1. Introduction
This paper addresses the challenge of classifying visual images based on human brain activity recorded through electroencephalogram (EEG) signals—a pattern recognition task where the goal is to determine what image category a person is viewing based solely on their neural responses. The electroencephalogram (EEG) signal, gathered from the scalp, directly reflects human brain activity over time. It is widely used as a non-invasive and easy-to-implement technique for classifying brain activity [
1]. EEG-based classifications have increasingly drawn the interest of brain science and neuroscience research teams and have been investigated in various areas such as emotion recognition [
2,
3], disease detection [
4,
5], brain–computer interface [
6,
7], semantic analysis [
8], etc.
In visual pattern recognition from brain signals, subjects view images from different categories (such as animals, objects, or scenes) and their EEG responses are recorded. The fundamental question we address is the following: Can we accurately predict which image category a person is viewing based only on their brain signals? This capability has significant implications for brain–computer interfaces, assistive technologies for individuals with motor disabilities, and understanding the neural basis of visual perception.
Recent advancements in EEG acquisition equipment, signal processing techniques, machine learning methods, and other related fields have significantly promoted research on image classification from brain signals. These advancements have shown that using EEG signals for content understanding, pattern recognition, and classification is not just feasible, but also universal. However, achieving robust image classification from EEG signals faces two critical challenges: (i) Most EEG datasets are relatively small due to the high cost of data gathering and the unique characteristics of EEG signals and (ii) EEG data are high-dimensional, which poses challenges in exploring EEG-based classification. For instance, utilizing deep models for EEG-based brain activity classification is tougher due to the need for sufficient training data. To address the issue of limited EEG data, methods such as data augmentation [
9], dimensionality reduction [
10], and database merging [
11] have been reported as potential solutions.
On the other hand, the scalp-recorded EEG signals contain spatiotemporal differences, including spatial and temporal perception differences. Specifically, the spatial perception difference of EEG signals is promoted by the distinctions in the size and shape of the subjects’ heads and the distinctions in the structure and function distribution of the subjects’ brains. In EEG signal recording, subjects of similar head sizes wear an electrode cap, including small, medium, and large. However, the specific measure and shape of these subjects’ heads are unique; this prompts the fact that the same electrode on an EEG cap cannot be ensured to fall at a similar scalp position of each subject. Thus, the distinctions in structure and function between subjects’ brains may lead to diverse functional information of the EEG signals collected by the same electrode. People ignore this spatial deviation and process EEG signals gathered from various subjects with a unified methodology. Moreover, the scale-recoded EEG signals contain time perception distinction between subjects and sessions. Various subjects have distinctive response speeds and response cycles for similar boosts. For one subject, the response speeds and cycles to the same stimuli may also be distinctive in various sessions because of the changing body state and intellectual level. The time perception difference expands the difficulty of feature extraction and feature selection on EEG data collected from multiple subjects [
12]. It reduces the generalization capability of the EEG-based classifier between individuals and tasks [
13]. For this issue, a few investigations have attempted to adjust EEG signals in the time domain and lessen the impact of time perception distinctions by the time-warping method [
1,
14].
To address the limitations of EEG signals and develop a unified spatiotemporal EEG-based classification model, we propose a comprehensive deep framework featuring context-based spatiotemporal disambiguation for EEG signal classification. Our contributions include: (i) utilizing dimensionality reduction based on brain asymmetry to mitigate sample size and dimensionality constraints; (ii) implementing a feature selection method to choose spatially representative channels that capture the principal components for the current task across all subjects; (iii) applying a time-warping method to align signals in the time domain effectively; and (iv) conducting extensive experiments to demonstrate that our deep framework outperforms existing state-of-the-art methods.
2. Related Work
Electroencephalographic signal analysis fundamentally operates through two distinct computational phases: feature extraction and pattern recognition [
15,
16]. The initial phase involves deriving meaningful characteristics from neural recordings through sophisticated signal processing and analytical methodologies. The subsequent phase encompasses the examination of extracted feature patterns utilizing appropriate pattern recognition algorithms. Before the widespread adoption of deep learning methodologies, electroencephalographic signal analysis predominantly relied on time-frequency domain characteristics, including power spectral density measurements and differential entropy calculations, which were obtained through conventional signal processing approaches. Classical pattern recognition and machine learning frameworks encompassed artificial neural networks, Naive Bayes classification algorithms, and support vector machine implementations. The emergence of deep learning technologies has catalyzed a paradigm shift, with numerous research groups within neuroscience and brain science disciplines actively exploring the implementation of advanced deep learning methodologies for electroencephalographic data comprehension and analysis. Contemporary research efforts are increasingly focused on developing comprehensive end-to-end architectures that seamlessly integrate feature extraction processes with classification or clustering operations, thereby eliminating the traditional separation between these computational stages and enabling more sophisticated neural signal interpretation capabilities.
