Capnophilic Lactic Fermentation from Thermotoga neapolitana: A Resourceful Pathway to Obtain Almost Enantiopure L-lactic Acid
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Culture Conditions
2.3. Chemical Analysis
2.4. Derivatization of Lactic Acid to the O-Trifluoroacetylated-(−)-Menthyl Ester
2.5. GC/MS Analysis
2.6. Bioinformatics
3. Results
3.1. Fermentation of T. Neapolitana Under CLF Conditions
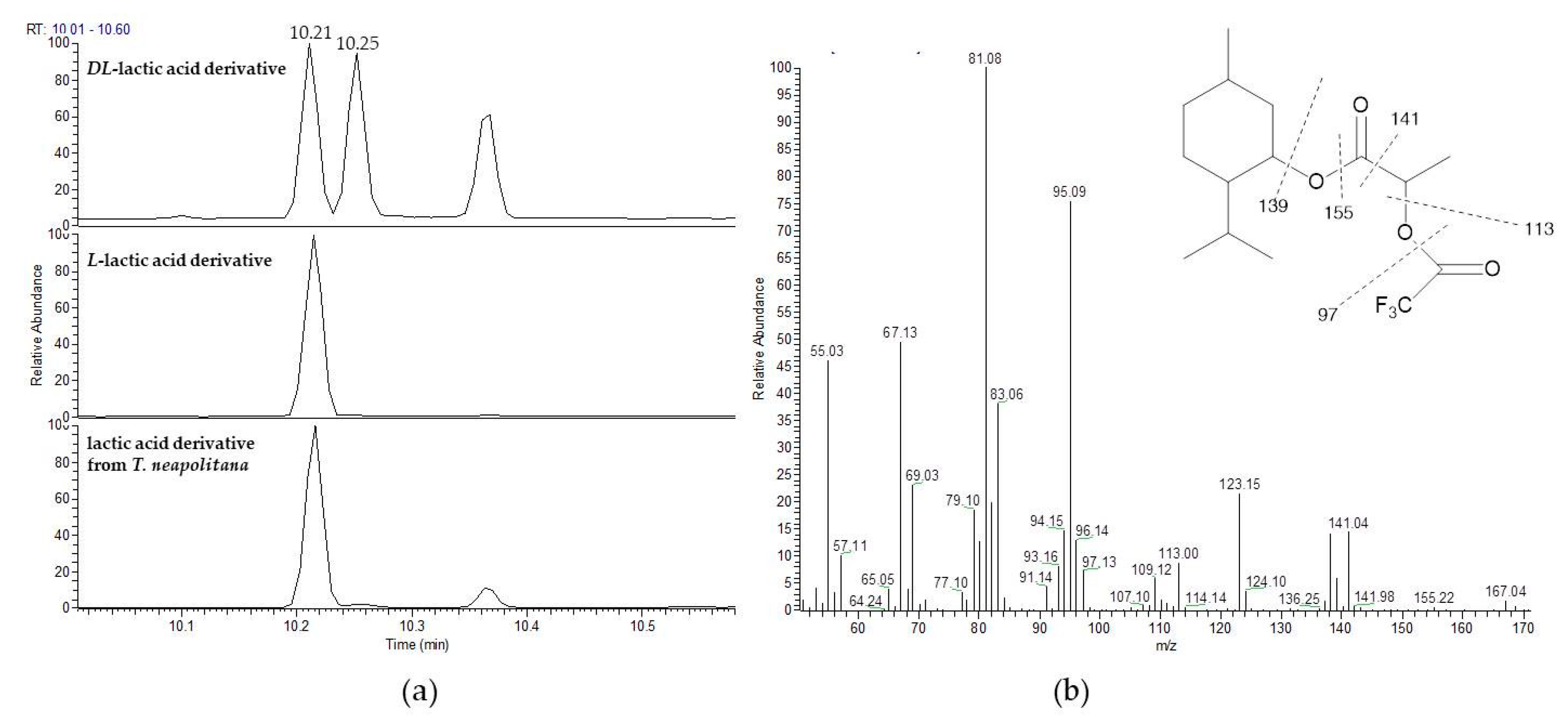
3.2. Stereochemistry Determination
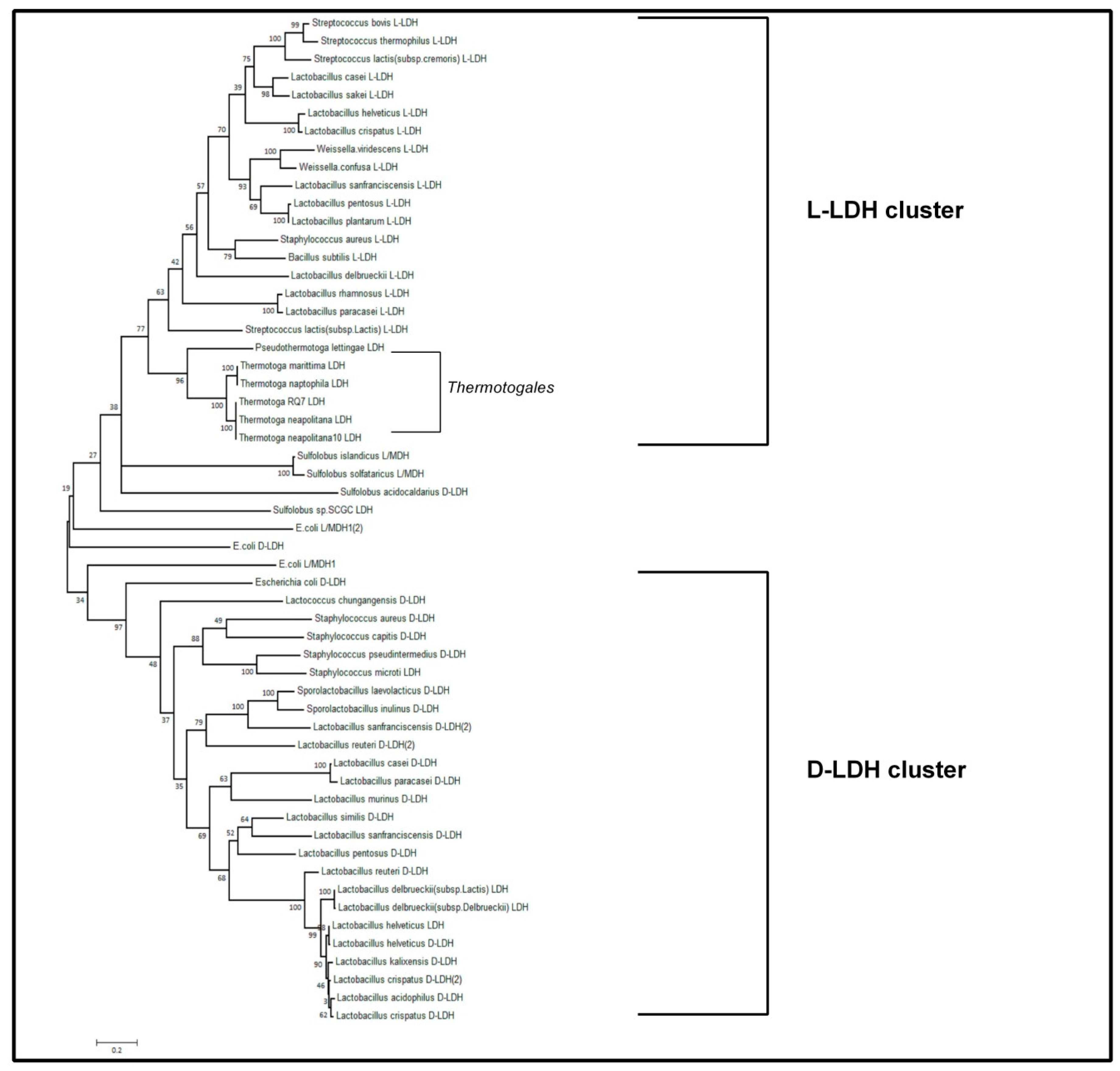
3.3. Bioinformatic Approach to Characterize the Lactate Dehydrogenase of T. Neapolitana (TnLDH)
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Juturu, V.; Wu, J.C. Microbial production of lactic acid: The latest development. Crit. Rev. Biotechnol. 2016, 36, 967–977. [Google Scholar] [CrossRef] [PubMed]
- Castillo Martinez, F.A.; Balciunas, E.M.; Salgado, J.M.; Domínguez González, J.M.; Converti, A.; de Souza Oliveira, R.P. Lactic acid properties, applications and production: A review. Trends Food Sci. Technol. 2013, 30, 70–83. [Google Scholar] [CrossRef]
- Garlotta, D. A litterature review of Poly(lactic acid). J. Polym. Environ. 2001, 9. [Google Scholar] [CrossRef]
- John, R.P.; Nampoothiri, K.M.; Pandey, A. Fermentative production of lactic acid from biomass: An overview on process developments and future perspectives. Appl. Microbiol. Biotechnol. 2007, 74, 524–534. [Google Scholar] [CrossRef]
- d’Ippolito, G.; Dipasquale, L.; Fontana, A. Recycling of Carbon Dioxide and Acetate as Lactic Acid by the Hydrogen-Producing Bacterium Thermotoga neapolitana. ChemSusChem 2014. [Google Scholar] [CrossRef] [PubMed]
- Dipasquale, L.; D’Ippolito, G.; Fontana, A. Capnophilic lactic fermentation and hydrogen synthesis by Thermotoga neapolitana: An unexpected deviation from the dark fermentation model. Int. J. Hydrog. Energy 2014. [Google Scholar] [CrossRef]
