Long Non-Coding RNAs in Cardiac and Pulmonary Fibroblasts and Fibrosis
Abstract
:1. Introduction
2. LncRNAs in Cardiac Fibrosis
3. LncRNAs in Pulmonary Fibrosis
4. Discussions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Liu, S.J.; Nowakowski, T.J.; Pollen, A.A.; Lui, J.H.; Horlbeck, M.A.; Attenello, F.J.; He, D.; Weissman, J.S.; Kriegstein, A.R.; Diaz, A.A.; et al. Single-cell analysis of long non-coding RNAs in the developing human neocortex. Genome Biol. 2016, 17, 67. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, X.; Chen, K.; Cuevas-Diaz Duran, R.; You, Y.; Sloan, S.A.; Zhang, Y.; Zong, S.; Cao, Q.; Barres, B.A.; Wu, J.Q. Comprehensive Identification of Long Non-coding RNAs in Purified Cell Types from the Brain Reveals Functional LncRNA in OPC Fate Determination. PLoS Genet. 2015, 11, e1005669. [Google Scholar] [CrossRef] [Green Version]
- Volders, P.J.; Anckaert, J.; Verheggen, K.; Nuytens, J.; Martens, L.; Mestdagh, P.; Vandesompele, J. LNCipedia 5: Towards a reference set of human long non-coding RNAs. Nucleic Acids Res. 2019, 47, D135–D139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Uszczynska-Ratajczak, B.; Lagarde, J.; Frankish, A.; Guigo, R.; Johnson, R. Towards a complete map of the human long non-coding RNA transcriptome. Nat. Rev. Genet. 2018, 19, 535–548. [Google Scholar] [CrossRef] [PubMed]
- Karakas, D.; Ozpolat, B. The Role of LncRNAs in Translation. Noncoding RNA 2021, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Sebastian-delaCruz, M.; Gonzalez-Moro, I.; Olazagoitia-Garmendia, A.; Castellanos-Rubio, A.; Santin, I. The Role of lncRNAs in Gene Expression Regulation through mRNA Stabilization. Noncoding RNA 2021, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, J.C.R.; Acuna, S.M.; Aoki, J.I.; Floeter-Winter, L.M.; Muxel, S.M. Long Non-Coding RNAs in the Regulation of Gene Expression: Physiology and Disease. Noncoding RNA 2019, 5, 17. [Google Scholar] [CrossRef] [Green Version]
- Marzec, J.; Nadadur, S. Inflammation resolution in environmental pulmonary health and morbidity. Toxicol. Appl. Pharmacol. 2022, 449, 116070. [Google Scholar] [CrossRef]
- Murtha, L.A.; Schuliga, M.J.; Mabotuwana, N.S.; Hardy, S.A.; Waters, D.W.; Burgess, J.K.; Knight, D.A.; Boyle, A.J. The Processes and Mechanisms of Cardiac and Pulmonary Fibrosis. Front. Physiol. 2017, 8, 777. [Google Scholar] [CrossRef] [Green Version]
- El Ayadi, A.; Jay, J.W.; Prasai, A. Current Approaches Targeting the Wound Healing Phases to Attenuate Fibrosis and Scarring. Int. J. Mol. Sci. 2020, 21, 1105. [Google Scholar] [CrossRef] [Green Version]
- Darby, I.A.; Hewitson, T.D. Fibroblast differentiation in wound healing and fibrosis. Int. Rev. Cytol. 2007, 257, 143–179. [Google Scholar] [CrossRef] [PubMed]
- Wynn, T.A.; Ramalingam, T.R. Mechanisms of fibrosis: Therapeutic translation for fibrotic disease. Nat. Med. 2012, 18, 1028–1040. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wynn, T.A. Cellular and molecular mechanisms of fibrosis. J. Pathol. 2008, 214, 199–210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.; Yang, X.; Gui, S.; Yang, F.; Cao, Z.; Cheng, R.; Xia, X.; Li, C. The Roles and Mechanisms of lncRNAs in Liver Fibrosis. Front. Pharmacol. 2021, 12, 779606. [Google Scholar] [CrossRef]
- He, Y.; Wang, W.; Jiang, P.; Yang, L.; Guo, Q.; Xiang, J.; Gao, Y.; Wang, Y.; Chen, R. Long Non-Coding RNAs in Oral Submucous Fibrosis: Their Functional Mechanisms and Recent Research Progress. J. Inflamm. Res. 2021, 14, 5787–5800. [Google Scholar] [CrossRef]
