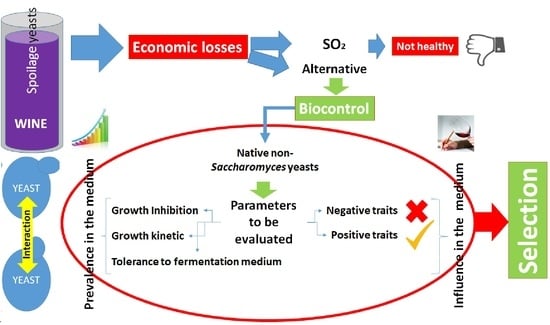
Selection of Native Non-Saccharomyces Yeasts with Biocontrol Activity against Spoilage Yeasts in Order to Produce Healthy Regional Wines
Abstract
1. Introduction
2. Materials and Methods
2.1. Microorganisms
2.2. Culture Media
2.3. Screening for Biocontrol Ability of Yeasts
2.4. Fermentative Performance
2.4.1. Growth Kinetics during Fermentation
2.4.2. Tolerance to Different Stress
2.4.3. Plate Assays
2.5. Data Analysis
3. Results and Discussion
3.1. Biocontrol Screening
3.2. Behavior of the Antagonistic Yeasts
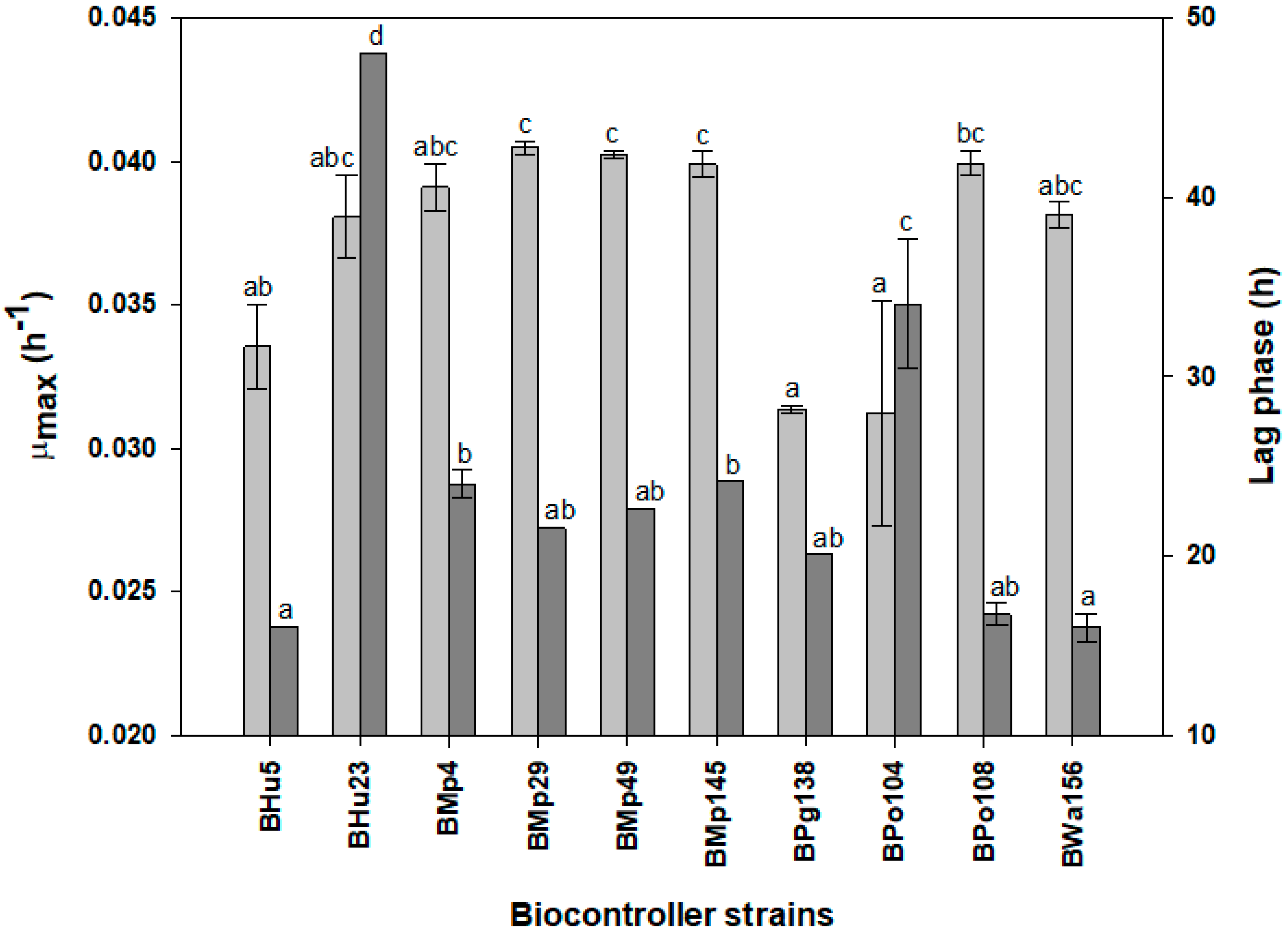
3.2.1. Kinetic Parameters
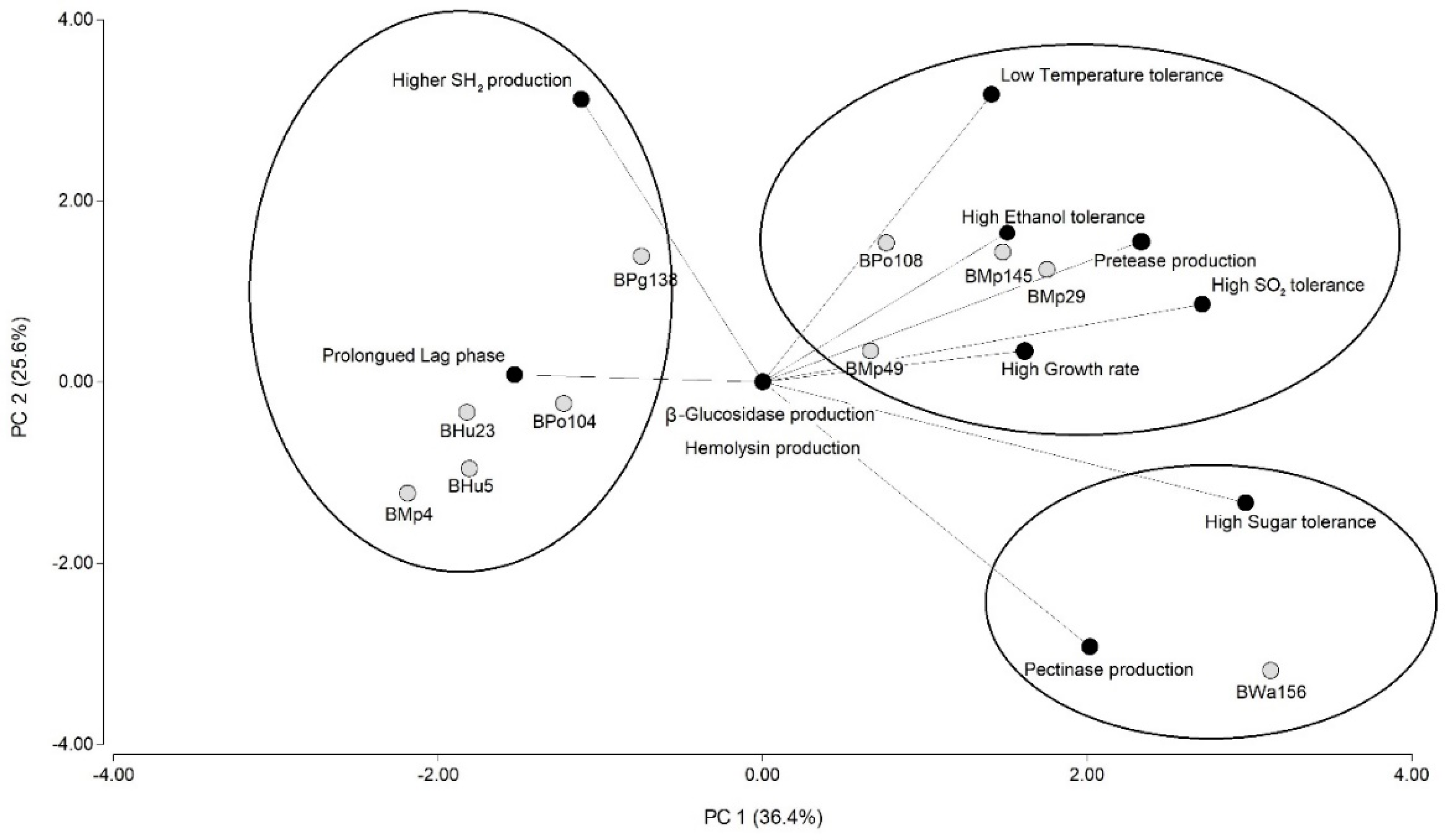
3.2.2. Enological Characterization of Yeasts: Tolerance to Molecular SO2, Ethanol, High Reducing Sugar Concentrations and Low Temperature
3.2.3. Enzyme and H2S Production
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ciani, M.; Capece, A.; Comitini, F.; Canonico, L.; Siesto, G.; Romano, P. Yeast interactions in inoculated wine fermentation. Front. Microbiol. 2016, 7, 555. [Google Scholar] [CrossRef] [PubMed]
- Mehlomakulu, N.N.; Setati, M.E.; Divol, B. Characterization of novel killer toxins secreted by wine-related non-Saccharomyces yeasts and their action on Brettanomyces spp. Int. J. Food Microbiol. 2014, 188, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Oelofse, A.; Lonvaud-Funel, A.; Du Toit, M. Molecular identification of Brettanomyces bruxellensis strains isolated from red wines and volatile phenol production. Food Microbiol. 2009, 26, 377–385. [Google Scholar] [CrossRef] [PubMed]
- Rojo, M.C.; López, F.A.; Lerena, M.C.; Mercado, L.; Torres, A.; Combina, M. Evaluation of different chemical preservatives to control Zygosaccharomyces rouxii growth in high sugar culture media. Food Control. 2015, 50, 349–355. [Google Scholar] [CrossRef]
- Stratford, M.; Steels, H.; Nebe-von-Caron, G.; Avery, S.V.; Novodvorska, M.; Archer, D.B. Population heterogeneity and dynamics in starter culture and lag phase adaptation of the spoilage yeast Zygosaccharomyces bailii to weak acid preservatives. Int. J. Food Microbiol. 2014, 181, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Fleet, G.H. Yeast Spoilage of Food and Beverages. In The Yeasts: A Taxonomic Study; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier Science: London, UK; Burlington, MA, USA, 2011; pp. 53–63. [Google Scholar]
- Escott, C.; del Fresno, J.M.; Loira, I.; Morata, A.; Suárez-Lepe, J.A. Zygosaccharomyces rouxii: Control Strategies and Applications in Food and Winemaking. Fermentation 2018, 4, 69. [Google Scholar] [CrossRef]
- Cejudo-Bastante, M.J.; Sonni, F.; Chinnici, F.; Versari, A.; Perez-Coello, M.S.; Riponi, C. Fermentation of sulphite-free white musts with added lysozyme and oenological tannins: Nitrogen consumption and biogenic amines composition of final wines. LWT-Food Sci. Technol. 2010, 43, 1501–1507. [Google Scholar] [CrossRef]
