1. Introduction
The projected growth of the global population and the simultaneous increase in demand for animal protein place significant pressure on traditional animal feed resources [
1]. The aquaculture industry, an essential part of global food security, still relies heavily on fishmeal as a primary protein source. This situation is highly unsustainable due to declining marine pelagic fish stocks, extreme price fluctuations, and serious environmental impacts [
2,
3]. This issue has led to a worldwide search for alternative, sustainable, and affordable protein ingredients for animal feeds by utilizing agro-industrial by-products [
4,
5].
Japan produces large amounts of organic waste with significant potential for bioconversion [
6,
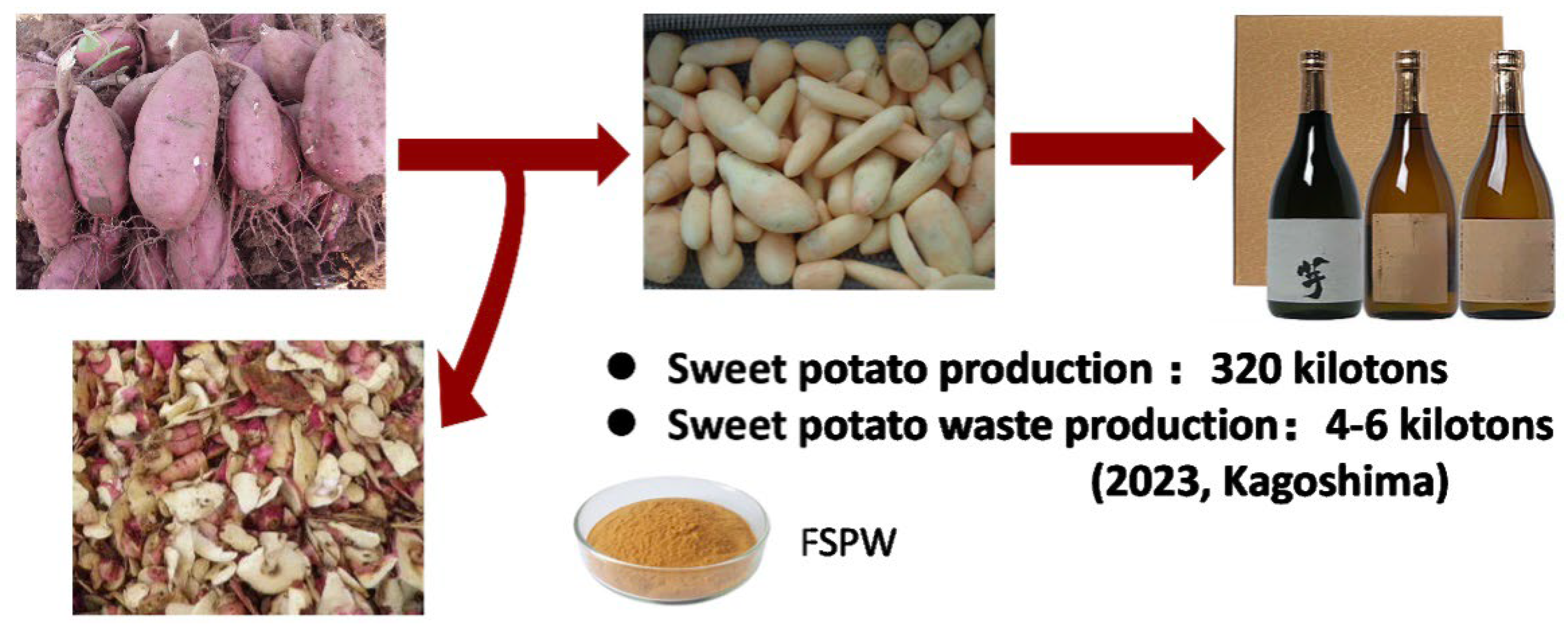
7]. In Kagoshima Prefecture, the heart of Japan’s shochu industry, about 300 kilotons of sweet potatoes are processed each year. This industrial activity generates over 4 kilotons of sweet potato waste (SPW) annually, mainly consisting of peels, rejected residues, and spilled tuber pieces (
Figure 1). SPW is a residual material rich in carbohydrates and bioactive compounds like polyphenols, making it an excellent substrate for biotechnological applications [
8]. While some of this waste is used as unprocessed feed by local farms, most of it is disposed of as industrial waste, posing environmental challenges and missing economic opportunities. These residues, abundant in carbohydrates and bioactive compounds such as polyphenols, are well-suited for biotechnological upgrading into high-value feed ingredients [
9].
