Using Genealogical Concordance and Coalescent-Based Species Delimitation to Assess Species Boundaries in the Diaporthe eres Complex
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Isolates
2.2. DNA Extraction and PCR Amplification
2.3. Phylogenetic Analyses
2.4. Pairwise Homoplasy Index Test and Phylogenetic Network Analysis
2.5. Species Delimitation Analyses
2.6. Population Genetic Diversity
2.7. Morphology of the Diaporthe eres Species Complex
3. Results
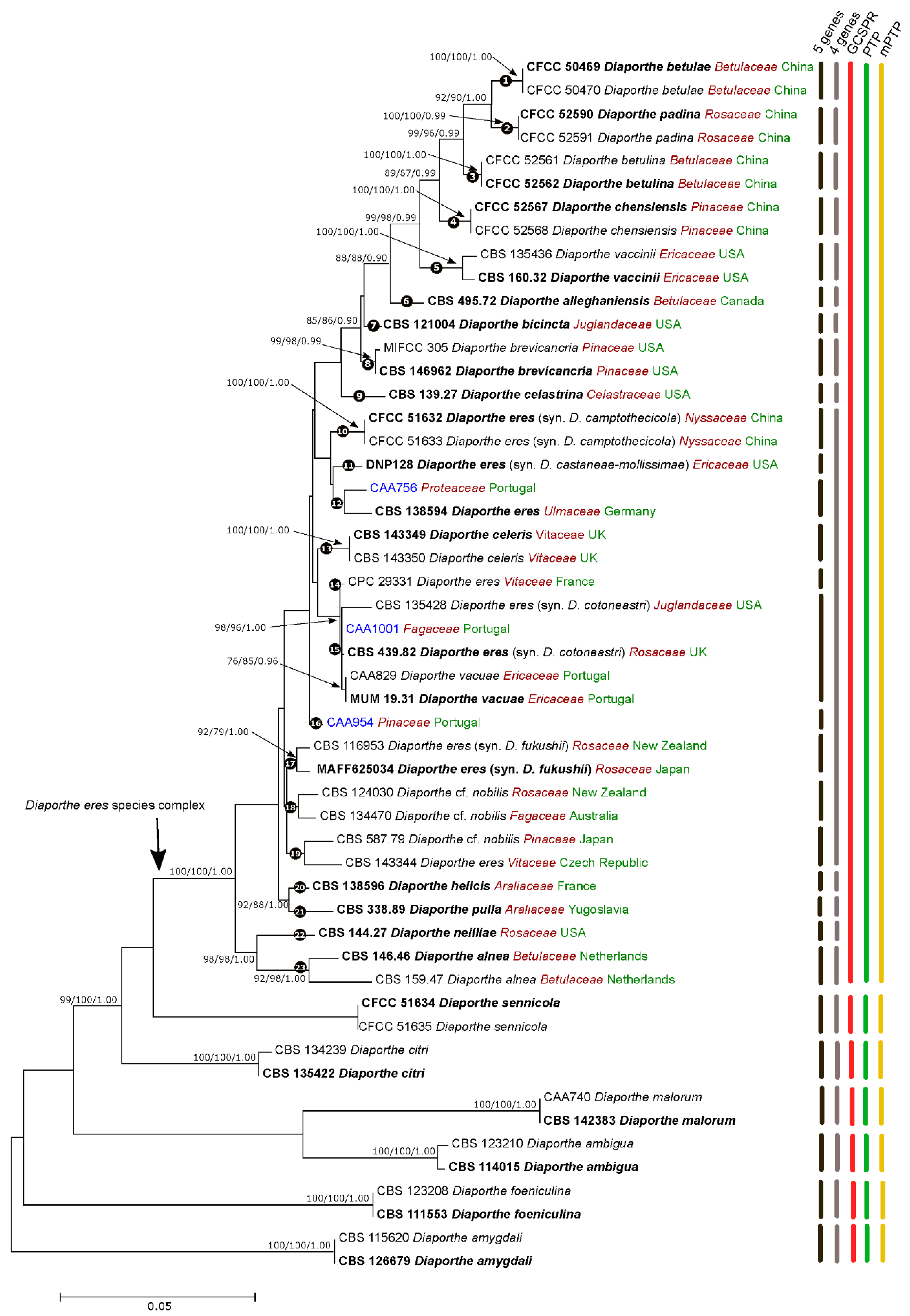
3.1. Phylogenetic Analyses and Informative Characters
3.2. Species Delimitation Based on the GCPSR Principle
3.3. Species Delimitation Based on Poisson Tree Processes
3.4. Pairwise Homoplasy Test and Phylogenetic Networks
3.5. Population Genetic Diversity
3.6. Morphology of the Diaporthe eres Species Complex
4. Taxonomy
- Diaporthe eres Nitschke, Pyrenomycetes Germanici 2:245 (1870)
- Basionym: Phoma oblonga Desm., Annales des Sciences Naturelles; Botanique, sér. 2, 22:218 (1853)
- =Diaporthe alnea Fuckel, Jahrbücher des Nassauischen Vereins für Naturkunde 23–24:207 (1870)
- =Diaporthe pulla Nitschke, Pyrenomycetes Germanici 2:249 (1870)
- =Diaporthe helicis Niessl, Verhandlungen des naturforschenden Vereines in Brünn 16:50 (1876)
- =Diaporthe nobilis Saccardo and Spegazzini, Michelia 1:386 (1878)
- =Diaporthe bicincta (Cooke and Peck) Sacc., Sylloge fungorum (Abellini) 1:622 (1882)
- =Diaporthe neilliae Peck, Annual Report on the New York State Museum of Natural History 39:52 (1887)
- =Diaporthe celastrina (Ellis and Barthol), The Journal of Mycology 8:173 (1902)
- ≡Phomopsis oblonga (Desm.) Traverso, Fl. ital. crypt., Pars 1: Fungi. Pyrenomycetae. Xylariaceae, Valsaceae, Ceratostomataceae: 248 (1906)
- =Phomopsis velata Sacc. Traverso, Fl. ital. crypt. (Florence) 2:248 (1906)
- =Diaporthe vaccinii Shear, United States Department of Agriculture 258:7 (1931)
- =Diaporthe alleghaniensis Arnold, Canadian Journal of Botany 45:787 (1967)
- ≡Diaporthe cotoneastri Udayanga, Crous and Hyde, Fungal Diversity 56:166 (2012)
- ≡Diaporthe castaneae-mollisimae Udayanga, Crous and Hyde, Fungal Diversity 56:166 (2012)
- =Diaporthe phragmitis Crous, Fungal Planet 283:219 (2014)
- =Diaporthe biguttusis Gao and Cai, Fungal Biology 119:300 (2015)
- =Diaporthe ellipicola Gao and Cai, Fungal Biology 119:300 (2015)
- =Diaporthe longicicola Gao and Cai, Fungal Biology 119:303 (2015)
- =Diaporthe mahothocarpus Gao and Cai, Fungal Biology 119:306 (2015)
- =Diaporthe betulae Tian and Fan, Phytotaxa 269:96 (2016)
- =Diaporthe maritima Tanney, Fungal Biology 120:1454 (2016)
- =Diaporthe camptothecicola Tian and Yang, Mycotaxon 132:595 (2017)
- =Diaporthe momicola Dissanayake, Li and Hyde, Mycosphere 8:541 (2017)
- =Diaporthe fukushii Dissanayake, Phillips and Hyde, Mycosphere 8:1130 (2017)
- =Diaporthe betulina Tian and Yang, Mycokeys 39:121 (2017)
- =Diaporthe chensiensis Tian and Yang, Mycokeys 39:127 (2017)
- =Diaporthe padina Tian and Yang, Mycokeys 39:137 (2017)
- =Diaporthe celeris Guarnaccia, Woodhall and Crous, Persoonia 40:146 (2018)
- =Diaporthe rosicola Wanasinghe, Jones and Hyde, Fungal Diversity 89:187 (2018)
- =Diaporthe vacuae Hilário, Santos and Alves, Mycologia 55:207 (2020)
- =Diaporthe brevicancria Sakalidis and Medina-Mora, Phytopathology (2020)
5. Discussion
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wingfield, M.J.; De Beer, Z.W.; Slippers, B.; Wingfield, B.D.; Groenewald, J.Z.; Lombard, L.; Crous, P.W. One fungus, one name promotes progressive plant pathology. Mol. Plant Pathol. 2012, 13, 604–613. [Google Scholar] [CrossRef]
- Hyde, K.D.; Jeewon, R.; Chen, Y.J.; Bhunjun, C.S.; Calabon, M.S.; Jiang, H.B.; Lin, C.-G.; Norphanphoun, C.; Sysouphathong, P.; Pem, D.; et al. The numbers of fungi: Is the descriptive curve flattening? Fungal Divers. 2020, 103, 219–271. [Google Scholar] [CrossRef]
- Lücking, R.; Hawksworth, D.L. Formal description of sequence-based voucherless Fungi: Promises and pitfalls, and how to resolve them. IMA Fungus 2018, 9, 143–165. [Google Scholar] [CrossRef]
- Gao, Y.H.; Liu, F.; Duan, W.; Crous, P.W.; Cai, L. Diaporthe is paraphyletic. IMA Fungus 2017, 8, 163–187. [Google Scholar] [CrossRef]
- Lücking, R.; Aime, M.C.; Robbertse, B.; Miller, A.N.; Ariyawansa, H.A.; Aoki, T.; Cardinali, G.; Crous, P.W.; Druzhinia, I.S.; Geiser, D.M.; et al. Unambiguous identification of fungi: Where do we stand and how accurate and precise is fungal DNA barcoding? IMA Fungus 2020, 11, 1–32. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.W.; Jacobson, D.J.; Kroken, S.; Kasuga, T.; Geiser, D.M.; Hibbett, D.S.; Fisher, M.C. Phylogenetic species recognition and species concepts in fungi. Fungal Genet. Biol. 2000, 31, 21–32. [Google Scholar] [CrossRef]
