Pathway-Guided Medium Engineering for Enhanced Prodiginine Production in Spartinivicinus ruber MCCC 1K03745T
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents
2.2. Bioinformatic Analysis
2.3. Fermentation Conditions
2.4. Sequential Design Strategy for Fermentation Optimization
2.4.1. Single-Factor Experiments for Preliminary Range Determination
2.4.2. Full Factorial Design and Steepest Ascent Experiments for Interaction Analysis and Region Exploration
2.4.3. Response Surface Methodology for Final Optimization
2.5. Extraction and Quantification of Prodiginine
2.6. Statistical Analysis
3. Results
3.1. Homology Analysis Reveals a Putative Prodiginine Biosynthetic Pattern in S. ruber
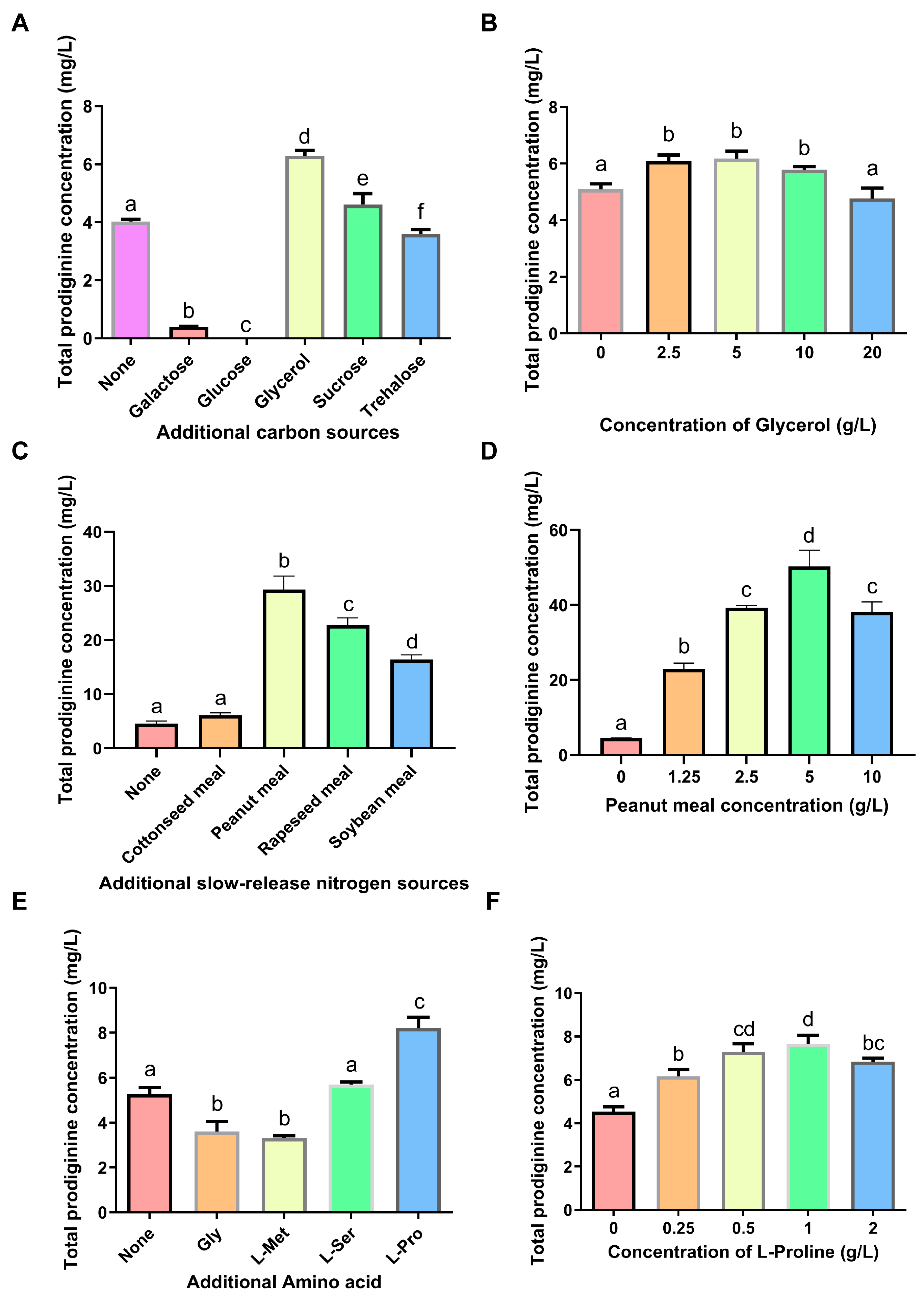
3.2. Effects of Single Factors on Prodiginine Production
3.3. Analysis of Main Effects and Interactions Using a Full Factorial Design
3.4. Approaching the Optimal Region via Steepest Ascent
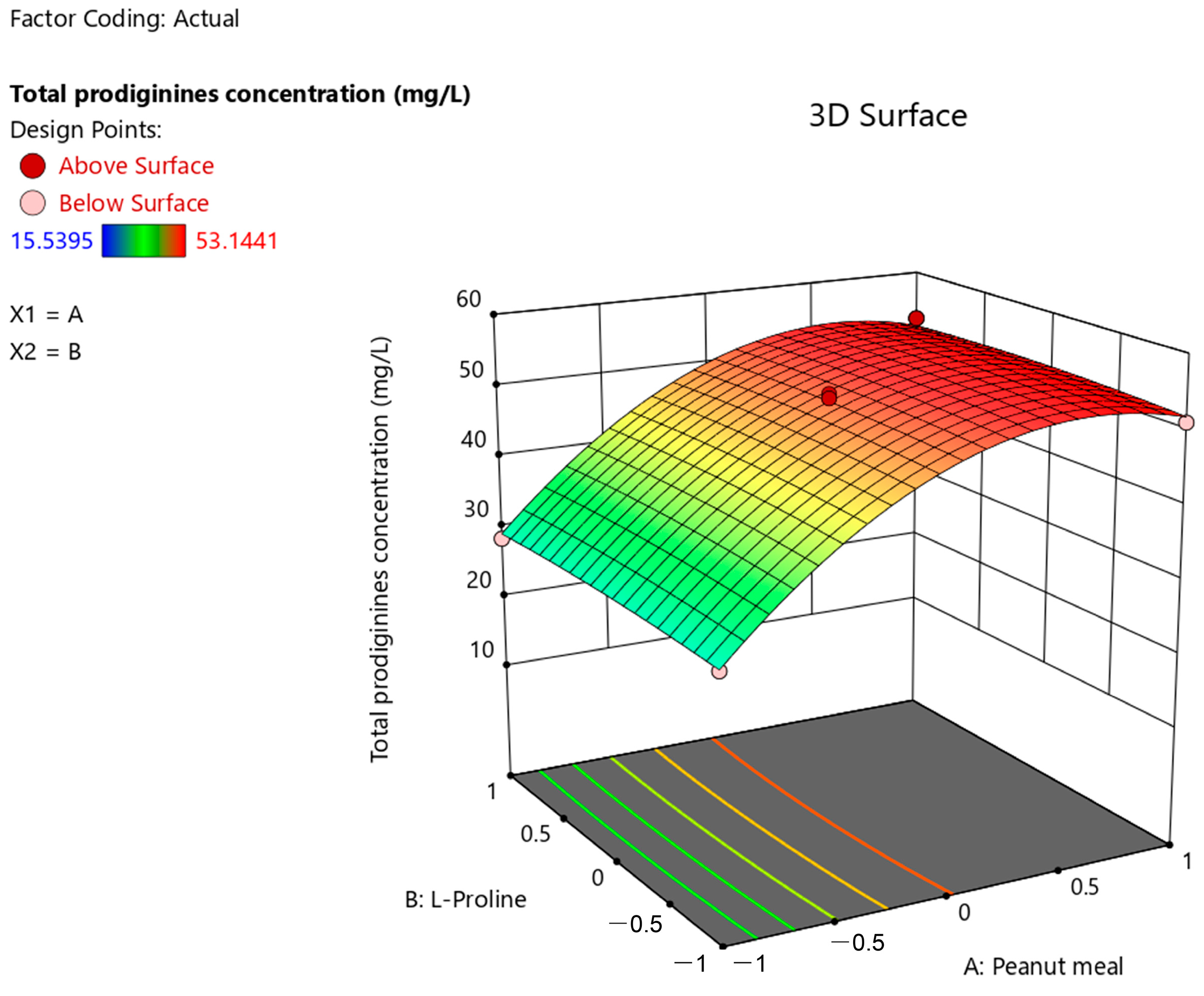
3.5. Optimization by Response Surface Methodology
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
Appendix A.1
| Enzymes in S. marcescens (UniProt Accession No.) | Homologs in S. ruber (NCBI Reference Sequence Accession No.) | Sequence Identities | Sequence Positives | Query Coverage | E Value |
|---|---|---|---|---|---|
| PigD (Q5W251) | prodigiosin MAP biosynthesis protein PigD (WP_163833139.1) | 51% | 68% | 90% | 0.0 |
| PigB (Q5W253) | NAD(P)/FAD-dependent oxidoreductase (WP_163833141.1) | 42% | 59% | 79% | 2e−140 |
| PigE (Q5W250) | prodigiosin MAP biosynthesis aminotransferase PigE (WP_163833138.1) | 59% | 76% | 100% | 0.0 |
Appendix A.2
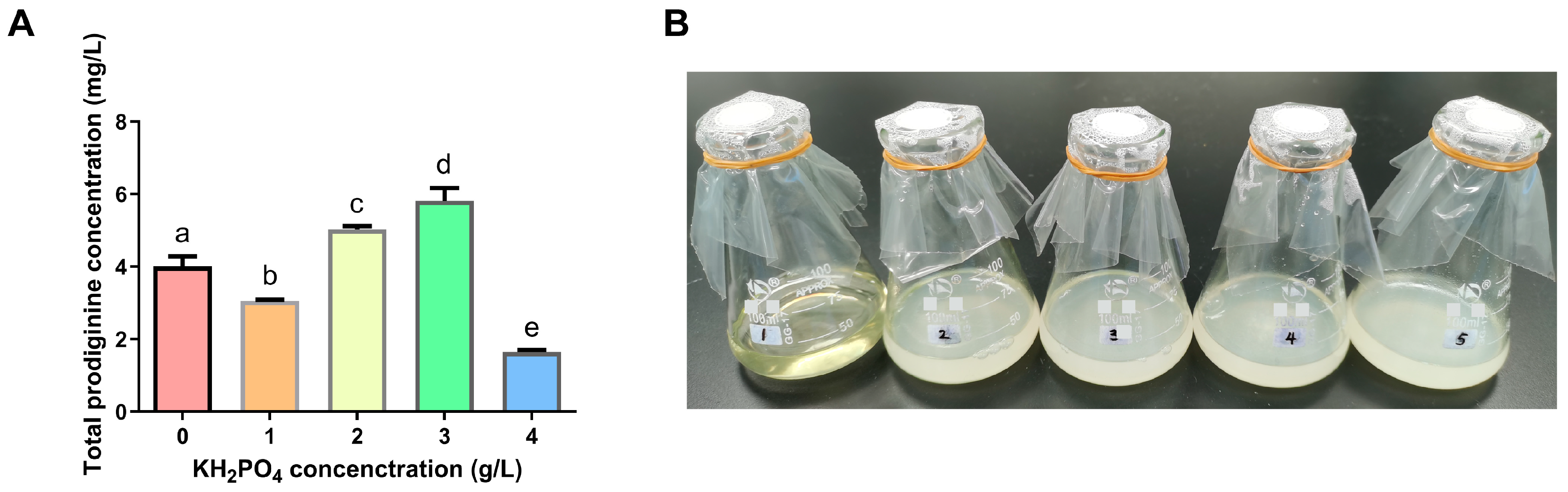
Appendix A.3
| Component | Amount (/L) | Grade | Supplier | Unit Price (CNY/g or mL) | Quotation Date | Cost (CNY/L) |
|---|---|---|---|---|---|---|
| Peptone | 11 g | - | Solarbio | 0.400 | March 2026 | 4.40 |
| Yeast extract | 1 g | - | Oxoid | 0.350 | March 2026 | 0.35 |
| Soybean oil | 5 mL | Food | Yihai Kerry | 0.018 | March 2026 | 0.09 |
| MgCl2·6H2O | 3 g | Analytical | Sinopharm | 0.068 | March 2026 | 0.20 |