- Ferrer-Gallego, R.; Puxeu, M.; Martín, L.; Nart, E.; Hidalgo, C.; Andorrà, I. Microbiological, Physical, and Chemical Procedures to Elaborate High-Quality SO2-Free Wines. In Grapes and Wines-Advances in Production, Processing, Analysis and Valorization; IntechOpen: Madrid, Spain, 2017; pp. 171–193. [Google Scholar]
- Swinnen, I.A.M.; Bernaerts, K.; Dens, E.J.; Geeraerd, A.H.; Van Impe, J.F. Predictive modelling of the microbial lag phase: A review. Int. J. Food Microbiol. 2004, 94, 137–159. [Google Scholar] [CrossRef]
- Comitini, F.; De Ingeniis, J.; Pepe, L.; Mannazzu, I.; Ciani, M. Pichia anomala and Kluyveromyces wickerhamii killer toxins as new tools against Dekkera/Brettanomyces spoilage yeasts. FEMS Microbiol. Lett. 2004, 238, 235–240. [Google Scholar] [CrossRef]
- Fleet, G.H. Wine yeasts for the future. FEMS Yeast Res. 2008, 8, 979–995. [Google Scholar] [CrossRef]
- Maturano, Y.P.; Mestre, M.V.; Kuchen, B.; Toro, M.E.; Mercado, L.A.; Vazquez, F.; Combina, M. Optimization of fermentation-relevant factors: A strategy to reduce ethanol in red wine by sequential culture of native yeasts. Int. J. Food Microbiol. 2019, 289, 40–48. [Google Scholar] [CrossRef] [PubMed]
- Oro, L.; Ciani, M.; Comitini, F. Antimicrobial activity of Metschnikowia pulcherrima on wine yeasts. J. Appl. Microbiol. 2014, 116, 1209–1217. [Google Scholar] [CrossRef] [PubMed]
- Drumonde-Neves, J.; Franco-Duarte, R.; Lima, T.; Schuller, D.; Pais, C. Yeast biodiversity in vineyard environments is increased by human intervention. PLoS ONE 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Pirt, S.J. Parameters of growth and analysis of growth data. In Principles of Microbe and Cell Cultivation; Blackwell Scientific Publications: Hoboken, NJ, USA, 1975; pp. 4–14. [Google Scholar]
- Duarte, A. Introducción a la Ingeniería Bioquímica; Universidad Nacional de Colombia: Bogotá, Colombia, 1998; p. 198. [Google Scholar]
- Stanbury, P.F.; Whitaker, A.; Hall, S.J. Microbial growth kinetics. In Principles of Fermentation Technology; Elsevier Science: Kent, UK, 2014; pp. 21–74. [Google Scholar]
- Albertin, W.; Zimmer, A.; Miot-Sertier, C.; Bernard, M.; Coulon, J.; Moine, V.; Colonna-Ceccaldi, B.; Bely, M.; Marullo, P.; Masneuf-Pomarede, I. Combined effect of the Saccharomyces cerevisiae lag phase and the non-Saccharomyces consortium to enhance wine fruitiness and complexity. Appl. Microbiol. Biotechnol. 2017, 101, 7603–7620. [Google Scholar] [CrossRef] [PubMed]
- Comitini, F.; Gobbi, M.; Domizio, P.; Romani, C.; Lencioni, L.; Mannazzu, I.; Ciani, M. Selected non-Saccharomyces wine yeasts in controlled multistarter fermentations with Saccharomyces cerevisiae. Food Microbiol. 2011, 28, 873–882. [Google Scholar] [PubMed]