- Liu, F.; Wang, M.; Damm, U.; Crous, P.W.; Cai, L. Species boundaries in plant pathogenic fungi: A Colletotrichum case study. BMC Evol. Biol. 2016, 16, 81. [Google Scholar] [CrossRef]
- Inderbitzin, P.; Robbertse, B.; Schoch, C.L. Species identification in plant-associated prokaryotes and fungi using DNA. Phytobiomes J. 2020, 4, 103–114. [Google Scholar] [CrossRef]
- Stewart, J.E.; Timmer, L.W.; Lawrence, C.B.; Pryor, B.M.; Peever, T.L. Discord between morphological and phylogenetic species boundaries: Incomplete lineage sorting and recombination results in fuzzy species boundaries in an asexual fungal pathogen. BMC Evol. Biol. 2014, 14, 38. [Google Scholar] [CrossRef]
- Achari, S.R.; Kaur, J.; Dinh, Q.; Mann, R.; Sawbridge, T.; Summerell, B.A.; Edwards, J. Phylogenetic relationship between Australian Fusarium oxysporum isolates and resolving the species complex using the multispecies coalescent model. BMC Genom. 2020, 21, 248. [Google Scholar] [CrossRef]
- Rannala, B.; Yang, Z. Species Delimitation. In Phylogenetics in the Genomic Era; Scornavacca, C., Delsuc, F., Galtier, N., Eds.; No commercial publisher; 2020; Chapter 5.5; pp. 1–18. Available online: https://hal.inria.fr/PGE (accessed on 12 September 2020).
- Carstens, B.C.; Knowles, L.L. Estimating species phylogeny from gene-tree probabilities despite incomplete lineage sorting: An example from Melanoplus grasshoppers. Syst. Biol. 2007, 56, 400–411. [Google Scholar] [CrossRef]
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef]
- Bustamante, D.E.; Oliva, M.; Leiva, S.; Mendoza, J.E.; Bobadilla, L.; Angulo, G.; Calderon, M.S. Phylogeny and species delimitations in the entomopathogenic genus Beauveria (Hypocreales, Ascomycota), including the description of B. peruviensis sp. nov. MycoKeys 2019, 58, 47–68. [Google Scholar] [CrossRef]
- Hilário, S.; Santos, L.; Alves, A. Diaporthe amygdali, a species complex or a complex species? Fungal Biol. 2021, 125, 505–518. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.; Yang, Q.; Bezerra, J.D.; Alvarez, L.V.; Tian, C. Diaporthe from walnut tree (Juglans regia) in China, with insight of the Diaporthe eres complex. Mycol. Prog. 2018, 17, 841–853. [Google Scholar] [CrossRef]
- Guarnaccia, V.; Groenewald, J.Z.; Woodhall, J.; Armengol, J.; Cinelli, T.; Eichmeier, A.; Ezra, D.; Fontaine, F.; Gramaje, D.; Gutierrez-Aguirregabiria, A.; et al. Diaporthe diversity and pathogenicity revealed from a broad survey of grapevine diseases in Europe. Persoonia 2018, 40, 135–153. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Fan, X.L.; Guarnaccia, V.; Tian, C.M. High diversity of Diaporthe species associated with dieback diseases in China, with twelve new species described. MycoKeys 2018, 39, 97–149. [Google Scholar] [CrossRef]
- Rehner, S.A.; Uecker, F.A. Nuclear ribosomal internal transcribed spacer phylogeny and host diversity in the coelomycete Phomopsis. Can. J. Bot. 1994, 72, 1666–1674. [Google Scholar] [CrossRef]
- Mostert, L.; Crous, P.W.; Kang, J.C.; Phillips, A.J.L. Species of Phomopsis and a Libertella sp. occurring on grapevines with specific reference to South Africa: Morphological, cultural, molecular and pathological characterization. Mycologia 2001, 93, 146–167. [Google Scholar]
- Gomes, R.R.; Glienke, C.; Videira, S.I.R.; Lombard, L.; Groenewald, J.Z.; Crous, P.W. Diaporthe: A genus of endophytic, saprobic and plant pathogenic fungi. Persoonia 2013, 31, 1–41. [Google Scholar] [CrossRef] [PubMed]
- Marin-Felix, Y.; Hernández-Restrepo, M.; Wingfield, M.J.; Akulov, A.; Carnegie, A.J.; Cheewangkoon, R.; Gramaje, D.; Groenewald, J.Z.; Guarnaccia, V.; Halleen, F.; et al. Genera of phytopathogenic fungi: GOPHY 2. Stud. Mycol. 2019, 92, 47–133. [Google Scholar] [CrossRef] [PubMed]
- Udayanga, D.; Castlebury, L.A.; Rossman, A.Y.; Chukeatirote, E.; Hyde, K.D. Insights into the genus Diaporthe: Phylogenetic species delimitation in the D. eres species complex. Fungal Divers. 2014, 67, 203–229. [Google Scholar] [CrossRef]
- Guo, Y.S.; Crous, P.W.; Bai, Q.; Fu, M.; Yang, M.M.; Wang, X.H.; Du, Y.M.; Hong, N.; Xu, W.X.; Wang, G.P. High diversity of Diaporthe species associated with pear shoot canker in China. Persoonia 2020, 45, 132–162. [Google Scholar] [CrossRef]
