Differential Physiological Prerequisites and Gene Expression Profiles of Conidial Anastomosis Tube and Germ Tube Formation in Colletotrichum gloeosporioides
Abstract
1. Introduction
2. Materials and Methods
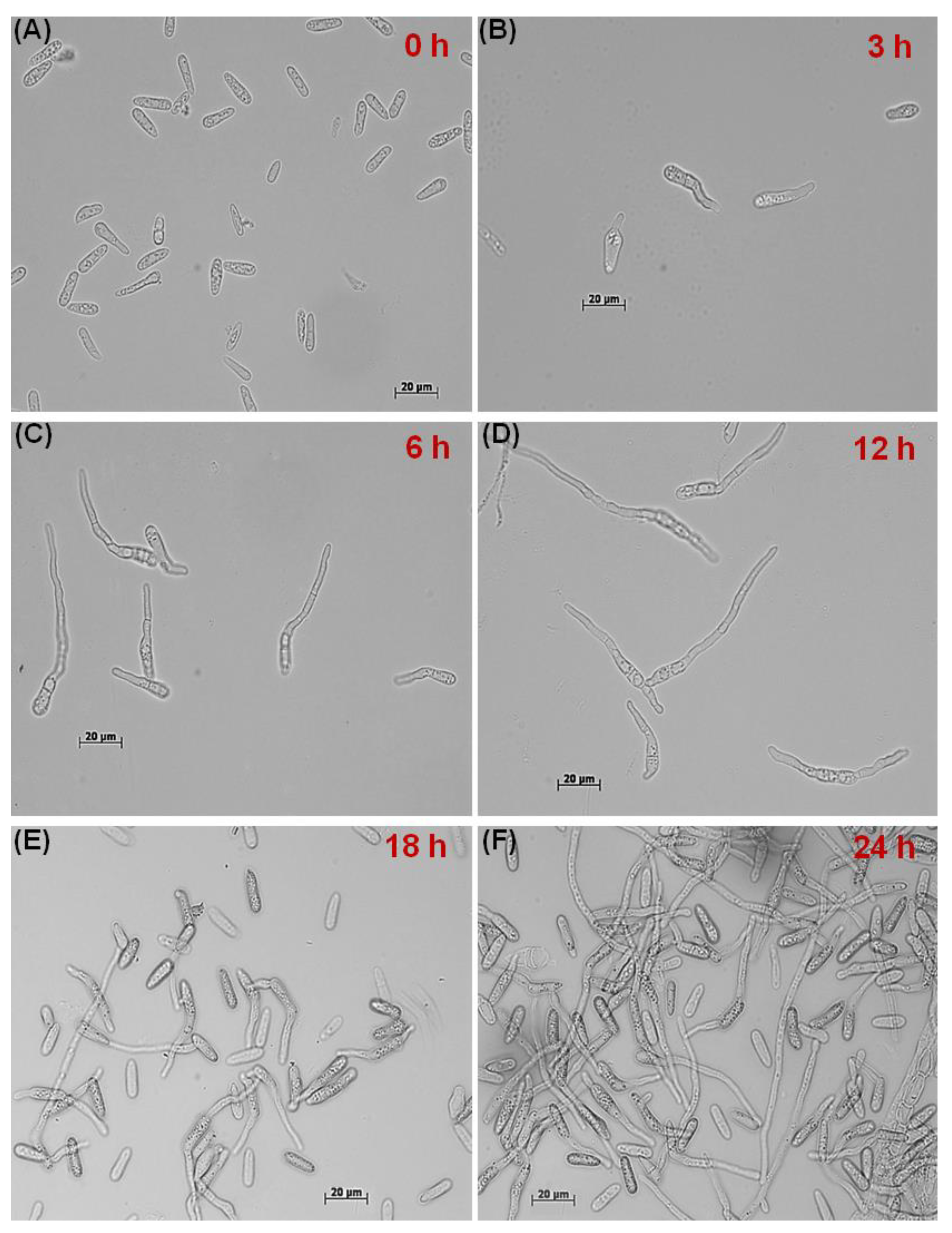
2.1. In Vitro Dynamics of GT Formation and CAT Fusion
2.2. Effects of Availability of Nutrients on GT and CAT Induction
2.3. Whole Transcriptome Analysis of Resting Conidia, GT and CAT
2.3.1. RNA Extraction, Quantification, and Qualification
2.3.2. Library Preparation and Transcriptome Sequencing
2.3.3. Quality Control, De Novo Assembly, and Sequence Clustering
2.3.4. Read Mapping to the Reference Genome and Differential Counting
2.3.5. Gene Ontology Enrichment, Differential Expression Analysis, and Functional Annotation
2.3.6. Transcription Factors and Secreted Protein Analysis
2.3.7. Data Availability
2.4. Real-Time qRT-PCR Validation
2.5. Statistical Analyses
3. Results
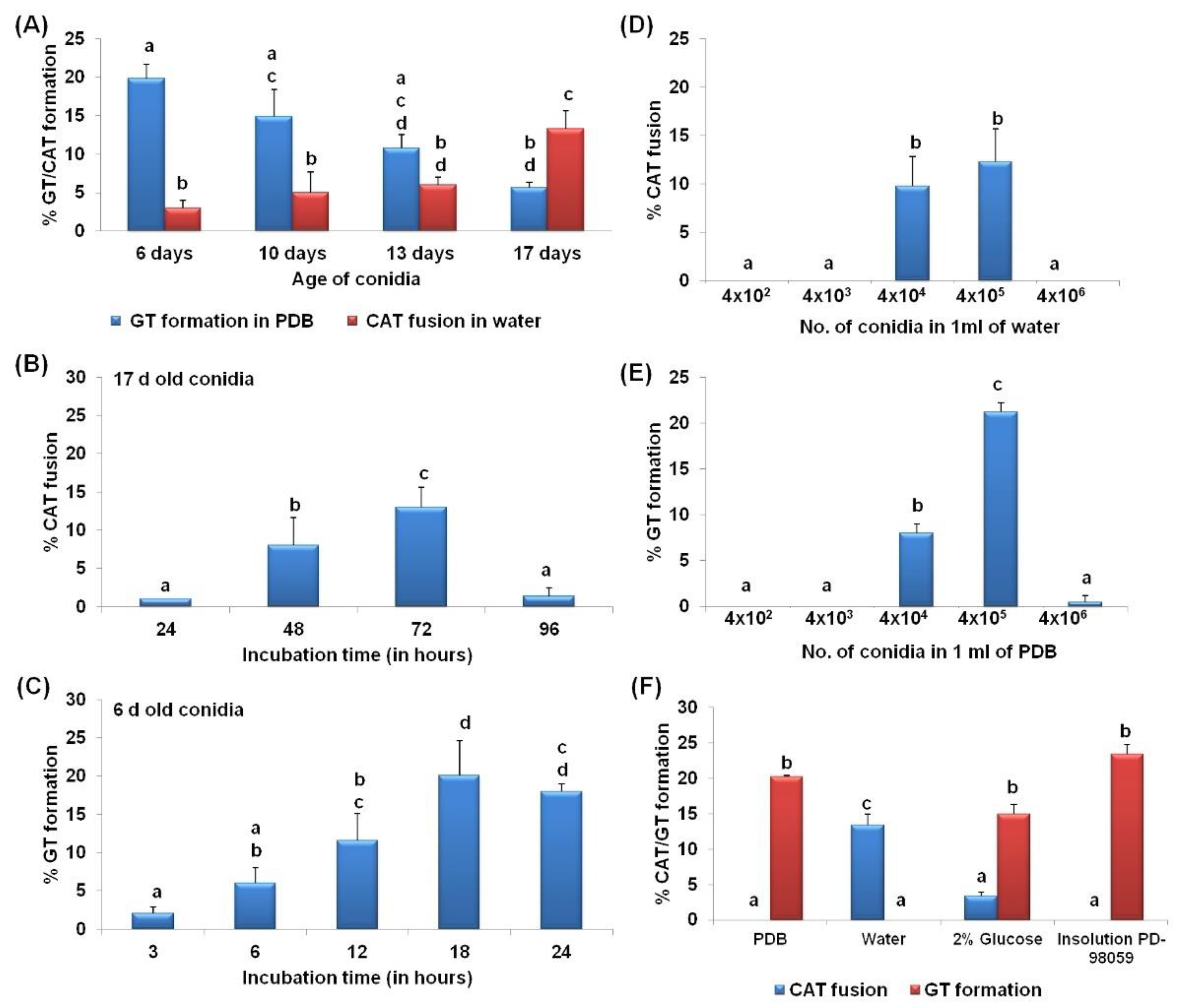
3.1. CAT Fusion and GT Formation Are Mutually Exclusive in C. gloeosporioides
3.2. CAT Fusion and GT Formation Are Dependent on the Conidial Density
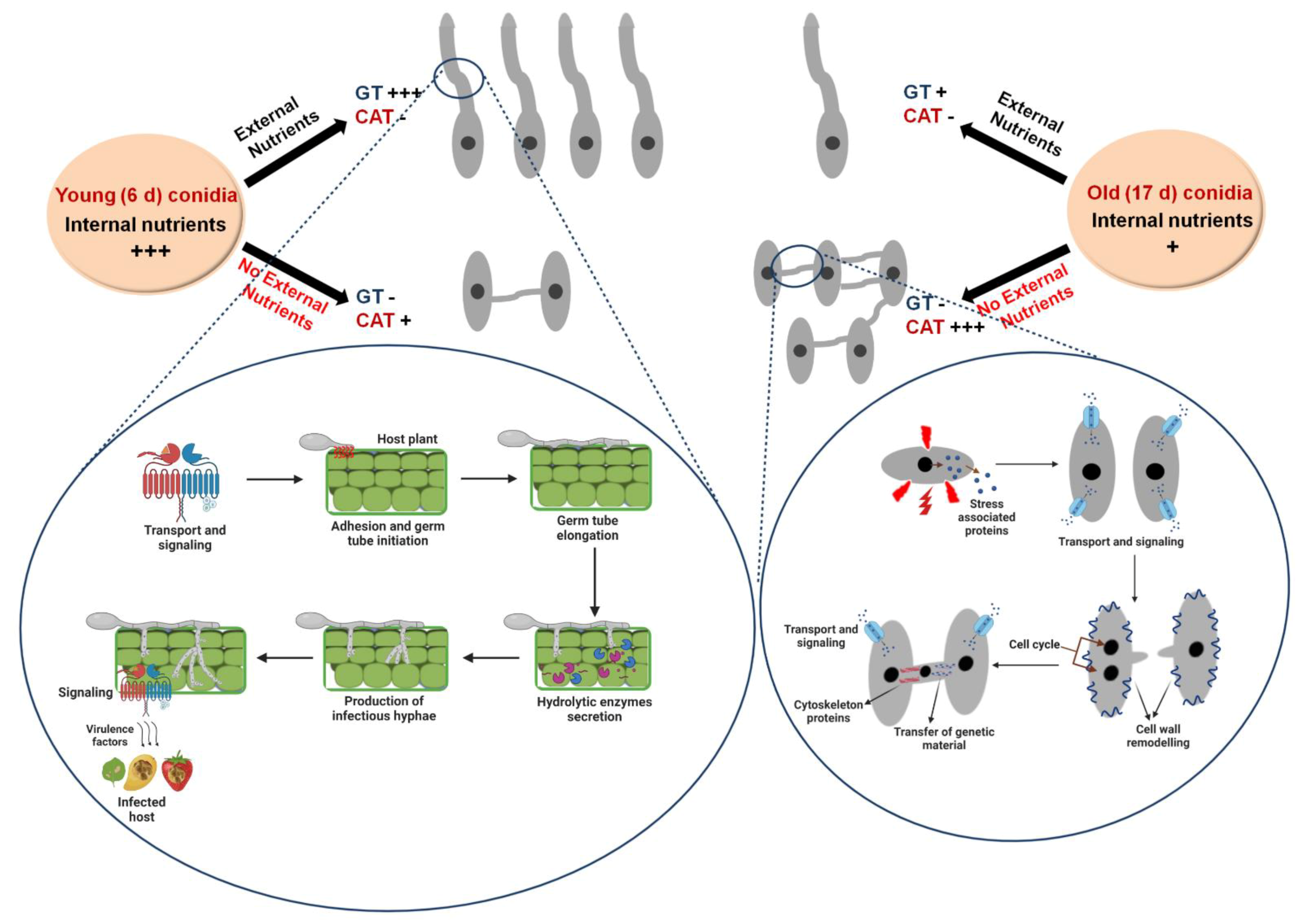
3.3. Differential Nutritional Requirements for CAT versus GT Formation
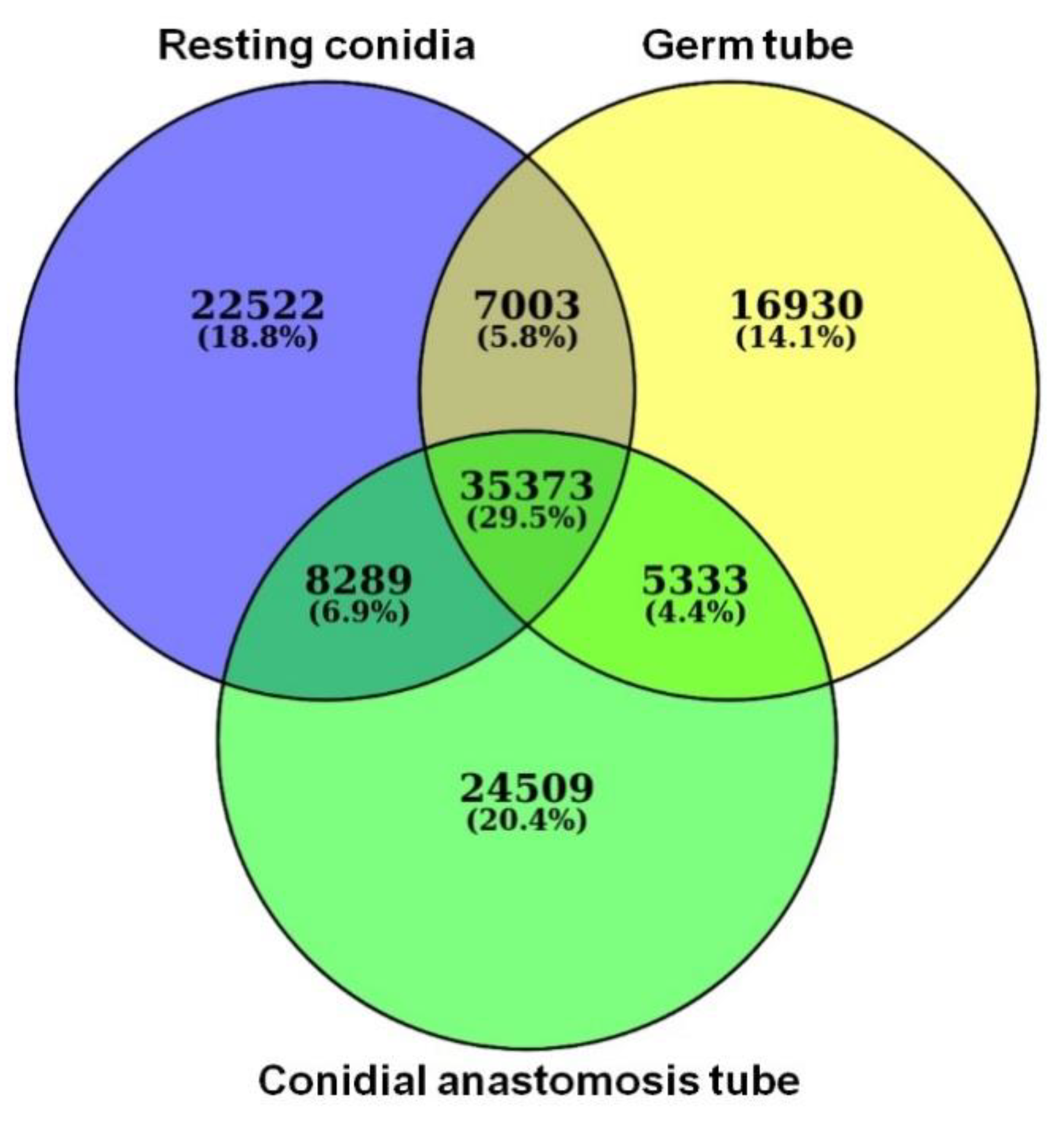
3.4. RNA-Seq Data Analysis
3.5. Gene Ontology Enrichment Analysis of GT and CAT
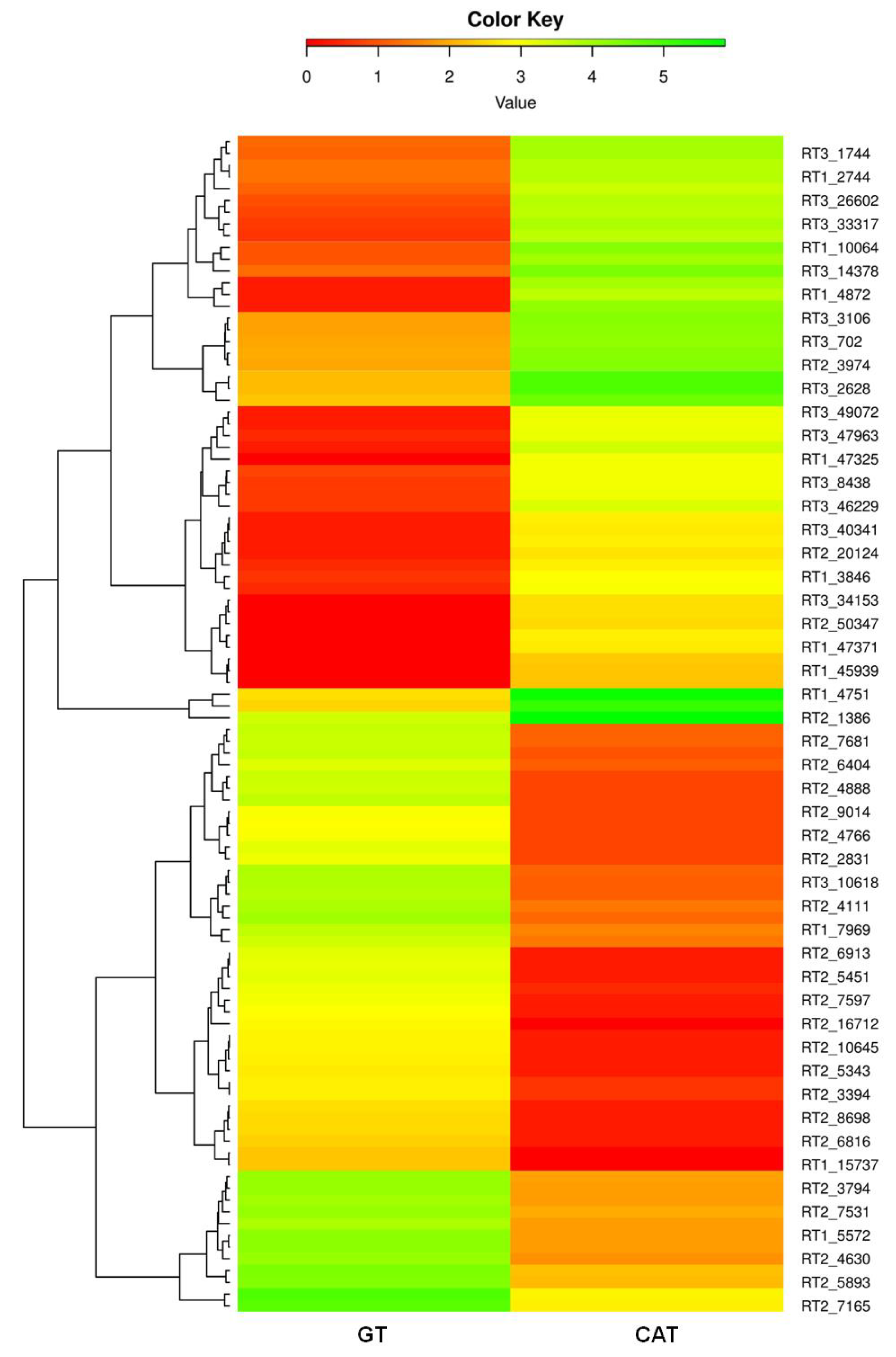
3.6. Identification and Functional Annotation of DEGs in GT and CAT
3.7. Transcription Factor Candidates Involved in GT Formation and CAT Fusion
3.8. Effector Candidates Secreted during GT Formation and CAT Fusion
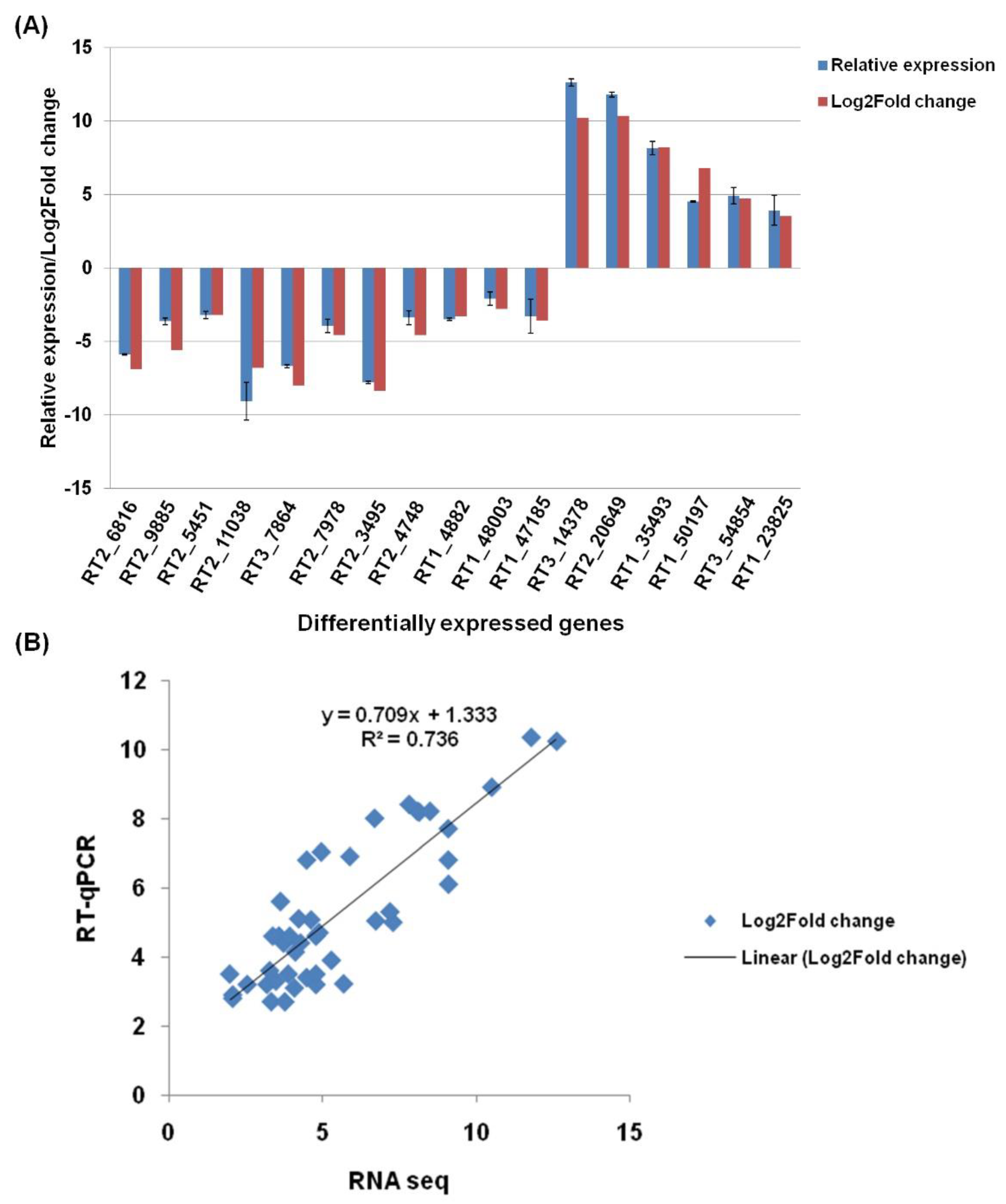
3.9. Real-Time qPCR Validation of Selected DEGs
4. Discussion
4.1. Differential Physiological Requirements for GT and CAT
4.2. Transcriptome Analysis of GT
4.3. Transcriptome Analyses of CAT
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Osherov, N.; May, G.S. The molecular mechanisms of conidial germination. FEMS Microbiol. Lett. 2001, 199, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Van Leeuwen, M.R.; Krijgsheld, P.; Bleichrodt, R.; Menke, H.; Stam, H.; Stark, J.; Wösten, H.A.B.; Dijksterhuis, J. Germination of conidia of Aspergillus niger is accompanied by major changes in RNA profiles. Stud. Mycol. 2013, 74, 59–70. [Google Scholar] [CrossRef] [PubMed]
- Read, N.D.; Lichius, A.; Shoji, J.Y.; Goryachev, A.B. Self-signalling and self-fusion in filamentous fungi. Curr. Opin. Microbiol. 2009, 12, 608–615. [Google Scholar] [CrossRef]