- Ghafouri-Fard, S.; Abak, A.; Talebi, S.F.; Shoorei, H.; Branicki, W.; Taheri, M.; Akbari Dilmaghani, N. Role of miRNA and lncRNAs in organ fibrosis and aging. Biomed. Pharmacother. 2021, 143, 112132. [Google Scholar] [CrossRef]
- Xia, W.; He, Y.; Gan, Y.; Zhang, B.; Dai, G.; Ru, F.; Jiang, Z.; Chen, Z.; Chen, X. Long Non-coding RNA: An Emerging Contributor and Potential Therapeutic Target in Renal Fibrosis. Front. Genet. 2021, 12, 682904. [Google Scholar] [CrossRef]
- Zhang, H.; Song, M.; Guo, J.; Ma, J.; Qiu, M.; Yang, Z. The function of non-coding RNAs in idiopathic pulmonary fibrosis. Open. Med. 2021, 16, 481–490. [Google Scholar] [CrossRef]
- Zhang, S.; Chen, H.; Yue, D.; Blackwell, T.S.; Lv, C.; Song, X. Long non-coding RNAs: Promising new targets in pulmonary fibrosis. J. Gene. Med. 2021, 23, e3318. [Google Scholar] [CrossRef]
- Guo, Y.; Wang, L.; Gou, R.; Tang, L.; Liu, P. Noncoding RNAs in peritoneal fibrosis: Background, Mechanism, and Therapeutic Approach. Biomed. Pharmacother. 2020, 129, 110385. [Google Scholar] [CrossRef]
- Frangogiannis, N.G. Cardiac fibrosis. Cardiovasc. Res. 2021, 117, 1450–1488. [Google Scholar] [CrossRef] [PubMed]
- Travers, J.G.; Kamal, F.A.; Robbins, J.; Yutzey, K.E.; Blaxall, B.C. Cardiac Fibrosis: The Fibroblast Awakens. Circ. Res. 2016, 118, 1021–1040. [Google Scholar] [CrossRef] [Green Version]
- Micheletti, R.; Plaisance, I.; Abraham, B.J.; Sarre, A.; Ting, C.C.; Alexanian, M.; Maric, D.; Maison, D.; Nemir, M.; Young, R.A.; et al. The long noncoding RNA Wisper controls cardiac fibrosis and remodeling. Sci. Transl. Med. 2017, 9, eaai9118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, F.; Fu, X.; Kataoka, M.; Liu, N.; Wang, Y.; Gao, F.; Liang, T.; Dong, X.; Pei, J.; Hu, X.; et al. Long noncoding RNA Cfast regulates cardiac fibrosis. Mol. Ther. Nucleic Acids 2021, 23, 377–392. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Nie, X.; Yuan, S.; Li, H.; Fan, J.; Li, C.; Sun, Y.; Zhao, Y.; Hou, H.; Wang, D.W.; et al. Circulating Long Non-coding RNA ENST00000507296 Is a Prognostic Indicator in Patients with Dilated Cardiomyopathy. Mol. Ther. Nucleic Acids 2019, 16, 82–90. [Google Scholar] [CrossRef] [Green Version]
- Choong, O.K.; Chen, C.Y.; Zhang, J.; Lin, J.H.; Lin, P.J.; Ruan, S.C.; Kamp, T.J.; Hsieh, P.C.H. Hypoxia-induced H19/YB-1 cascade modulates cardiac remodeling after infarction. Theranostics 2019, 9, 6550–6567. [Google Scholar] [CrossRef]
- Piccoli, M.T.; Gupta, S.K.; Viereck, J.; Foinquinos, A.; Samolovac, S.; Kramer, F.L.; Garg, A.; Remke, J.; Zimmer, K.; Batkai, S.; et al. Inhibition of the Cardiac Fibroblast-Enriched lncRNA Meg3 Prevents Cardiac Fibrosis and Diastolic Dysfunction. Circ. Res. 2017, 121, 575–583. [Google Scholar] [CrossRef]
- Ge, Z.; Yin, C.; Li, Y.; Tian, D.; Xiang, Y.; Li, Q.; Tang, Y.; Zhang, Y. Long noncoding RNA NEAT1 promotes cardiac fibrosis in heart failure through increased recruitment of EZH2 to the Smad7 promoter region. J. Transl. Med. 2022, 20, 7. [Google Scholar] [CrossRef]
- Hao, K.; Lei, W.; Wu, H.; Wu, J.; Yang, Z.; Yan, S.; Lu, X.A.; Li, J.; Xia, X.; Han, X.; et al. LncRNA-Safe contributes to cardiac fibrosis through Safe-Sfrp2-HuR complex in mouse myocardial infarction. Theranostics 2019, 9, 7282–7297. [Google Scholar] [CrossRef]