- Chaisiri, C.; Liu, X.; Lin, Y.; Fu, Y.; Zhu, F.; Luo, C. Phylogenetic and haplotype network analyses of Diaporthe eres species in China based on sequences of multiple loci. Biology 2021, 10, 179. [Google Scholar] [CrossRef]
- Lombard, L.; van Leeuwen, G.C.M.; Guarnaccia, V.; Polizii, G.; van Rijswick, P.C.J.; Rosendahl, C.H.M.; Gabler, J.; Crous, P.W. Diaporthe species associated with Vaccinium, with specific reference to Europe. Phytopathol. Mediterr. 2014, 53, 287–299. [Google Scholar]
- Ali, S.; Renderos, W.; Bevis, E.; Hebb, J.; Abbasi, P.A. Diaporthe eres causes stem cankers and death of young apple rootstocks in Canada. Can. J. Plant Pathol. 2020, 42, 218–227. [Google Scholar] [CrossRef]
- Lopes, A.F.; Batista, E.; Hilário, S.; Santos, L.; Alves, A. Occurrence of Diaporthe species in Eucalyptus globulus, Pinus pinaster and Quercus suber in Portugal. For. Pathol. 2021, 51, e12674. [Google Scholar]
- Hilário, S.; Amaral, I.; Gonçalves, F.M.; Lopes, A.; Santos, L.; Alves, A. Diversity and pathogenicity of Diaporthe species on blueberry plants in Portugal, with description of 4 new species. Mycologia 2020, 112, 293–308. [Google Scholar] [CrossRef]
- Möller, E.M.; Bahnweg, G.; Sandermann, H.; Geiger, H.H. A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fungi, fruit bodies, and infected plant tissues. Nucleic Acids Res. 1992, 20, 6115–6116. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Alves, A.; Correia, A.; Luque, J.; Phillips, A.J.L. Botryosphaeria corticola, sp. nov. on Quercus species, with notes and description of Botryosphaeria stevensii and its anamorph, Diplodia mutila. Mycologia 2004, 96, 598–613. [Google Scholar] [CrossRef]
- Alves, A.; Crous, P.W.; Correia, A.; Phillips, A.J.L. Morphological and molecular data reveal cryptic speciation in Lasiodiplodia theobromae. Fungal Divers. 2008, 28, 1–13. [Google Scholar]
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous Ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol. Phylogenet. Evol. 1997, 7, 103–116. [Google Scholar] [CrossRef]
- Crous, P.W.; Groenewald, J.Z.; Risede, J.M.; Hywe, N.L. Calonectria species and their Cylindrocladium anamorphs: Species with sphaeropedunculate vesicles. Stud. Mycol. 2004, 50, 415–430. [Google Scholar]
- Carbone, I.; Kohn, L.M. A method for designing primer set for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Higgins, D.G. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Vaydia, G.; Lohman, D.J.; Meier, R. SequenceMatrix: Concatenation software for the last assembly of multi-gene datasets with character set and codon information. Cladistics 2011, 27, 171–180. [Google Scholar] [CrossRef]
- Swofford, D.L. PAUP* Phylogenetic Analysis Using Parsimony, (*and Other Methods); Version 4.0 b10; Sinauer Associates: Sunderland, MA, USA, 2003. [Google Scholar]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Page, R.D.M. TreeView: An application to display phylogenetic trees on personal computers. CABIOS 1996, 12, 357–358. [Google Scholar]
- Rodríguez, F.; Oliver, J.L.; Marín, A.; Medina, J.R. The general stochastic model of nucleotide substitutions. J. Theor. Biol. 1990, 142, 485–501. [Google Scholar] [CrossRef]
- Bruen, T.C. A simple and robust statistical test for detecting the presence of recombination. Genetics 2005, 172, 2665–2681. [Google Scholar] [CrossRef] [PubMed]
- Huson, D.H.; Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 2006, 23, 254–267. [Google Scholar] [CrossRef]
- Kapli, P.; Lutteropp, S.; Zhang, J.; Kobert, K.; Pavlidis, P.; Stamatakis, A.; Flouri, T. Multi-rate Poisson tree processes for single-locus species delimitation under maximum likelihood and Markov chain Monte Carlo. Bioinformatics 2017, 33, 1630–1638. [Google Scholar] [CrossRef] [PubMed]
- Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 1989, 123, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Murtagh, F.; Legendre, P. Ward’s hierarchical agglomerative clustering method: Which algorithms implement Ward’s criterion? J. Classif. 2014, 31, 274–295. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. Available online: https://www.R-project.org/ (accessed on 14 January 2021).
- Galili, T. Dendextend: An R package for visualizing, adjusting, and comparing trees of hierarchical clustering. Bioinformatics 2015, 31, 3718–3720. [Google Scholar] [CrossRef]
- Farr, D.F.; Rossman, A.Y. Fungal Databases, U.S. National Fungus Collections, ARS, USDA. 2021. Available online: https://nt.ars-grin.gov/fungaldatabases/ (accessed on 8 April 2021).