- Kurian, S.M.; Di Pietro, A.; Read, N.D. Live-cell imaging of conidial anastomosis tube fusion during colony initiation in Fusarium oxysporum. PLoS ONE 2018, 13, e0195634. [Google Scholar] [CrossRef] [PubMed]
- Roca, M.G.; Arlt, J.; Jeffree, C.E.; Read, N.D. Cell biology of conidial anastomosis tubes in Neurospora crassa. Eukaryot. Cell 2005, 4, 911–919. [Google Scholar] [CrossRef] [PubMed]
- Roca, M.G.; Davide, L.C.; Mendes-Costa, M.C.; Wheals, A. Conidial anastomosis tubes in Colletotrichum. Fungal Genet. Biol. 2003, 40, 138–145. [Google Scholar] [CrossRef]
- Roca, M.G.; Read, N.D.; Wheals, A.E. Conidial anastomosis tubes in filamentous fungi. FEMS Microbiol. Lett. 2005, 249, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, A.E.; Velho, A.C.; Stadnik, M.J. Formation of conidial anastomosis tubes and melanization of appressoria are antagonistic processes in Colletotrichum spp. from apple. Eur. J. Plant Pathol. 2016, 146, 497–506. [Google Scholar] [CrossRef]
- Shahi, S.; Beerens, B.; Bosch, M.; Linmans, J.; Rep, M. Nuclear dynamics and genetic rearrangement in heterokaryotic colonies of Fusarium oxysporum. Fungal Genet. Biol. 2016, 91, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, F.H.; Souza, E.A.; Read, N.D.; Roca, M.G. Live-cell imaging of conidial fusion in the bean pathogen, Colletotrichum lindemuthianum. Fungal Biol. 2010, 114, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Mehta, N.; Baghela, A. Quorum sensing-mediated inter-specific conidial anastomosis tube fusion between Colletotrichum gloeosporioides and C. siamense. IMA Fungus 2021, 12, 1–33. [Google Scholar] [CrossRef]
- Nordzieke, D.E.; Sanken, A.; Antelo, L.; Raschke, A.; Deising, H.B.; Pöggeler, S. Specialized infection strategies of falcate and oval conidia of Colletotrichum graminicola. Fungal Genet. Biol. 2019, 133, 103276. [Google Scholar] [CrossRef]
- Ishikawa, F.H.; Souza, E.A.; Shoji, J.Y.; Connolly, L.; Freitag, M.; Read, N.D.; Roca, M.G. Heterokaryon incompatibility is suppressed following conidial anastomosis tube fusion in a fungal plant pathogen. PLoS ONE 2012, 7, e31175. [Google Scholar] [CrossRef]
- Roca, M.G.; Davide, L.C.; Davide, L.M.C.; Mendes-Costa, M.C.; Schwan, R.F.; Wheals, A.E. Conidial anastomosis fusion between Colletotrichum species. Mycol. Res. 2004, 108, 1320–1326. [Google Scholar] [CrossRef]