However, using sweet potato peel residue directly in high-performance animal diets, especially for monogastric species, is heavily limited by its natural biochemical qualities. Nutritional details are summarized in
Table 1. As shown, the raw material is typical lignocellulosic biomass, characterized by a low crude protein content (about 6.0% on a dry matter basis) and a high amount of complex carbohydrates, including starch and dietary fiber [
4,
5]. Importantly, it contains high levels of anti-nutritional factors (ANFs), mainly trypsin inhibitors. Sporamin, the primary storage protein in sweet potatoes, functions as a strong trypsin inhibitor, greatly hindering proteolytic digestion in monogastric animals [
10,
11,
12]. Therefore, an effective valorization strategy must address three main challenges: increasing protein content, breaking down the tough lignocellulosic matrix, and removing ANFs.
Solid-state fermentation (SSF) is a highly effective and cost-efficient method for upgrading agro-industrial waste [
13,
14]. However, the complex and resistant nature of sweet potato peel requires an advanced fermentation approach that goes beyond using a single microorganism. A two-stage sequential fermentation process employing
Aspergillus oryzae (
A. oryzae) and
Lactobacillus plantarum (
L. plantarum) offers a biochemically synergistic pathway, allowing for sequential lignocellulosic breakdown followed by rapid microbial biosynthesis [
15,
16,
17,
18].
Aspergillus oryzae, a GRAS (Generally Recognized as Safe) fungus essential to the shochu industry, is used in the first aerobic stage because of its extensive secretion of hydrolytic enzymes (such as amylases, cellulases, and proteases) that decompose the lignocellulosic matrix and convert complex carbohydrates into sugars [
15,
19]. This initial hydrolysis produces a nutrient-rich pool of simple sugars and free amino acids for the second anaerobic stage, where
Lactobacillus plantarum, a potent lactic acid bacterium, quickly ferments the released sugars. This second step is vital for further breaking down anti-nutritional factors (ANFs), lowering the substrate pH to prevent spoilage, and introducing beneficial probiotic microorganisms to the final product [
20,
21].
Table 1.
Projected comprehensive nutritional composition of sweet potato peel residue (dry matter (DM) basis).
Table 1.
Projected comprehensive nutritional composition of sweet potato peel residue (dry matter (DM) basis).
| Category | Component | Content Range (DM Basis) | Key Notes & Challenges | References |
|---|
| Proximate Analysis | Crude Protein | 5.6–8.8% | Low content, primarily composed of anti-nutritional protein (sporamin). | [12,22,23] |
| | Crude Fat/Ether Extract | 1.7–4.5% | Low content, but can be transformed via fungal fermentation. | [22,23] |
| | Crude Fiber | 2.0–5.0% | Traditional measure; actual dietary fiber is much higher, limiting digestion. | [22,23] |
| | Ash | 4.0–7.0% | Indicates a rich source of minerals. | [22,23,24] |
| Carbohydrate Profile | Starch | 19.4–52.0% | Key energy source, but physically encapsulated by the cell wall. | [4,22,25] |
| | Total Dietary Fiber | >21% (up to 60%) | Includes cellulose, hemicellulose, and lignin; a key target for degradation. | [26] |
| | Cellulose | ~20.2–31.2% | Structural polysaccharide requiring cellulase for degradation. | [22,26] |
| | Hemicellulose | ~11.4–15.2% | Structural polysaccharide requiring hemicellulase for degradation. | [4,22,25] |
| | of which: Lignin | ~10.0–16.9% | Recalcitrant phenolic polymer, a major barrier to enzymatic hydrolysis. | [27,28] |
| Anti-Nutritional Factors | Trypsin Inhibitors | High | Core limiting factor; severely inhibits protein digestion. | [12,25,26] |
| | Phytate | ~755 mg/100 g | Chelates minerals, reducing their bioavailability. | [22] |
| | Tannins | ~533 mg/100 g | Reduces protein digestibility and affects palatability. | [22] |
Optimizing a biological system governed by highly non-linear, multi-variable interactions remains a significant challenge [
20,
