany, 2017; Volume 6. [Google Scholar]
- Heidenreich, S.; Gross, H.; Bär, M. Bayesian approach to the statistical inverse problem of scatterometry: Comparison of three surrogate models. Int. J. Uncertain. Quantif. 2015, 5, 511–526. [Google Scholar] [CrossRef]
- Heidenreich, S.; Gross, H.; Bär, M. Bayesian approach to determine critical dimensions from scatterometric measurements. Metrologia 2018, 55, S201. [Google Scholar]
- Rynn, J.A.; Cotter, S.L.; Powell, C.E.; Wright, L. Surrogate accelerated Bayesian inversion for the determination of the thermal diffusivity of a material. Metrologia 2019, 56, 015018. [Google Scholar]
- Weissenbrunner, A.; Fiebach, A.; Schmelter, S.; Bär, M.; Thamsen, P.U.; Lederer, T. Simulation-based determination of systematic errors of flow meters due to uncertain inflow conditions. Flow Meas. Instrum. 2016, 52, 25–39. [Google Scholar] [CrossRef]
- Sudret, B. Global sensitivity analysis using polynomial chaos expansions. Reliab. Eng. Syst. Saf. 2008, 93, 964–979. [Google Scholar] [CrossRef]
- Loewe, A.; Krueger, M.W.; Platonov, P.G.; Holmqvist, F.; Dössel, O.; Seemann, G. Left and Right Atrial Contribution to the P-wave in Realistic Computational Models. In Lecture Notes in Computer Science; Springer: Cham, Switzerland, 2015; Volume 9126, pp. 439–447. [Google Scholar] [CrossRef]
- Courtemanche, M.; Ramirez, R.J.; Nattel, S. Ionic mechanisms underlying human atrial action potential properties: Insights from a mathematical model. Am. J. Physiol.-Heart Circ. Physiol. 1998, 275, H301–H321. [Google Scholar] [CrossRef] [PubMed]
- Odille, F.; Liu, S.; van Dam, P.; Felblinger, J.; Odille, F.; Liu, S.; van Dam, P.; Felblinger, J. Statistical Variations of Heart Orientation in Healthy Adults. In Proceedings of the 2017 Computing in Cardiology Conference (CinC), Rennes, France, 24–27 September 2017; Volume 44. [Google Scholar] [CrossRef]
- Stenroos, M.; Mäntynen, V.; Nenonen, J. A Matlab library for solving quasi-static volume conduction problems using the boundary element method. Comput. Methods Programs Biomed. 2007, 88, 256–263. [Google Scholar] [CrossRef] [PubMed]
- Schuler, S.; Tate, J.D.; Oostendorp, T.F.; MacLeod, R.S.; Dössel, O. Spatial Downsampling of Surface Sources in the Forward Problem of Electrocardiography. In Proceedings of the Functional Imaging and Modeling of the Heart; Coudière, Y., Ozenne, V., Vigmond, E., Zemzemi, N., Eds.; Lecture Notes in Computer Science; Springer International Publishing: Cham, Switzerland, 2019; Volume 11504, pp. 29–36. [Google Scholar] [CrossRef]
- Saltelli, A. Global sensitivity analysis: An introduction. In Proceedings of the 4th International Conference on Sensitivity Analysis of Model Output (SAMO 2004), Santa Fe, NM, USA, 8–11 March 2004. [Google Scholar]
- Ge, Q.; Menendez, M. Extending Morris method for qualitative global sensitivity analysis of models with dependent inputs. Reliab. Eng. Syst. Saf. 2017, 162, 28–39. [Google Scholar] [CrossRef]
- Sobol, I.M. Sensitivity analysis for non-linear mathematical models. Math. Model. Comput. Exp. 1993, 1, 407–414. [Google Scholar]
- Ye, M.; Hill, M. Chapter 10—Global Sensitivity Analysis for Uncertain Parameters, Models, and Scenarios. In Sensitivity Analysis in Earth Observation Modelling; Petropoulos, G.P., Srivastava, P.K., Eds.; Elsevier: Amsterdam, The Netherlands, 2017; pp. 177–210. [Google Scholar] [CrossRef]
- Morris, M.D. Factorial Sampling Plans for Preliminary Computational Experiments. Technometrics 1991, 33, 161–174. [Google Scholar] [CrossRef]
- Campolongo, F.; Cariboni, J.; Saltelli, A. An effective screening design for sensitivity analysis of large models. Environ. Model. Softw. 2007, 22, 1509–1518. [Google Scholar] [CrossRef]
- Sobol’, I. Global sensitivity indices for nonlinear mathematical models and their Monte Carlo estimates. Math. Comput. Simul. 2001, 55, 271–280. [Google Scholar] [CrossRef]
- Wiener, N. The homogeneous chaos. Am. J. Math. 1938, 60, 897–936. [Google Scholar] [CrossRef]
- Xiu, D.; Karniadakis, G.E. The Wiener–Askey Polynomial Chaos for Stochastic Differential Equations. SIAM J. Sci. Comput. 2002, 24, 619–644. [Google Scholar] [CrossRef]
- Ernst, O.G.; Mugler, A.; Starkloff, H.J.; Ullmann, E. On the convergence of generalized polynomial chaos expansions. ESAIM Math. Model. Numer. Anal. 2012, 46, 317–339. [Google Scholar] [CrossRef]
- Kaintura, A.; Dhaene, T.; Spina, D. Review of Polynomial Chaos-Based Methods for Uncertainty Quantification in Modern Integrated Circuits. Electronics 2018, 7, 30. [Google Scholar] [CrossRef]
- Farchmin, N.; Hammerschmidt, M.; Schneider, P.I.; Wurm, M.; Bodermann, B.; Bär, M.; Heidenreich, S. Efficient Bayesian inversion for shape reconstruction of lithography masks. J. Micro/Nanolith. MEMS MOEMS 2020, 19, 024001. [Google Scholar] [CrossRef]
- Saltelli, A. Making Best Use of Model Evaluations to Compute Sensitivity Indices. Comput. Phys. Commun. 2002, 145, 280–297. [Google Scholar] [CrossRef]
- Oakley, J.E.; O’Hagan, A. Probabilistic sensitivity analysis of complex models: A Bayesian approach. J. R. Stat. Soc. Ser. B (Stat. Methodol.) 2004, 66, 751–769. [Google Scholar] [CrossRef]
- Farchmin, N. Adaptive and Non-Intrusive Uncertainty Quantication for High-Dimensional Parametric PDEs. Ph.D. Thesis, TU Berlin, Berlin, Germany, 2022. [Google Scholar]
- Farchmin, N. PyThia Uncertainty Quantification Toolbox. Vers.: 2.0. 2021. Available online: https://gitlab1.ptb.de/pythia/pythia/ (accessed on 15 September 2022).