- Luo, S.; Zhang, M.; Wu, H.; Ding, X.; Li, D.; Dong, X.; Hu, X.; Su, S.; Shang, W.; Wu, J.; et al. SAIL: A new conserved anti-fibrotic lncRNA in the heart. Basic Res. Cardiol. 2021, 116, 15. [Google Scholar] [CrossRef]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cifuentes-Rojas, C.; Hernandez, A.J.; Sarma, K.; Lee, J.T. Regulatory interactions between RNA and polycomb repressive complex 2. Mol. Cell 2014, 55, 171–185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davidovich, C.; Wang, X.; Cifuentes-Rojas, C.; Goodrich, K.J.; Gooding, A.R.; Lee, J.T.; Cech, T.R. Toward a consensus on the binding specificity and promiscuity of PRC2 for RNA. Mol. Cell 2015, 57, 552–558. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davidovich, C.; Zheng, L.; Goodrich, K.J.; Cech, T.R. Promiscuous RNA binding by Polycomb repressive complex 2. Nat. Struct. Mol. Biol. 2013, 20, 1250–1257. [Google Scholar] [CrossRef] [Green Version]
- Kaneko, S.; Son, J.; Shen, S.S.; Reinberg, D.; Bonasio, R. PRC2 binds active promoters and contacts nascent RNAs in embryonic stem cells. Nat. Struct. Mol. Biol. 2013, 20, 1258–1264. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Goodrich, K.J.; Gooding, A.R.; Naeem, H.; Archer, S.; Paucek, R.D.; Youmans, D.T.; Cech, T.R.; Davidovich, C. Targeting of Polycomb Repressive Complex 2 to RNA by Short Repeats of Consecutive Guanines. Mol. Cell 2017, 65, 1056–1067.e1055. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Liu, F.; Zhou, L.Y.; Long, B.; Yuan, S.M.; Wang, Y.; Liu, C.Y.; Sun, T.; Zhang, X.J.; Li, P.F. The long noncoding RNA CHRF regulates cardiac hypertrophy by targeting miR-489. Circ. Res. 2014, 114, 1377–1388. [Google Scholar] [CrossRef] [Green Version]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Chakraborty, C.; Sharma, A.R.; Sharma, G.; Lee, S.S. Therapeutic advances of miRNAs: A preclinical and clinical update. J. Adv. Res. 2021, 28, 127–138. [Google Scholar] [CrossRef]
- Kotowska-Zimmer, A.; Pewinska, M.; Olejniczak, M. Artificial miRNAs as therapeutic tools: Challenges and opportunities. Wiley Interdiscip. Rev. RNA 2021, 12, e1640. [Google Scholar] [CrossRef]
- Zhou, H.; Tang, W.; Yang, J.; Peng, J.; Guo, J.; Fan, C. MicroRNA-Related Strategies to Improve Cardiac Function in Heart Failure. Front. Cardiovasc. Med. 2021, 8, 773083. [Google Scholar] [CrossRef] [PubMed]
- Pottier, N.; Cauffiez, C.; Perrais, M.; Barbry, P.; Mari, B. FibromiRs: Translating molecular discoveries into new anti-fibrotic drugs. Trends Pharmacol. Sci. 2014, 35, 119–126. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Tang, C.; Huang, B.; Gu, L.; Zhou, J.; Mo, Z.; Liu, C.; Liu, Y. LncRNA H19 Drives Proliferation of Cardiac Fibroblasts and Collagen Production via Suppression of the miR-29a-3p/miR-29b-3p-VEGFA/TGF-beta Axis. Mol. Cells 2022, 45, 122–133. [Google Scholar] [CrossRef] [PubMed]
- Kriegel, A.J.; Liu, Y.; Fang, Y.; Ding, X.; Liang, M. The miR-29 family: Genomics, cell biology, and relevance to renal and cardiovascular injury. Physiol. Genom. 2012, 44, 237–244. [Google Scholar] [CrossRef]
- Zhu, Y.; Feng, Z.; Jian, Z.; Xiao, Y. Long noncoding RNA TUG1 promotes cardiac fibroblast transformation to myofibroblasts via miR29c in chronic hypoxia. Mol. Med. Rep. 2018, 18, 3451–3460. [Google Scholar] [CrossRef] [Green Version]
- Guo, Y.; Sun, Z.; Chen, M.; Lun, J. LncRNA TUG1 Regulates Proliferation of Cardiac Fibroblast via the miR-29b-3p/TGF-beta1 Axis. Front. Cardiovasc. Med. 2021, 8, 646806. [Google Scholar] [CrossRef]
- Zhu, C.; Zhang, H.; Wei, D.; Sun, Z. Silencing lncRNA GAS5 alleviates apoptosis and fibrosis in diabetic cardiomyopathy by targeting miR-26a/b-5p. Acta. Diabetol. 2021, 58, 1491–1501. [Google Scholar] [CrossRef]
- Han, Y.; Wu, N.; Xia, F.; Liu, S.; Jia, D. Long noncoding RNA GAS5 regulates myocardial ischemiareperfusion injury through the PI3K/AKT apoptosis pathway by sponging miR5325p. Int. J. Mol. Med. 2020, 45, 858–872. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.C.; Xia, L.; Jiang, Y.; Wu, D.Q.; Liu, S.C.; Zhou, X.N.; Zhang, F.X. Effect of lncRNA GAS5 on rats with acute myocardial infarction through regulating miR-21. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 8573–8579. [Google Scholar] [CrossRef]