- Domizio, P.; Romani, C.; Lencioni, L.; Comitini, F.; Gobbi, M.; Mannazzu, I.; Ciani, M. Outlining a future for non-Saccharomyces yeasts: Selection of putative spoilage wine strains to be used in association with Saccharomyces cerevisiae for grape juice fermentation. Int. J. Food Microbiol. 2011, 147, 170–180. [Google Scholar] [CrossRef] [PubMed]
- Maturano, Y.P.; Rodriguez, L.A.; Toro, M.E.; Nally, M.C.; Vallejo, M.; Castellanos de Figueroa, L.I.; Combina, M.; Vazquez, F. Multi-enzyme production by pure and mixed cultures of Saccharomyces and non-Saccharomyces yeasts during wine fermentation. Int. J. Food Microbiol. 2012, 155, 43–50. [Google Scholar] [CrossRef]
- Mestre, V.; Maturano, Y.P.; Combina, M.; Mercado, L.A.; Toro, M.E.; Vazquez, F. Selection of non-Saccharomyces yeasts to be used in grape musts with high alcoholic potential: A strategy to obtain wines with reduced ethanol content. FEMS Yeast Res. 2017, 17. [Google Scholar] [CrossRef]
- Sturm, M.E.; Assof, M.; Fanzone, M.; Martinez, C.; Ganga, M.A.; Jofré, V.; Ramirez, M.L.; Combina, M. Relation between coumarate decarboxylase and vinylphenol reductase activity with regard to the production of volatile phenols by native Dekkera bruxellensis strains under ‘wine-like’ conditions. Int. J. Food Microbiol. 2015, 206, 51–55. [Google Scholar] [CrossRef]
- Esteve-Zarzoso, B.; Belloch, C.; Uruburu, F.; Querol, A. Identification of yeasts by RFLP analysis of the 5.8 S rRNA gene and the two ribosomal internal transcribed spacers. Int. J. Syst. Evol. Microbiol. 1999, 49, 329–337. [Google Scholar] [CrossRef]
- Santos, A.; San Mauro, M.; Bravo, E.; Marquina, D. PMKT2, a new killer toxin from Pichia membranifaciens, and its promising biotechnological properties for control of the spoilage yeast Brettanomyces bruxellensis. Microbiology 2009, 155, 624–634. [Google Scholar] [CrossRef] [PubMed]
- Vazquez, F.; Figueroa, L.; Toro, M. Enological characteristics of yeasts. In Methods in Biotechnology: Food Microbiology Protocols; Spencer, J.F.T., Spencer, A.L., Eds.; Humana Press: Totowa, NJ, USA, 2001; Volume 14, pp. 297–306. [Google Scholar]
- Monod, J. Recherches ser la Croissance des Cultures Bactériennes, 2nd ed.; Hermann: Paris, France, 1942. [Google Scholar]
- Lodge, R.M.; Hinshelwood, C.N. Physicochemical aspects of bacterial growth. Part IX. The lag phase of Bac. lactis aerogenes. J. Chem. Soc. 1943, 51, 213–219. [Google Scholar] [CrossRef]
- Sudraud, P.; Chauvet, S. Activite antilevure de l’anhydride sulfureux moleculaire. Conn. Vigne Vin. 1985, 19, 31–40. [Google Scholar] [CrossRef]
- Ribéreau-Gayon, J.; Peynaud, E.; Ribéreau-Gayon, P.; Sudraud, P. Sciences et Techniques du Vin. In Clarification et Stabilisation. Matériels et Installation; Dunod: Paris, France, 1977; Volume 4. [Google Scholar]
- Strauss, M.L.A.; Jolly, N.P.; Lambrechts, M.G.; Van Rensburg, P. Screening for the production of extracellular hydrolytic enzymes by non-Saccharomyces wine yeasts. J. Appl. Microbiol. 2001, 91, 182–190. [Google Scholar] [CrossRef]