- De Silva, D.D.; Crous, P.W.; Ades, P.K.; Hyde, K.D.; Taylor, P.W.J. Life styles of Colletotrichum species and implications for plant biosecurity. Fungal Biol. Rev. 2017, 31, 155–168. [Google Scholar] [CrossRef]
- Araujo, L.; Stadnik, M.J. Multiple appressoria and conidial anastomoses tubes in the infectious process of Colletotrichum gloeosporioides in apple trees. Bragantia 2013, 72, 180–183. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Götz, S.; García-Gómez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Petersen, T.N.; Brunak, S.; Von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef]
- Emanuelsson, O.; Brunak, S.; von Heijne, G.; Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protoc. 2007, 2, 953. [Google Scholar] [CrossRef]
- Möller, S.; Croning, M.D.R.; Apweiler, R. Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 2001, 17, 646–653. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef]
- Rudd, J.J.; Kanyuka, K.; Hassani-Pak, K.; Derbyshire, M.; Andongabo, A.; Devonshire, J.; Lysenko, A.; Saqi, M.; Desai, N.M.; Powers, S.J.; et al. Transcriptome and metabolite profiling of the infection cycle of Zymoseptoria tritici on wheat reveals a biphasic interaction with plant immunity involving differential pathogen chromosomal contributions and a variation on the hemibiotrophic lifestyle def. Plant Physiol. 2015, 167, 1158–1185. [Google Scholar] [CrossRef]
- Schefe, J.H.; Lehmann, K.E.; Buschmann, I.R.; Unger, T.; Funke-Kaiser, H. Quantitative real-time RT-PCR data analysis: Current concepts and the novel “gene expression’s C T difference” formula. J. Mol. Med. 2006, 84, 901–910. [Google Scholar] [CrossRef] [PubMed]
- Oliva, R.; Win, J.; Raffaele, S.; Boutemy, L.; Bozkurt, T.O.; Chaparro-Garcia, A.; Segretin, M.E.; Stam, R.; Schornack, S.; Cano, L.M.; et al. Recent developments in effector biology of filamentous plant pathogens. Cell. Microbiol. 2010, 12, 705–715. [Google Scholar] [CrossRef] [PubMed]
- Leu, L.S. Anastomosis in Venturia Inaequalis; University of Wisconsin: Madison, WI, USA, 1967. [Google Scholar]
- Fischer-Harman, V.; Jackson, K.J.; Muñoz, A.; Shoji, J.Y.; Read, N.D. Evidence for tryptophan being a signal molecule that inhibits conidial anastomosis tube fusion during colony initiation in Neurospora crassa. Fungal Genet. Biol. 2012, 49, 896–902. [Google Scholar] [CrossRef] [PubMed]