Recent investigations have demonstrated the substantial potential of deep learning architectures in electroencephalographic signal analysis across diverse neurological applications. Kulasingham et al. implemented dual deep learning methodologies, specifically deep belief networks and stacked autoencoder architectures, for detecting P300 event-related potentials within guilty knowledge test paradigms [
17]. Recent research by Zhou et al. (2025) [
18] investigated cognitive load recognition using EEG signals during complex simulated flight missions, addressing the gap between simplistic laboratory tasks and real-world operational environments. Their study employed the Multi-Attribute Task Battery (MATB) to induce three levels of cognitive load (low, medium, high) across multiple sessions, collecting both EEG data and behavioral metrics from 36 participants. The researchers compared traditional machine learning approaches (PSD features with SVM) against several convolutional neural network architectures, including shallow CNN, deep CNN, EEGNet, EEGNex, and EEGTCN. Their findings revealed that simpler CNN models, particularly the shallow CNN, achieved superior performance (up to 83% accuracy) in within-subject cognitive load classification compared to more complex architectures [
18]. Wang et al. presented an innovative electroencephalographic motor imagery classification system based on long short-term memory network architectures, incorporating one-dimensional aggregate approximation techniques to achieve enhanced signal representation capabilities [
19]. Furthermore, Gao et al. constructed a specialized electroencephalographic spatial-temporal convolutional neural network (ESTCNN) that strengthens temporal dependency modeling for individual electrode channels while simultaneously improving spatial feature extraction mechanisms, resulting in substantial performance improvements for driver fatigue detection applications [
20]. Liu et al. addressed the critical issue of reliability assessment in automated EEG-based epileptic seizure detection by proposing an Evidential Multi-view Learning (EML) framework. Recognizing that traditional seizure detection methods focus primarily on accuracy while neglecting decision reliability, their approach integrates multiple feature representations (temporal, spectral, and temporal-spectral views) to mitigate noise effects inherent in EEG signals. By dynamically weighting views based on their confidence levels during the fusion process and incorporating a multi-view common graph to maintain cross-view consistency, the method achieved 99.05% accuracy on the CHB-MIT dataset, outperforming state-of-the-art approaches [
21]. Li et al. demonstrated the clinical utility of transfer learning methodologies in constructing convolutional neural network models, establishing their effectiveness for objective, precise, and expeditious diagnosis of mild depressive disorders [
22]. Zhang et al. proposed innovative cascade and parallel convolutional recurrent neural network architectures for precise detection of voluntary human movements through effective spatiotemporal feature extraction from unprocessed electroencephalographic signals [
23]. Additionally, Tan et al. engineered a hybrid brain–computer interface rehabilitation support system utilizing combined convolutional neural network and recurrent neural network methodologies for electroencephalographic signal classification, integrating video-based electroencephalographic analysis with optical flow processing techniques [
24].
Deep learning has significantly enhanced EEG-based emotion recognition in recent years, a crucial aspect of EEG analysis. Numerous deep learning techniques have been introduced to accelerate progress and expand the application of this field. Wang et al. introduced a hierarchical spatial learning transformer (HSLT) model for EEG-based emotion recognition that addresses the limitations of conventional approaches in capturing long-range spatial dependencies across electrodes and brain regions. Their method organizes EEG electrodes into nine anatomically based brain region clusters and employs a two-stage transformer architecture: electrode-level encoders that integrate information within individual brain regions, followed by brain-region-level encoding that captures inter-regional dependencies. Through subject-independent experiments on the DEAP and MAHNOB-HCI databases, they achieved accuracies of 65.75% and 66.51% for arousal and valence classification, respectively, on DEAP, with comparable results on MAHNOB-HCI [
25]. Li et al. developed a bi-hemisphere domain adversarial neural network (BiDANN) model featuring a global and two local domain discriminators that adversarially interact with a classifier to extract distinct emotional features for each hemisphere, achieving leading performance on the SEED emotion recognition database [
26]. Luo et al. introduced a novel Wasserstein generative adversarial network domain adaptation (WGANDA) framework to address domain shift issues in cross-subject EEG-based emotion recognition, demonstrating that this framework significantly surpasses existing domain adaptation methods on two public EEG datasets for emotion recognition [
27].
In addition, deep learning methods are becoming increasingly vital in EEG-based multimedia content analysis [
28,
29,
30,
31]. Spampinato et al. introduced an RNN-based approach to learn descriptors for visual stimuli-evoked EEG data, mapping CNN image features to EEG features to predict image classes accurately [
32]. Xue et al. developed a hybrid local–global neural network architecture for EEG-based visual classification that processes raw signals without requiring handcrafted frequency-domain features. Their framework introduces several innovative components: a reweight module that adaptively learns electrode importance across subjects rather than relying on fixed spatial coordinates, a local feature extraction module combining one-dimensional temporal convolutions with residual connections to capture both simple and complex signal patterns, and a global transformer block for modeling long-range temporal dependencies. For high sampling rate scenarios, they incorporated a feature fusion module to handle the increased dimensionality effectively. The model was extensively validated across five public datasets at multiple sampling rates (62.5 Hz, 125 Hz, and 250 Hz), achieving state-of-the-art performance including 55.93% accuracy on the EEG72 6-class task and 32.24% on the 72-class task, surpassing previous methods that relied on combined spectral-temporal features [
33]. Research has demonstrated the potential for generating multimedia content information from EEG data. Kavasidis et al. proposed a method for creating images from visually evoked brain signals recorded via EEG [
34], using variational autoencoders (VAE) and generative adversarial networks (GAN) to produce semantically coherent images. Additionally, Tirupattur et al. utilized adversarial learning to reveal that EEG signals encode cues from thoughts, generating semantically relevant visualizations [
35].
The advancement of deep learning technology has contributed to the progress of EEG classification, particularly in tasks like EEG-based visual content analysis. However, compared to the progress seen in computer vision [
36,
37,
38,
39], EEG analysis still faces several limitations. These include EEG data’s limited availability, EEG datasets’ high dimensionality, and the spatiotemporal variations in EEG signals. Further research is needed to address these issues.