- Tao, H.; Zhang, J.G.; Qin, R.H.; Dai, C.; Shi, P.; Yang, J.J.; Deng, Z.Y.; Shi, K.H. LncRNA GAS5 controls cardiac fibroblast activation and fibrosis by targeting miR-21 via PTEN/MMP-2 signaling pathway. Toxicology 2017, 386, 11–18. [Google Scholar] [CrossRef]
- Zhou, Z.; Chen, J.; Huang, Y.; Liu, D.; Chen, S.; Qin, S. Long Noncoding RNA GAS5: A New Factor Involved in Bone Diseases. Front. Cell Dev. Biol. 2021, 9, 807419. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Xia, F.; Wang, Y.; Zhu, L.; Li, Y.; Jia, D.; Gao, Y.; Shi, U.; Zhang, C.; He, Y.; et al. The Role of LncRNA TUG1 in Obesity-Related Diseases. Mini Rev. Med. Chem. 2022, 22, 1305–1313. [Google Scholar] [CrossRef] [PubMed]
- Filippova, E.A.; Fridman, M.V.; Burdennyy, A.M.; Loginov, V.I.; Pronina, I.V.; Lukina, S.S.; Dmitriev, A.A.; Braga, E.A. Long Noncoding RNA GAS5 in Breast Cancer: Epigenetic Mechanisms and Biological Functions. Int. J. Mol. Sci. 2021, 22, 6810. [Google Scholar] [CrossRef]
- Tan, X.; Jiang, H.; Fang, Y.; Han, D.; Guo, Y.; Wang, X.; Gong, X.; Hong, W.; Tu, J.; Wei, W. The essential role of long non-coding RNA GAS5 in glioma: Interaction with microRNAs, chemosensitivity and potential as a biomarker. J. Cancer 2021, 12, 224–231. [Google Scholar] [CrossRef] [PubMed]
- Lambrou, G.I.; Hatziagapiou, K.; Zaravinos, A. The Non-Coding RNA GAS5 and Its Role in Tumor Therapy-Induced Resistance. Int. J. Mol. Sci. 2020, 21, 7633. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Qi, Y.; Qu, J.; Gai, L.; Shi, Y.; Yuan, C. Pathophysiological Functions of the lncRNA TUG1. Curr. Pharm. Des. 2020, 26, 688–700. [Google Scholar] [CrossRef]
- Zhou, H.; Sun, L.; Wan, F. Molecular mechanisms of TUG1 in the proliferation, apoptosis, migration and invasion of cancer cells. Oncol. Lett. 2019, 18, 4393–4402. [Google Scholar] [CrossRef] [Green Version]
- Wijsenbeek, M.; Cottin, V. Spectrum of Fibrotic Lung Diseases. N. Engl. J. Med. 2020, 383, 958–968. [Google Scholar] [CrossRef]
- Hung, C.F. Origin of Myofibroblasts in Lung Fibrosis. Curr. Tissue Microenviron. Rep. 2020, 1, 155–162. [Google Scholar] [CrossRef]
- Yang, J.; Antin, P.; Berx, G.; Blanpain, C.; Brabletz, T.; Bronner, M.; Campbell, K.; Cano, A.; Casanova, J.; Christofori, G.; et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2020, 21, 341–352. [Google Scholar] [CrossRef] [Green Version]
- Reinert, T.; Baldotto, C.S.d.R.; Nunes, F.A.P.; Scheliga, A.A.d.S. Bleomycin-Induced Lung Injury. J. Cancer Res. 2013, 2013, 480608. [Google Scholar] [CrossRef] [Green Version]
- Moeller, A.; Ask, K.; Warburton, D.; Gauldie, J.; Kolb, M. The bleomycin animal model: A useful tool to investigate treatment options for idiopathic pulmonary fibrosis? Int. J. Biochem. Cell Biol. 2008, 40, 362–382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, J.; Jin, T.; Su, W.; Guo, Y.; Niu, Z.; Guo, J.; Li, L.; Wang, J.; Ma, L.; Yu, T.; et al. The long non-coding RNA PFI protects against pulmonary fibrosis by interacting with splicing regulator SRSF1. Cell Death Differ. 2021, 28, 2916–2930. [Google Scholar] [CrossRef]
- Savary, G.; Dewaeles, E.; Diazzi, S.; Buscot, M.; Nottet, N.; Fassy, J.; Courcot, E.; Henaoui, I.S.; Lemaire, J.; Martis, N.; et al. The Long Noncoding RNA DNM3OS Is a Reservoir of FibromiRs with Major Functions in Lung Fibroblast Response to TGF-beta and Pulmonary Fibrosis. Am. J. Respir. Crit. Care Med 2019, 200, 184–198. [Google Scholar] [CrossRef] [PubMed]