- Palma-Guerrero, J.; Huang, I.C.; Jansson, H.B.; Salinas, J.; Lopez-Llorca, L.V.; Read, N.D. Chitosan permeabilizes the plasma membrane and kills cells of Neurospora crassa in an energy dependent manner. Fungal Genet. Biol. 2009, 46, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Zhou, Z.; Wu, J.; Ji, Z.; Zhang, J. Comparative transcriptome analysis reveals significant differences in gene expression between appressoria and hyphae in Colletotrichum gloeosporioides. Gene 2018, 670, 63–69. [Google Scholar] [CrossRef]
- Mims, C.W.; Richardson, E.A.; Clay, R.P.; Nicholson, R.L. Ultrastructure of conidia and the conidium aging process in the plant pathogenic fungus Colletotrichum graminicola. Int. J. Plant Sci. 1995, 156, 9–18. [Google Scholar] [CrossRef]
- Zhang, L.; Huang, X.; He, C.; Zhang, Q.Y.; Zou, X.; Duan, K.; Gao, Q. Novel fungal pathogenicity and leaf defense strategies are revealed by simultaneous transcriptome analysis of Colletotrichum fructicola and strawberry infected by this fungus. Front. Plant Sci. 2018, 9, 434. [Google Scholar] [CrossRef] [PubMed]
- Tucker, S.L.; Talbot, N.J. Surface attachment and pre-penetration stage development by plant pathogenic fungi. Annu. Rev. Phytopathol. 2001, 39, 385–417. [Google Scholar] [CrossRef] [PubMed]
- Ohtaki, S.; Maeda, H.; Takahashi, T.; Yamagata, Y.; Hasegawa, F.; Gomi, K.; Nakajima, T.; Abe, K. Novel hydrophobic surface binding protein, HsbA, produced by Aspergillus oryzae. Appl. Environ. Microbiol. 2006, 72, 2407–2413. [Google Scholar] [CrossRef]
- Sun, C.B.; Suresh, A.; Deng, Y.Z.; Naqvi, N.I. A multidrug resistance transporter in Magnaporthe is required for host penetration and for survival during oxidative stress. Plant Cell 2006, 18, 3686–3705. [Google Scholar] [CrossRef]
- Gupta, A.; Chattoo, B.B. Functional analysis of a novel ABC transporter ABC4 from Magnaporthe grisea. FEMS Microbiol. Lett. 2008, 278, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Stefanato, F.L.; Abou-Mansour, E.; Buchala, A.; Kretschmer, M.; Mosbach, A.; Hahn, M.; Bochet, C.G.; Métraux, J.P.; Schoonbeek, H.J. The ABC transporter BcatrB from Botrytis cinerea exports camalexin and is a virulence factor on Arabidopsis thaliana. Plant J. 2009, 58, 499–510. [Google Scholar] [CrossRef]