3. Methodology
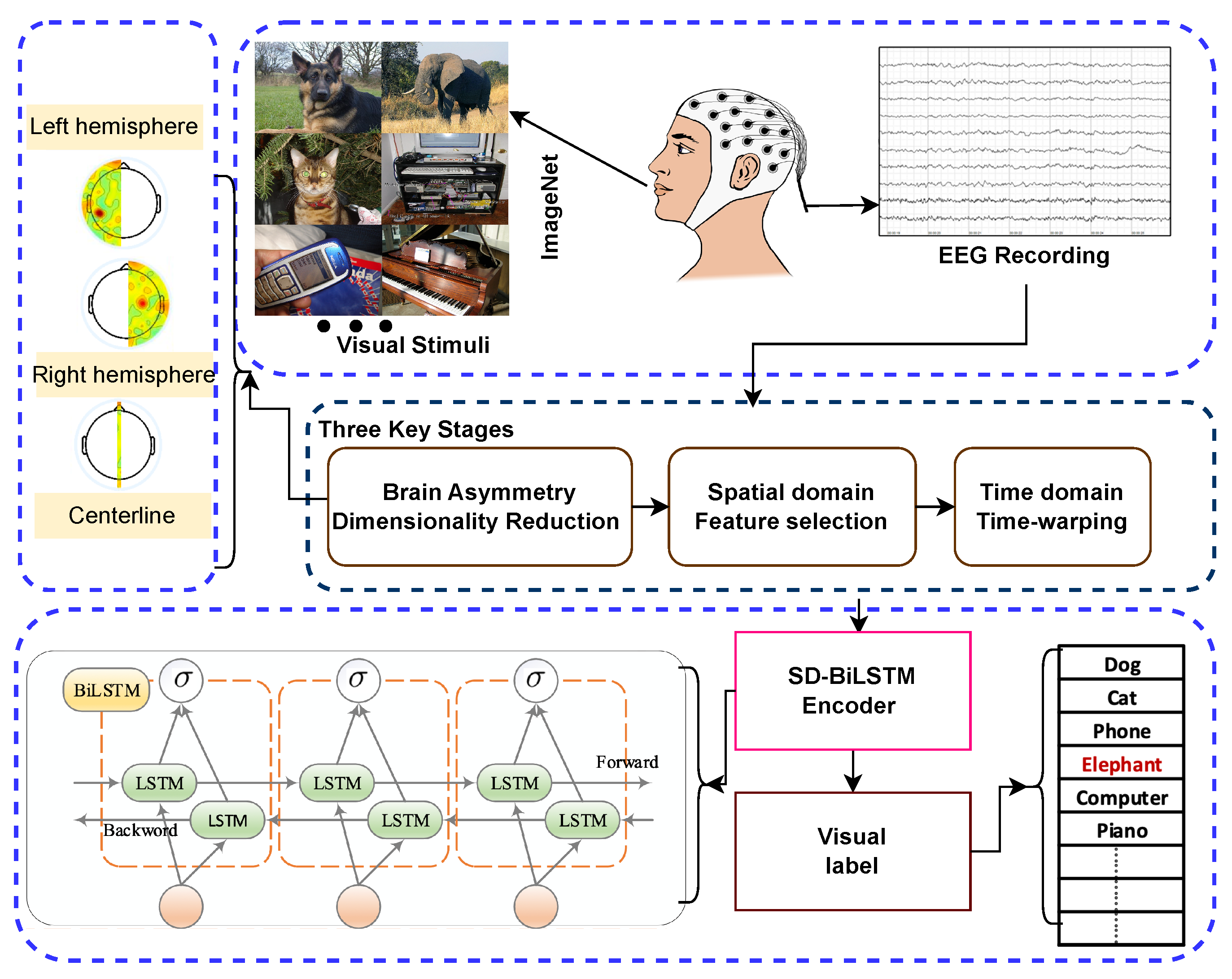
Based on the comprehensive survey of previous studies, a deep framework driven by spatiotemporal disambiguation (SD) for classifying visual images based on EEG recordings (SD-BiLSTM) is proposed. Our approach enables accurate prediction of which image category from ImageNet a subject is viewing based on their brain signals. The structure of the proposed Spatiotemporal Disambiguation BiLSTM (SD-BiLSTM) framework, along with the training and testing phases, is depicted in
Figure 1. Our approach comprises three stages:
Dimensionality Reduction Based on Brain Asymmetry: This step reduces data complexity and addresses sample size limitations.
Feature Selection of Spatially Efficient Channels: We select channels that effectively represent principal components for the task.
Time-Warping: This aligns signals in the time domain for consistency.
Figure 1.
Architecture of the proposed Spatiotemporal Disambiguation BiLSTM (SD-BiLSTM) deep framework.
Figure 1.
Architecture of the proposed Spatiotemporal Disambiguation BiLSTM (SD-BiLSTM) deep framework.
Multi-channel EEG signals are recorded while subjects view images, and the data are divided into training and test subsets. During the training phase, the classifier uses the training data to generate the model. In the testing phase, this model predicts the visual labels of the test subset.
3.1. Brain Asymmetry-Based Dimensionality Reduction
In information theory, machine learning, and statistics, dimensionality reduction involves reducing data dimensions to obtain a set of principal features. This helps with data compression, visualization, redundancy elimination, simplification, and model training and testing optimization. When dealing with multi-channel EEG signals, reducing the number of channels can not only remove noise [
40] but also lessen the impact of the EEG data shortage, thus enhancing the generalization capability of EEG-based models [
10].
In this study, we implement dimensionality reduction on EEG signals based on brain asymmetry. Brain asymmetry refers to the brain’s selective specialization of certain neural activities or cognitive processes in either the right or left hemisphere [
41]. Research has shown that brain asymmetry can provide valuable features for EEG-based tasks. For example, Ahmed et al. used statistical features from EEG data of the left and right brains to explore the correlation between brain hemispheres and emotions [
42]. Aris et al. proposed the Asymmetry Score feature to investigate brain activity in relaxed and non-relaxed states [
43]. Additionally, Koolen et al. investigated the quantification of interhemispheric synchrony (IHS) in neonatal EEG using the activation synchrony index (ASI), addressing the limitations of traditional visual assessment methods that lack objective, quantifiable definitions [
44].
