Bacillus subtilis Genome Reduction Improves Surfactin Production
Abstract
1. Introduction
2. Materials and Methods
2.1. DNA Sequences and Cloning
2.2. Bacterial Strains and Growth Conditions
2.3. B. subtilis Strain Engineering
2.4. Analytical Methods
2.5. Data Analysis
3. Results
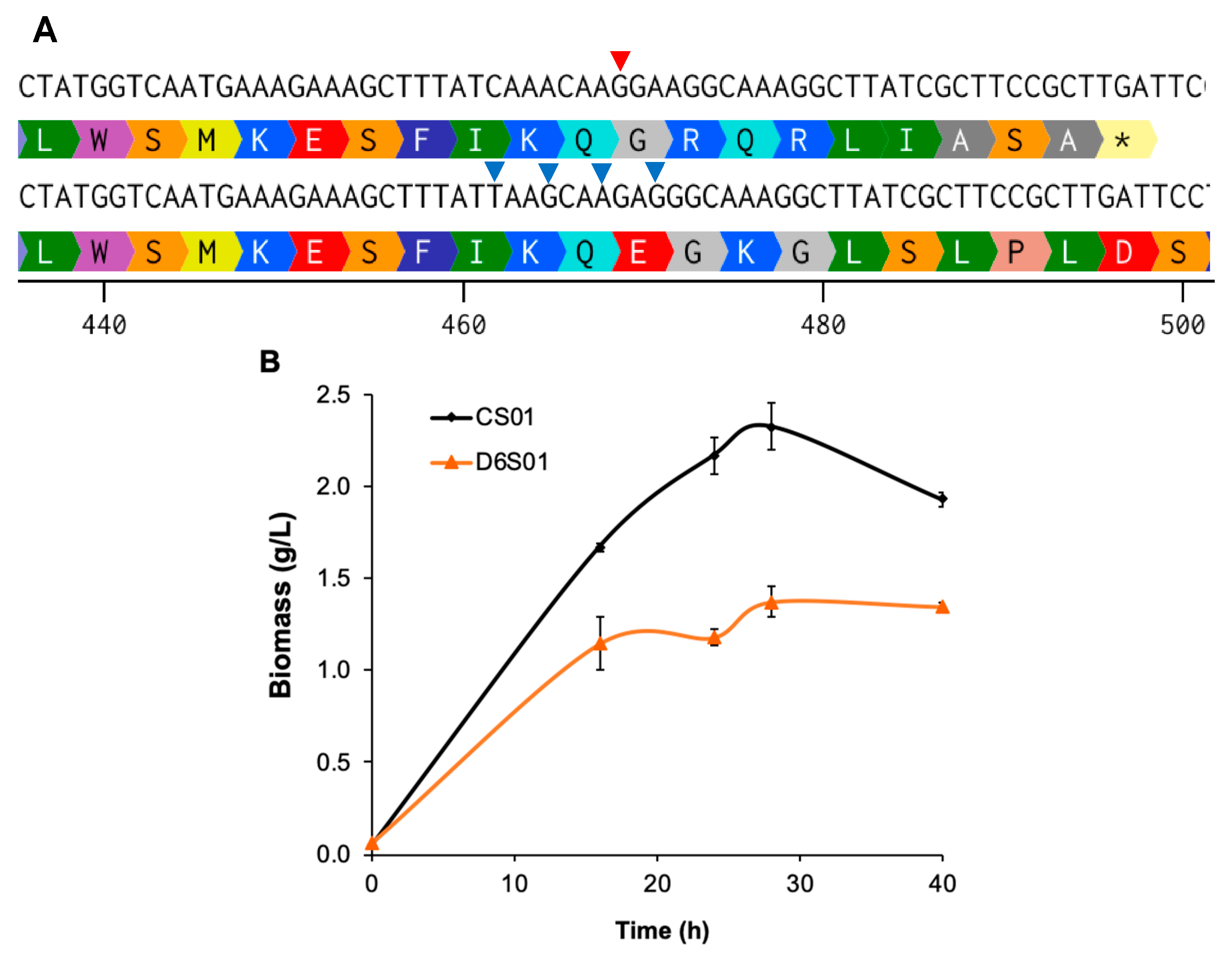
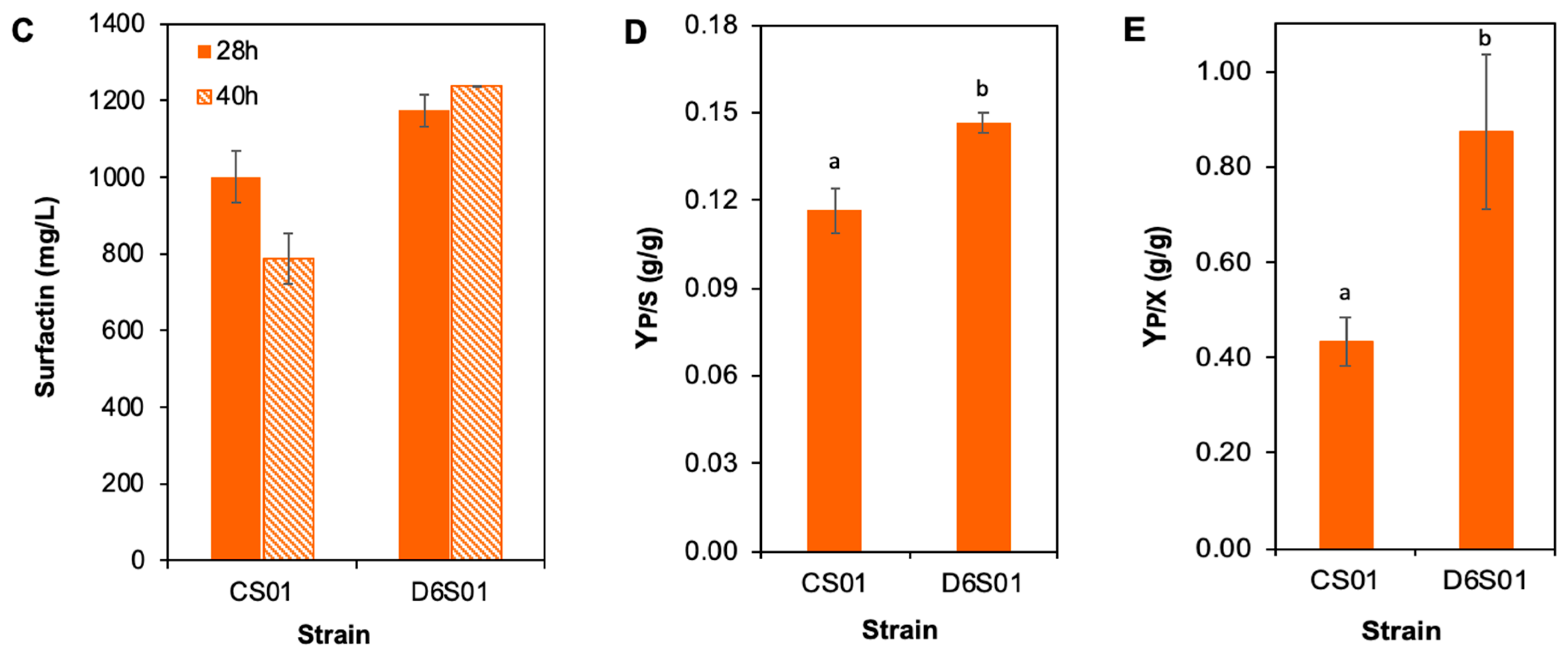
3.1. Genome-Reduced B. subtilis Performs Better than Its Parental Strain
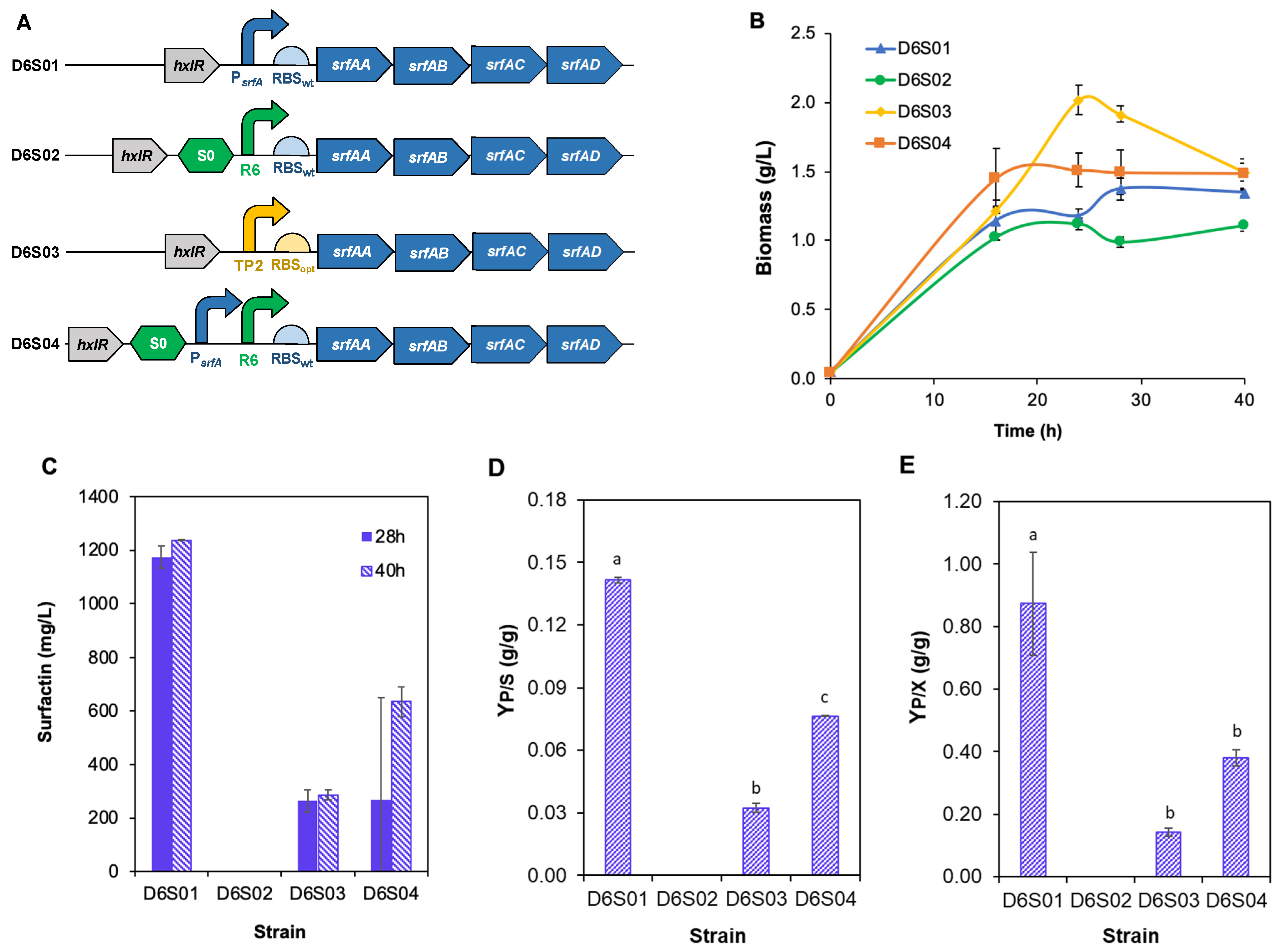
3.2. Promoter Editing
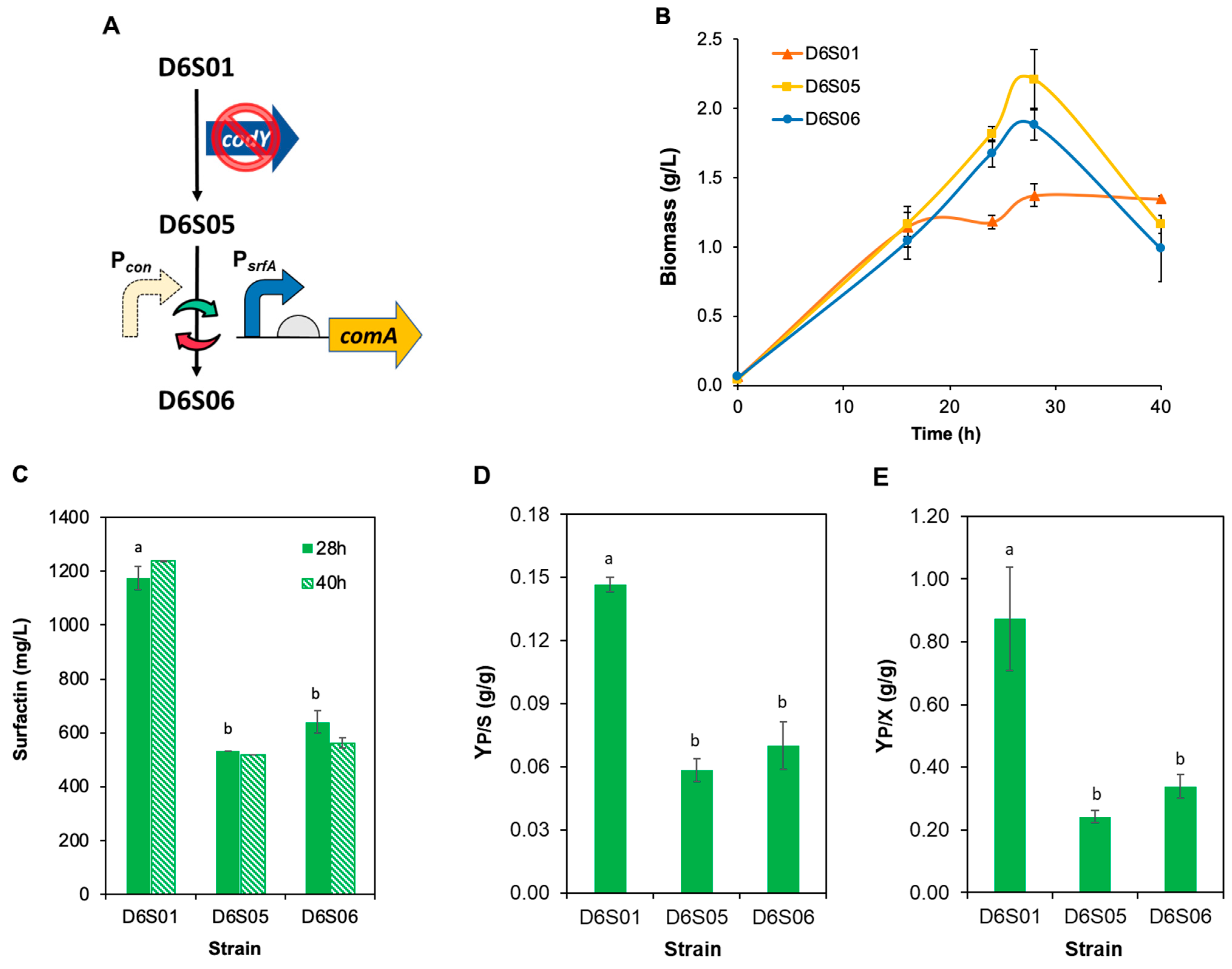
3.3. Engineering Transcription Regulators
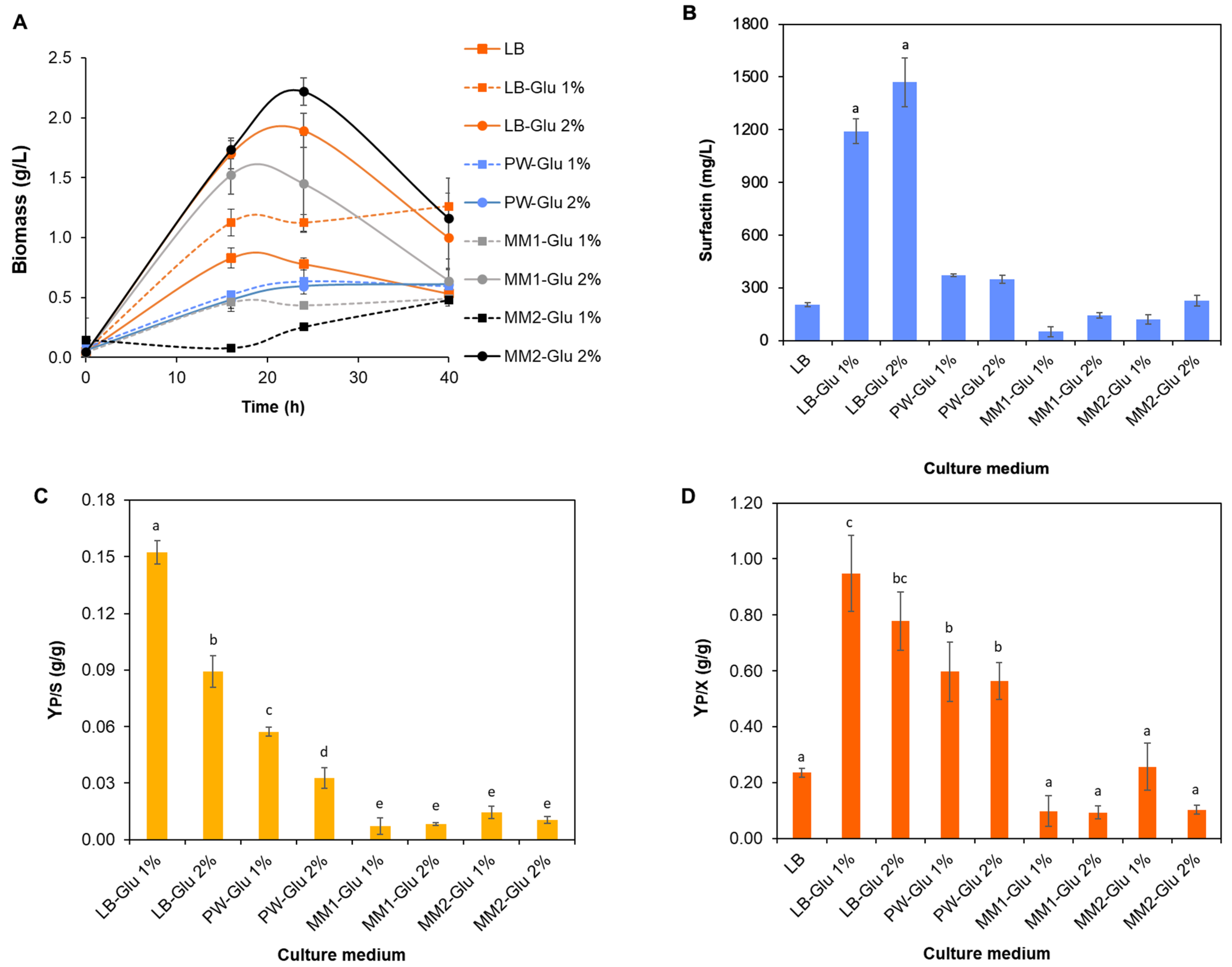
3.4. Culture Medium Optimization
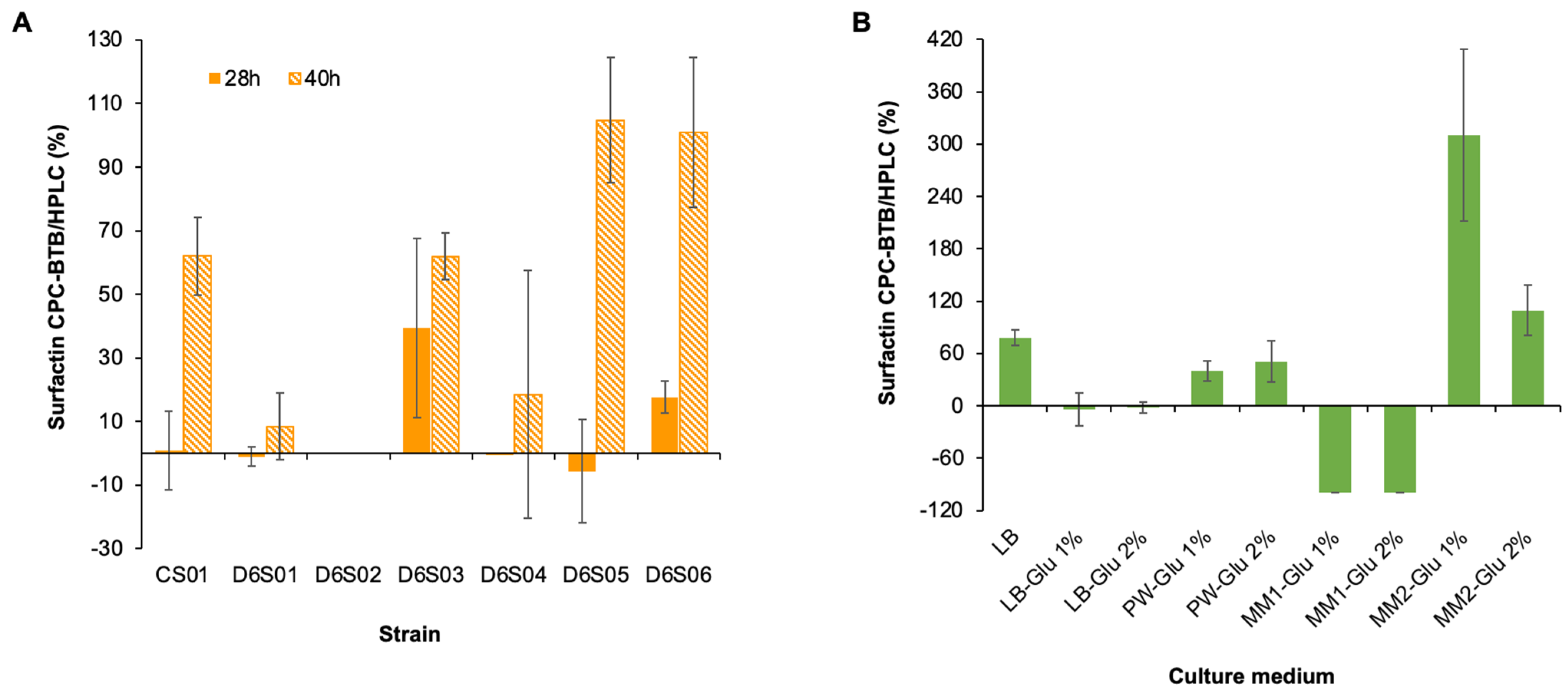
3.5. The CPC–BTB Method Is Suitable for Interference from the Culture Medium
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
Appendix A
Appendix A.1. gRNA Sequences
| ID | Target | Sequence (5′-3′) | Reference |
|---|---|---|---|
| GS1 | sfp | AAAGCTTTATCAAACAAGGA | This study |
| GS2 | PsrfA | AAAGATTGAACGCAGCAGTT | This study |
| GS3 | R6 | AATAGGTAAGGATAAAGAGA | This study |
| GS4 | codY | ACCCAAGGAGTTTCCCTCTG | This study |
| GS5 | comA | ACTAGTGATTGATGACCATC | This study |
Appendix A.2. Donor DNA Sequences
| DNA Part * | Description | Reference |
|---|---|---|
| S0-R6 | Transcription autoinduction device | [40] |
| TP2 | Constitutive dual promoter | [39] |
| RBSopt | Optimized RBS | [39] |
| PsrfA | Wild-type inducible promoter, amplified from the B. subtilis Δ6 genome | This study |
| RBSwt | Wild-type RBS associated with PsrfA, amplified from the B. subtilis Δ6 genome | This study |
| DS1 | Donor DNA 1 for sfp point mutations | This study |
| DS2 | Donor DNA 2 for substitution of PsrfA for S0-R6 to control srfA operon expression | This study |
| DS3 | Donor DNA 3 for substitution of PsrfA-RBSwt for TP2-RBSopt to control srfA operon expression | This study |
| DS4 | Donor DNA 4 for insertion of PsrfA between S0 and R6 to control srfA operon expression | This study |
| DS5 | Donor DNA 5 for codY deletion | This study |
| DS6 | Donor DNA 6 for insertion of PsrfA-RBSwt to control comA expression | This study |
Appendix A.3. Culture Medium Optimization: Reducing Sugars Quantification
| Medium | Initial Reducing Sugars (g L−1) | Final Reducing Sugars (g L−1) | Consumed Reducing Sugars (g L−1) | Surfactin (g L−1) |
|---|---|---|---|---|
| LB | 0.16 ± 0.00 | 0.11 ± 0.02 | 0.04 ± 0.01 | 0.20 ± 0.01 |
| LB-Glu 1% | 9.26 ± 0.03 | 0.28 ± 0.05 | 7.82 ± 0.14 | 1.19 ± 0.07 |