- Fernandez-Salomäo, T.; Amorim, A.C.; Chaves-Alves, V.; Coelho, J.; Silva, D.; Araujo, E. Isolation of pectinase hyperproducing mutants of Penicillium expansum. Rev. Microbiol. 1996, 27, 15–18. [Google Scholar]
- Manns, J.M.; Mosser, D.M.; Buckley, H.R. Production of a hemolytic factor by Candida albicans. Infect. Immun. 1994, 62, 5154–5156. [Google Scholar]
- Menezes, A.G.T.; Ramos, C.L.; Cenzi, G.; Melo, D.S.; Dias, D.R.; Schwan, R.F. Probiotic Potential, Antioxidant Activity, and Phytase Production of Indigenous Yeasts Isolated from Indigenous Fermented Foods. Probiotics Antimicrob. Prot. 2019. [Google Scholar] [CrossRef]
- Iglewsky, B.H. Medical Microbiology, 4th ed.; Chapter 27: Pseudomonas; Baron, S., Ed.; University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996. [Google Scholar]
- Luo, G.; Samaranayake, L.P.; Yau, J.Y. Candida species exhibit differential in vitro hemolytic activities. J. Clin. Microbiol. 2001, 39, 2971–2974. [Google Scholar] [CrossRef]
- Massera, A.; Assof, M.; Sturm, M.E.; Sari, S.; Jofré, V.; Cordero-Otero, R.; Combina, M. Selection of indigenous Saccharomyces cerevisiae strains to ferment red musts at low temperature. Ann. Microbiol. 2012, 62, 367–380. [Google Scholar] [CrossRef]
- Liu, G.L.; Chi, Z.; Wang, G.Y.; Wang, Z.P.; Li, Y.; Chi, Z.M. Yeast killer toxins, molecular mechanisms of their action and their applications. Crit. Rev. Biotechnol. 2013, 35, 222–234. [Google Scholar] [CrossRef]
- De Ingeniis, J.; Raffaelli, N.; Ciani, M.; Mannazzu, I. Pichia anomala DBVPG 3003 secretes a ubiquitin-like protein that has antimicrobial activity. Appl. Environ. Microbiol. 2009, 75, 1129–1134. [Google Scholar] [CrossRef] [PubMed]
- Starmer, W.T.; Lachance, M.A. Yeast Ecology. In The Yeasts: A Taxonomic Study; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier Science: London, UK; Burlington, MA, USA, 2011; pp. 65–83. [Google Scholar]
- De Vries, S.; von Dahlen, J.K.; Schnake, A.; Ginschel, S.; Schulz, B.; Rose, L.E. Broad-spectrum inhibition of Phytophthora infestans by fungal endophytes. FEMS Microbiol. Ecol. 2018, 94, fiy037. [Google Scholar] [CrossRef] [PubMed]
- Odds, F.C.; Brown, A.J.P.; Gow, N.A.R. Antifungal agents: Mechanisms of action. Trends Microbiol. 2003, 11, 272–279. [Google Scholar] [CrossRef]
- Ginovart, M.; Prats, C.; Portell, X.; Silbert, M. Exploring the lag phase and growth initiation of a yeast culture by means of an individual-based model. Food Microbial. 2011, 28, 810–817. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Combina, M.; Elía, A.; Mercado, L.; Catania, C.; Ganga, A.; Martinez, C. Dynamics of indigenous yeast populations during spontaneous fermentation of wines from Mendoza, Argentina. Int. J. Food Microbiol. 2005, 99, 237–243. [Google Scholar] [CrossRef] [PubMed]