- Zhou, Z.; Wu, J.; Wang, M.; Zhang, J. ABC protein CgABCF2 is required for asexual and sexual development, appressorial formation and plant infection in Colletotrichum gloeosporioides. Microb. Pathog. 2017, 110, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Gan, P.; Ikeda, K.; Irieda, H.; Narusaka, M.; O’Connell, R.J.; Narusaka, Y.; Takano, Y.; Kubo, Y.; Shirasu, K. Comparative genomic and transcriptomic analyses reveal the hemibiotrophic stage shift of Colletotrichum fungi. New Phytol. 2013, 197, 1236–1249. [Google Scholar] [CrossRef]
- Muszewska, A.; Stepniewska-Dziubinska, M.M.; Steczkiewicz, K.; Pawlowska, J.; Dziedzic, A.; Ginalski, K. Fungal lifestyle reflected in serine protease repertoire. Sci. Rep. 2017, 7, 1–2. [Google Scholar] [CrossRef]
- Le Govic, Y.; Papon, N.; Le Gal, S.; Bouchara, J.P.; Vandeputte, P. Non-ribosomal Peptide Synthetase Gene Clusters in the Human Pathogenic Fungus Scedosporium apiospermum. Front. Microbiol. 2019, 10, 2062. [Google Scholar] [CrossRef]
- Kleemann, J.; Rincon-Rivera, L.J.; Takahara, H.; Neumann, U.; van Themaat, E.V.L.; van der Does, H.C.; Hacquard, S.; Stüber, K.; Will, I.; Schmalenbach, W.; et al. Sequential delivery of host-induced virulence effectors by appressoria and intracellular hyphae of the phytopathogen Colletotrichum higginsianum. PLoS Pathog. 2012, 8, e1002643. [Google Scholar] [CrossRef]
- Fischer, M.S.; Glass, N.L. Communicate and fuse: How filamentous fungi establish and maintain an interconnected mycelial network. Front. Microbiol. 2019, 10, 619. [Google Scholar] [CrossRef]
- Cano-Domínguez, N.; Bowman, B.; Peraza-Reyes, L.; Aguirre, J. Neurospora crassa NADPH Oxidase NOX-1 Is Localized in the Vacuolar System and the Plasma Membrane. Front. Microbiol. 2019, 10, 1–14. [Google Scholar] [CrossRef]
- Nordzieke, D.E.; Fernandes, T.R.; El Ghalid, M.; Turrà, D.; Di Pietro, A. NADPH oxidase regulates chemotropic growth of the fungal pathogen Fusarium oxysporum towards the host plant. New Phytol. 2019, 224, 1600–1612. [Google Scholar] [CrossRef]
- Vangalis, V.; Knop, M.; Typas, M.A.; Papaioannou, I.A. Establishment of conidial fusion in the asexual fungus Verticillium dahliae as a useful system for the study of non-sexual genetic interactions. Curr. Genet. 2021, 67, 471–485. [Google Scholar] [CrossRef] [PubMed]
- Roca, G.M.; Weichert, M.; Siegmund, U.; Tudzynski, P.; Fleißner, A. Germling fusion via conidial anastomosis tubes in the grey mould Botrytis cinerea requires NADPH oxidase activity. Fungal Biol. 2012, 116, 379–387. [Google Scholar] [CrossRef]