29,
30]. Traditional statistical techniques, such as the Response Surface Methodology (RSM) utilizing a Central Composite Design, depend on second-order polynomial equations that often fail to capture the extreme non-linearities and local optima inherent in dynamic microbial ecosystems [
31,
32]. To address this, advanced machine learning architectures like Artificial Neural Networks (ANNs) and Random Forest (RF) algorithms paired with heuristic optimization methods are increasingly preferred [
33,
34,
35]. ANNs excel at modeling complex, non-linear relationships in high-dimensional datasets, while Differential Evolution (DE) algorithms efficiently explore extensive parameter spaces to identify global optima [
18,
19,
20]. However, bioprocessing datasets often face the “small N, large P” problem, where the number of experimental samples (N) is too small compared to the high-dimensional complexity of the parameters (P). While gradient-based deep learning methods within ANNs are highly effective at modeling complex relationships, they need large amounts of data to avoid catastrophic overfitting [
36]. On the other hand, ensemble learning techniques like RF use bootstrap aggregating (bagging) across decision trees, enabling them to handle small sample sizes effectively, reduce individual biases, and lower overall model variance [
37].
This optimized fermentation is expected to create an ideal thermodynamic and biochemical environment for maximum nutritional enrichment. We anticipate that crude protein levels will increase significantly, fiber will decrease, and the levels of key anti-nutritional factors will be almost eliminated through enzymatic degradation [
10,
12,
38,
39]. The final product is expected to be a synbiotic matrix consisting of probiotics (living
L. plantarum), prebiotics (partially hydrolyzed dietary fibers that support gut health), and postbiotics (bioactive metabolites produced during fermentation, such as organic acids, enzymes, and peptides). This combination is expected to improve nutrient utilization and biochemical quality [
40,
41,
42,
43,
44].
Therefore, this study is based on the scientific hypothesis that a two-stage, co-culture solid-state fermentation of sweet potato peel waste can be modeled and optimized using an ANN or RF algorithm to produce a high-quality, nutritionally enhanced ingredient. The main objectives are: (1) to systematically gather experimental data on the fermentation process using a multi-variable design; (2) to develop and validate a predictive ANN or RF model for fermentation outcomes with robust cross-validation techniques; and (3) to utilize a Constrained Differential Evolution (CDE) algorithm to find the optimal process parameters that maximize nutritional value and functionality, followed by laboratory validation. The innovative aspect of this study lies in combining an Augmented Experimental Design (AED) with a sequential bioprocess, enabling strong ensemble learning on a highly resistant agricultural waste stream.
2. Materials and Methods
2.1. Substrate and Chemicals
Sweet potato waste (SPW), mainly consisting of peels and unused pieces of tuberous roots (Ipomoea batatas L.), was sourced from a local shochu distillery in Kagoshima, Japan. The raw material had a moisture content of 77% and a crude protein content of 5.95 ± 0.12% on a dry matter (DM) basis, as measured by the Nutritional Biochemistry and Feed Chemistry Laboratory at the Faculty of Agriculture, Kagoshima University (Kagoshima, Japan).
Ammonium sulfate ((NH4)2SO4) of analytical grade was obtained from Wako Pure Chemical Industries, Ltd. (Osaka, Japan) and used as the supplementary nitrogen source. Potato Dextrose Agar was obtained from Nissui Pharmaceutical Co., Ltd. (Tokyo, Japan), Polysorbate 80 (Tween 80), o-phthaldialdehyde (OPA), BAPNA Substrate, Folin-Ciocalteu Reagent, DPPH Radical, Gallic Acid/Tannic Acid were obtained from Nacalai Tesque, Inc. (Kyoto, Japan), and all other chemicals and reagents employed in this study were of analytical or HPLC grade.