- Rivero, D.; Berná, L.; Stefanini, I.; Baruffini, E.; Bergerat, A.; Csikász-Nagy, A.; De Filippo, C.; Cavalieri, D. Hsp12p and PAU genes are involved in ecological interactions between natural yeast strains. Environ. Microbiol. 2015, 17, 3069–3081. [Google Scholar] [CrossRef] [PubMed]
- Nally, M.C. Función del carácter killer en la competencia por el dominio del substrato: Caracterización de mutantes de cepas de levadura Saccharomyces cerevisiae. In Tesis de Licenciatura en Biología; Universidad Nacional de San Juan: San Juan, Argentina, 2005. [Google Scholar]
- Auling, G.; Bellgardt, K.H.; Diekmann, M.; Thoma, M. Dynamics of the growth in batch and continuous culture of Saccharomyces cerevisiae during a shift from aerobiosis to anaerobiosis and reverse. Appl. Microbiol. Biotechnol. 1984, 19, 353–357. [Google Scholar] [CrossRef]
- Verduyn, C.; Postma, E.; Scheffers, W.A.; van Dijken, J.P. Physiology of Saccharomyces cerevisiae in anaerobic glucoselimited chemostat cultures. J. Gen. Microbiol. 1990, 136, 395–403. [Google Scholar] [CrossRef] [PubMed]
- Jain, V.K. Relationship between energy metabolism and growth. I. Glucose dependence of the exponential growth rate of Saccharomyces cerevisiae. Arch. Microbiol. 1970, 72, 252–259. [Google Scholar]
- Andorrà, I.; Martín, L.; Nart, E.; Puxeu, M.; Hidalgo, C.; Ferrer-Gallego, R. Effect of grape juice composition and nutrient supplementation on the production of sulfur dioxide and carboxylic compounds by Saccharomyces cerevisiae. Aust. J. Grape Wine Res. 2018, 24, 260–266. [Google Scholar] [CrossRef]
- Milheiro, J.; Filipe-Ribeiro, L.; Vilela, A.; Cosme, F.; Nunes, F.M. 4-Ethylphenol, 4-ethylguaiacol and 4-ethylcatechol in red wines: Microbial formation, prevention, remediation and overview of analytical approaches. Crit. Rev. Food Sci. Nutr. 2017, 59, 1367–1391. [Google Scholar] [CrossRef] [PubMed]
- Jolly, N.P.; Augustyn, O.P.H.; Pretorius, I.S. The role and use of non-Saccharomyces yeasts in wine production. S. Afr. J. Enol. Vitic. 2006, 27, 15–29. [Google Scholar] [CrossRef]
- Pretorius, I.S. Tailoring wine yeast for the new millennium: Novel approaches to the ancient art of winemaking. Yeast 2000, 16, 675–729. [Google Scholar] [CrossRef]
- Wang, H.; Hu, Z.; Long, F.; Guo, C.; Niu, C.; Yuan, Y.; Yue, T. Combined effect of sugar content and pH on the growth of a wild strain of Zygosaccharomyces rouxii and time for spoilage in concentrated apple juice. Food Control. 2016, 59, 298–305. [Google Scholar] [CrossRef]
- Molina, A.M.; Swiegers, J.H.; Varela, C.; Pretorius, I.S.; Agosin, E. Influence of wine fermentation temperature on the synthesis of yeast-derived volatile aroma compounds. Appl. Microbiol. Biotechnol. 2007, 77, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Charoenchai, C.; Fleet, G.H.; Henschke, P.A.; Todd, B.E.N. Screening of non-Saccharomyces wine yeasts for the presence of extracellular hydrolytic enzymes. Aust. J. Grape Wine Res. 1997, 3, 2–8. [Google Scholar] [CrossRef]