- Petti, A.A.; Crutchfield, C.A.; Rabinowitz, J.D.; Botstein, D. Survival of starving yeast is correlated with oxidative stress response and nonrespiratory mitochondrial function. Proc. Natl. Acad. Sci. USA 2011, 108, E1089–E1098. [Google Scholar] [CrossRef]
- Jakubowski, W.; Biliński, T.; Bartosz, G. Oxidative stress during aging of stationary cultures of the yeast Saccharomyces cerevisiae. Free Radic. Biol. Med. 2000, 28, 659–664. [Google Scholar] [CrossRef]
- Roca, M.G.; Kuo, H.C.; Lichius, A.; Freitag, M.; Read, N.D. Nuclear dynamics, mitosis, and the cytoskeleton during the early stages of colony initiation in Neurospora crassa. Eukaryot. Cell 2010, 9, 1171–1183. [Google Scholar] [CrossRef][Green Version]
- Leeder, A.C.; Jonkers, W.; Li, J.; Louise Glass, N. Early colony establishment in Neurospora crassa requires a MAP kinase regulatory network. Genetics 2013, 195, 883–898. [Google Scholar] [CrossRef] [PubMed]
- Dekhang, R.; Wu, C.; Smith, K.M.; Lamb, T.M.; Peterson, M.; Bredeweg, E.L.; Ibarra, O.; Emerson, J.M.; Karunarathna, N.; Lyubetskaya, A.; et al. The Neurospora transcription factor ADV-1 transduces light signals and temporal information to control rhythmic expression of genes involved in cell fusion. G3 Genes Genomes Genet. 2017, 7, 129–142. [Google Scholar] [CrossRef] [PubMed]
- Maddi, A.; Dettman, A.; Fu, C.; Seiler, S.; Free, S.J. WSC-1 and HAM-7 are MAK-1 MAP kinase pathway sensors required for cell wall integrity and hyphal fusion in Neurospora crassa. PLoS ONE 2012, 7, e42374. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Palma-Guerrero, J.; Hall, C.R.; Kowbel, D.; Welch, J.; Taylor, J.W.; Brem, R.B.; Glass, N.L. Genome Wide Association Identifies Novel Loci Involved in Fungal Communication. PLoS Genet. 2013, 9, e1003669. [Google Scholar] [CrossRef]
- Takeshita, N.; Evangelinos, M.; Zhou, L.; Serizawa, T.; Somera-Fajardo, R.A.; Lu, L.; Takaya, N.; Nienhaus, G.U.; Fischer, R. Pulses of Ca2+ coordinate actin assembly and exocytosis for stepwise cell extension. Proc. Natl. Acad. Sci. USA 2017, 114, 5701–5706. [Google Scholar] [CrossRef]


| Transcript ID | Gene Name | Possible Functions | GT | CAT |
|---|---|---|---|---|
| Cell wall degrading enzymes | ||||
| RT2_5451 | Pectin lyase | Degrades pectin | 1712 | 2 |
| RT2_4111 | Glycosyl hydrolase family 76 | Degrades cellulose, hemicellulose, and lignin found in plant cell wall | 7920 | 24 |