- d’Ippolito, G.; Dipasquale, L.; Vella, F.M.; Romano, I.; Gambacorta, A.; Cutignano, A.; Fontana, A. Hydrogen metabolism in the extreme thermophile Thermotoga neapolitana. Int. J. Hydrog. Energy 2010. [Google Scholar] [CrossRef]
- Pradhan, N.; Dipasquale, L.; d’Ippolito, G.; Panico, A.; Lens, P.N.L.; Esposito, G.; Fontana, A. Hydrogen production by the thermophilic bacterium Thermotoga neapolitana. Int. J. Mol. Sci. 2015, 16, 12578–12600. [Google Scholar] [CrossRef]
- Dipasquale, L.; Pradhan, N.; d’Ippolito, G.; Fontana, A. Potential of Hydrogen Fermentative Pathways in Marine Thermophilic Bacteria: Dark Fermentation and Capnophilic Lactic Fermentation in Thermotoga and Pseudothermotoga Species. Grand Challenges in Marine Biotechnology. In Grand Challenges in Biology and Biotechnology; Rampelotto, P., Trincone, A., Eds.; Springer: Cham, Switzerland, 2018; pp. 217–235. [Google Scholar]
- Pradhan, N.; Dipasquale, L.; d’Ippolito, G.; Fontana, A.; Panico, A.; Pirozzi, F.; Lens, P.N.L.; Esposito, G. Model development and experimental validation of capnophilic lactic fermentation and hydrogen synthesis by Thermotoga neapolitana. Water Res. 2016, 99, 225–234. [Google Scholar] [CrossRef]
- Luongo, V.; Palma, A.; Rene, E.; Fontana, A.; Pirozzi, F.; Esposito, G.; Lens, P.L. Lactic acid recovery from a simplified mimic of Thermotoga neapolitana fermentation broth using ion exchange resins in batch and fixed-bed reactors. Sep. Sci. Technol. 2018. [Google Scholar] [CrossRef]
- Pradhan, N.; Dipasquale, L.; d’Ippolito, G.; Panico, A.; Lens, P.N.L.; Esposito, G.; Fontana, A. Hydrogen and lactic acid synthesis by the wild-type and a laboratory strain of the hyperthermophilic bacterium Thermotoga neapolitana DSMZ 4359T under capnophilic lactic fermentation conditions. Int. J. Hydrog. Energy 2017. [Google Scholar] [CrossRef]