- Du, Z.; Fan, X.L.; Hyde, K.D.; Yang, Q.; Liang, Y.M.; Tian, C.M. Phylogeny and morphology reveal two new species of Diaporthe from Betula spp. in China. Phytotaxa 2016, 269, 90–102. [Google Scholar] [CrossRef][Green Version]
- Gao, Y.H.; Su, Y.Y.; Sun, W.; Cai, L. Diaporthe species occurring on Lithocarpus glabra in China, with descriptions of five new species. Fungal Biol. 2015, 119, 295–309. [Google Scholar] [CrossRef]
- Sakalidis, M.; Medina-Mora, C.M.; Shin, K.; Fulbright, D.W. Characterization of Diaporthe spp. associated with spruce decline on Colorado blue spruce in Michigan. Phytopathology 2021, 111, 509–520. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Fan, X.L.; Du, Z.; Liang, Y.M.; Tian, C.M. Diaporthe camptothecicola sp. nov. on Camptotheca acuminata in China. Mycotaxon 2017, 132, 591–601. [Google Scholar] [CrossRef]
- Gao, Y.H.; Sun, W.; Su, Y.Y.; Cai, L. Three new species of Phomopsis in Gutianshan nature reserve in China. Mycol. Prog. 2014, 13, 111–121. [Google Scholar] [CrossRef]
- Tanney, J.B.; McMullin, D.R.; Green, B.D.; Miller, J.D.; Seifert, K.A. Production of antifungal and antiinsectan metabolites by the Picea endophyte Diaporthe maritima sp. nov. Fungal Biol. 2016, 120, 1448–1457. [Google Scholar] [CrossRef] [PubMed]
- Dissanayake, A.J.; Zhang, W.; Liu, M.; Hyde, K.D.; Zhao, W.S.; Li, X.H.; Yan, J.Y. Diaporthe species associated with peach tree dieback in Hubei, China. Mycosphere 2017, 8, 533–549. [Google Scholar]
- Li, Y.; Tan, P.; Zhao, D.G. Diaporthe nobilis, a new record on Camellia sinensis in Guizhou Province, China. Mycosphere 2017, 8, 1–8. [Google Scholar] [CrossRef]
- Crous, P.W.; Wingfield, M.J.; Schumacher, R.K.; Summerell, B.A.; Giraldo, A.; Gené, J.; Guarro, J.; Wanasinghe, D.N.; Hyde, K.D.; Camporesi, E.; et al. Fungal Planet description sheets: 281–319. Persoonia 2014, 33, 212–289. [Google Scholar] [CrossRef]
- Wanasinghe, D.N.; Phukhamsakda, C.; Hyde, K.D.; Jeewon, R.; Lee, H.B.; Jones, E.G.; Tibpromma, S.; Tennakoon, D.S.; Jayasiri, S.C.; Gafforov, Y.; et al. Fungal diversity notes 709–839: Taxonomic and phylogenetic contributions to fungal taxa with an emphasis on fungi on Rosaceae. Fungal Divers. 2018, 89, 1–236. [Google Scholar] [CrossRef]
- Farr, D.F.; Castlebury, L.A.; Rossman, A.Y. Morphological and molecular characterization of Phomopsis vaccinii and additional isolates of Phomopsis from blueberry and cranberry in the eastern United States. Mycologia 2002, 94, 494–504. [Google Scholar] [CrossRef]
- Santos, L.; Alves, A.; Alves, R. Evaluating multi-locus phylogenies for species boundaries determination in the genus Diaporthe. Peer J. 2017, 5, e3120. [Google Scholar] [CrossRef]
- Udayanga, D.; Liu, X.; McKenzie, E.H.C.; Chukeatirote, E.; Bahkali, A.H.A.; Hyde, K.D. The genus Phomopsis: Biology, applications, species concepts and names of common phytopathogens. Fungal Divers. 2011, 50, 189–225. [Google Scholar] [CrossRef]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W.; Fungal Barcoding Consortium. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef]
- Crouch, J.A.; Clarke, B.B.; Hillman, B.I. What is the value of ITS sequence data in Colletotrichum systematics and species diagnosis? A case study using the falcate-spored graminicolous Colletotrichum group. Mycologia 2009, 101, 648–656. [Google Scholar] [CrossRef] [PubMed]
- Kubatko, L.S.; Degnan, J.H. Inconsistency of phylogenetic estimates from concatenated data under coalescence. Syst. Biol. 2007, 56, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Steenkamp, E.T.; Wingfield, M.J.; McTaggart, A.R.; Wingfield, B.D. Fungal species and their boundaries matter—Definitions, mechanisms and practical implications. Fungal Biol. Rev. 2018, 32, 104–116. [Google Scholar] [CrossRef]
- Gazis, R.; Rehner, S.; Chaverri, P. Species delimitation in fungal endophyte diversity studies and its implications in ecological and biogeographic inferences. Mol. Ecol. 2011, 20, 3001–3013. [Google Scholar] [CrossRef] [PubMed]
- Mendez-Harclerode, F.M.; Strauss, R.E.; Fulhorst, C.F.; Milazzo, M.L.; Ruthven, D.C.; Bradley, R.D. Molecular evidence for high levels of intrapopulation genetic diversity in woodrats (Neotoma micropus). J. Mammal. 2007, 88, 360–370. [Google Scholar] [CrossRef]
- Manawasinghe, I.S.; Dissanayake, A.; Liu, M.; Wanasinghe, D.; Xu, J.; Zhao, W.; Zhang, W.; Zhou, Y.; Hyde, K.D.; Brooks, S.; et al. High genetic diversity and species complexity of Diaporthe associated with grapevine dieback in China. Front. Microbiol. 2019, 10, 1936. [Google Scholar] [CrossRef]
- Boluda, C.G.; Rico, V.J.; Divakar, P.K.; Nadyeina, O.; Myllys, L.; McMullin, R.T.; Zamora, J.C.; Scheidegger, C.; Hawksworth, D.L. Evaluating methodologies for species delimitation: The mismatch between phenotypes and genotypes in lichenized fungi (Bryoria sect. Implexae, Parmeliaceae). Persoonia 2019, 42, 75–100. [Google Scholar] [CrossRef]
- Pons, J.; Barraclough, T.; Gomez-Zurita, J.; Cardoso, A.; Duran, D.; Hazell, S.; Kamoun, S.; Sumlin, W.; Vogler, A. Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Syst. Biol. 2006, 55, 595–609. [Google Scholar] [CrossRef]
- Matute, D.R.; Sepúlveda, V.E. Fungal species boundaries in the genomics era. Fungal Genet. Biol. 2019, 131, 103249. [Google Scholar] [CrossRef] [PubMed]
- Ho, S.Y.; Duchêne, S. Molecular-clock methods for estimating evolutionary rates and timescales. Mol. Ecol. 2014, 23, 5947–5965. [Google Scholar] [CrossRef] [PubMed]
- Berbee, M.L.; Taylor, J.W. Dating the molecular clock in fungi—How close are we? Fungal Biol. Rev. 2010, 24, 1–16. [Google Scholar] [CrossRef]
- Tang, C.Q.; Humphreys, A.M.; Fontaneto, D.; Barraclough, T.G.; Paradis, E. Effects of phylogenetic reconstruction method on the robustness of species delimitation using single-locus data. Methods Ecol. Evol. 2014, 5, 1086–1094. [Google Scholar] [CrossRef]