- Senavirathna, L.K.; Liang, Y.; Huang, C.; Yang, X.; Bamunuarachchi, G.; Xu, D.; Dang, Q.; Sivasami, P.; Vaddadi, K.; Munteanu, M.C.; et al. Long Noncoding RNA FENDRR Inhibits Lung Fibroblast Proliferation via a Reduction of beta-Catenin. Int. J. Mol. Sci. 2021, 22, 8536. [Google Scholar] [CrossRef]
- Huang, C.; Liang, Y.; Zeng, X.; Yang, X.; Xu, D.; Gou, X.; Sathiaseelan, R.; Senavirathna, L.K.; Wang, P.; Liu, L. Long Noncoding RNA FENDRR Exhibits Antifibrotic Activity in Pulmonary Fibrosis. Am. J. Respir. Cell Mol. Biol. 2020, 62, 440–453. [Google Scholar] [CrossRef]
- Chen, L.; Yang, Y.; Yue, R.; Peng, X.; Yu, H.; Huang, X. Exosomes derived from hypoxia-induced alveolar epithelial cells stimulate interstitial pulmonary fibrosis through a HOTAIRM1-dependent mechanism. Lab. Investig. 2022. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, H.; Chen, H.; Li, H.; Xu, P.; Liu, B.; Zhang, Q.; Lv, C.; Song, X. ATF3 -activated accelerating effect of LINC00941/lncIAPF on fibroblast-to-myofibroblast differentiation by blocking autophagy depending on ELAVL1/HuR in pulmonary fibrosis. Autophagy 2022, 1–20. [Google Scholar] [CrossRef]
- Song, X.; Xu, P.; Meng, C.; Song, C.; Blackwell, T.S.; Li, R.; Li, H.; Zhang, J.; Lv, C. lncITPF Promotes Pulmonary Fibrosis by Targeting hnRNP-L Depending on Its Host Gene ITGBL1. Mol. Ther. 2019, 27, 380–393. [Google Scholar] [CrossRef] [Green Version]
- Huang, Z.W.; Tian, L.H.; Yang, B.; Guo, R.M. Long Noncoding RNA H19 Acts as a Competing Endogenous RNA to Mediate CTGF Expression by Sponging miR-455 in Cardiac Fibrosis. DNA Cell Biol. 2017, 36, 759–766. [Google Scholar] [CrossRef]
- Zhang, B.F.; Jiang, H.; Chen, J.; Hu, Q.; Yang, S.; Liu, X.P.; Liu, G. LncRNA H19 ameliorates myocardial infarction-induced myocardial injury and maladaptive cardiac remodelling by regulating KDM3A. J. Cell Mol. Med. 2020, 24, 1099–1115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, Y.; He, R.; An, J.; Deng, P.; Huang, L.; Yang, W. The effect of H19-miR-29b interaction on bleomycin-induced mouse model of idiopathic pulmonary fibrosis. Biochem. Biophys. Res. Commun. 2016, 479, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Mu, X.; Wang, H.; Li, H. Silencing of long noncoding RNA H19 alleviates pulmonary injury, inflammation, and fibrosis of acute respiratory distress syndrome through regulating the microRNA-423-5p/FOXA1 axis. Exp. Lung Res. 2021, 47, 183–197. [Google Scholar] [CrossRef] [PubMed]
- Lu, Q.; Guo, Z.; Xie, W.; Jin, W.; Zhu, D.; Chen, S.; Ren, T. The lncRNA H19 Mediates Pulmonary Fibrosis by Regulating the miR-196a/COL1A1 Axis. Inflammation 2018, 41, 896–903. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Han, L.; Yan, W.; Ji, X.; Han, R.; Yang, J.; Yuan, J.; Ni, C. miR-489 inhibits silica-induced pulmonary fibrosis by targeting MyD88 and Smad3 and is negatively regulated by lncRNA CHRF. Sci. Rep. 2016, 6, 30921. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mo, R.; Li, J.; Chen, Y.; Ding, Y. lncRNA GAS5 promotes pyroptosis in COPD by functioning as a ceRNA to regulate the miR2233p/NLRP3 axis. Mol. Med. Rep. 2022, 26, 1–11. [Google Scholar] [CrossRef]
- Che, H.; Wang, Y.; Li, H.; Li, Y.; Sahil, A.; Lv, J.; Liu, Y.; Yang, Z.; Dong, R.; Xue, H.; et al. Melatonin alleviates cardiac fibrosis via inhibiting lncRNA MALAT1/miR-141-mediated NLRP3 inflammasome and TGF-beta1/Smads signaling in diabetic cardiomyopathy. FASEB J. 2020, 34, 5282–5298. [Google Scholar] [CrossRef]