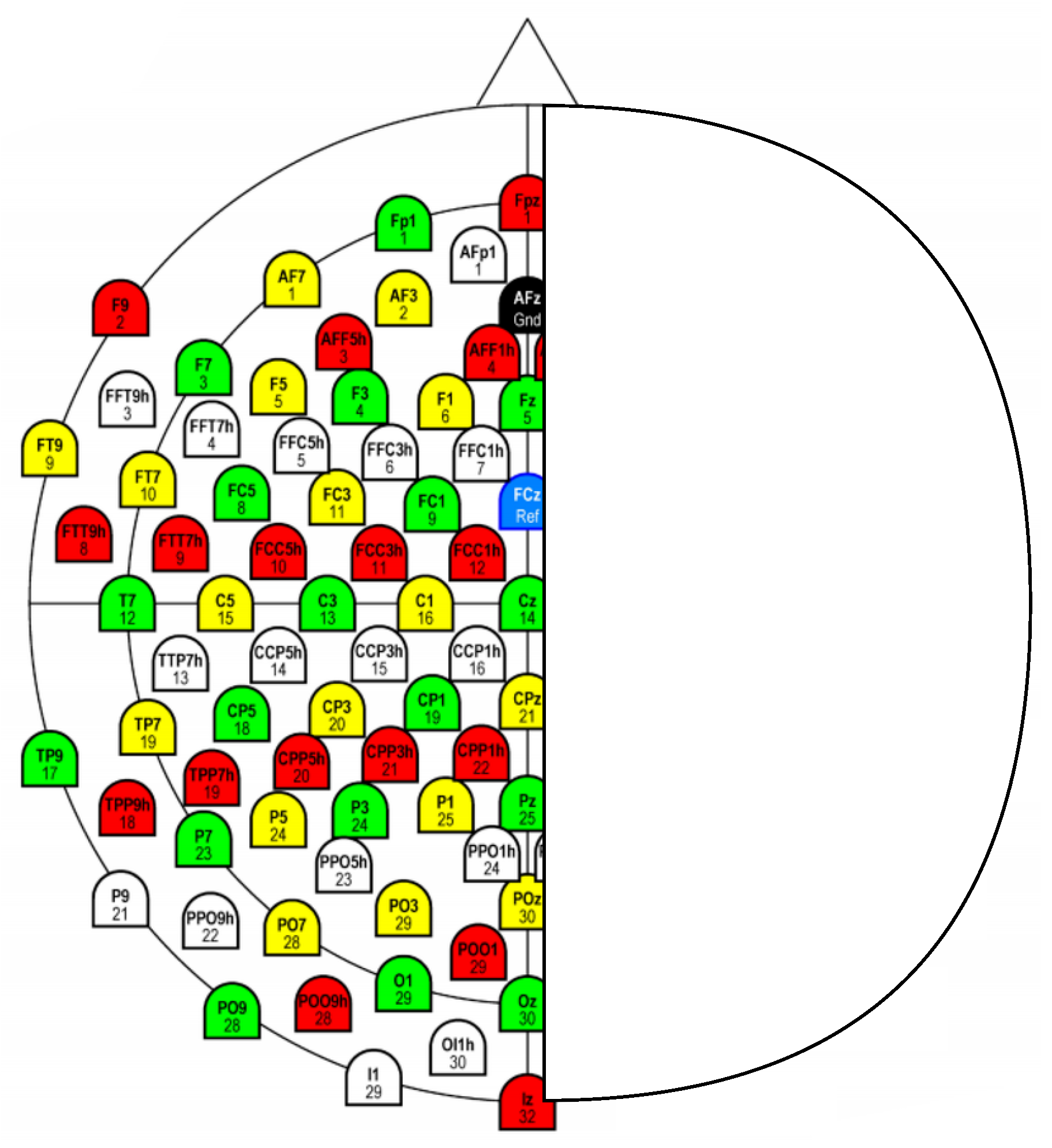
Our approach integrates dimension reduction with brain asymmetry by applying dimensionality reduction to EEG signals. Specifically, we retain signals from electrodes at the centerline position and subtract the signal from electrodes in the right hemisphere from their corresponding counterparts in the left hemisphere. For example, using the 128-channel EEG cap of the international 10–20 system, we focus on eight centerline electrodes (Fpz, Fz, Cz, CPz, Pz, POz, Oz, and Iz) and sixty pairs of brain-symmetric electrodes, as illustrated in
Figure 2.
3.2. Channel Selection for Spatial Representation
This investigation utilizes channel selection methodologies to determine the most informative EEG electrodes that efficiently encapsulate the fundamental characteristics of the experimental paradigm across all participants. As depicted in
Figure 1, the training dataset encompasses EEG recordings from all subjects. The implementation employs the Fisher score feature selection approach to identify spatially informative channels from the training dataset and corresponding class labels. The Fisher score represents a feature selection methodology designed to determine the most discriminative attributes for classification problems [
45]. This technique assesses each feature dimension through computation of the between-class and within-class variance ratio. The primary goal involves selecting attributes that maximize inter-class separation while minimizing intra-class dispersion.
Between-class Variance (): Quantifies the variance across different classes, representing the degree of class separation.
Within-class Variance (): Quantifies the variance within individual classes, indicating class cohesion.
The Fisher score for the
jth dimensional attribute is expressed as
An elevated Fisher score value indicates that the attribute possesses substantial discriminative capability, establishing it as an optimal candidate for class differentiation within the dataset. Through ranking attributes according to their Fisher score values, one can identify the most efficient attributes for enhancing classification performance. The between-class variance is computed as follows:
where
L represents the number of classes in the training dataset,
N denotes the total sample count,
indicates the sample count for class
l,
represents the mean value for class
l, and
denotes the global mean. The within-class variance is determined as
where
represents the value of the
jth dimension for sample
z. A dimension exhibiting robust classification capability should demonstrate elevated between-class variance and reduced within-class variance. Consequently, a higher Fisher score indicates stronger classification correlation for that dimension. To select multiple dimensions, one can rank the Fisher score and select those with superior values.
3.3. Time-Warping
In temporal signal processing, temporal alignment, Time-Warping, constitutes a frequently utilized approach for synchronizing peaks in dual spectra through time axis adjustments [
46]. We implement temporal alignment to accommodate temporal perception variations between sessions and subjects by synchronizing EEG signals that exhibit temporal deviations. Specifically, we utilize spatially informative channels identified through feature selection on the training dataset as the reference signal for temporal alignment. For each electrode within the EEG sample, we employ the temporal alignment algorithm to synchronize the electrode’s signal with the nearest spatially informative channel. It should be emphasized that the spatially informative channels identified from the training dataset are directly utilized as the alignment reference for the testing dataset. Our methodology employs the Dynamic Temporal Warping (DTW) algorithm to synchronize the EEG signals. Let us provide a concise overview of this algorithm.
Dynamic Temporal Warping (DTW) represents a distinguished algorithm for evaluating similarity between dual temporal sequences that may vary in velocity or duration [
47]. It achieves this through determining an optimal correspondence, termed the warping trajectory.
We consider dual time series, and , with lengths and . DTW attempts to identify the warping trajectory that minimizes the accumulated distance between corresponding points in these sequences.
The warping trajectory
comprises index pairs
, where
, and
R represents the trajectory length, satisfying
Each pair corresponds point in with point in . The trajectory begins at and terminates at , ensuring complete traversal of both sequences. The indices u and v progress monotonically, preserving sequence ordering.