- Jeger, M.; Bragard, C.; Caffier, D.; Candresse, T.; Chatzivassiliou, E.; Dehnen-Schmutz, K.; Gilioli, G.; Grégoire, J.C.; Miret, J.; MacLeod, A.; et al. Pest risk assessment of Diaporthe vaccinii for the EU territory. EFSA J. 2017, 15, 1–185. [Google Scholar]
- Cardinaals, J.; Wenneker, M.; Voogd, J.G.B.; van Leeuwen, G.C.M. Pathogenicity of Diaporthe spp. on two blueberry cultivars (Vaccinium corymbosum). EPPO Bull. 2018, 48, 128–134. [Google Scholar] [CrossRef]




| Species | Strain 1 | Host | Country | GenBank Accession | ||||
|---|---|---|---|---|---|---|---|---|
| ITS | TEF1-α | TUB2 | HIS3 | CAL | ||||
| D. ambigua | CBS 114015 | Pyrus communis | South Africa | KC343010 | KC343736 | KC343978 | KC343494 | KC343252 |
| CBS 123210 | Foeniculum vulgare | Portugal | KC343012 | KC343738 | KC343980 | KC343496 | KC343254 | |
| D. amygdali | CBS 126679 | Prunus dulcis | Portugal | KC343022 | KC343748 | KC343990 | KC343506 | KC343264 |
| CBS 115620 | Prunus persica | USA | KC343020 | KC343746 | KC343988 | KC343504 | KC343262 | |
| D. citri | CBS 134239 | Citrus cinensis | USA | KC357553 | KC357522 | KC357456 | MF418280 | KC357488 |
| CBS 135422 | Citrus sp. | USA | KC843311 | KC843071 | KC843187 | MF418281 | KC843157 | |
| D. eres | CPC 29331 | Vitis vinifera | France | MG281034 | MG281555 | MG281207 | KC343631 | KC343389 |
| CBS 138594 | Ulmus laevis | Germany | KJ210529 | KJ210550 | KJ420799 | KC343637 | KC343395 | |
| CBS 143344 | Vitis vinifera | Czech Republic | MG281020 | MG281541 | MG281193 | MG281366 | MG281715 | |
| CAA756 | Banksia sp. | Portugal | MW040531 | MW052385 | MW091320 | MW052384 | MW091319 | |
| CAA954 | Pinus pinaster | Portugal | MN190309 | MT309431 | MT309457 | MT309440 | MT309448 | |
| CAA1001 | Quercus suber | Portugal | MT237172 | MT309432 | MT309458 | MT309441 | MT309449 | |
| D. eres (syn. D. alnea) | CBS 146.46 | Alnus sp. | The Netherlands | KC343008 | KC343734 | KC343976 | KC343492 | KC343250 |
| CBS 159.47 | Alnus sp. | The Netherlands | KC343009 | KC343735 | KC343977 | KC343493 | KC343251 | |
| D. eres (syn. D. alleghaniensis) | CBS 495.72 | Betula alleghaniensis | Canada | FJ889444 | GQ250298 | KC843228 | KC343491 | KC343249 |
| D. eres (syn. D. betulae) | CFCC 50469 | Betula platyphylla | China | KT732950 | KT733016 | KT733020 | KT732999 | KT732997 |
| CFCC 50470 | Betula platyphylla | China | KT732951 | KT733017 | KT733021 | KT733000 | KT732998 | |
| D. eres (syn. D. betulina) | CFCC 52562 | Betula platyphylla | China | MH121497 | MH121539 | MH121579 | MH121457 | MH121421 |
| CFCC 52561 | Betula platyphylla | China | MH121496 | MH121538 | MH121578 | MH121456 | MH121421 | |
| D. eres (syn. D. bicincta) | CBS 121004 | Juglans sp. | USA | KC343134 | KC343860 | KC344102 | KC343618 | KC343376 |
| D. eres (syn. D. biguttusis) | CGMCC 3.17081 | Lithocarpus glabra | China | KF576282 | KF576257 | KF576307 | - | - |
| D. eres (syn. D. brevicancria) | CBS 146962 | Picea pungens | USA | MN136180 | MN136153 | MN136190 | MN136178 | MN136129 |
| MIFCC 305 | Picea glauca | USA | MN136184 | MN136151 | MN136188 | MN136176 | MN136127 | |
| D. eres (syn. Diaporthe cf. nobilis) | CBS 124030 | Malus pumila | New Zealand | KC343149 | KC343875 | KC344117 | KC343633 | KC343391 |
| CBS 134470 | Castanea sativa | Australia | KC343146 | KC343872 | KC344114 | KC343630 | KC343388 | |
| CBS 587.79 | Pinus pentaphylla | Japan | KC343153 | KC343879 | KC344121 | KC343637 | KC343395 | |
| D. eres (syn. D. camptothecicola) | CFCC 51632 | Camptotheca acuminata | China | KY203726 | KY228887 | KY228893 | KY228881 | KY228877 |
| CFCC 51633 | Camptotheca acuminata | China | KY203727 | KY228888 | KY228894 | KY228882 | KY228878 | |
| D. eres (syn. D. castaneae-mollissimae) | DNP128 | Vaccinium corymbosum | China | KC763096 | KJ210561 | KJ420801 | KJ420852 | KJ435040 |
| D. eres (syn. D. celeris) | CBS 143349 | Vitis vinifera | UK | MG281017 | MG281538 | MG281190 | MG281363 | MG281712 |
| CBS 143350 | Vitis vinifera | UK | MG281018 | MG281539 | MG281191 | MG281364 | MG281713 | |
| D. eres (syn. D. celastrina) | CBS 139.27 | Celastrus sp. | USA | KC343047 | KC343773 | KC344015 | KC343531 | KC343289 |
| D. eres (syn. D. chensiensis) | CFCC 52567 | Abies chensiensis | China | MH121502 | MH121544 | MH121584 | MH121462 | MH121426 |
| CFCC 52568 | Abies chensiensis | China | MH121503 | MH121545 | MH121585 | MH121463 | MH121427 | |
| D. eres (syn. D. cotoneastri) | CBS 135428 | Juglans cinerea | USA | KC843328 | KC843121 | KC843229 | KC343630 | KC343388 |
| CBS 439.82 | Cotoneaster sp. | UK | KC343090 | KC343816 | KC344058 | KC343574 | KC343332 | |
| D. eres (syn. D. ellipicola) | CGMCC 3.17084 | Lithocarpus glabra | China | KF576270 | KF576245 | KF576291 | - | - |
| D. eres (syn. D. fukushii) | CBS 116953 | Pyrus pyrifolia | New Zealand | KC343147 | KC343873 | KC344115 | KC343631 | KC343389 |
| CBS 116954 | Pyrus pyrifolia | Japan | JQ807469 | JQ807418 | KJ420819 | KJ420868 | KJ435023 | |
| D. eres (syn. D. helicis) | CBS 138596 | Hedera helix | France | KJ210538 | KJ210559 | KJ420828 | KJ420875 | KJ435043 |
| D. eres (syn. D. longicicola) | CGMCC 3.17089 | Lithocarpus glabra | China | KF576267 | KF576242 | KF576291 | - | - |
| CGMCC 3.17090 | Lithocarpus glabra | China | KF576268 | KF576243 | KF576292 | - | - | |
| D. eres (syn. D. mahothocarpus) | CGMCC 3.15181 | Lithocarpus glabra | China | KC153096 | KC153087 | KF576312 | - | - |
| D. eres (syn. D. maritima) | DAOMC 250563 | Picea rubens | Canada | KU552027 | KU552022 | KU574616 | - | - |
| D. eres (syn. D. momicola) | MFLUCC 16-0113 | Prunus persica | China | KU557563 | KU557631 | KU557587 | - | KU557611 |