2.2. Substrate Pre-Treatment
The SPW was dried in a forced-air oven at 60 °C for 8 h to reduce moisture content to about 10–12% for stable storage. The dried SPW was then ground into a fine powder using a pulverizer (DD-2-3.7, Makino MFG Co., Tokyo, Japan) and sifted through a 40-mesh sieve. For each fermentation run, the ground SPW powder underwent high-pressure steam pre-treatment to break down the lignocellulosic structure and improve enzymatic accessibility. The powder was autoclaved at 103.4 kPa (around 121 °C) for 20 min and then cooled to room temperature [
45].
2.3. Microorganisms and Inoculum Preparation
2.3.1. Microbial Strains
Aspergillus oryzae (strain RIB 40) was obtained from the NITE Biological Resource Center (Chiba, Japan). Lactobacillus plantarum (strain JCM 1149) was procured from the Japan Collection of Microorganisms, RIKEN BioResource Research Center (Tsukuba, Japan).
2.3.2. Starter Culture Preparation
The
A. oryzae starter culture was prepared by inoculating the strain onto Potato Dextrose Agar (PDA) slants (Nissui Pharmaceutical Co., Ltd., Tokyo, Japan) and incubating at 30 °C for 5–7 days until abundant sporulation was observed. Spores were harvested by washing the slant surface with 10 mL of sterile 0.1% (
v/
v) Tween 80 solution. The resulting spore suspension was filtered through sterile glass wool, and the spore concentration was determined using a hemocytometer (Erma Inc., Tokyo, Japan) and adjusted to 1 × 10
8 spores/mL with sterile distilled water [
20].
The L. plantarum inoculum was prepared by transferring a loopful of the stock culture into 50 mL of De Man, Rogosa, and Sharpe (MRS) broth (Becton, Dickinson and Company, Franklin Lakes, NJ, USA). The culture was incubated anaerobically at 37 °C for 24 h. We then placed this activated culture in fresh MRS broth and incubated it for 18 h under the same conditions to ensure the cells were in the late logarithmic growth phase. We used centrifugation (5000× g, 10 min, 4 °C) to collect the bacterial cells, washed them twice with sterile physiological saline, and then resuspended them in saline to a final concentration of 1 × 1010 Colony Forming Units (CFU)/mL. The standard plate count method on MRS agar was used to check the number of living cells.
2.4. Two-Stage Solid-State Fermentation (SSF) Process
The fermentation was carried out in two steps using 500 mL erlenmeyer flasks (AGC Techno Glass Co., Ltd., Shizuoka, Japan) with 50 g (dry matter basis) of the pre-treated SPW powder.
Stage 1: Aerobic Fungal Fermentation: The initial moisture level of the substrate (ranging continuously between 55.0% and 80.0%) was strictly controlled. The exact volume of aqueous
A. oryzae inoculum, the volumetric solution of the nitrogen supplement (ammonium sulfate), and additional sterile distilled water were mathematically balanced using mass-balance equations against the known dry matter weight to achieve the precise target moisture percentage. The flasks were sealed with breathable cotton plugs and incubated aerobically under the conditions described in
Table 2.
Stage 2: Anaerobic Bacterial Fermentation: A true sequential co-culture approach was used. No intermediate sterilization was done before bacterial inoculation in Stage 2, allowing the primary fungal strain to stay biologically active until it was naturally inhibited by the rapidly decreasing pH and oxygen depletion [
46]. The substrate was inoculated with the specific volumetric
L. plantarum suspension needed to reach the target CFU/g, sealed to create anaerobic conditions, and incubated (
Table 2).
2.5. Modeling and Optimization Strategy
A comparative study using two main machine learning (ML) algorithms, the Artificial Neural Network (ANN) and the Random Forest (RF), was carried out to model and optimize the complex, non-linear fermentation process. The aim was to determine which model could most accurately predict the five main fermentation outcomes (
Table 3) using the eight input process variables listed in
Table 2.