- Bautista-Ortín, A.B.; Fernández-Fernández, J.I.; López-Roca, J.M.; Gómez-Plaza, E. The effects of enological practices in anthocyanins, phenolic compounds and wine colour and their dependence on grape characteristics. J. Food Compos. Anal. 2007, 20, 546–552. [Google Scholar] [CrossRef]
- Blanco, P.; Sieiro, C.; Villa, T.G. Production of pectic enzymes in yeasts. FEMS Microbiol. Lett. 1999, 175, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Apel, A.R.; Ouellet, M.; Szmidt-Middleton, H.; Keasling, J.D.; Mukhopadhyay, A. Evolved hexose transporter enhances xylose uptake and glucose/xylose co-utilization in Saccharomyces cerevisiae. Sci. Rep. 2016, 6, 19512. [Google Scholar] [CrossRef]
- Masoud, W.; Jespersen, L. Pectin degrading enzymes in yeasts involved in fermentation of Coffea arabica in East Africa. Int. J. Food Microbiol. 2006, 110, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Martos, M.A.; Butiuk, A.P.; Rojas, N.L.; Hours, R.A. Cultivo por lote de Wickerhamomyces anomalus en un biorreactor a escala laboratorio para la producción de una poligalacturonasa. Rev. Colomb. Biotecnol. 2014, 16, 68–73. [Google Scholar] [CrossRef]
- Schmidt, S.; Zimmermann, S.Y.; Tramsen, L.; Koehl, U.; Lehrnbecher, T. Natural killer cells and antifungal host response. Clin. Vaccine Immunol. 2013, 20, 452–458. [Google Scholar] [CrossRef] [PubMed][Green Version]
| Species | N° of Isolates | Strain Nomenclature |
|---|---|---|
| Candida apis | 1 | BCa80 |
| Candida cantarelli | 1 | BCca78 |
| Candida catenulata | 1 | BCct79 |
| Candida diversa | 1 | BCd75 |
| Candida famata | 4 | BCf84 |
| Candida intermedia | 1 | BCi85 |
| Candida membranifaciens | 3 | BCm69, BCm70, BCm71 |
| Candida pararugosa | 1 | BCp73 |
| Candida rugosa | 1 | BCr81 |
| Candida sake | 6 | BCsa74, BCsa82, BCsa83, BCsa86, BCsa88, BCsa95 |
| Candida steatolytica | 1 | BCse76 |
| Candida stellata | 1 | BCst68 |
| Clavispora lusitaniae | 1 | BCl157 |
| Cryptococcus albidus | 1 | BCra158 |
| Debaryomyces hansenii | 5 | BDb150, BDb152, BDb153, BDb154, BDb155 |
| Debaryomycesvanrijiae | 1 | BDv151 |
| Hanseniaspora guilliermondii | 6 | BHg42, BHg44, BHg45, BHg46, BHg47, BHg48 |
| Hanseniaspora osmophila | 1 | BHo51 |
| Hanseniaspora uvarum | 27 | BHu1, BHu2, BHu3, BHu5, BHu8, BHu9, BHu10, BHu11, BHu12, BHu13, BHu17, BHu18, BHu19, BHu20, BHu21, BHu23, BHu24, BHu26, BHu27, BHu28, BHu30, BHu31, BHu32, BHu38, BHu40, BHu41 |
| Hanseniaspora vineae | 2 | BHv43, BHv50 |
| Issatchenkia orientalis | 1 | BIo160 |
| Metschnikowia pulcherrima | 6 | BMp4, BMp29, BMp49, BMp151, BMp144, BMp145 |
| Pichia fabianii | 1 | BPf127 |
| Pichia guilliermondii | 1 | BPg138 |
| Pichia kluyveri | 3 | BPkl130, BPkl131, BPkl133 |