| LB-Glu 2% | 26.09 ± 1.20 | 7.29 ± 0.45 | 16.47 ± 1.06 | 1.47 ± 0.14 |
| PW-Glu 1% | 7.83 ± 0.17 | 0.31 ± 0.13 | 6.50 ± 0.36 | 0.37 ± 0.01 |
| PW-Glu 2% | 23.23 ± 2.18 | 11.71 ± 2.30 | 10.82 ± 1.76 | 0.35 ± 0.02 |
| MM1-Glu 1% | 8.08 ± 0.04 | 0.54 ± 0.24 | 6.96 ± 0.02 | 0.05 ± 0.03 |
| MM1-Glu 2% | 23.76 ± 0.57 | 6.68 ± 0.94 | 17.08 ± 0.94 | 0.14 ± 0.02 |
| MM2-Glu 1% | 8.42 ± 0.32 | 0.07 ± 0.01 | 8.32 ± 0.04 | 0.12 ± 0.03 |
| MM2-Glu 2% | 24.30 ± 0.89 | 0.07 ± 0.02 | 21.79 ± 1.01 | 0.23 ± 0.03 |
References
- Reuß, D.R.; Altenbuchner, J.; Mäder, U.; Rath, H.; Ischebeck, T.; Sappa, P.K.; Thürmer, A.; Guérin, C.; Nicolas, P.; Steil, L.; et al. Large-Scale Reduction of the Bacillus subtilis Genome: Consequences for the Transcriptional Network, Resource Allocation, and Metabolism. Genome Res. 2017, 27, 289–299. [Google Scholar] [CrossRef]
- Choe, D.; Cho, S.; Kim, S.C.; Cho, B.K. Minimal Genome: Worthwhile or Worthless Efforts toward Being Smaller? Biotechnol. J. 2016, 11, 199–211. [Google Scholar] [CrossRef]
- Aguilar Suárez, R.; Kohlstedt, M.; Öktem, A.; Neef, J.; Wu, Y.; Ikeda, K.; Yoshida, K.I.; Altenbuchner, J.; Wittmann, C.; van Dijl, J.M. Metabolic Profile of the Genome-Reduced Bacillus subtilis Strain IIG-Bs-27-39: An Attractive Chassis for Recombinant Protein Production. ACS Synth. Biol. 2024, 13, 2199–2214. [Google Scholar] [CrossRef] [PubMed]
- Geissler, M.; Kühle, I.; Morabbi Heravi, K.; Altenbuchner, J.; Henkel, M.; Hausmann, R. Evaluation of Surfactin Synthesis in a Genome Reduced Bacillus subtilis Strain. AMB Express 2019, 9, 84. [Google Scholar] [CrossRef]
- Yan, P.; Wu, Y.; Yang, L.; Wang, Z.; Chen, T. Engineering Genome-Reduced Bacillus subtilis for Acetoin Production from Xylose. Biotechnol. Lett. 2018, 40, 393–398. [Google Scholar] [CrossRef]
- Arima, K.; Kakinuma, A.; Tamura, G. Surfactin, a Crystalline Peptidelipid Surfactant Produced by Bacillus subtilis: Isolation, Characterization and Its Inhibition of Fibrin Clot Formation. Biochem. Biophys. Res. Commun. 1968, 31, 488–494. [Google Scholar] [CrossRef]
- Abdel-Mawgoud, A.M.; Aboulwafa, M.M.; Hassouna, N.A.H. Characterization of Surfactin Produced by Bacillus subtilis Isolate BS5. Appl. Biochem. Biotechnol. 2008, 150, 289–303. [Google Scholar] [CrossRef]
- Meena, K.R.; Kanwar, S.S. Lipopeptides as the Antifungal and Antibacterial Agents: Applications in Food Safety and Therapeutics. BioMed Res. Int. 2015, 2015. [Google Scholar] [CrossRef]