- Inoue, Y.; Shinka, T.; Ohse, M.; Ikawa, H.; Kuhara, T. Application of optical isomer analysis by diastereomer derivatization GC/MS to determine the condition of patients with short bowel syndrome. J. Chromatogr. B 2006, 838, 37–42. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef]
- Lim, L.T.; Auras, R.; Rubino, M. Processing technologies for poly(lactic acid). Prog. Polym. Sci. 2008, 33, 820–852. [Google Scholar] [CrossRef]
- Jardini, A.; Martinez, G.; Lasprilla, A.; Filho, A.L.; Rubens, M. Poly-lactic acid synthesis for application in biomedical devices—A review. Biotechnol. Adv. 2012, 30, 321–328. [Google Scholar]
- Niu, D.; Tian, K.; Prior, B.A.; Wang, M.; Wang, Z.; Lu, F.; Singh, S. Highly efficient L-lactate production using engineered Escherichia coli with dissimilar temperature optima for L-lactate formation and cell growth. Microb. Cell Fact. 2014, 13, 1–11. [Google Scholar] [CrossRef]
- Adams, M.J.; Buehner, M.; Chandrasekhar, K.; Ford, G.C.; Hackert, M.L.; Liljas, A.; Taylor, S.S. Structure-Function Relationships in Lactate Dehydrogenase (amino-acid sequence/crystallographic structure). PNAS 1973, 70, 1968–1972. [Google Scholar] [CrossRef]
- Garmyn, D.; Ferain, T.; Bernard, N.; Hols, P.; Delcour, J. Cloning, nucleotide sequence, and transcriptional analysis of the Pediococcus acidilactici L-(+)-lactate dehydrogenase gene. Appl. Environ. Microbiol. 1995, 61, 266–272. [Google Scholar]
- Taguchi, H.; Ohta, T. Unusual amino acid substitution in the anion-binding site of Lactobacillus plantarum non-allosteric L-lactate dehydrogenase. Eur. J. Biochem. 1992, 209, 993–998. [Google Scholar] [CrossRef]