The accumulated distance along
, designated as
, is calculated as
Here, denotes the distance between corresponding points and , commonly computed using the Euclidean distance.
To establish the optimal warping trajectory, DTW minimizes
. This is typically accomplished through dynamic programming, which iteratively computes the accumulated distance
for subsequences of
and
up to indices
u and
v:
The value represents the minimal distance along the optimal trajectory, effectively quantifying the similarity between the time series.
4. Experimental Results
The performance assessment of our proposed spatiotemporal disambiguation methodology is conducted using a publicly accessible electroencephalogram dataset specifically designed for visual stimulus classification tasks [
32,
48]. This comprehensive dataset encompasses 12,000 neural signal recordings collected from six participants during visual perception experiments. The experimental protocol involved presenting participants with carefully curated image stimuli selected from ImageNet, organized into 40 distinct categorical classes, with each class containing 50 representative images.
Neural activity acquisition was accomplished using a high-density electrode array configuration consisting of 128 recording channels equipped with active, low-impedance sensing elements (actiCAP 128Ch system). The data acquisition system operated at a sampling frequency of 1000 Hz to ensure adequate temporal resolution for capturing rapid neural dynamics. Following standard preprocessing procedures and artifact removal protocols to eliminate contaminated signal segments, the resulting dataset contains individual trial recordings with a duration of 440 milliseconds each.
The complete dataset underwent systematic partitioning into three distinct subsets to facilitate robust model evaluation: the training partition comprises 80% of the available data (9191 samples), while both validation and testing partitions each contain 10% of the total samples (1127 samples, respectively). To ensure experimental integrity and prevent data leakage, a subject-wise and stimulus-wise splitting strategy was implemented, whereby all neural responses recorded from individual participants for specific visual stimuli were exclusively assigned to a single data partition.
To evaluate our model’s performance, we used the study by Spampinato et al. [
32] as a reference, which utilized the same dataset and LSTM encoder. Our research investigates the effectiveness of various model configurations by comparing classification performance. Our classifier is a BiLSTM model, and we compared its results with the LSTM model reported by Spampinato et al. [
32]. We also examined the impact of different data processing methods, such as dimensionality reduction based on brain asymmetry, selection of spatially representative channels, and time-warping with spatiotemporal disambiguation. Additionally, we compared our time-warping algorithm, DTW, with the Fast Parametric Time Warping (Fast-PTW) algorithm proposed by Wehrens et al. [
49] for aligning chromatograms.
In the upcoming sections, we validate the effectiveness of spatiotemporal disambiguation in two steps. First, we quantify the contributions of three key processing stages: dimensionality reduction, feature selection, and time-warping. Then, we quantify the contribution of the proposed spatiotemporal disambiguation framework to EEG-based visual classification. Our model includes a BiLSTM model with five layers: the input layer, BiLSTM layer, dense fully connected layer, soft-max layer, and output layer. We follow the specific parameter settings of Spampinato et al. [
32]. All statistical experiments are repeated five times, and the average results are reported.
4.1. Evaluations of Three Key Stages
In this phase of experiments, we first compared the precision rates of EEG-based visual classification using LSTM and BiLSTM models before and after dimensionality reduction based on brain asymmetry. The experimental results are reported in
Table 1. The precision rate of the original 128-channel dataset on LSTM was 83.41%. However, we found that dimension reduction based on brain asymmetry not only reduced the data dimension and removed redundancies but also obtained effective asymmetry features and improved the performance of visual classification.
After dimensionality reduction, the input data dimension was reduced to 68 as seen in