- Cosmina, P.; Rodriguez, F.; de Ferra, F.; Grandi, G.; Perego, M.; Venema, G.; van Sinderen, D. Sequence and Analysis of the Genetic Locus Responsible for Surfactin Synthesis in Bacillus subtilis. Mol. Microbiol. 1993, 8, 821–831. [Google Scholar] [CrossRef]
- Koglin, A.; Löhr, F.; Bernhard, F.; Rogov, V.V.; Frueh, D.P.; Strieter, E.R.; Mofid, M.R.; Güntert, P.; Wagner, G.; Walsh, C.T.; et al. Structural Basis for the Selectivity of the External Thioesterase of the Surfactin Synthetase. Nature 2008, 454, 907–911. [Google Scholar] [CrossRef]
- Zhi, Y.; Wu, Q.; Xu, Y. Genome and Transcriptome Analysis of Surfactin Biosynthesis in Bacillus Amyloliquefaciens MT45. Sci. Rep. 2017, 7, 40976. [Google Scholar] [CrossRef]
- Finking, R.; Marahiel, M.A. Biosynthesis of Nonribosomal Peptides. Annu. Rev. Microbiol. 2004, 58, 453–488. [Google Scholar] [CrossRef] [PubMed]
- Hamdache, A.; Azarken, R.; Lamarti, A.; Aleu, J.; Collado, I.G. Comparative Genome Analysis of Bacillus spp. and Its Relationship with Bioactive Nonribosomal Peptide Production. Phytochem. Rev. 2013, 12, 685–716. [Google Scholar] [CrossRef]
- Reuter, K. Crystal Structure of the Surfactin Synthetase-Activating Enzyme Sfp: A Prototype of the 4’-Phosphopantetheinyl Transferase Superfamily. EMBO J. 1999, 18, 6823–6831. [Google Scholar] [CrossRef] [PubMed]
- Hamoen, L.W.; Venema, G.; Kuipers, O.P. Controlling Competence in Bacillus subtilis: Shared Use of Regulators. Microbiology 2003, 149, 9–17. [Google Scholar] [CrossRef]
- Auchtung, J.M.; Lee, C.A.; Grossman, A.D. Modulation of the ComA-Dependent Quorum Response in Bacillus subtilis by Multiple Rap Proteins and Phr Peptides. J. Bacteriol. 2006, 188, 5273–5285. [Google Scholar] [CrossRef]
- Serror, P.; Sonenshein, A.L. CodY Is Required for Nutritional Repression of Bacillus subtilis Genetic Competence. J. Bacteriol. 1996, 178, 5910–5915. [Google Scholar] [CrossRef]
- Miras, M.; Dubnau, D. A DegU-P and DegQ-Dependent Regulatory Pathway for the K-State in Bacillus subtilis. Front. Microbiol. 2016, 7, 1868. [Google Scholar] [CrossRef]
- Hayashi, K.; Ohsawa, T.; Kobayashi, K.; Ogasawara, N.; Ogura, M. The H2O2 Stress-Responsive Regulator PerR Positively Regulates SrfA Expression in Bacillus subtilis. J. Bacteriol. 2005, 187, 6659–6667. [Google Scholar] [CrossRef]
- Qi, X.; Liu, W.; He, X.; Du, C. A Review on Surfactin: Molecular Regulation of Biosynthesis. Arch. Microbiol. 2023, 205, 313. [Google Scholar] [CrossRef]