| total | 5.04 |
| Component | Amount (/L) | Grade | Supplier | Unit Price (CNY/g or mL) | Quotation Date | Cost (CNY/L) |
|---|---|---|---|---|---|---|
| Peptone | 5 g | - | Solarbio | 0.400 | March 2026 | 2.00 |
| Yeast extract | 1 g | - | Oxoid | 0.350 | March 2026 | 0.35 |
| Peanut meal | 7.611 g | Fermentation | Hongrun Baoshun | 0.080 | March 2026 | 0.61 |
| L-Proline | 0.695 g | Analytical | Aladdin | 0.260 | March 2026 | 0.18 |
| total | 3.14 |
Appendix A.4

References
- Williamson, N.R.; Fineran, P.C.; Leeper, F.J.; Salmond, G.P. The biosynthesis and regulation of bacterial prodiginines. Nat. Rev. Microbiol. 2006, 4, 887–899. [Google Scholar] [CrossRef]
- Llagostera, E.; Soto-Cerrato, V.; Joshi, R.; Montaner, B.; Gimenez-Bonafe, P.; Perez-Tomas, R. High cytotoxic sensitivity of the human small cell lung doxorubicin-resistant carcinoma (GLC4/ADR) cell line to prodigiosin through apoptosis activation. Anticancer Drugs 2005, 16, 393–399. [Google Scholar] [CrossRef] [PubMed]
- Elahian, F.; Moghimi, B.; Dinmohammadi, F.; Ghamghami, M.; Hamidi, M.; Mirzaei, S.A. The anticancer agent prodigiosin is not a multidrug resistance protein substrate. DNA Cell Biol. 2013, 32, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Soto-Cerrato, V.; Llagostera, E.; Montaner, B.; Scheffer, G.L.; Perez-Tomas, R. Mitochondria-mediated apoptosis operating irrespective of multidrug resistance in breast cancer cells by the anticancer agent prodigiosin. Biochem. Pharmacol. 2004, 68, 1345–1352. [Google Scholar] [CrossRef]
- Lin, X.; Dong, L.; Miao, Q.; Huang, Z.; Wang, F. Cycloheptylprodigiosin from marine bacterium Spartinivicinus ruber MCCC 1K03745T induces a novel form of cell death characterized by Golgi disruption and enhanced secretion of cathepsin D in non-small cell lung cancer cell lines. Eur. J. Pharmacol. 2024, 974, 176608. [Google Scholar] [CrossRef]
- Huang, Z.; Dong, L.; Lai, Q.; Liu, J. Spartinivicinus ruber gen. nov., sp. nov., a Novel Marine Gammaproteobacterium Producing Heptylprodigiosin and Cycloheptylprodigiosin as Major Red Pigments. Front. Microbiol. 2020, 11, 2056. [Google Scholar] [CrossRef]