Figure 2, and the precision rates of LSTM and BiLSTM improved to 88.82% and 91.75% respectively, which are better than the results of the original 128-channel data on LSTM and BiLSTM (83.41% and 87.93%). Additionally, we observed that the classification precision rates on BiLSTM were better than those on LSTM, indicating that EEG signals contain effective bidirectional features that can help improve classification accuracy.
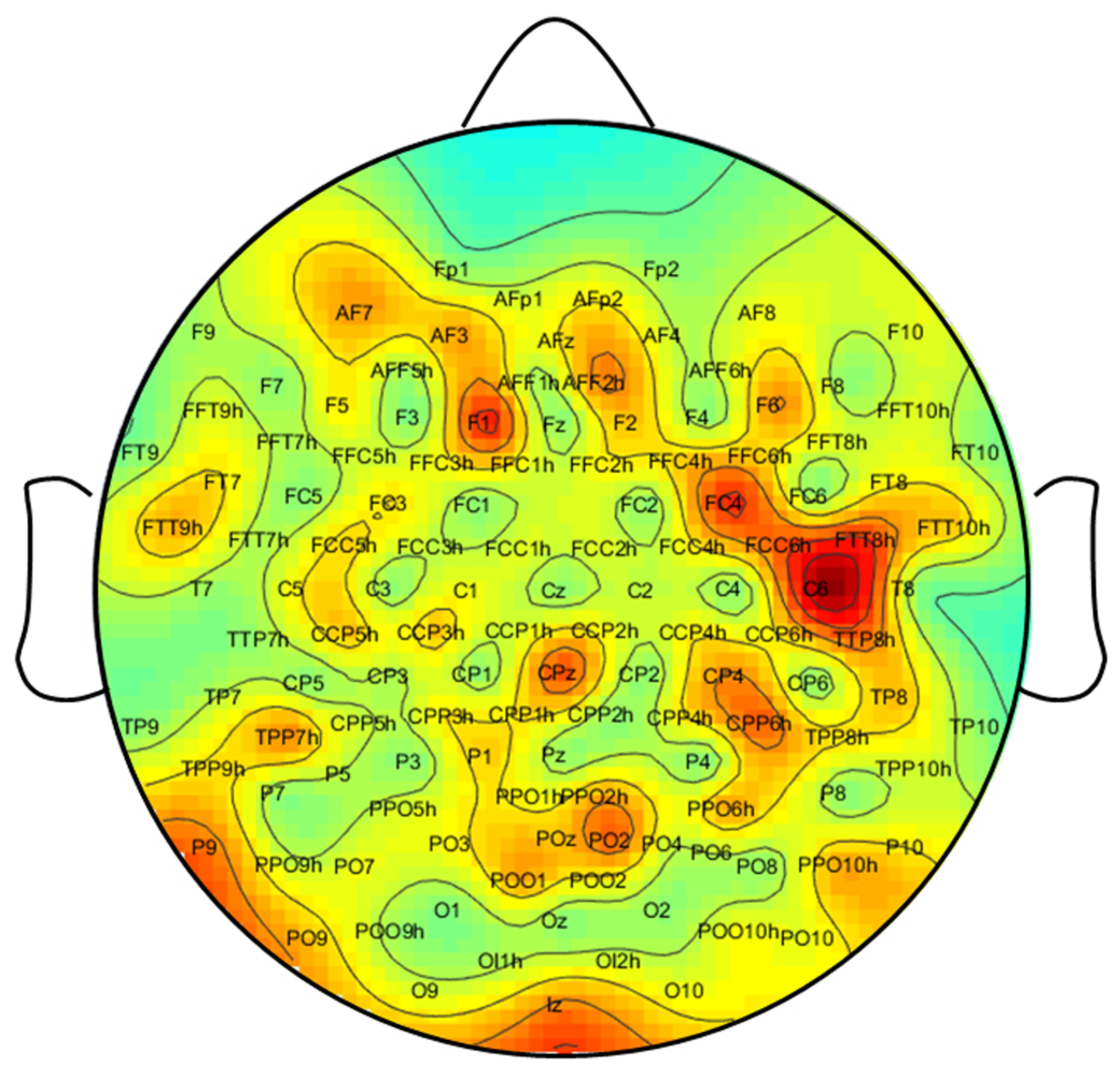
In order to investigate how effectively selecting specific spatially representative channels impacts performance, we conducted experiments on LSTM and BiLSTM models. To achieve this, we employed the Fisher score feature selection method to rank the 128 channels according to their classification ability. The distribution of the Fisher score for each channel is illustrated in
Figure 3. This figure reveals variations in the visual classification capabilities of each channel, as well as functional differences between the left and right hemispheres. For instance, channels FC4 and C6 on the right hemisphere have higher Fisher scores than those on the left hemisphere.
In this phase of the experiments, we selected the top 10, 32, and 96 high-efficiency spatially representative channels based on the Fisher score sorting as the input for the classifiers. The precision rates for visual classification are provided in
Table 2. We achieved the highest accuracy of 89.53% using LSTM with 96 channels and 90.06% using BiLSTM with 32 channels, which are higher than the results obtained with the original 128-channel data on LSTM and BiLSTM (83.41% and 87.93%, respectively).
In this phase of the experiments, we investigate the impact of the time-warping method on the original 128-channel data using LSTM and BiLSTM. After sorting based on the Fisher score, we selected 32 and 64 high-efficiency spatial channels. Subsequently, we applied the DTW algorithm to align each channel’s signal with the nearest spatially representative channel for each channel in an EEG sample. The resulting warped 128-channel EEG data were then used as input for the models.
Table 3 shows the precision rates of visual classification for the warped 128-channel data using LSTM and BiLSTM. The results demonstrate improved precision rates for visual classification after time warping. Specifically, the best accuracy of 92.10% was achieved on BiLSTM with time warping using 64 spatially efficient channels, in comparison to the accuracy of 83.41% for LSTM and 87.93% for BiLSTM using the original 128-channel data.
4.2. Experimental Results of Spatiotemporal Disambiguation
We conducted experiments to introduce and evaluate three key methods: dimension reduction based on brain asymmetry, selection of spatially efficient public channels, and time-warping. Our findings demonstrate that, in most cases, BiILSTM-based models yield better results than LSTM-based models. In this phase of experiments, we aimed to combine the aforementioned three key stages and propose a novel spatiotemporal-driven disambiguation deep framework for EEG-based brain activity classifications.
Initially, we extracted 68-dimensional asymmetry features from the original 128-channel signals using dimension reduction based on brain asymmetry. We then used the Fisher score to select 32 spatially representative channels from these 68 asymmetry features. Subsequently, we applied the DTW algorithm to align the signals of each channel with the nearest spatially representative channel. The aligned 68-channel features serve as input for the BiLSTM model. Additionally, in this phase of experiments, we compared the effectiveness of the Fast-PTW algorithm with the DTW algorithm. The experimental results are shown in
Table 4.
Table 4 summarizes the experimental results for the classification precisions of the proposed deep framework using two time-warping algorithms. The findings indicate that spatiotemporal disambiguation significantly enhances the performance of EEG-based visual classification. The precision rates achieved by the spatiotemporal disambiguation method on the BiLSTM model surpass the best results obtained from using the three key steps independently (90.06%, 91.75%, and 92.10%). By applying spatiotemporal disambiguation with the DTW algorithm, we achieved the highest precision rate of 94.23%, which exceeds the result obtained with Fast-PTW (93.35%).