- Chumsakul, O.; Takahashi, H.; Oshima, T.; Hishimoto, T.; Kanaya, S.; Ogasawara, N.; Ishikawa, S. Genome-Wide Binding Profiles of the Bacillus subtilis Transition State Regulator AbrB and Its Homolog Abh Reveals Their Interactive Role in Transcriptional Regulation. Nucleic Acids Res. 2011, 39, 414–428. [Google Scholar] [CrossRef] [PubMed]
- Westers, H.; Dorenbos, R.; Van Dijl, J.M.; Kabel, J.; Flanagan, T.; Devine, K.M.; Jude, F.; Séror, S.J.; Beekman, A.C.; Darmon, E.; et al. Genome Engineering Reveals Large Dispensable Regions in Bacillus subtilis. Mol. Biol. Evol. 2003, 20, 2076–2090. [Google Scholar] [CrossRef] [PubMed]
- Reuß, D.R.; Thürmer, A.; Daniel, R.; Quax, W.J.; Stülke, J. Complete Genome Sequence of Bacillus subtilis subsp. subtilis Strain Δ6. Genome Announc. 2016, 4, 10-1128. [Google Scholar] [CrossRef] [PubMed]
- Vargas-Bautista, C.; Rahlwes, K.; Straight, P. Bacterial Competition Reveals Differential Regulation of the pks Genes by Bacillus subtilis. J. Bacteriol. 2014, 196, 717–728. [Google Scholar] [CrossRef]
- Wu, Q.; Zhi, Y.; Xu, Y. Systematically Engineering the Biosynthesis of a Green Biosurfactant Surfactin by Bacillus subtilis 168. Metab. Eng. 2019, 52, 87–97. [Google Scholar] [CrossRef]
- Altenbuchner, J. Editing of the Bacillus subtilis Genome by the CRISPR-Cas9 System. Appl. Environ. Microbiol. 2016, 82, 5421–5427. [Google Scholar] [CrossRef]
- Park, J.; Bae, S.; Kim, J.-S. Cas-Designer: A web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 2015, 31, 24, 4014–4016. [Google Scholar] [CrossRef]
- Yanisch-Perron, C.; Vieira, J.; Messing, J. Improved M13 Phage Cloning Vectors and Host Strains: Nucleotide Sequences of the M13mpl8 and PUC19 Vectors. Gene 1985, 33, 103–119. [Google Scholar] [CrossRef]
- Bron, S.; Meima, R.; van Dijl, J.M.; Wipat, A.; Harwood, C.R. Molecular Biology and Genetics of Bacillus Species. In Manual of Industrial Microbiology and Biotechnology; Demain, A.L., Davies, J.E., Atlas, R.M., Eds.; American Society for Microbiology: Washington, DC, USA, 1999; pp. 392–416. ISBN 9781555811280. [Google Scholar]
- Miller, G.L. Use of Dinitrosalicylic Acid Reagent for Determination of Reducing Sugar. Anal. Chem. 1959, 31, 426–428. [Google Scholar] [CrossRef]
- de Lima, J.G.O.; Veríssimo, N.V.P.; de Azevedo Lima, C.; Picheli, F.P.; de Paula, A.V.; Santos-Ebinuma, V.D.C. Improvement of Torularhodin Production by Rhodotorula Glutinis through the Stimulation of Physicochemical Stress and Application of the Bioproduct as an Additive in the Food Industry. Bioprocess. Biosyst. Eng. 2025, 48, 543–563. [Google Scholar] [CrossRef]