- Lin, X.; Wu, P.; Huang, Y.; Dai, C. Optimization of Fermentation Parameters and Medium Composition for Producing Prodiginines from Marine Bacterium Spartinivicinus ruber MCCC 1K03745T. Fermentation 2025, 11, 629. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Altschul, S.F.; Wootton, J.C.; Gertz, E.M.; Agarwala, R.; Morgulis, A.; Schaffer, A.A.; Yu, Y.K. Protein database searches using compositionally adjusted substitution matrices. FEBS J. 2005, 272, 5101–5109. [Google Scholar] [CrossRef]
- Gouet, P.; Robert, X.; Courcelle, E. ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 2003, 31, 3320–3323. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Liu, D.; Xing, W.; Tang, C.; Feng, X.; Zhang, J. Simulating fatty acid autoxidation and exploring the related volatiles formation mechanism. LWT 2024, 214, 117083. [Google Scholar] [CrossRef]
- Luo, Q.; Smith, D.P. Global cancer burden: Progress, projections, and challenges. Lancet 2025, 406, 1536–1537. [Google Scholar] [CrossRef]
- Zhao, C.; Qiu, S.; He, J.; Peng, Y.; Xu, H.; Feng, Z.; Huang, H.; Du, Y.; Zhou, Y.; Nie, Y. Prodigiosin impairs autophagosome-lysosome fusion that sensitizes colorectal cancer cells to 5-fluorouracil-induced cell death. Cancer Lett. 2020, 481, 15–23. [Google Scholar] [CrossRef]
- McCoy, F.; Hurwitz, J.; McTavish, N.; Paul, I.; Barnes, C.; O’Hagan, B.; Odrzywol, K.; Murray, J.; Longley, D.; McKerr, G.; et al. Obatoclax induces Atg7-dependent autophagy independent of beclin-1 and BAX/BAK. Cell Death Dis. 2010, 1, e108. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, C.; Takemoto, H.; Kuno, K.; Yamamoto, D.; Tsubura, A.; Kamata, K.; Hirata, H.; Yamamoto, A.; Kano, H.; Seki, T.; et al. Cycloprodigiosin hydrochloride, a new H+/Cl− symporter, induces apoptosis in human and rat hepatocellular cancer cell lines in vitro and inhibits the growth of hepatocellular carcinoma xenografts in nude mice. Hepatology 1999, 30, 894–902. [Google Scholar] [CrossRef]