- Yeswanth, S.; Nanda Kumar, Y.; Venkateswara Prasad, U.; Swarupa, V.; Koteswara rao, V.; Venkata Gurunadha Krishna Sarma, P. Cloning and characterization of l-lactate dehydrogenase gene of Staphylococcus aureus. Anaerobe 2013, 24, 43–48. [Google Scholar] [CrossRef]
| Growth Parameters | Yields (mol/mol glucose) | |||||
|---|---|---|---|---|---|---|
| OD 540 nm | Dry Biomass (g/L) | Glucose Consumption (g/L) | H2 | Acetic Acid | Lactic Acid | AL/AA |
| 0.79 ± 0.06 | 0.290 ± 0.03 | 3.5 ± 0.2 | 2.5 ± 0.3 | 1.14 ± 0.05 | 0.7 ± 0.02 | 0.58 |
| Peak Area | Relative AREA % | % e.e. | |
|---|---|---|---|
| L-Lactate | 44618648.45 | 97.6 | 95.2 |
| D-Lactate | 1088407.05 | 2.4 | - |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nuzzo, G.; Landi, S.; Esercizio, N.; Manzo, E.; Fontana, A.; d’Ippolito, G. Capnophilic Lactic Fermentation from Thermotoga neapolitana: A Resourceful Pathway to Obtain Almost Enantiopure L-lactic Acid. Fermentation 2019, 5, 34. https://doi.org/10.3390/fermentation5020034
Nuzzo G, Landi S, Esercizio N, Manzo E, Fontana A, d’Ippolito G. Capnophilic Lactic Fermentation from Thermotoga neapolitana: A Resourceful Pathway to Obtain Almost Enantiopure L-lactic Acid. Fermentation. 2019; 5(2):34. https://doi.org/10.3390/fermentation5020034
Chicago/Turabian StyleNuzzo, Genoveffa, Simone Landi, Nunzia Esercizio, Emiliano Manzo, Angelo Fontana, and Giuliana d’Ippolito. 2019. "Capnophilic Lactic Fermentation from Thermotoga neapolitana: A Resourceful Pathway to Obtain Almost Enantiopure L-lactic Acid" Fermentation 5, no. 2: 34. https://doi.org/10.3390/fermentation5020034
APA StyleNuzzo, G., Landi, S., Esercizio, N., Manzo, E., Fontana, A., & d’Ippolito, G. (2019). Capnophilic Lactic Fermentation from Thermotoga neapolitana: A Resourceful Pathway to Obtain Almost Enantiopure L-lactic Acid. Fermentation, 5(2), 34. https://doi.org/10.3390/fermentation5020034