- Geissler, M.; Oellig, C.; Moss, K.; Schwack, W.; Henkel, M.; Hausmann, R. High-Performance Thin-Layer Chromatography (HPTLC) for the Simultaneous Quantification of the Cyclic Lipopeptides Surfactin, Iturin A and Fengycin in Culture Samples of Bacillus Species. J. Chromatogr. B Analyt Technol. Biomed. Life Sci. 2017, 1044–1045, 214–224. [Google Scholar] [CrossRef] [PubMed]
- Koim-Puchowska, B.; Kłosowski, G.; Mikulski, D.; Menka, A. Evaluation of Various Methods of Selection of B. subtilis Strains Capable of Secreting Surface-Active Compounds. PLoS ONE 2019, 14, e0225108. [Google Scholar] [CrossRef] [PubMed]
- Mohd Isa, M.H.; Shamsudin, N.H.; Al-Shorgani, N.K.N.; Alsharjabi, F.A.; Kalil, M.S. Evaluation of Antibacterial Potential of Biosurfactant Produced by Surfactin-Producing Bacillus Isolated from Selected Malaysian Fermented Foods. Food Biotechnol. 2020, 34, 1–24. [Google Scholar] [CrossRef]
- Yang, H.; Yu, H.; Shen, Z. A Novel High-Throughput and Quantitative Method Based on Visible Color Shifts for Screening Bacillus subtilis THY-15 for Surfactin Production. J. Ind. Microbiol. Biotechnol. 2015, 42, 1139–1147. [Google Scholar] [CrossRef]
- Sun, H.; Bie, X.; Lu, F.; Lu, Y.; Wu, Y.; Lu, Z. Enhancement of Surfactin Production of Bacillus subtilis FmbR by Replacement of the Native Promoter with the Pspac Promoter. Can. J. Microbiol. 2009, 55, 1003–1006. [Google Scholar] [CrossRef]
- Jiao, S.; Li, X.; Yu, H.; Yang, H.; Li, X.; Shen, Z. In Situ Enhancement of Surfactin Biosynthesis in Bacillus subtilis Using Novel Artificial Inducible Promoters. Biotechnol. Bioeng. 2017, 114, 832–842. [Google Scholar] [CrossRef]
- Coutte, F.; Leclère, V.; Béchet, M.; Guez, J.S.; Lecouturier, D.; Chollet-Imbert, M.; Dhulster, P.; Jacques, P. Effect of Pps Disruption and Constitutive Expression of SrfA on Surfactin Productivity, Spreading and Antagonistic Properties of Bacillus subtilis 168 Derivatives. J. Appl. Microbiol. 2010, 109, 480–491. [Google Scholar] [CrossRef]
- Willenbacher, J.; Mohr, T.; Henkel, M.; Gebhard, S.; Mascher, T.; Syldatk, C.; Hausmann, R. Substitution of the Native SrfA Promoter by Constitutive Pveg in Two B. subtilis Strains and Evaluation of the Effect on Surfactin Production. J. Biotechnol. 2016, 224, 14–17. [Google Scholar] [CrossRef]
- Liu, D.; Mao, Z.; Guo, J.; Wei, L.; Ma, H.; Tang, Y.; Chen, T.; Wang, Z.; Zhao, X. Construction, Model-Based Analysis, and Characterization of a Promoter Library for Fine-Tuned Gene Expression in Bacillus subtilis. ACS Synth. Biol. 2018, 7, 1785–1797. [Google Scholar] [CrossRef]