Both models were developed using Python (v3.10) with the scikit-learn library [
47]. The model with the highest predictive accuracy on unseen test data was then selected as the objective function for a global optimization algorithm.
2.5.1. Data Preprocessing and Leakage Prevention
The experimental dataset (N = 80) was loaded into a pandas (version 2.3.3) DataFrame [
48]. To strictly prevent data leakage, all preprocessing steps were applied only to the training subset within each cross-validation fold and then used on the test subset.
Log-Transformation: To normalize the distributions of the input variables (A. oryzae Inoculum, L. plantarum Inoculum) and the output variable (Viable LAB Count) more normally, we log10-transformed them.
Data Partitioning: The complete dataset was evaluated using a repeated k-fold cross-validation strategy (5-fold, repeated 3 times with different random seeds) to ensure statistical robustness and stability in model comparisons.
Feature Scaling: StandardScaler was used to standardize the input and output features for the ANN model only (mean = 0, std = 1). This step is essential for the ANN model to perform well, but is not necessary for the RF model [
47,
49,
50].
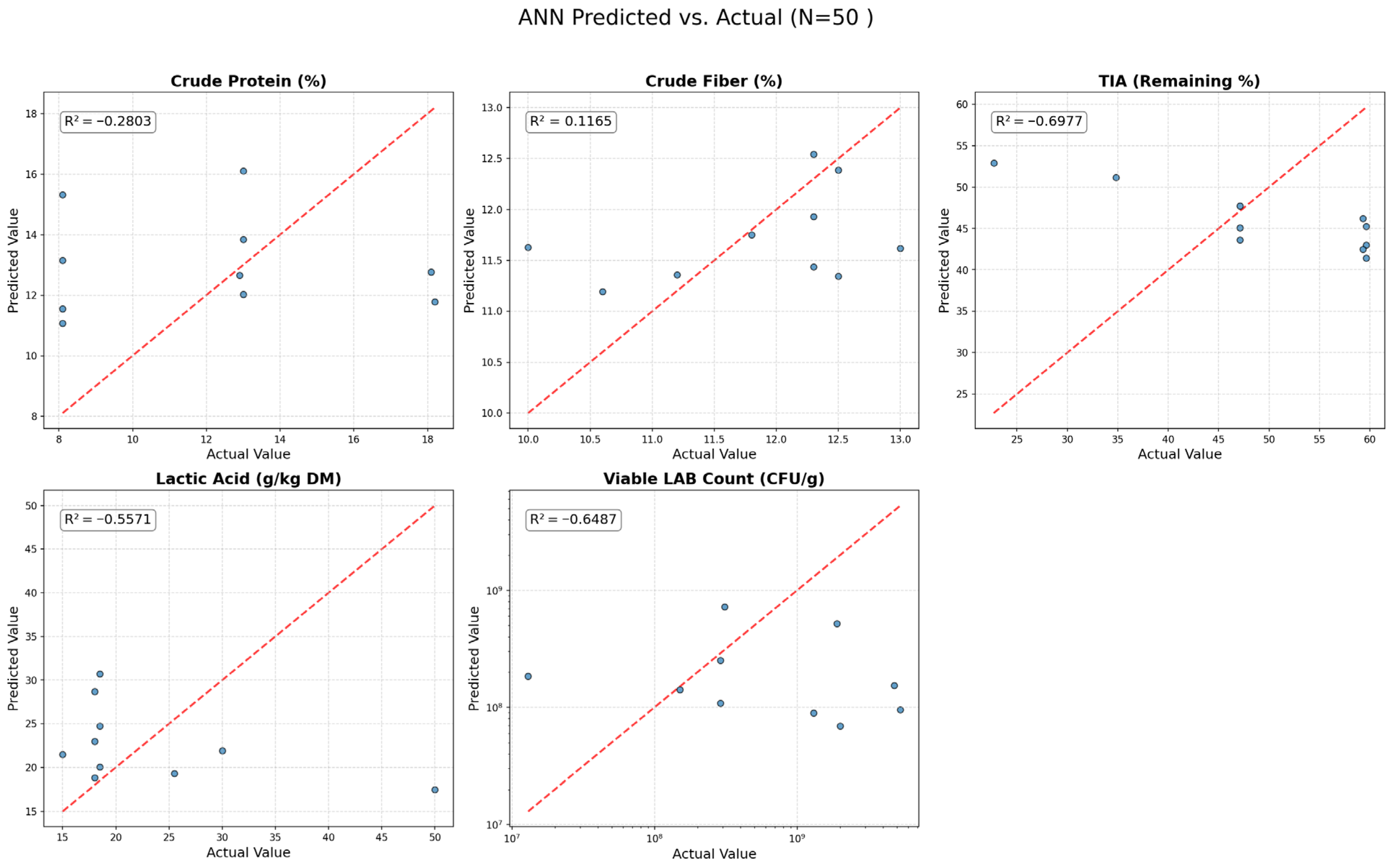
2.5.2. Model 1: Artificial Neural Network (ANN)
Scikit-learn’s MLPRegressor (Multi-layer Perceptron) was used to develop the ANN model.