- Oseledets, I.V. Tensor-train decomposition. SIAM J. Sci. Comput. 2011, 33, 2295–2317. [Google Scholar] [CrossRef]
- Homma, T.; Saltelli, A. Importance measures in global sensitivity analysis of nonlinear models. Reliab. Eng. Syst. Saf. 1996, 52, 1–17. [Google Scholar] [CrossRef]
- Gillette, K.; Gsell, M.A.; Nagel, C.; Bender, J.; Winkler, B.; Williams, S.E.; Bär, M.; Schäffter, T.; Dössel, O.; Plank, G.; et al. MedalCare-XL: 16,900 healthy and pathological 12 lead ECGs obtained through electrophysiological simulations. arXiv 2022, arXiv:2211.15997. [Google Scholar]
- Nagel, C.; Pilia, N.; Loewe, A.; Dössel, O. Quantification of Interpatient 12-lead ECG Variabilities within a Healthy Cohort. Curr. Dir. Biomed. Eng. 2020, 6, 493–496. [Google Scholar] [CrossRef]








| Model Parameter | Small Variation | Large Variation |
|---|---|---|
| CV (bulk tissue) | ||
| CV (interatrial connections) | ||
| translation X | [−10 mm, 10 mm] | [−20 mm, 20 mm] |
| translation Y | [−10 mm, 10 mm] | [−20 mm, 20 mm] |
| translation Z | [−10 mm, 10 mm] | [−20 mm, 20 mm] |
| angle X | [, ] | [, ] |
| angle Y | [, ] | [, ] |
| angle Z | [, ] | [, ] |
| I | II | III | aVL | aVR | aVF | V1 | V2 | V3 | V4 | V5 | V6 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | |
| 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | |
| 0.106 | 0.136 | 0.003 | 0.177 | 0.032 | 0.038 | 0.162 | 0.176 | 0.165 | 0.119 | 0.157 | 0.146 | |
| 0.092 | 0.119 | 0.001 | 0.155 | 0.027 | 0.032 | 0.250 | 0.283 | 0.321 | 0.340 | 0.252 | 0.049 | |
| 0.346 | 0.039 | 0.167 | 0.236 | 0.294 | 0.037 | 0.103 | 0.139 | 0.075 | 0.024 | 0.015 | 0.034 | |
| 0.032 | 0.154 | 0.090 | 0.082 | 0.045 | 0.144 | 0.110 | 0.124 | 0.185 | 0.189 | 0.172 | 0.154 | |
| 0.202 | 0.378 | 0.710 | 0.062 | 0.497 | 0.713 | 0.006 | 0.016 | 0.020 | 0.031 | 0.064 | 0.318 | |
| 0.176 | 0.138 | 0.016 | 0.231 | 0.079 | 0.024 | 0.127 | 0.102 | 0.171 | 0.249 | 0.286 | 0.236 | |
| 0.954 | 0.963 | 0.988 | 0.943 | 0.974 | 0.986 | 0.758 | 0.840 | 0.937 | 0.952 | 0.946 | 0.938 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Winkler, B.; Nagel, C.; Farchmin, N.; Heidenreich, S.; Loewe, A.; Dössel, O.; Bär, M. Global Sensitivity Analysis and Uncertainty Quantification for Simulated Atrial Electrocardiograms. Metrology 2023, 3, 1-28. https://doi.org/10.3390/metrology3010001
Winkler B, Nagel C, Farchmin N, Heidenreich S, Loewe A, Dössel O, Bär M. Global Sensitivity Analysis and Uncertainty Quantification for Simulated Atrial Electrocardiograms. Metrology. 2023; 3(1):1-28. https://doi.org/10.3390/metrology3010001
Chicago/Turabian StyleWinkler, Benjamin, Claudia Nagel, Nando Farchmin, Sebastian Heidenreich, Axel Loewe, Olaf Dössel, and Markus Bär. 2023. "Global Sensitivity Analysis and Uncertainty Quantification for Simulated Atrial Electrocardiograms" Metrology 3, no. 1: 1-28. https://doi.org/10.3390/metrology3010001
APA StyleWinkler, B., Nagel, C., Farchmin, N., Heidenreich, S., Loewe, A., Dössel, O., & Bär, M. (2023). Global Sensitivity Analysis and Uncertainty Quantification for Simulated Atrial Electrocardiograms. Metrology, 3(1), 1-28. https://doi.org/10.3390/metrology3010001