| Pichia kudriavzevii | 5 | BPku128, BPku129, BPku132, BPku134, BPku135 |
| Pichia manshurica | 1 | BPm125 |
| Pichia membranifaciens | 1 | BPm136 |
| Pichia occidentalis | 21 | BPo96, BPo97, BPo98, BPo100, BPo101, BPo102, BPo104, BPo108, BPo110, BPo111, BPo112, BPo113, BPo114, BPo115, BPo116, BPo117, BPo120, BPo121, BPo122, BPo123, BPo124 |
| Starmerella bacillaris | 12 | BSb52, BSb53, BSb54, BSb55, BSb56, BSb57, BSb58, BSb59, BSb62, BSb63, BSb66, BSb67 |
| Torulaspora delbrueckii | 3 | BTd147, BTd148, BTd149 |
| Wickerhamomyces anomalus | 1 | BWa156 |
| TOTAL | 122 |
| Yeast specie | Isolate | BBb1 | BBb11 | BBb20 | BBb29 | BZr4 | BZr6 | BZr9 | BZr10 | BSc114 | Intraspecific | Intraspecific | Total |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| B. bruxellensis | Z. rouxii | inhibition | |||||||||||
| inhibition | inhibition | ||||||||||||
| Candida intermedia | BCi85 | − | − | + | − | + | − | − | − | − | 0.25 | 0.25 | 0.25 |
| Candida membranifaciens | BCm70 | − | − | − | + | − | − | − | − | − | 0.25 | 0 | 0.13 |
| Candida sake | BCs88 | − | − | − | − | + | + | − | − | − | 0 | 0.5 | 0.25 |
| Candida sake | BCs95 | − | − | + | − | + | + | − | − | − | 0.25 | 0.5 | 0.38 |
| Hanseniaspora uvarum | BHu5 | + | − | + | + | + | + | − | − | − | 0.75 | 0.5 | 0.63 |
| Hanseniaspora uvarum | BHu18 | − | − | − | − | + | + | + | − | − | 0 | 0.75 | 0.38 |
| Hanseniaspora uvarum | BHu23 | − | − | − | − | + | + | + | + | − | 0 | 1 | 0.5 |
| Hanseniaspora uvarum | BHu27 | + | + | − | − | − | − | + | − | − | 0.5 | 0.25 | 0.38 |
| Hanseniaspora uvarum | BHu31 | − | − | − | + | − | + | − | + | − | 0.25 | 0.5 | 0.38 |
| Hanseniaspora uvarum | BHu32 | − | − | + | − | − | + | − | + | − | 0.25 | 0.5 | 0.38 |
| Issatchenkia orientalis | BIo160 | − | − | − | − | − | + | − | − | − | 0 | 0.25 | 0.13 |
| Metschnikowia pulcherrima | BMp4 | − | + | + | + | + | + | + | − | − | 0.75 | 0.75 | 0.75 |
| Metschnikowia pulcherrima | BMp29 | + | − | + | − | − | + | + | − | − | 0.5 | 0.5 | 0.5 |
| Metschnikowia pulcherrima | BMp49 | + | + | + | + | + | + | + | − | − | 1 | 0.75 | 0.88 |
| Metschnikowia pulcherrima | BMp145 | − | − | + | − | + | + | + | + | − | 0.25 | 1 | 0.63 |
| Metschnikowia pulcherrima | BMp151 | − | − | − | − | + | − | − | − | − | 0 | 0.25 | 0.13 |
| Pichia occidentalis | BPo104 | − | − | + | − | + | + | + | − | − | 0.25 | 0.75 | 0.5 |
| Pichia occidentalis | BPo108 | − | − | + | − | + | + | + | + | − | 0.25 | 1 | 0.63 |
| Pichia occidentalis | BPo120 | − | − | − | − | − | − | − | + | − | 0 | 0.25 | 0.13 |
| Pichia guilliermondii | BPg138 | + | + | + | + | + | − | − | − | − | 1 | 0.25 | 0.63 |
| Starmerella bacillaris | BSb57 | − | − | + | − | + | + | − | − | − | 0.25 | 0.5 | 0.38 |
| Starmerella bacillaris | BSb58 | − | − | + | − | − | − | − | − | − | 0.25 | 0 | 0.13 |