- Corrêa, G.G.; Lins, M.R.D.C.R.; Silva, B.F.; de Paiva, G.B.; Zocca, V.F.B.; Ribeiro, N.V.; Picheli, F.P.; Mack, M.; Pedrolli, D.B. A Modular Autoinduction Device for Control of Gene Expression in Bacillus subtilis. Metab. Eng. 2020, 61, 326–334. [Google Scholar] [CrossRef]
- Sonenshein, A.L. Control of Key Metabolic Intersections in Bacillus subtilis. Nat. Rev. Microbiol. 2007, 5, 917–927. [Google Scholar] [CrossRef] [PubMed]
- Shivers, R.P.; Sonenshein, A.L. Activation of the Bacillus subtilis Global Regulator CodY by Direct Interaction with Branched-Chain Amino Acids. Mol. Microbiol. 2004, 53, 599–611. [Google Scholar] [CrossRef] [PubMed]
- Debarbouille, M.; Gardan, R.; Arnaud, M.; Rapoport, G. Role of BkdR, a Transcriptional Activator of the SigL-Dependent Isoleucine and Valine Degradation Pathway in Bacillus subtilis. J. Bacteriol. 1999, 181, 2059–2066. [Google Scholar] [CrossRef]
- Comella, N.; Grossman, A.D. Conservation of Genes and Processes Controlled by the Quorum Response in Bacteria: Characterization of Genes Controlled by the Quorum-Sensing Transcription Factor ComA in Bacillus subtilis. Mol. Microbiol. 2005, 57, 1159–1174. [Google Scholar] [CrossRef]
- Xia, L.; Hou, Z.; Zhu, F.; Wen, J. Enhancing Surfactin Production in Bacillus subtilis: Insights from Proteomic Analysis of Nitrate-Induced Overproduction and Strategies for Combinatorial Metabolic Engineering. Bioresour. Technol. 2024, 397, 130499. [Google Scholar] [CrossRef]
- Martin-Verstraete, I.; Dkbarbouilk, M.; Klier, A.; Rapoport, G. Levanase Operon of Bacillus subtilis Includes a Fructose-Specific Phosphotransferase System Regulating the Expression of the Operon. J. Mol. Biol. 1990, 214, 657–671. [Google Scholar] [CrossRef]
- Wang, Q.; Yu, H.; Wang, M.; Yang, H.; Shen, Z. Enhanced Biosynthesis and Characterization of Surfactin Isoforms with Engineered Bacillus subtilis through Promoter Replacement and Vitreoscilla Hemoglobin Co-Expression. Process Biochem. 2018, 70, 36–44. [Google Scholar] [CrossRef]
- Wang, C.; Cao, Y.; Wang, Y.; Sun, L.; Song, H. Enhancing Surfactin Production by Using Systematic CRISPRi Repression to Screen Amino Acid Biosynthesis Genes in Bacillus subtilis. Microb. Cell Fact. 2019, 18, 90. [Google Scholar] [CrossRef]