| RT2_6404 | Mandelate racemase/muconate lactonizing enzyme domain-containing protein | Involved in the breakdown of lignin-derived aromatics | 1848 | 13 |
| RT2_6816 | Fungal cellulose binding domain-containing protein | Active in plant cell-wall hydrolysis | 254 | 2 |
| Host-fungus interaction and germination | ||||
| RT2_16712 | Quinone reductase | Host-fungus interaction & infection | 761 | 1 |
| RT1_10305 | Hydrophobin 2 | Involved in adhesion | 9563 | 16 |
| RT1_4917 | Cerato-platanin | Role during fungus-plant interactions and infection | 16,867 | 62 |
| RT2_5893 | GPI-anchored cell wall beta-1,3-endoglucanase EglC | Role in germination | 21,332 | 143 |
| RT3_7789 | Hydrophobic surface binding protein A | Increase the hydrophobicity of conidia, aerial hyphae, and fruiting bodies | 86,745 | 621 |
| Transport and signaling | ||||
| RT2_6913 | Fg-gap repeat protein | Important for ligand binding | 1571 | 2 |
| RT2_4144 | Phosphate-repressible phosphate permease/transporter | Membrane transport proteins | 1556 | 2 |
| RT2_4587 | Transmembrane amino acid transporter | Ubiquitin dependent endocytosis and signaling responses | 3406 | 6 |
| RT1_15737 | MFS monosaccharide transporter | Membrane transport proteins | 182 | 1 |
| RT2_4766 | High affinity methionine permease | Cysteine and methionine transport across plasma membrane | 1027 | 6 |
| RT3_9669 | Uso1/p115 like vesicle tethering protein | Intracellular protein transport | 24,227 | 173 |
| Virulence | ||||
| RT2_3663 | Peptidase S41 family protein | Involved in pathogenesis | 4430 | 6 |
| RT2_27038 | nrps-like protein | Involved in virulence | 1299 | 3 |
| RT2_3495 | Polysaccharide deacetylase | Involved in virulence | 3561 | 10 |
| RT2_10908 | FGGY-family pentulose kinase | Involved in virulence | 625 | 2 |
| RT1_5572 | Acetylornithine aminotransferase | Required for growth, conidiogenesis, and pathogenicity | 17,456 | 64 |
| RT2_5240 | GMC oxidoreductase | Role in pathogenicity | 324 | 2 |
| RT2_2131 | GNAT family acetyltransferase | Role in chitin metabolism of fungi development and pathogenicity | 10,433 | 68 |
| RT2_7531 | CAP-22 protein | Expressed in the conidium during appressorium formation | 13,700 | 100 |
| RT1_7969 | NmrA family transcriptional regulator | Required for the invasive virulence | 4225 | 33 |
| RT1_7368 | Cyanide hydratase | Catalyzes the hydration of cyanide to formamide. It may be necessary for plant pathogenic fungi in infection of cyanogenic plants | 2902 | 23 |
 .