| MFLUCC 16-0114 | Prunus persica | China | KU557564 | KU557632 | KU557588 | - | KU557612 | |
| D. eres (syn. D. neilliae) | CBS 144.27 | Sapiraea sp. | USA | KC343144 | KC343870 | KC344112 | KC343628 | KC343386 |
| D. eres (syn. D. padina) | CFCC 52590 | Prunus padus | China | MH121525 | MH121567 | MH121604 | MH121483 | MH121443 |
| CFCC 52591 | Malus domestica | China | MH121526 | MH121568 | MH121605 | MH121484 | MH121444 | |
| D. eres (syn. D. phragmitis) | CBS 138897 | Phragmites australis | China | KP004445 | - | KP004507 | KP004503 | - |
| D. eres (syn. D. pulla) | CBS 338.89 | Hedera helix | Yugoslavia | KC343152 | KC343878 | KC344120 | KC343636 | KC343394 |
| D. eres (syn. D. rosicola) | MFLU 17-0646 | Rosa sp. | UK | MG828895 | MG829270 | MG843877 | - | MG829274 |
| D. eres (syn. D. vaccinii) | CBS 160.32 | Vaccinium macrocarpon | USA | AF317578 | GQ250326 | KC344196 | KC343712 | KC343470 |
| CBS 135436 | Vaccinium corymbosum | USA | AF317570 | JQ807380 | KC843225 | KJ420877 | KC849457 | |
| D. eres (syn. D. vacuae) | CAA829 | Vaccinium corymbosum | Portugal | MK792306 | MK828077 | MK837928 | MK871446 | MK883832 |
| MUM 19.31 | Vaccinium corymbosum | Portugal | MK792309 | MK828080 | MK837931 | MK871449 | MK883834 | |
| D. foeniculina | CBS 123208 | Foeniculum vulgare | Portugal | KC343104 | KC343830 | KC344072 | KC343588 | KC343346 |
| CBS 111553 | Foeniculum vulgare | Spain | KC343101 | KC343827 | KC344069 | KC343585 | KC343343 | |
| D. malorum | CAA740 | Malus domestica | Portugal | KY435642 | KY435629 | KY435670 | KY435650 | KY435660 |
| CBS 142383 | Malus domestica | Portugal | KY435638 | KY435627 | KY435668 | KY435648 | KY435658 | |
| D. sennicola | CFCC 51634 | Senna bicapsularis | China | KY203722 | KY228883 | KY228889 | KY228879 | KY228873 |
| CFCC 51635 | Senna bicapsularis | China | KY203723 | KY228883 | KY228889 | KY228879 | KY228873 | |
| Character Status Summary | Loci and Combined Alignments | ||||||
|---|---|---|---|---|---|---|---|
| ITS | TEF1-α | TUB2 | HIS3 | CAL | 4 loci | 5 loci | |
| Total characters | 537 | 387 | 430 | 488 | 522 | 1827 | 2364 |
| Invariable characters | 434 | 183 | 273 | 342 | 307 | 1104 | 1539 |
| Informative characters (%) | 92 (17%) | 184 (47%) | 143 (33%) | 116 (23%) | 200 (38%) | 649 (35%) | 735 (31%) |
| Uninformative characters | 11 | 20 | 14 | 30 | 15 | 74 | 90 |
| Tree length (TL) | 206 | 366 | 255 | 280 | 356 | 1358 | 1647 |
| Consistency index (CI) | 0.6359 | 0.7486 | 0.7686 | 0.7036 | 0.8090 | 0.7018 | 0.6594 |
| Homoplasy index (HI) | 0.3641 | 0.2514 | 0.2314 | 0.2964 | 0.1910 | 0.2982 | 0.3406 |
| Retention index (RI) | 0.8727 | 0.8892 | 0.8959 | 0.8683 | 0.9037 | 0.8554 | 0.8311 |
| Rescaled consistency index (RC) | 0.5549 | 0.6657 | 0.6886 | 0.6109 | 0.7311 | 0.6003 | 0.5480 |
| Nucleotide substitution models 1 | K2 + G | HKY | K2 + G | KKY + G | T92 + G | TN93 + G | TN93 + G |
| Loci | Number of Haplotypes (h) | Polymorphic Sites (S) | Haplotype Diversity (hd) | Nucleotide Diversity (π) | Tajima’s D Test |
|---|---|---|---|---|---|
| ITS | 18 | 35 | 0.948 | 0.025 | 0.6097 |
| TEF1-α | 19 | 43 | 0.966 | 0.019 | −1.6281 |
| TUB2 | 17 | 43 | 0.943 | 0.022 | −0.7208 |
| HIS3 | 13 | 44 | 0.882 | 0.021 | −1.0077 |
| CAL | 12 | 13 | 0.921 | 0.009 | −0.3513 |
| 5 loci 1 | 25 | 178 | 0.990 | 0.019 | −0.7359 |
| 4 loci 2 | 24 | 143 | 0.988 | 0.018 | −1.0881 |
| Species | Conidiomata | Conidiophores and Conidiogenous Cells | Alpha Conidia (µm) | Beta Conidia (µm) | References |
|---|---|---|---|---|---|
| D. alleghaniensis Arnold 1967 * | Conidiomata small, with a conical shape, 200–250 µm diameter | Conidiophores 9–15 × 1–2 μm, hyaline, unbranched, ampulliform, cylindrical to sub-cylindrical. Conidiogenous phiailidic, cylindrical, slightly tapering towards apex. | Alpha conidia 7–9 × 3–4 μm, aseptate, hyaline, smooth, ovate to ellipsoidal, biguttulate or multiguttulate, base sub-truncate. | Not observed | [23] |
| D. alnea Fuckel 1870 * =Phomopsis alnea | Pycnidia with 100–200 μm diameter, globose to subglobose, embedded in tissue, erumpent at maturity | Conidiophores 9–16 × 1–2 μm, hyaline, unbranched, ampulliform, cylindrical to sub-cylindrical. Conidiogenous cells phiailidic, cylindrical, tapering towards the apex. | Alpha conidia 8–10 × 2–3 μm, aseptate, hyaline, smooth, ellipsoidal, biguttulate or multiguttulate, base subtruncate. | Not observed | [23] |
| D. betulae Tian and Fan 2016 * | Conidiomatal stromata immersed, erumpent, separate, conical, with a single locule | Conidiophores reduced to phiailidic conidiogenous cells hyaline, straight or slightly curved. | Alpha conidia hyaline, ellipsoidal, aseptate, smooth, biguttulate, 8.5–11.5 × 3.5–4.5 μm. | Not observed | [55] |
| D. betulina Tian and Yang 2018 * | Conidiomata pycnidial, conical, immersed and erumpent through the bark surface, 290–645 μm diameter | Conidiophores 12.5–17.5 × 1.5–2 μm, cylindrical, hyaline, phiailidic, branched, straight or slightly curved. | Alpha conidia hyaline, aseptate, ellipsoidal to fusiform, biguttulate, acute at both ends, 8–10 × 2.5–3 μm. | Beta conidia hyaline, aseptate, filiform, straight, eguttulate, tapering towards one apex, 26–32.5 × 1 µm | [18] |
| D. bicincta Cooke and Peck 1882 * | Pycnidia with 200–300 μm diameter, globose, erumpent at maturity, conidial cirrhus extruding from ostiole | Conidiophores 7–12 × 1–2 μm, hyaline, smooth, unbranched, cylindrical to sub-cylindrical. Conidiogenous cells 0.5–1 μm diameter, phiailidic, cylindrical, tapering towards the apex. | Alpha conidia 9–12 × 2–3.5 μm, aseptate, hyaline, ovate to ellipsoidal, biguttulate or multiguttulate, base subtruncate. | Not observed | [23] |