- Yao, M.Y.; Zhang, W.H.; Ma, W.T.; Liu, Q.H.; Xing, L.H.; Zhao, G.F. Long non-coding RNA MALAT1 exacerbates acute respiratory distress syndrome by upregulating ICAM-1 expression via microRNA-150-5p downregulation. Aging 2020, 12, 6570–6585. [Google Scholar] [CrossRef]
- Yan, W.; Wu, Q.; Yao, W.; Li, Y.; Liu, Y.; Yuan, J.; Han, R.; Yang, J.; Ji, X.; Ni, C. MiR-503 modulates epithelial-mesenchymal transition in silica-induced pulmonary fibrosis by targeting PI3K p85 and is sponged by lncRNA MALAT1. Sci. Rep. 2017, 7, 11313. [Google Scholar] [CrossRef] [Green Version]
- Dai, H.; Zhao, N.; Liu, H.; Zheng, Y.; Zhao, L. LncRNA Nuclear-Enriched Abundant Transcript 1 Regulates Atrial Fibrosis via the miR-320/NPAS2 Axis in Atrial Fibrillation. Front. Pharmacol. 2021, 12, 647124. [Google Scholar] [CrossRef]
- Liu, Y.; Lu, F.A.; Wang, L.; Wang, Y.F.; Wu, C.F. Long noncoding RNA NEAT1 promotes pulmonary fibrosis by regulating the microRNA4553p/SMAD3 axis. Mol. Med. Rep. 2021, 23, 1. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Hu, S.; Yu, J.; Li, W.; Li, P.; Huang, H. CREB1 transcription-activated lncRNA PVT1 promotes cardiac fibrosis via miR-145/HCN1 axis. Int. J. Cardiol. 2022, 353, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Dai, Q.; Hong, Y.; Li, J. PVT1 knockdown inhibited the biological behavior of LPS-induced cardiac fibroblasts by regulating miR-24. Genes. Genom. 2021, 43, 1003–1009. [Google Scholar] [CrossRef] [PubMed]
- Cao, F.; Li, Z.; Ding, W.M.; Yan, L.; Zhao, Q.Y. LncRNA PVT1 regulates atrial fibrosis via miR-128-3p-SP1-TGF-beta1-Smad axis in atrial fibrillation. Mol. Med. 2019, 25, 7. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Sun, W.; Pan, H.; Yuan, J.; Xu, Q.; Xu, T.; Li, P.; Cheng, D.; Liu, Y.; Ni, C. LncRNA-PVT1 activates lung fibroblasts via miR-497-5p and is facilitated by FOXM1. Ecotoxicol. Environ. Saf. 2021, 213, 112030. [Google Scholar] [CrossRef]
- Ilieva, M.; Miller, H.E.; Agarwal, A.; Paulus, G.K.; Madsen, J.H.; Bishop, A.J.R.; Kauppinen, S.; Uchida, S. FibroDB: Expression Analysis of Protein-Coding and Long Non-Coding RNA Genes in Fibrosis. Noncoding RNA 2022, 8, 13. [Google Scholar] [CrossRef]
- Ramanathan, M.; Porter, D.F.; Khavari, P.A. Methods to study RNA-protein interactions. Nat. Methods 2019, 16, 225–234. [Google Scholar] [CrossRef]
- Hosen, M.R.; Militello, G.; Weirick, T.; Ponomareva, Y.; Dassanayaka, S.; Moore, J.B.T.; Doring, C.; Wysoczynski, M.; Jones, S.P.; Dimmeler, S.; et al. Airn Regulates Igf2bp2 Translation in Cardiomyocytes. Circ. Res. 2018, 122, 1347–1353. [Google Scholar] [CrossRef]
| Name of LncRNA | Study Model | Function | Reference |
|---|---|---|---|
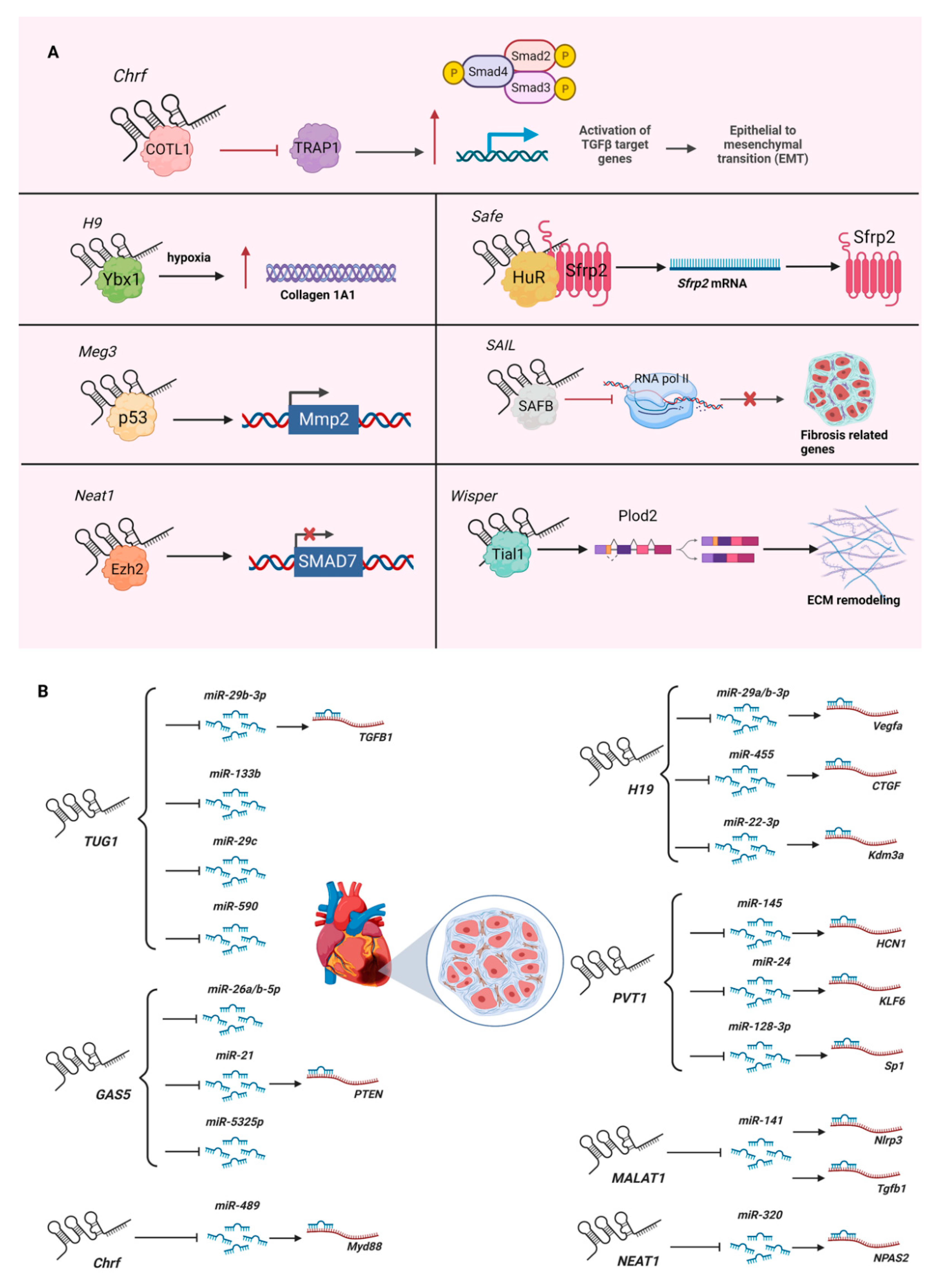
| Cfast (cardiac fibroblast-associated transcript; also known as AK048087) | Knockdown by siRNA in primary neonatal murine cardiac fibroblasts; knockdown by lentiviral shRNA in infarcted (permanent ligation of left anterior descending coronary artery) or hypertrophied murine hearts (injection of isoproterenol). | Binds COTL1 (coactosin-like F-actin-binding protein 1) to competitively inhibit its interaction with TRAP1 (TNF receptor-associated protein 1), which enhances TGF-β signaling by augmenting SMAD2/SMAD4 complex formation. | [24] |
| H19 (H19 imprinted maternally expressed transcript) | Overexpression by AAV9 (adeno-associated virus serotype 9) in infarcted murine hearts (permanent ligation of left anterior descending coronary artery) or intraperitoneal injection into postnatal 8–12-day-old mice; myocardial infarction in H19-knockout mice generated using the CRISPR/Cas9 system; knockdown by siRNA or overexpression in primary adult and neonatal murine cardiac fibroblasts, hiPSC-CFs (human-induced-pluripotent-stem-cell-derived cardiac fibroblasts), and NIH3T3 cells. | Binds Ybx1 (Y box protein 1) under hypoxia to cause de-repression of collagen 1A1 expression. | [26] |