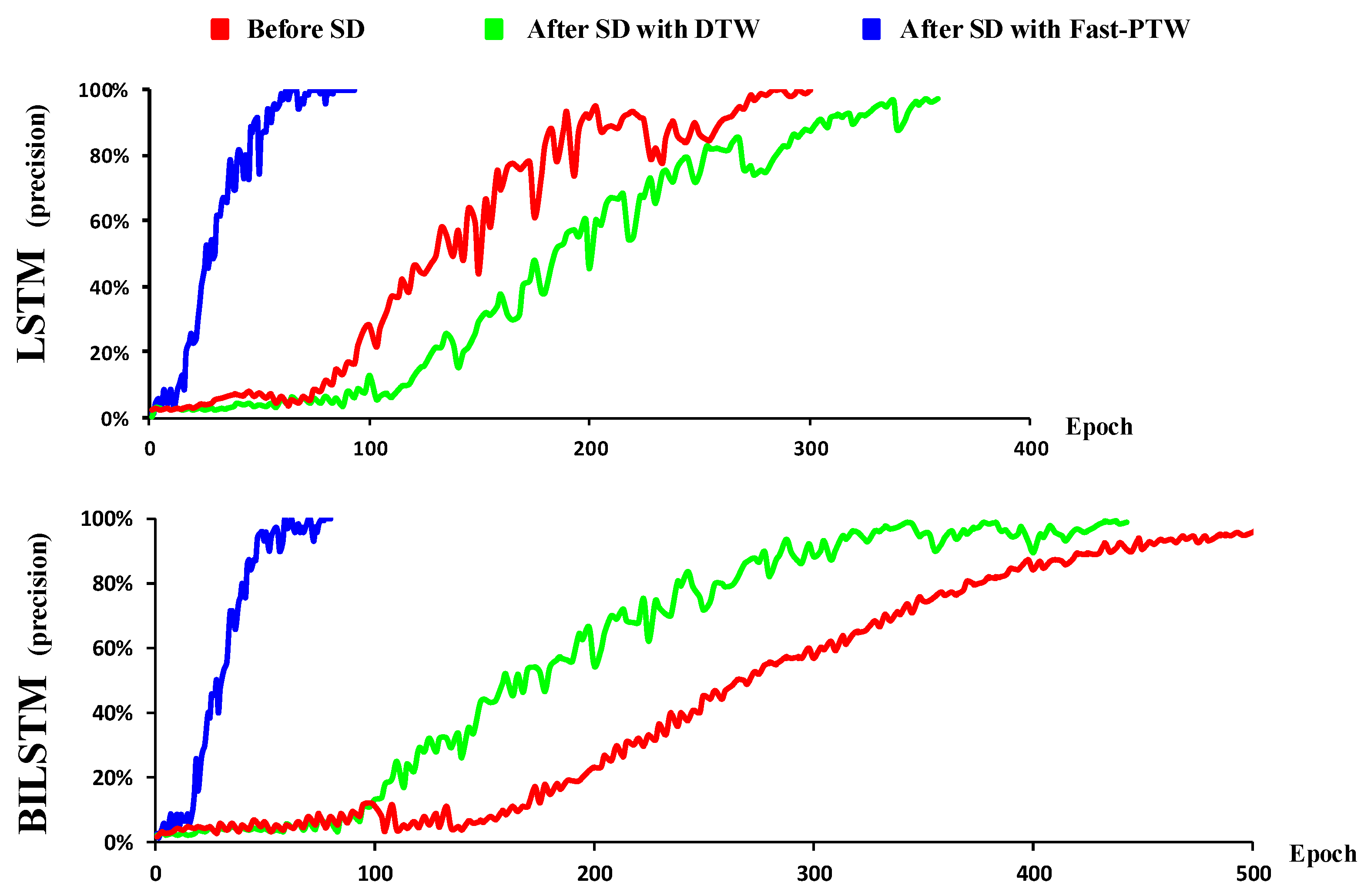
Furthermore, we found that the proposed spatiotemporal disambiguation framework quickens the convergence speed of the models during the training stage. It was observed that the convergence speed of the LSTM model after spatiotemporal disambiguation with the DTW algorithm is faster than without it (
Figure 4). For the BiLSTM model, both the convergence speeds after applying spatiotemporal disambiguation with the DTW algorithm and the Fast-PTW algorithm are faster than those without spatiotemporal disambiguation. These results indicate that the proposed spatiotemporal disambiguation method significantly reduces the data’s inconsistency, making the EEG data more consistent in both time and space dimensions. This not only improves classification performance but also accelerates the learning process of the model and reduces the requirements for the amount of training data for EEG-related tasks.
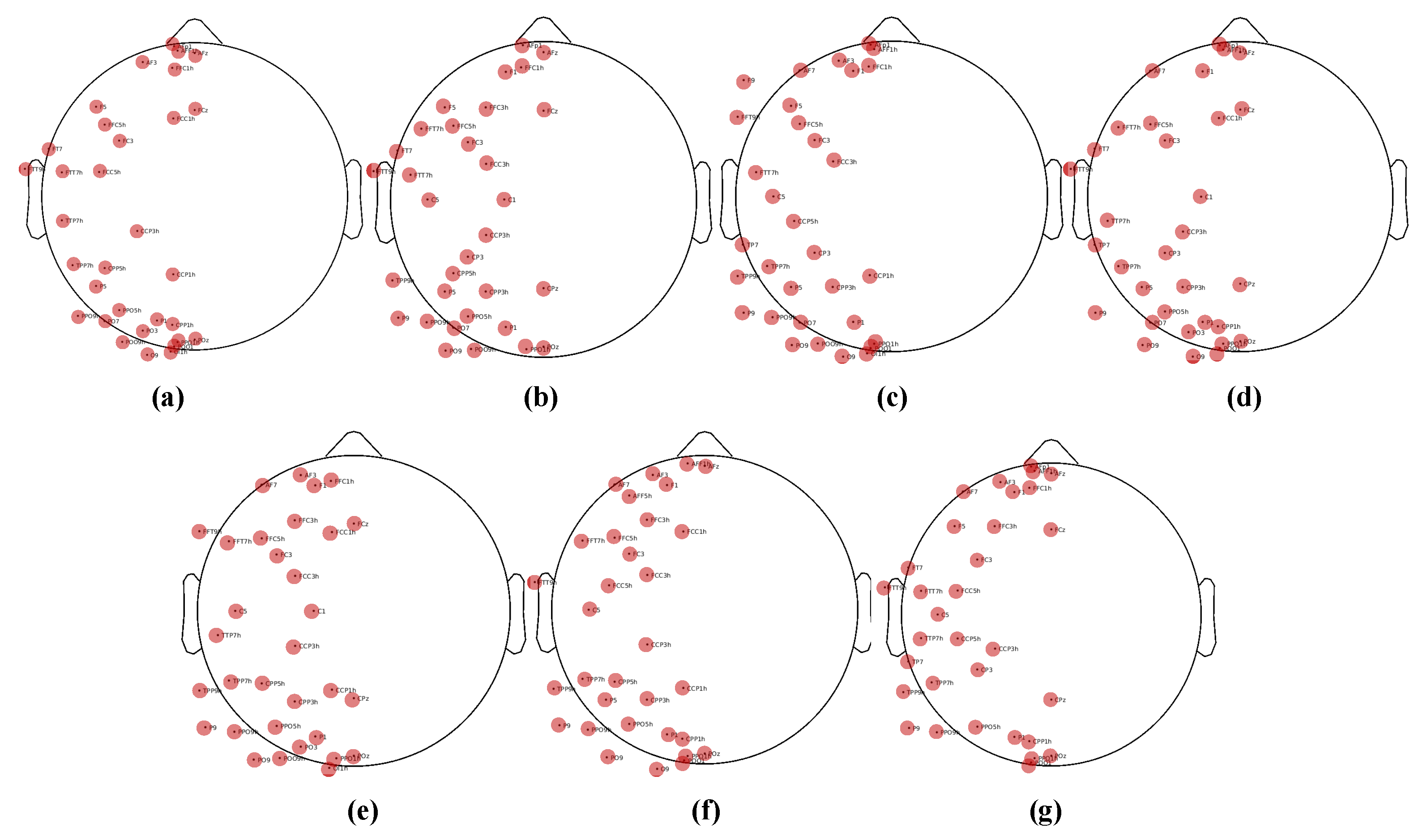
In addition to the classification results, we also provide an analysis based on feature selection. As previously described, we obtained 68-dimensional features from the original 128-channel signals by reducing dimensionality based on the human brain’s lateralization effect. We then used the Fisher score to identify 32 spatially representative channels. Are these selected channels consistent for each subject? To illustrate this, in
Figure 5, we display each subject’s physical locations on the top 32 channels. It is important to note that
Figure 5a–f correspond to each individual subject, while
Figure 5g displays the physical locations of the combined EEG data from all subjects, with features selected based on the Fisher score. It is clear that the selected channels across different subjects are consistent with each other. This demonstrates that there are some similarities between different subjects, and our method effectively captures this similarity.