- Wang, M.; Yu, H.; Shen, Z. Antisense RNA-Based Strategy for Enhancing Surfactin Production in Bacillus subtilis TS1726 via Overexpression of the Unconventional Biotin Carboxylase II to Enhance ACCase Activity. ACS Synth. Biol. 2019, 8, 251–256. [Google Scholar] [CrossRef]
- Aguilar Suárez, R.; Stülke, J.; van Dijl, J.M. Less Is More: Toward a Genome-Reduced Bacillus Cell Factory for “Difficult Proteins”. ACS Synth. Biol. 2019, 8, 99–108. [Google Scholar] [CrossRef]
- Wenzel, M.; Altenbuchner, J. Development of a Markerless Gene Deletion System for Bacillus subtilis Based on the Mannose Phosphoenolpyruvate-Dependent Phosphotransferase System. Microbiology 2015, 161, 1942–1949. [Google Scholar] [CrossRef]
- Liu, F.-F.; Liu, Y.-F.; Qiao, Y.-W.; Guo, Y.-Z.; Kuang, F.-Y.; Lin, X.-Q.; Liu, J.-F.; Yang, S.-Z.; Zhang, H.-Z.; Sand, W.; et al. Improvement Surfactin Production by Substitution of Promoters in Bacillus subtilis TD7. Appl. Environ. Biotechnol. 2021, 6, 31–41. [Google Scholar] [CrossRef]
- Coutte, F.; Niehren, J.; Dhali, D.; John, M.; Versari, C.; Jacques, P. Modeling Leucine’s Metabolic Pathway and Knockout Prediction Improving the Production of Surfactin, a Biosurfactant from Bacillus subtilis. Biotechnol. J. 2015, 10, 1216–1234. [Google Scholar] [CrossRef]
- Dhali, D.; Coutte, F.; Arias, A.A.; Auger, S.; Bidnenko, V.; Chataigné, G.; Lalk, M.; Niehren, J.; de Sousa, J.; Versari, C.; et al. Genetic Engineering of the Branched Fatty Acid Metabolic Pathway of Bacillus subtilis for the Overproduction of Surfactin C14 Isoform. Biotechnol. J. 2017, 12, 1600574. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Liu, Y.; Lin, F.; Lu, Z.; Lu, Y. CodY, ComA, DegU and Spo0A Controlling Lipopeptides Biosynthesis in Bacillus amyloliquefaciens FmbJ. J. Appl. Microbiol. 2021, 131, 1289–1304. [Google Scholar] [CrossRef]
- Su, Y.; Gao, L.; Li, C.; Wang, L.; Zhou, H.; Zhang, C.; Xia, X. Regulation Mechanism and Bioactivity Characteristic of Surfactin Homologues with C14 and C15 Fatty Acid Chains. Microb. Cell Fact. 2024, 23, 94. [Google Scholar] [CrossRef]
| Plasmid | Characteristics | Reference |
|---|---|---|
| pJOE8999 | Base plasmid: pUCori, rep pE194ts, kanR, PmanP-cas9, PvanP-lacPOZ’-sgRNA | [26] |
| pEBScas9.1 | pJOE8999, PvanP-gRNA_1, sfp * | This study |
| pEBScas9.2 | pJOE8999, PvanP-gRNA_2, hxlR*-S0-R6-srfAA * | This study |
| pEBScas9.3 | pJOE8999, PvanP-gRNA_2, hxlR*-TP2-srfAA * | This study |
| pEBScas9.4 | pJOE8999, PvanP-gRNA_3, S0*-PsrfA-R6-srfAA * | This study |
| pEBScas9.5 | pJOE8999, PmanP-cas9, PvanP-gRNA_4, hslU *-flgB * | This study |