- Yamamoto, D.; Uemura, Y.; Tanaka, K.; Nakai, K.; Yamamoto, C.; Takemoto, H.; Kamata, K.; Hirata, H.; Hioki, K. Cycloprodigiosin hydrochloride, H+/CL− symporter, induces apoptosis and differentiation in HL-60 cells. Int. J. Cancer 2000, 88, 121–128. [Google Scholar] [CrossRef]
- Wang, J.; Liu, H.; Zhu, L.; Wang, J.; Luo, X.; Liu, W.; Ma, Y. Prodigiosin from Serratia marcescens in Cockroach Inhibits the Proliferation of Hepatocellular Carcinoma Cells through Endoplasmic Reticulum Stress-Induced Apoptosis. Molecules 2022, 27, 7281. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.Y.; Chen, N.F.; Kuo, H.M.; Yang, S.N.; Sung, C.S.; Sung, P.J.; Wen, Z.H.; Chen, W.F. Prodigiosin stimulates endoplasmic reticulum stress and induces autophagic cell death in glioblastoma cells. Apoptosis 2018, 23, 314–328. [Google Scholar] [CrossRef] [PubMed]
- Montaner, B.; Perez-Tomas, R. The cytotoxic prodigiosin induces phosphorylation of p38-MAPK but not of SAPK/JNK. Toxicol. Lett. 2002, 129, 93–98. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; He, S.; Zhang, X.; Gao, Q.; Liu, Y.; Liu, L. Structures, biosynthesis, and bioactivities of prodiginine natural products. Appl. Microbiol. Biotechnol. 2022, 106, 7721–7735. [Google Scholar] [CrossRef]
- Su, W.T.; Tsou, T.Y.; Liu, H.L. Response surface optimization of microbial prodigiosin production from Serratia marcescens. J. Taiwan Inst. Chem. Eng. 2011, 42, 217–222. [Google Scholar] [CrossRef]
- Chen, W.C.; Yu, W.J.; Chang, C.C.; Chang, J.S.; Huang, S.H.; Chang, C.H.; Chen, S.Y.; Chien, C.C.; Yao, C.L.; Chen, W.M.; et al. Enhancing production of prodigiosin from Serratia marcescens C3 by statistical experimental design and porous carrier addition strategy. Biochem. Eng. J. 2013, 78, 93–100. [Google Scholar] [CrossRef]
- Li, X.P.; Zhang, G.J.; Zhu, T.J.; Li, D.H.; Gu, Q.Q. Strain and Culture Medium Optimization for Production Enhancement of Prodiginines from Marine-Derived Streptomyces sp. GQQ-10. J. Ocean. Univ. China 2012, 11, 361–365. [Google Scholar] [CrossRef]