| Meg3 (maternally expressed 3) | Knockdown by GapmeR in primary adult murine cardiac fibroblasts or hypertrophied (transverse aortic constriction) murine hearts. | Interacts with P53 to regulate the expression of Mmp2 (matrix metallopeptidase 2). | [27] |
| Neat1 (nuclear paraspeckle assembly transcript 1) | Knockdown by siRNA or overexpression by adenovirus in primary neonatal murine cardiac fibroblasts treated with TGF-β1. | Binds Ezh2 to recruit it to the promoter of Smad7 to inhibit its expression. | [28] |
| Safe (AK137033) | Knockdown by shRNA or CRISPR/Cas9-mediated knockout in primary adult murine cardiac fibroblasts treated with TGF-β; knockdown by lentiviral shRNA or overexpression by lentivirus in infarcted murine hearts (permanent ligation of left anterior descending coronary artery). | Promotes the Safe-Sfrp2-HuR (Elavl1, ELAV (embryonic lethal, abnormal vision)-like 1 (Hu antigen R)) complex-mediated Sfrp2 (secreted frizzled related protein 2) mRNA stability and protein expression. | [29] |
| SAIL (scaffold attachment factor B interacting lncRNA; Gm19522 (predicted gene, 19522)) | Knockdown by siRNA or overexpression by plasmids in primary neonatal murine cardiac fibroblasts or human cardiac fibroblasts treated with TGF-β1; overexpression by Ad5 (adenovirus serotype 5) in infarcted murine hearts (permanent ligation of left anterior descending coronary artery). | Binds with SAFB (scaffold attachment factor B) to block its access to RNA pol II (RNA polymerase II), and reduces the transcription of fibrosis-related genes. | [30] |
| Wisper (Wisp2 super-enhancer-associated RNA) | Knockdown by GapmeR in primary neonatal and adult murine cardiac fibroblasts, primary adult murine lung fibroblasts, primary human cardiac fibroblasts, and infarcted murine hearts (permanent ligation of left anterior descending coronary artery); overexpression by CRISPR-on in P19CL6 cells. | Interacts with Tial1 (Tia1 cytotoxic granule-associated RNA-binding protein-like 1, also known as TIAR) to regulate alternative splicing of Plod2 (procollagen lysine, 2-oxoglutarate 5-dioxygenase 2) to stabilize the extracellular matrix. | [23] |
| Name of LncRNA | Study Model | Function | Reference |
|---|---|---|---|
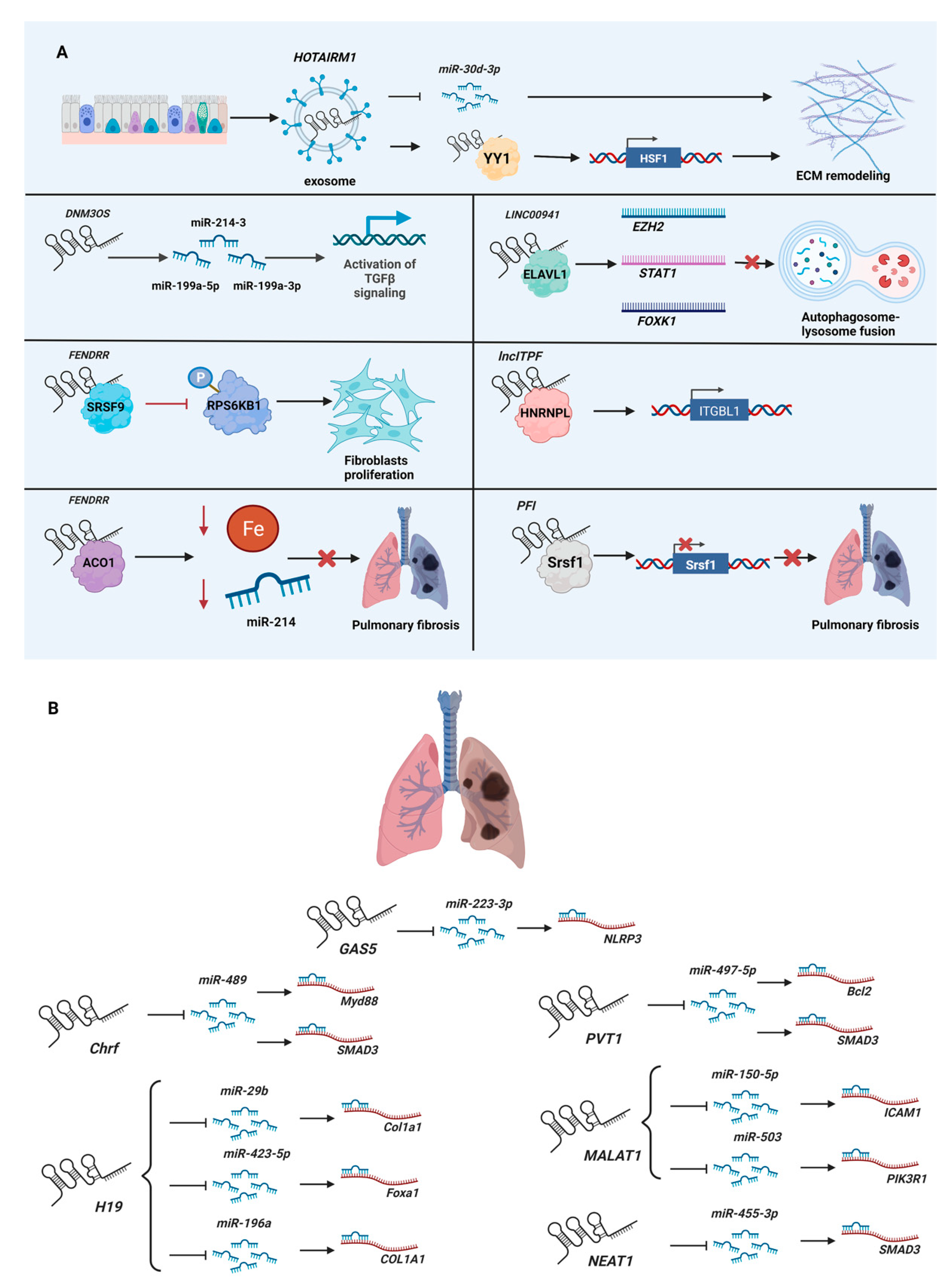
| DNM3OS (DNM3 opposite strand/antisense RNA) | Knockdown by GapmeR in the human lung fibroblastic cell line MRC5 and a murine bleomycin-induced lung fibrosis model. | Encodes three fibromiRs (miR-199a-5p/3p and miR-214-3), which regulate SMAD and non-SMAD components of TGF-β signaling. | [64] |
| FENDRR (FOXF1 adjacent non-coding developmental regulatory RNA) | Knockdown by lentiviral shRNA in human lung fibroblastic cell line LL29; overexpression by adenovirus in a murine asbestos-induced lung fibrosis model. | Binds SRSF9 (serine- and arginine-rich splicing factor 9) to inhibit the phosphorylation of RPS6KB1 (ribosomal protein S6 kinase B1, also known as PS6K), thereby suppressing the proliferation of fibroblasts. | [65] |
| FENDRR (FOXF1 adjacent non-coding developmental regulatory RNA) | Knockdown by lentiviral shRNA in the human lung fibroblastic cell lines HFL1 and LL29; overexpression by adenovirus in a murine bleomycin-induced lung fibrosis model. | Interacts with ACO1 (aconitase 1, also known as IRP1) to decrease cellular iron concentration and sequester pro-fibrotic miR-214 to reduce pulmonary fibrosis. | [66] |