Architecture: Five separate 8-input-1-output (MISO) models were trained for each of the five responses, rather than a single 8-input-5-output (MIMO) model. This prevents gradient vanishing or exploding caused by output variables with different scales. Each network consists of an input layer (8 neurons), two hidden layers (with 12 and 6 neurons), and an output layer (1 neuron) (an 8-12-6-1 structure).
Training: The ‘adam’ solver was used with a high L2 regularization (alpha = 0.1) and a learning rate of 0.001 [
51]. This strong regularization was a deliberate choice to force the model to find only the most generalizable patterns and combat the severe overfitting typically seen when training ANNs on small datasets. A limited hyperparameter grid search confirmed that the fundamental lack of data density (N = 80) for an 8-dimensional space caused the network to fail regardless of tuning.
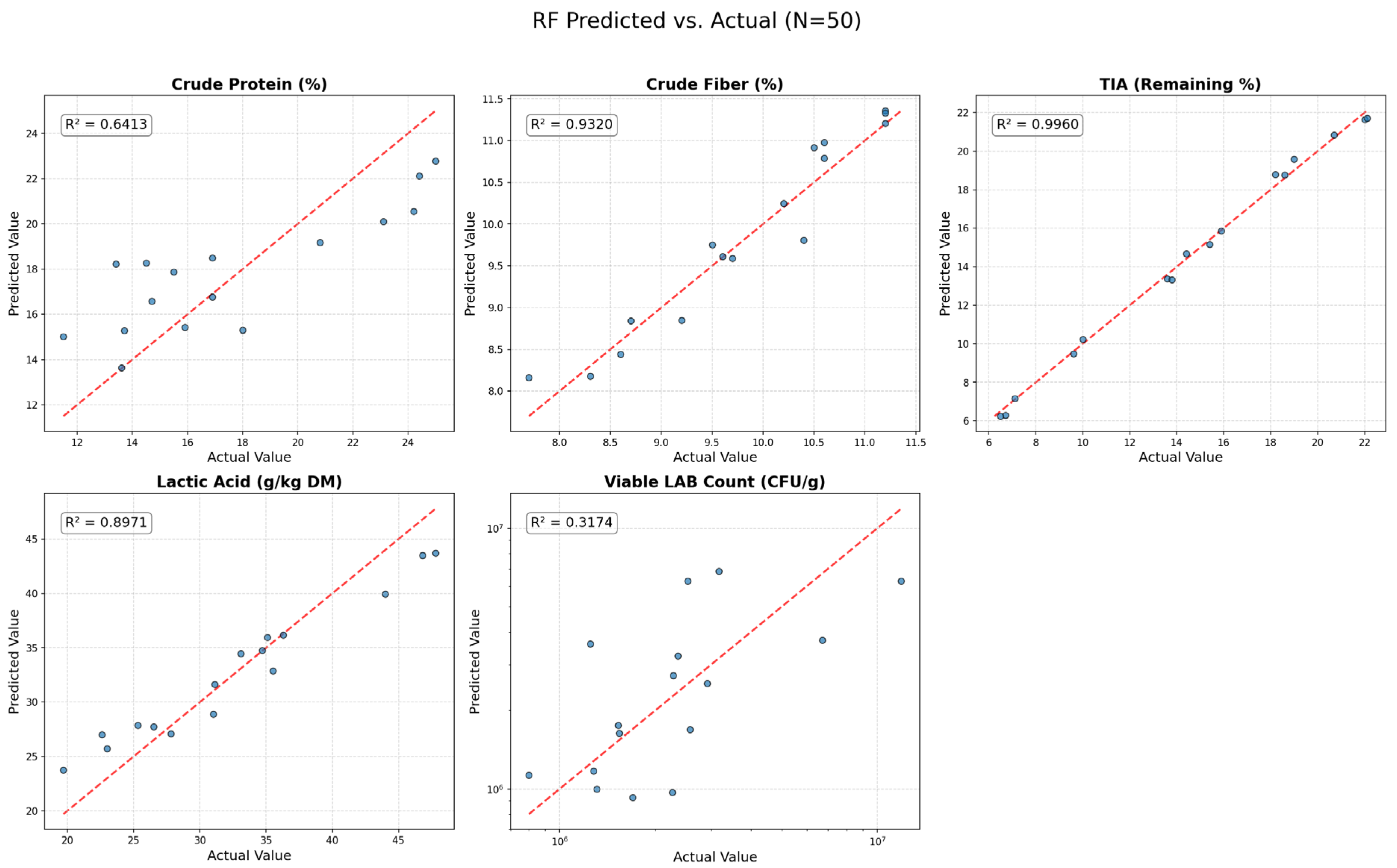
2.5.3. Model 2: Random Forest (RF)
We used the RandomForestRegressor from scikit-learn to make the comparative RF model [
47,
52].
Architecture & Theory: RF is an ensemble largetitude of learning that builds a number of decision trees (n_estimators = 100) during training [
53]. RF creates 100 “weak” trees, each trained on a different random subset of the data (“bagging”).
Prediction and Uncertainty: The final prediction is the average of all 100 trees’ predictions. This averaging mitigates “noise” and errors from individual trees, enabling the model to capture complex, non-linear relationships without overfitting. Furthermore, predictive uncertainty at any given coordinate in the parameter space was estimated by calculating the standard deviation (SD) of predictions across all 100 individual trees, thereby generating visual confidence intervals [
54].
Training: For a direct performance comparison, the model was trained on the same k-fold cross-validation splits as the ANN. To get the best predictive accuracy, key hyperparameters (max depth = 5, min samples leaf = 2) were tuned.