- Giri, A.V.; Anandkumar, N.; Muthukumaran, G.; Pennathur, G. A novel medium for the enhanced cell growth and production of prodigiosin from Serratia marcescens isolated from soil. BMC Microbiol. 2004, 4, 11. [Google Scholar] [CrossRef]
- Wang, X.; Cui, Z.; Zhang, Z.; Zhao, J.; Liu, X.; Meng, G.; Zhang, J.; Zhang, J. Two-Step Optimization for Improving Prodigiosin Production Using a Fermentation Medium for Serratia marcescens and an Extraction Process. Fermentation 2024, 10, 85. [Google Scholar] [CrossRef]
- Qadri, S.M.; Williams, R.P. Induction of prodigiosin biosynthesis after shift-down in temperature of nonproliferating cells of Serratia marcescens. Appl. Microbiol. 1972, 23, 704–709. [Google Scholar] [CrossRef] [PubMed]
- Qadri, S.M.; Williams, R.P. Role of methionine in biosynthesis of prodigiosin by Serratia marcescens. J. Bacteriol. 1973, 116, 1191–1198. [Google Scholar] [CrossRef]
- Wei, Y.H.; Yu, W.J.; Chen, W.C. Enhanced undecylprodigiosin production from Serratia marcescens SS-1 by medium formulation and amino-acid supplementation. J. Biosci. Bioeng. 2005, 100, 466–471. [Google Scholar] [CrossRef] [PubMed]
- Siva, R.; Subha, K.; Bhakta, D.; Ghosh, A.R.; Babu, S. Characterization and enhanced production of prodigiosin from the spoiled coconut. Appl. Biochem. Biotechnol. 2012, 166, 187–196. [Google Scholar] [CrossRef]
- Lim, D.V.; Qadri, S.M.; Nichols, C.; Williams, R.P. Biosynthesis of prodigiosin by non-proliferating wild-type Serratia marcescens and mutants deficient in catabolism of alanine, histidine, and proline. J. Bacteriol. 1977, 129, 124–130. [Google Scholar] [CrossRef]
- Williams, R.P.; Scott, R.H.; Lim, D.V.; Qadri, S.M. Macromolecular syntheses during biosynthesis of prodigiosin by Serratia marcescens. Appl. Environ. Microbiol. 1976, 31, 70–77. [Google Scholar] [CrossRef]
- Kim, C.H.; Kim, S.W.; Hong, S.I. An integrated fermentation–separation process for the production of red pigment by Serratia sp. KH-95. Process Biochem. 1999, 35, 485–490. [Google Scholar] [CrossRef]