4.3. Comparative Analysis with State-of-the-Art Methods
To comprehensively evaluate the efficacy of our proposed spatiotemporal disambiguation framework, extensive comparative experiments against established benchmark methods in EEG-based visual classification were conducted. This comparative analysis serves to validate the practical advantages of our approach by directly measuring classification performance against existing state-of-the-art methodologies.
Our experimental protocol utilized the same publicly available ImageNet-EEG dataset [
32,
48] employed throughout this study, ensuring consistent evaluation conditions across all methods. The comparative analysis encompasses five representative approaches from the literature: the foundational RNN-based model proposed by Spampinato et al. [
32], Transformer-based Methods, including Vision Transformer (ViT) adapted for EEG [
50] and Transformer with Positional Encoding [
51], advanced Siamese network architectures [
48], multimodal integration networks [
30], the CogniNet framework [
52], and traditional RS-LDA methodology [
53].
Table 5 presents the comparative classification accuracies achieved by our spatiotemporal disambiguation framework against these established benchmarks. Our proposed SD-BiLSTM framework, incorporating the complete spatiotemporal disambiguation pipeline, achieves a classification accuracy of 94.23%, demonstrating substantial improvements over existing approaches.
The experimental results reveal several noteworthy observations. While contemporary deep learning approaches such as Siamese networks (93.70%) and multimodal networks (94.10%) achieve competitive performance levels, our spatiotemporal disambiguation framework demonstrates consistent superiority. This improvement can be attributed to the systematic addressing of fundamental EEG signal challenges through our three-stage approach: brain asymmetry-based dimensionality reduction effectively captures lateralization features while managing data complexity, Fisher score-based channel selection identifies spatially informative electrodes across subjects, and DTW-based temporal alignment resolves inter-subject and inter-session timing variations.
The substantial performance gap between our method and the baseline RNN approach (82.90%) underscores the importance of addressing spatiotemporal variability in EEG signals. Traditional methods that process raw EEG data without explicit disambiguation suffer from the inherent signal inconsistencies across subjects and sessions. Similarly, the limited performance of classical machine learning approaches like RS-LDA (13.00%) highlights the necessity of deep learning architectures for capturing complex neural patterns in high-dimensional EEG data.
While Vision Transformer adapted for EEG signals achieves competitive performance (91.85%), it falls short of our method. The standard Transformer with positional encoding performs slightly better (92.18%), likely due to better temporal modeling. However, both Transformer variants struggle with the limited training data available in EEG datasets, as they typically require larger datasets to reach optimal performance.
These comparative results confirm that our spatiotemporal disambiguation framework not only achieves state-of-the-art classification accuracy but also provides a principled approach to handling the fundamental challenges in EEG signal analysis. The consistent improvements across different evaluation metrics suggest that our methodology successfully enhances the generalization capability of EEG-based classification systems, making them more robust for practical applications in brain–computer interfaces and neurological assessment tasks.
5. Conclusions
Electroencephalographic signal analysis for classification applications possesses a strong neurophysiological foundation. Nevertheless, this research domain encounters substantial obstacles, including insufficient dataset availability, excessive feature dimensionality, and inherent spatiotemporal signal variability. To overcome these limitations, we developed a comprehensive neural signal classification framework that prioritizes spatiotemporal signal disambiguation techniques. Our approach systematically addresses data complexity through hemisphere-based asymmetry analysis for dimensional reduction, effectively mitigating complications arising from high-dimensional feature spaces and constrained sample sizes. Subsequently, we implemented an advanced channel selection methodology to isolate the most discriminative electrodes that effectively capture spatial signal characteristics across participants. Finally, we deployed a sophisticated temporal alignment algorithm that synchronizes signal sequences derived from these optimally selected channels, thereby resolving temporal inconsistencies observed across different recording sessions and individual subjects. Comprehensive experimental validation conducted on an established electroencephalographic visual classification benchmark dataset confirmed that our spatiotemporal disambiguation approach significantly improves classification accuracy. The systematic evaluation reveals that each methodological component contributes meaningfully to overall performance enhancement while simultaneously facilitating accelerated model convergence during the training phase.
Future research directions encompass extending this framework to diverse real-world applications in neural signal interpretation and brain activity analysis tasks. Furthermore, promising opportunities exist for advancing the spatiotemporal disambiguation methodology through multimodal integration approaches, specifically combining electroencephalographic recordings, which provide superior temporal resolution, with functional magnetic resonance imaging modalities that offer enhanced spatial precision. Such integration could potentially yield more comprehensive and robust neural signal analysis capabilities.