2.5.4. Optimization by Differential Evolution
Based on the comparative results (detailed in
Section 3), the trained RF model was confirmed as the only reliable predictor for the non-linear responses. Therefore, the validated RF model was used as the objective function for a Constrained Differential Evolution (CDE) algorithm, a robust and dependable global optimizer implemented in scipy.Optimize library [
55]. To ensure practical biological viability, the DE algorithm was programmed to maximize the expected Crude Protein (%) output, subject to strict boundary constraints: Trypsin Inhibitor Activity (TIA) ≤ 15% and Lactic Acid ≤ 30 g/kg.
2.6. Experimental Design: A Mixed Method for Machine Learning
Training and testing robust machine learning models require a comprehensive experimental dataset. Traditional Response Surface Methodology (RSM) designs, such as the Central Composite Design (CCD), work well for optimizing simple quadratic models with few experimental runs. However, their sparsity (N < 50) severely limits the effectiveness of data-intensive models like ANNs. To create a dataset that allows a fair comparison between ANN and RF, we used an Augmented Experimental Design (AED) with 80 runs. The original 50 experimental runs from the CCD provided a strong foundation, and 30 additional runs were strategically added to increase dataset density. These new runs were based on the original ranges but rounded to practical testing values (e.g., 0.5 °C increments, 1-h increments). This approach enabled a more thorough exploration of the parameter space.
2.7. Section and Preparation
At the end of each fermentation run, the fermented sweet potato waste (FSPW) was collected. A portion was immediately used for pH measurement and microbial analysis. The rest was lyophilized (freeze-dried), ground in a rotary mill to pass through a 0.2 mm mesh sieve, and stored at −20 °C for further chemical analysis. Unfermented SPW was processed in the same way to serve as the control.