- Domrose, A.; Klein, A.S.; Hage-Hulsmann, J.; Thies, S.; Svensson, V.; Classen, T.; Pietruszka, J.; Jaeger, K.E.; Drepper, T.; Loeschcke, A. Efficient recombinant production of prodigiosin in Pseudomonas putida. Front. Microbiol. 2015, 6, 972. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Lee, J.S.; Park, Y.K.; Kim, J.F.; Jeong, H.; Oh, T.K.; Kim, B.S.; Lee, C.H. Biosynthesis of antibiotic prodiginines in the marine bacterium Hahella chejuensis KCTC 2396. J. Appl. Microbiol. 2007, 102, 937–944. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Kim, Y.S.; Park, S.; Kim, J.; Kang, S.J.; Lee, M.H.; Ryu, S.; Choi, J.M.; Oh, T.K.; Yoon, J.H. Exceptional production of both prodigiosin and cycloprodigiosin as major metabolic constituents by a novel marine bacterium, Zooshikella rubidus S1-1. Appl. Environ. Microbiol. 2011, 77, 4967–4973. [Google Scholar] [CrossRef]




| Factors | Low Level (−1) | High Level (+1) |
|---|---|---|
| A-peanut meal (g/L) | 1 | 3 |
| B-glycerol (g/L) | 0 | 5 |
| C-L-proline (g/L) | 0 | 0.5 |
| Run | A- Peanut Meal | B- Glycerol | C- L-Proline | Prodiginine Concentration (mg/L) |
|---|---|---|---|---|
| 1 | +1 | −1 | −1 | 39.26 |
| 2 | +1 | −1 | +1 | 46.53 |
| 3 | −1 | −1 | +1 | 23.13 |
| 4 | +1 | +1 | −1 | 41.41 |
| 5 | −1 | +1 | −1 | 18.55 |
| 6 | +1 | +1 | +1 | 45.03 |
| 7 | −1 | +1 | +1 | 25.61 |
| 8 | −1 | −1 | −1 | 18.80 |
| Step | Peanut Meal (g/L) | L-Proline (g/L) |
|---|---|---|
| 1 | 3 | 0.500 |
| 2 | 4 | 0.567 |
| 3 | 5 | 0.633 |
| 4 | 6 | 0.700 |
| 5 | 7 | 0.767 |
| 6 | 8 | 0.833 |
| Level | Factors | |
|---|---|---|
| A-Peanut Meal (g/L) | B-L-Proline (g/L) | |
| −1.41421 | 0.55 | 0.337 |
| −1 | 2 | 0.433 |
| 0 | 5.5 | 0.667 |
| +1 | 9 | 0.900 |
| +1.41421 | 10.45 | 0.997 |
| Run | Factors | Prodiginine Concentration (mg/L) | |
|---|---|---|---|
| A-Peanut Meal | B-L-Proline | ||
| 1 | 0 | +1.41421 | 49.57 |
| 2 | 0 | 0 | 51.87 |
| 3 | +1 | −1 | 51.03 |
| 4 | +1 | +1 | 53.14 |
| 5 | 0 | 0 | 51.26 |
| 6 | 0 | 0 | 50.08 |
| 7 | 0 | −1.41421 | 50.10 |
| 8 | −1.41421 | 0 | 15.54 |
| 9 | 0 | 0 | 49.61 |
| 10 | 0 | 0 | 51.31 |
| 11 | −1 | −1 | 28.69 |
| 12 | −1 | +1 | 28.44 |