| HOTAIRM1 (HOXA transcript antisense RNA, myeloid-specific 1) | Exosomes isolated from the murine lung epithelial cell line MLE-12 and co-cultured with primary murine lung fibroblasts; overexpressed by lentivirus in the murine lung epithelial cell line MLE-12, extracted exosomes, and then injected exosomes and lentivirus into a murine bleomycin-induced lung fibrosis model. | Released via exosomes from alveolar epithelial cells to lung fibroblasts, where it sequesters miR-30d-3p and recruits YY1 (YY1 transcription factor) to upregulate HSF1 (heat shock transcription factor 1), thereby promoting extracellular matrix remodeling. | [67] |
| LINC00941 (long intergenic non-protein coding RNA 941, also known as lncIAPF) | Knockdown by siRNA and overexpression by adenovirus in the human lung fibroblast cell line MRC-5; overexpression by adenovirus in the murine fibroblast cell line L929; overexpression by adenovirus in a murine bleomycin-induced lung fibrosis model. | Formed an RNA–protein complex with ELAVL1 to inhibit autophagosome fusion with a lysosome by controlling the stability of EZH2, STAT1 (signal transducer and activator of transcription 1), and FOXK1 (forkhead box K1) mRNAs. | [68] |
| lncITPF (lncRNA regulates its host gene Itgbl1 during pulmonary fibrogenesis, also known as MRAK053938) | Knockdown by siRNA and overexpression in the human lung fibroblast cell line MRC-5; knockdown by lentiviral shRNA in a murine bleomycin-induced lung fibrosis model. | Binds HNRNPL (heterogeneous nuclear ribonucleoprotein L) to epigenetically regulate its host gene, ITGBL1 (integrin subunit beta-like 1). | [69] |
| PFI (pulmonary fibrosis inhibitor, also known as NONMMUT060091) | Knockdown by LncRNA Smart Silencer and overexpression in primary murine lung fibroblasts; PFI transgenic (TG-PFI) mice with a murine bleomycin-induced lung fibrosis model. | Binds Srsf1 (serine and arginine-rich splicing factor 1) to repress its expression and pro-fibrotic activity. | [63] |
| Name of lncRNA | Cardiac Fibrosis | References | Pulmonary Fibrosis | References |
|---|---|---|---|---|
| Chrf (cardiac hypertrophy-related factor) | miR-489/Myd88 | [37] | miR-489/Myd88 & Smad3 | [75] |
| GAS5 (growth arrest specific 5) | miR-21/PTEN | [50] | miR-223-3p/NLRP3 | [76] |
| H19 (H19 imprinted maternally expressed transcript) | miR-455/CTGF; miRNA-22-3p/Kdm3a | [70]; [71] | miR-29b/Col1a1; miR-423-5p/Foxa1; miR-196a/COL1A1 | [72]; [73]; [74] |
| MALAT1 (metastasis-associated lung adenocarcinoma transcript 1) | miR-141/Nlrp3 & Tgfb1 | [77] | miR-150-5p/ICAM1; miR-503/PIK3R1 | [78]; [79] |
| NEAT1 (nuclear paraspeckle assembly transcript 1) | miR-320/NPAS2 | [80] | miR-455-3p/SMAD3 | [81] |
| PVT1 (Pvt1 oncogene | miR-145/HCN1; miR-24/KLF6; miR-128-3p/Sp1 | [82]; [83]; [84] | miR-497-5p/Bcl2 & Smad3 | [85] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ilieva, M.; Uchida, S. Long Non-Coding RNAs in Cardiac and Pulmonary Fibroblasts and Fibrosis. Non-Coding RNA 2022, 8, 53. https://doi.org/10.3390/ncrna8040053
Ilieva M, Uchida S. Long Non-Coding RNAs in Cardiac and Pulmonary Fibroblasts and Fibrosis. Non-Coding RNA. 2022; 8(4):53. https://doi.org/10.3390/ncrna8040053
Chicago/Turabian StyleIlieva, Mirolyuba, and Shizuka Uchida. 2022. "Long Non-Coding RNAs in Cardiac and Pulmonary Fibroblasts and Fibrosis" Non-Coding RNA 8, no. 4: 53. https://doi.org/10.3390/ncrna8040053
APA StyleIlieva, M., & Uchida, S. (2022). Long Non-Coding RNAs in Cardiac and Pulmonary Fibroblasts and Fibrosis. Non-Coding RNA, 8(4), 53. https://doi.org/10.3390/ncrna8040053