| D. biguttusis Gao and Cai 2015 * | Pycnidial conidiomata, dark brown, globose clustered, 79–227 μm diameter | Conidiophores 11.5–27.1 × 1.4–2.3 μm, cylindrical, single to multi-septate, densely aggregated, slightly tapering towards the apex. | Alpha conidia hyaline, biguttulate, fusiform or oval, with both ends obtuse, 5.9–8.5 × 1.9–2.6 μm. | Beta conidia hyaline, aseptate, filiform, hamate, eguttulate, tapering towards both ends, 28.1–37.9 × 1.3–2.0 μm | [56] |
| D. brevicancria Sakalidis and Medina-Mora 2021 * | Pycnidia dark brown to black, emersed in host tissue, solitary or aggregated, often with creamy yellow conidial cirrhus, 236–368 μm diameter | Conidiophores hyaline, reduced to conidiogenous cells phiailidic, and narrowing towards the apex, 7.1–17.5 × 1.0–2.4 μm. | Alpha conidia, hyaline, aseptate, oblong to ellipsoid, often biguttulate. with a sub-truncated base, 4.4–8.6 × 1.3–3.3 µm. | Beta conidia aseptate, hyaline, smooth, mostly convex at one end, hooked, 12.4–27.4 × 0.9–2.1 µm | [57] |
| D. camptothecicola Tiang and Yang 2017 | Conidiomatal pycnidia immersed or slightly erumpent through bark surface, sparse, globose to ovoid, with 560 μm diameter | Conidiophores (8.3–)12.5–15.8(−17.0) × 0.9–1.2 μm hyaline, unbranched, smooth, cylindrical, straight or slightly curved, conidiogenous cells enteroblastic, phiailidic. | Alpha conidia hyaline, aseptate, oblong, biguttulate, (4.6–)5.5–7.0(−7.5) × 1.5–1.8 μm. | Beta conidia hyaline, aseptate, filiform with obtuse ends, 19.5–28.3 × 1.0 μm | [58] |
| D. celastrina Ellis and Barthol 1902 * | Pycnidia with 200–300 μm diameter, globose, embedded in tissue, erumpent at maturity conidial cirrhus extruding from ostiole | Conidiophores 7–21 × 1–2 μm, hyaline, smooth, unbranched, ampulliform, cylindrical. Conidiogenous cells 0.5–1 μm diam, phiailidic, cylindrical, terminal, slightly tapering towards apex. | Alpha conidia 9–12 × 2–3.5 μm, aseptate, hyaline, ellipsoidal, biguttulate, multiguttulate, or eguttulate, base subtruncate. | Not observed | [23] |
| D. celeris Guarnaccia, Woodhall and Crous 2018 * | Conidiomata pycnidial, globose or irregular, solitary, erumpent, dark brown to black, 350–650 μm diameter, with yellowish translucent to brown conidial cirrhus | Conidiophores hyaline, smooth, unbranched, cylindrical, straight, 5–18 × 1–3 μm. Conidiogenous cells phiailidic, hyaline, cylindrical, 5–8 × 1–2 μm, tapering towards the apex. | Alpha conidia, aseptate, fusiform, hyaline, mono- to biguttulate and acutely rounded at both ends, 5.5–7.5 × 2–3 μm. | Beta conidia hyaline, eguttulate, filiform, curved, tapering towards both ends, 16–22.5 × 1–2 μm | [17] |
| D. chensiensis Tian and Yang 2018 * | Conidiomata pycnidial, immersed in bark, slightly erumpent discoid, ostiolate, 200–325 μm diameter | Conidiophores 8.5–13 × 2–3 μm, cylindrical, hyaline, phiailidic, unbranched, straight or slightly curved, tapering towards the apex. | Alpha conidia hyaline, aseptate, smooth, ellipsoidal, biguttulate, rounded at both ends, 6.5–11 × 2–2.2 μm. | Beta conidia present on the host only, hyaline, eguttulate, smooth, filiform, 21–28.5 × 0.8–1.1 μm | [18] |
| D. ellipicola Gao and Cai 2015 | Pycnidial conidiomata, globose, 141–338 µm diameter, erumpent, single or clustered, extruding yellowish translucent conidial droplets from the ostioles | Conidiophores cylindrical, branched, septate, hyaline, 12–22.4 × 1.1–2 µm, phiailidic, cylindrical, straight, slightly tapering towards the apex. | Alpha conidia 6–8.7 × 2–3 µm, aseptate, hyaline, smooth, biguttulate, oval, ellipsoid rounded at both ends. | Beta conidia 23.4–35.5 × 1.4–2 µm, hyaline, curved | [56] |
| D. eres Nitschke 1870 =D. castaneae-mollisimae =D. cotoneastri =D. fukushi | Pycnidia with 200–250 μm diameter, globose, embedded in tissue, erumpent at maturity, often with yellowish, conidial cirrhus extruding from ostiole | Conidiophores 10–15 × 2–3 μm, hyaline, smooth, unbranched, ampulliform, straight to sinuous. Conidiogenous cells 0.5–1 μm diameter, phiailidic, cylindrical, slightly tapering towards the apex. | Alpha conidia (6–)6.5–8.5(−9) × 3–4 μm, aseptate, hyaline, smooth, ovate to ellipsoidal, often biguttulate, base subtruncate. | Beta conidia 18–29 × 1–1.5 μm, aseptate, hyaline, smooth, fusiform to hooked, base sub-truncate | [23] |
| D. helicis Niessl 1876 * =D. nitschkei | Pycnidia with 200–300 μm diameter, globose, embedded in tissue, erumpent at maturity, often with white conidial cirrhus extruding from ostiole | Conidiophores (6–)8–15 (16.5) × 1–2 μm, hyaline, smooth, unbranched, ampulliform, cylindrical to clavate. Conidiogenous cells phiailidic, cylindrical, tapering towards the apex. | Alpha conidia (5.5–)6–8(9.5) × 2.5–3.5 μm, aseptate, hyaline, smooth, cylindrical to ellipsoidal, biguttulate or multiguttulate, base subtruncate. | Not observed | [23] |
| D. longicicola Gao and Cai 2015 | Conidiomata pycnidial, globose to subglobose, 500–750 µm diameter, dark brown to black, covered with white mycelium | Conidiophores 14.1–22.5 × 1.3–2 µm, hyaline, branched, densely aggregated, cylindrical, tapering towards the apex. | Alpha conidia 5.3–10.4 × 1.5–3.1 µm, with two big guttulate or 2–3 small guttulate, hyaline, ellipsoid or clavate, with one end obtuse and the other end acute and elongate. | Beta conidia filiform, hyaline, hamate or curved, aseptate, 25–32.2 × 1.2–2 µm | [56] |
| D. mahothocarpus Gao and Cai 2015 =Phomopsis mahothocarpus | Conidiomata globose, 200–350 μm diameter, ostiolate, deeply embedded in culture, aggregated in clusters | Conidiophores 15.5–21.8 × 1.6–2.2 μm, cylindrical, hyaline, branched, septate, straight or slightly curved. | Alpha conidia 5.5–8.0 × 1.8–2.9 μm, hyaline, aseptate, oval or fusiform, usually with one guttule at each end. | Beta conidia 21.1–28.5 × 1.2–1.9 μm, aseptate, filiform, hyaline, curved, eguttulate, with obtuse ends | [59] |