| 13 | +1.41421 | 0 | 47.11 |
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value | |
|---|---|---|---|---|---|---|
| Model | 995.63 a | 4 | 248.91 | 1666.39 | <0.0001 | significant |
| A-peanut meal | 927.41 | 1 | 927.41 | 6208.82 | <0.0001 | significant |
| B-glycerol | 1.03 | 1 | 1.03 | 6.88 | 0.0789 | |
| C-L-proline | 62.09 | 1 | 62.09 | 415.66 | 0.0003 | significant |
| ABC | 5.11 | 1 | 5.11 | 34.20 | 0.0100 | significant |
| Residual | 0.4481 | 3 | 0.1494 | |||
| Cor Total | 996.08 | 7 |
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value | |
|---|---|---|---|---|---|---|
| Model | 1725.13 a | 4 | 431.28 | 518.40 | <0.0001 | significant |
| A-peanut meal | 1051.11 | 1 | 1051.11 | 1263.44 | <0.0001 | significant |
| B-L-proline | 0.1551 | 1 | 0.1551 | 0.1865 | 0.6773 | |
| A2 | 670.19 | 1 | 670.19 | 805.57 | <0.0001 | significant |
| B2 | 2.17 | 1 | 2.17 | 2.61 | 0.1446 | |
| Residual | 6.66 | 8 | 0.8319 | |||
| Lack of Fit | 3.09 | 4 | 0.7734 | 0.8685 | 0.5527 | not significant |
| Pure Error | 3.56 | 4 | 0.8905 | |||
| Cor Total | 1731.79 | 12 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Lin, X.; Xiao, L.; Xiao, J.; Dai, C. Pathway-Guided Medium Engineering for Enhanced Prodiginine Production in Spartinivicinus ruber MCCC 1K03745T. Fermentation 2026, 12, 192. https://doi.org/10.3390/fermentation12040192
Lin X, Xiao L, Xiao J, Dai C. Pathway-Guided Medium Engineering for Enhanced Prodiginine Production in Spartinivicinus ruber MCCC 1K03745T. Fermentation. 2026; 12(4):192. https://doi.org/10.3390/fermentation12040192
Chicago/Turabian StyleLin, Xiaosi, Liping Xiao, Jingru Xiao, and Congjie Dai. 2026. "Pathway-Guided Medium Engineering for Enhanced Prodiginine Production in Spartinivicinus ruber MCCC 1K03745T" Fermentation 12, no. 4: 192. https://doi.org/10.3390/fermentation12040192
APA StyleLin, X., Xiao, L., Xiao, J., & Dai, C. (2026). Pathway-Guided Medium Engineering for Enhanced Prodiginine Production in Spartinivicinus ruber MCCC 1K03745T. Fermentation, 12(4), 192. https://doi.org/10.3390/fermentation12040192