| Wickerhamomyces anomalus | BWa156 | − | − | + | + | + | + | − | − | − | 0.5 | 0.5 | 0.5 |
| Molecular SO2 (ppm) | BHu5 | BHu23 | BMp4 | BMp29 | BMp49 | BMp145 | BPg138 | BPo104 | BPo108 | BWa156 | BSc114 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 (Control) | + | + | + | + | + | + | + | + | + | + | + |
| 0.1 | + | + | − | + | + | + | + | + | + | + | + |
| 0.15 | + | + | − | + | + | + | + | + | + | + | + |
| 0.2 | + | + | − | + | + | + | + | + | + | + | + |
| 0.3 | + | + | − | + | + | + | + | + | + | + | + |
| 0.4 | − | − | − | + | + | + | + | + | + | + | + |
| Ethanol (% v/v) | |||||||||||
| 0 (Control) | + | + | + | + | + | + | + | + | + | + | + |
| 8 | + | + | + | + | + | + | + | + | + | + | + |
| 10 | + | + | + | + | + | + | + | − | + | + | + |
| 12 | − | − | − | + | + | − | − | − | − | − | + |
| 14 | − | − | − | − | − | − | − | − | − | − | + |
| High Sugar concentration (°Brix) | |||||||||||
| 21 (Control) | + | + | + | + | + | + | + | + | + | + | + |
| 30 | − | − | − | + | − | + | − | − | − | + | + |
| Low Temperature (°C) | |||||||||||
| 25 (Control) | + | + | + | + | + | + | + | + | + | + | + |
| 15 | − | + | + | + | + | + | + | + | + | + | + |
| Isolate | Positive Traits | Negative Trait | ||
|---|---|---|---|---|
| Protease | Pectinase | β-Glucosidase | H2S production | |
| BHu5 | + | − | − | 3 |
| BHu23 | + | − | − | 3 |
| BMp4 | − | − | − | 3 |
| BMp29 | + | − | − | 3 |
| BMp49 | + | − | − | 3 |
| BMp145 | + | − | − | 3 |
| BPg138 | + | − | − | 4 |
| BPo104 | + | − | − | 3 |
| BPo108 | + | − | − | 4 |
| BWa156 | + | + | − | 2 |
| BSc114 | − | − | − | 2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kuchen, B.; Maturano, Y.P.; Mestre, M.V.; Combina, M.; Toro, M.E.; Vazquez, F. Selection of Native Non-Saccharomyces Yeasts with Biocontrol Activity against Spoilage Yeasts in Order to Produce Healthy Regional Wines. Fermentation 2019, 5, 60. https://doi.org/10.3390/fermentation5030060
Kuchen B, Maturano YP, Mestre MV, Combina M, Toro ME, Vazquez F. Selection of Native Non-Saccharomyces Yeasts with Biocontrol Activity against Spoilage Yeasts in Order to Produce Healthy Regional Wines. Fermentation. 2019; 5(3):60. https://doi.org/10.3390/fermentation5030060
Chicago/Turabian StyleKuchen, Benjamín, Yolanda Paola Maturano, María Victoria Mestre, Mariana Combina, María Eugenia Toro, and Fabio Vazquez. 2019. "Selection of Native Non-Saccharomyces Yeasts with Biocontrol Activity against Spoilage Yeasts in Order to Produce Healthy Regional Wines" Fermentation 5, no. 3: 60. https://doi.org/10.3390/fermentation5030060
APA StyleKuchen, B., Maturano, Y. P., Mestre, M. V., Combina, M., Toro, M. E., & Vazquez, F. (2019). Selection of Native Non-Saccharomyces Yeasts with Biocontrol Activity against Spoilage Yeasts in Order to Produce Healthy Regional Wines. Fermentation, 5(3), 60. https://doi.org/10.3390/fermentation5030060