| D. maritima Tanney 2016 * | Conidiomata pycnidial, globose to subglobose, unilocular/multilocular, aggregated, dark brown to black, ostiolate, up to 300 μm diameter, with yellowish conidial mass | Conidiogenous cells phiailidic, subcylindrical to ampulliform, straight to sinuous, cylindrical or slightly tapering towards the apex, (8.5–)9–12.5(–16) × 2–3 μm. | Alpha conidia aseptate, hyaline, smooth, oblong to fusiform or ellipsoidal, apex rounded, base subtruncate, bi- to multiguttulate (10–)11–12.5(–13.5) × (3–)3.5–4 μm. | Beta conidia aseptate, hyaline, smooth, straight to hamiform or uncinate 29–40 × 1–2 μm | [60] |
| D. momicola Dissanayake, Li and Hyde 2017 | Conidiomata up to 350 μm diameter, solitary or in groups with black cylindrical ostiolate necks, subglobose | Conidiophores reduced to conidiogenous cells. | Alpha conidia 6.5–9.5 × 1.5–2 μm, hyaline, smooth, biguttulate, fusiform to oval, tapered at both ends, cylindrical to ellipsoidal. | Beta conidia 20–32 × 1–1.5 μm, scattered among the alpha conidia | [61] |
| D. nobilis Saccardo and Spegazzini 1878 | Conidiomata pycnidial, scattered to confluent, uniloculate, dark brown to black, broadly spherical to flattened, 650–700 μm high and 400–500 μm wide | Conidiophores thin walled, brown, vertically aligned, multicellular, 2–6 μm wide, elongate. Conidiogenous cells formed at the apex of the conidiophores cylindric, straight or curved. | Alpha conidia 7–9 × 3–5 μm, aseptate, cylindrical or ellipsoidal, obtuse at both ends, hyaline, generally biguttulate. | Beta conidia 20–30 × 0.3–0.8 μm, filiform, blunt at one end, pointed and usually curved at the other, hyaline, one-celled | [62] |
| D. padina Tian and Yang 2017 * | Conidiomata pycnidial, immersed in bark, scattered, slightly erumpent, light brown, one ostiole, 330–520 μm diameter | Conidiophores 5.5–12.5 × 1–1.5 μm, hyaline, unbranched, cylindrical, straight or slightly curved. | Alpha conidia hyaline, aseptate, ellipsoidal to fusiform, eguttulate, 7–8 × 1.5–2 μm. | Beta conidia hyaline, filiform, straight or hamate, eguttulate, base truncate, 21–24 × 1 µm | [18] |
| D. phragmitis Crous 2014 * | Conidiomata pycnidial, globose, up to 250 µm diameter, black, erumpent, exuding creamy conidial droplets from central ostioles | Conidiophores hyaline, smooth, septate, rarely branched, densely aggregated, cylindrical, 20–30 × 3–4 µm. Conidiogenous cells phiailidic, cylindrical, terminal, intercalary. | Alpha conidia aseptate, hyaline, smooth, multi- or bi-guttulate, fusoid to ellipsoid, tapering towards both ends, base subtruncate, 6–8.5 × 2–3 µm. | Not observed | [63] |
| D. pulla Nitschke 1870 * =Phoma pulla =Phomopsis pulla | Pycnidia with 200–300 μm diameter, globose, embedded in tissue, erumpent at maturity, black stromata, with bright yellow conidial cirrhus | Conidiophores 10–25 × 1–2 μm, hyaline, unbranched, cylindrical to clavate. Conidiogenous cells phiailidic, cylindrical, slightly tapering towards the apex. | Alpha conidia (6–)6.5–7.5 (8) × (2–) 2.5–3.5(−4) μm, aseptate, hyaline, smooth, cylindrical to ellipsoidal, biguttulate or multi-guttulate, base subtruncate. | Not observed | [23] |
| D. rosicola Wanasinghe, Jones and Hyde 2018 * | Conidiomata pycnidial, 120–160 µm diameter, solitary, semi-immersed, unilocular, globose, dark brown, ostiolate | Conidiophores hyaline, smooth, unbranched, cylindrical, straight to sinuous. Conidiogenous cells phiailidic, cylindrical, slightly tapering towards the apex. | Alpha conidia 7–9.5 × 2.4–3 µm, hyaline, biguttulate, fusiform or oval, both ends, obtuse. | Beta conidia 12–22 × 1.2–1.6 µm, hyaline, aseptate, filiform, tapering towards both ends | [64] |
| D. vaccinii Shear 1931 * =Phomopsis vaccinii | Conidiomata superficial, scattered, black, spherical to irregular, uniloculate, with ostiole circular, exuding white to yellowish cirrhus | Conidiogenous cells enteroblastic, phiailidic, with conidiophores short, 1–2 septa or multiseptate, branched. | Alpha conidia 5.9–11.3 × 2.1–3.9 μm, hyaline, fusiform, straight, guttulate, aseptate. | Beta conidia hyaline, filiform, straight or curved, eguttulate, aseptate | [65] |
| D. vacuae Hilário, Santos and Alves 2020 * | Pycnidial conidiomata, brown to black, broadly spherical, covered in white mycelium, with yellowish conidial cirrhus extruding from ostiole | Conidiophores reduced to conidiogenous cells, hyaline, smooth and straight to sinuous, broadening in the base, slightly tapering toward the apex (10.9 ± 2.2 × 1.8 ± 0.3) μm. | Alpha conidia infrequent, hyaline, smooth, cylindrical, 9.3 ± 1.1 × 2.6 ± 0.3 μm. | Beta conidia hyaline, 1-celled, smooth, filiform, frequently hooked in apical part, apex acute, 27.4 ± 2.3 × 1.6 ± 0.2 μm | [29] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hilário, S.; Gonçalves, M.F.M.; Alves, A. Using Genealogical Concordance and Coalescent-Based Species Delimitation to Assess Species Boundaries in the Diaporthe eres Complex. J. Fungi 2021, 7, 507. https://doi.org/10.3390/jof7070507
Hilário S, Gonçalves MFM, Alves A. Using Genealogical Concordance and Coalescent-Based Species Delimitation to Assess Species Boundaries in the Diaporthe eres Complex. Journal of Fungi. 2021; 7(7):507. https://doi.org/10.3390/jof7070507
Chicago/Turabian StyleHilário, Sandra, Micael F. M. Gonçalves, and Artur Alves. 2021. "Using Genealogical Concordance and Coalescent-Based Species Delimitation to Assess Species Boundaries in the Diaporthe eres Complex" Journal of Fungi 7, no. 7: 507. https://doi.org/10.3390/jof7070507
APA StyleHilário, S., Gonçalves, M. F. M., & Alves, A. (2021). Using Genealogical Concordance and Coalescent-Based Species Delimitation to Assess Species Boundaries in the Diaporthe eres Complex. Journal of Fungi, 7(7), 507. https://doi.org/10.3390/jof7070507