2.8. Analytical Methods
2.8.1. pH and Organic Acid Analysis
The pH of the samples was measured using a digital pH meter after homogenizing 10 g of the sample with 90 mL of deionized water. The concentrations of lactic acid and acetic acid were determined by High-Performance Liquid Chromatography (HPLC) (Shimadzu Corp., Kyoto, Japan), equipped with a UV detector at 210 nm and an Aminex HPX-87H column (Bio-Rad, Hercules, CA, USA), using 5 mM H2SO4 as the mobile phase.
2.8.2. Nutritional Analysis
The proximate composition (dry matter, crude protein, crude fat, crude fiber, and ash) of both SPW and FSPW samples was determined using the standard procedures of AOAC International [
56]. 1 Crude protein was measured via the Kjeldahl method (N × 6.25). Total amino acid profiles were analyzed with an automated amino acid analyzer (Hitachi L-8900, Hitachi, Tokyo, Japan) after acid hydrolysis with 6 M HCl at 110 °C for 24 h.
2.8.3. Degree of Protein Hydrolysis (DH)
The DH, defined as the percentage of cleaved peptide bonds, was measured using the o-phthaldialdehyde (OPA) spectrophotometric method as described in our previous research [
57]. Absorbance was taken at 340 nm.
2.8.4. Anti-Nutritional Factors (ANFs) Assays
TIA was measured according to the AOCS Official Method, which is based on the inhibition of trypsin hydrolysis of Nα-Benzoyl-L-arginine 4-nitroanilide hydrochloride (BAPNA) [
56]. Phytic acid content was determined spectrophotometrically based on the pink-colored complex formed between ferric iron and 2,2′-bipyridine. Total tannins were quantified using the Folin–Ciocalteu method with tannic acid as a standard.
2.8.5. Total Phenolic Content (TPC) and Antioxidant Activity
TPC was measured using the Folin–Ciocalteu reagent method, with results expressed as gallic acid equivalents (GAE). Antioxidant activity was evaluated through the DPPH (2,2-diphenyl-1-picrylhydrazyl) radical scavenging assay.
2.9. Statistical Analysis
The experimental design and RSM analysis utilized the “rsm” package in R (version 4.2.1). Implementations of ANN and RF were performed using Python (v3.10), with code available in the
Supplementary Files. All experiments were conducted in triplicate, except for in vitro validation runs, which had
n = 6, and data were presented as mean ± standard deviation (SD). Machine learning performance metrics were averaged across multiple k-fold validation splits, expressed as mean ± SD. Differences between treatment means were tested using a one-way ANOVA and Tukey’s test, with a significance level of
p < 0.05. The performance of the ANN model was assessed through the coefficient of determination (R
2) and the F-test.
4. Conclusions
This study demonstrated a robust, data-driven framework using Random Forest algorithms and a two-stage solid-state fermentation process to improve the nutritional profile of sweet potato waste significantly. Through repeated k-fold cross-validation, we confirmed that ensemble learning greatly exceeds Artificial Neural Networks in handling the complex, sparse datasets typical of bioprocess engineering. The application of a Constrained Differential Evolution optimizer successfully identified an optimal biological strategy, which was strictly validated through wet-lab experiments. The optimized fermentation resulted in a highly digestible bio-matrix with 24.6% crude protein, a 123.9% increase in essential amino acids, and substantial reductions in anti-nutritional factors. Additionally, the fermentation process led to a comprehensive biochemical enhancement, marked by a significant increase in the degree of protein hydrolysis, total phenolic content, and antioxidant activity, along with the effective breakdown of phytic acid and tannins. Importantly, the bioprocess produced a safe and stable product by quickly lowering pH through the synergistic accumulation of lactic and acetic acids, while also increasing crude fat and ash levels. Although future in vivo trials are needed to verify physiological effects on animals, this research offers the shochu industry a scientifically solid, scalable blueprint to convert environmental waste into a functionally improved, sustainable protein resource.