| pEBScas9.6 | pJOE8999, PvanP-gRNA_5, comP *-PsrfA-comA | This study |
| Strain | Characteristics | Reference |
|---|---|---|
| E. coli JM109 | recA1, endA1, gyrA96, thi-1, hsdR17(rK−, mk+), mcrA, supE44, gyrA96, relA1, λ−, Δ(lac-proAB), F’(traD36, proAB +, lacIq, (ΔlacZ)M15) | [28] |
| E. coli TOP10 | mcrA, Δ(mrr-hsdRMS-mcrBC), Phi80lacZ(del)M15, ΔlacX74, deoR, recA1, araD139, Δ(ara-leu)7697, galU, galK, rpsL(SmR), endA1, nupG | Invitrogen |
| B. subtilis ATCC 21332 | Wild-type, surfactin producer | BGSCID 3A22 * |
| B. subtilis 168 | B. subtilis Marburg 168, trpC2 | BGSCID 1A1 * |
| B. subtilis Δ6 | Genome-reduced B. subtilis 168 trpC2, Δskin, ICEBs1(0), Δprophage1, Δprophage3, ΔSPbeta, ΔPBSX, Δpks::CmR | BGSID 1A1299 * |
| B. subtilis CS01 | B. subtilis 168 derivate sfp+ | This study |
| B. subtilis D6S01 | B. subtilis Δ6 derivate sfp+ | This study |
| B. subtilis D6S02 | B. subtilis D6S01 derivate sfp+, S0-R6-srfAABCD | This study |
| B. subtilis D6S03 | B. subtilis D6S01 derivate sfp+, TP2-srfAABCD | This study |
| B. subtilis D6S04 | B. subtilis D6S02 derivate sfp+, S0-PsrfA-R6-srfAABCD | This study |
| B. subtilis D6S05 | B. subtilis D6S01 derivate sfp+, ΔcodY | This study |
| B. subtilis D6S06 | B. subtilis D6S05 derivate sfp+, ΔcodY, PsrfA-comA | This study |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zocca, V.F.B.; Barban, F.d.O.; Kundlatsch, G.E.; Tavares, L.F.; Picheli, F.P.; Moura, A.C.d.S.; Pedrolli, D.B. Bacillus subtilis Genome Reduction Improves Surfactin Production. Fermentation 2025, 11, 508. https://doi.org/10.3390/fermentation11090508
Zocca VFB, Barban FdO, Kundlatsch GE, Tavares LF, Picheli FP, Moura ACdS, Pedrolli DB. Bacillus subtilis Genome Reduction Improves Surfactin Production. Fermentation. 2025; 11(9):508. https://doi.org/10.3390/fermentation11090508
Chicago/Turabian StyleZocca, Vitória Fernanda Bertolazzi, Fabiane de Oliveira Barban, Guilherme Engelberto Kundlatsch, Leonardo Ferro Tavares, Flávio Pereira Picheli, Adriana Candido da Silva Moura, and Danielle Biscaro Pedrolli. 2025. "Bacillus subtilis Genome Reduction Improves Surfactin Production" Fermentation 11, no. 9: 508. https://doi.org/10.3390/fermentation11090508
APA StyleZocca, V. F. B., Barban, F. d. O., Kundlatsch, G. E., Tavares, L. F., Picheli, F. P., Moura, A. C. d. S., & Pedrolli, D. B. (2025). Bacillus subtilis Genome Reduction Improves Surfactin Production. Fermentation, 11(9), 508. https://doi.org/10.3390/fermentation11090508