.| Transcript ID | Gene Name | Possible Function | CAT | GT |
|---|---|---|---|---|
| Stress associated genes | ||||
| RT1_6087 | Taurine catabolism dioxygenase | Utilization of taurine under sulfate starvation | 8376 | 2 |
| RT3_33317 | Pyridine nucleotide-disulfide | Involved in cellular oxidative stress response | 7093 | 5 |
| RT3_14378 | Formate dehydrogenase | Role under hypoxia stress response | 24,057 | 20 |
| RT3_26602 | NADH dehydrogenase | Role in oxidative stress | 8132 | 14 |
| RT1_5411 | Oxalate decarboxylase family bicupin | Role in stress response | 8913 | 17 |
| RT3_3106 | Maleylacetate reductase | Has an oxidoreductase activity | 19,971 | 75 |
| RT1_51991 | Proline dehydrogenase | Role in osmotic, drought, and salinity stress | 213 | 1 |
| RT1_52730 | O-acetyl homo serine amino carboxypropyl transferase | Involved in oxidative stress | 773 | 14 |
| RT1_51711 | Iron sulfur cluster assembly protein 1 | Involved in oxidative stress | 79 | 2 |
| RT1_51144 | bZIP transcription factor | Responses to oxidative stress | 266 | 21 |
| RT1_59783 | HhH-GPD superfamily base excision DNA repair protein | Involve in stress and hormone signaling | 24 | 2 |
| RT1_47091 | Ribulose-phosphate 3-epimerase | Cellular response to oxidative stress | 34 | 3 |
| RT3_1744 | NADPH cytochrome p450 | Role in redox signaling | 5805 | 8 |
| RT1_54531 | Stress response regulator SrrA | Involved in stress signal transduction | 62 | 2 |
| RT1_20193 | Dipeptidase A | Role in fungal growth and heat shock stress | 13 | 2 |
| Cell wall, membrane transport, and signaling | ||||
| RT1_11804 | MFS transporter | Membrane transport proteins | 14,522 | 2 |
| RT1_871 | Transmembrane amino acid transporter | Promoting endocytosis and triggering signaling responses | 19,654 | 95 |
| RT3_59479 | Peptidoglycan-N-acetylglucosamine deacetylase | Involved in cell wall morphogenesis and remodeling | 201 | 6 |
| RT1_49642 | Alcohol dehydrogenase GroES-like domain | Functions in protein folding and intercellular signaling | 59 | 2 |
| RT3_50435 | Flippase-like domain-containing protein | Pheromone response, cytoskeletal dynamics, cell division, lipid metabolism, and lipid signaling | 76 | 3 |
| RT3_62264 | Acetyl-CoA carboxylase biotin carboxylase subunit | Fatty acid synthesis pathway | 20 | 1 |
| RT3_59549 | Glycosyltransferase family 2 protein | Cell wall synthesis and other functions | 79 | 5 |
| Cytoskeleton, cell cycle, and cell rescue | ||||
| RT1_7001 | MreB/Mbl protein | Constituents of the eukaryotic cytoskeleton (tubulin, actin) | 2907 | 2 |
| RT3_59288 | F-box domain protein | Involved in cell division cycle, glucose sensing, and stress response | 1332 | 2 |
| RT3_43400 | DNA repair protein | Protects cells from DNA damage-induced genome instability | 597 | 1 |
| RT1_35493 | Zn 2cys6 transcription factor | Involved in meiosis, stress response, and pleiotropic drug resistance | 582 | 2 |
| RT3_50458 | MutS domain V | DNA mismatch repair protein | 562 | 3 |
| RT1_51248 | Actin | Major components of the cytoskeleton | 4419 | 131 |
| Pathogenicity associated proteins | ||||
| RT1_2872 | Tetraspanin 10 | Small integral membrane proteins, required for pathogenicity | 17,772 | 71 |
| RT3_8438 | Integral membrane protein | Required for pathogenicity | 1046 | 5 |
 .
.| TF Family Name | Number of TF Families | |
|---|---|---|
| GT Formation | CAT Fusion | |
| APSES | 16 | 14 |
| AraC type | 509 | 443 |
| AT-rich interaction region | 8 | 13 |
| BED-type predicted | 4 | 2 |
| bHLH | 35 | 38 |
| Bromodomain transcription factor | 9 | 8 |
| bZIP | 109 | 130 |
| C2H2 zinc finger | 590 | 665 |
| CCHC-type | 142 | 179 |
| DHHC-type | 34 | 43 |
| Forkhead | 9 | 15 |
| GATA type zinc finger | 54 | 53 |
| Grainyhead/CP2 | 5 | 4 |
| GRF-type | 12 | 7 |
| Helix-turn-helix type 3 | 3 | 4 |
| HMG | 228 | 264 |
| Homeobox | 24 | 16 |
| Homeodomain-like | 221 | 226 |
| HTH | 1 | 2 |
| LSD1-type | 13 | 18 |
| MADS-box | 31 | 27 |
| MIZ-type | 13 | 10 |
| Myb | 60 | 62 |
| NDT80/PhoG like DNA-binding | 4 | 5 |
| Negative transcriptional regulator | 7 | 8 |
| NF-X1-type | 8 | 17 |
| Not1 | 7 | 10 |
| OB-fold | 500 | 540 |
| p53-like transcription factor | 9 | 12 |
| Psq | 6 | 4 |
| Rad18-type putative | 28 | 30 |
| ssDNA-binding transcriptional regulator | 16 | 20 |
| TEA/ATTS | 20 | 24 |
| Transcription factor jumonji | 9 | 12 |
| Transcription factor TFIIS | 13 | 11 |
| Tubby transcription factors | 10 | 9 |
| UAF complex subunit Rrn10 | 3 | 4 |
| Zn2Cys6 | 2057 | 2134 |
 .
.| Transcript ID | Protein Names | Possible Functions |
|---|---|---|
| Hydrolytic enzymes | ||
| RT2_4955.p1 | Acetyl esterase | Degradation of hemicelluloses and pectin |
| RT2_12928.p2 | Alkaline phosphatase | Role in P mobilization from organic substrates under P starvation conditions |
| RT2_9649.p1 | Amidohydrolase | Type of hydrolase that acts upon amide bonds |
| RT2_4824.p1 | Berberine bridge enzyme | It is a cellobiose oxidase that degrades the carboxymethylcellulose (CMC), xylan, and lignin |
| RT2_14227.p1 | Beta-1,3-endoglucanase | Role in cell wall softening |
| RT2_10729.p1 | Endo-chitosanase | Role in degradation of the deacetylated portion of chitin in the fungal cell wall |
| RT2_9596.p1 | Extracellular cellulase | Degrades cellulose and some other related polysaccharides |
| RT2_1942.p1 | Glycosyl hydrolase family 15 | Degrades cellulose, hemicellulose, and lignin found in plant cell wall |
| RT2_15512.p2 | Glycosyl hydrolase family 28 | Degrades cellulose, hemicellulose, and lignin found in plant cell wall |
| RT2_618.p1 | Glycosyl hydrolase family 31 | Degrades cellulose, hemicellulose, and lignin found in plant cell wall |
| RT2_12327.p1 | Xylosidase arabinofuranosidase | Role in xylan degradation |
| Adhesion, germination, and hyphal development | ||
| RT2_1963.p1 | Acid trehalase | Provides energy during conidial germination |
| RT2_6726.p1 | Carbonic anhydrase | Role in fruiting body development and ascospore germination |
| RT2_2892.p2 | Endoglucanase 3 | Role in germination |
| RT2_10852.p2 | Fungal hydrophobin | Role in adhesion |
| RT2_5434.p1 | Glycosyltransferase sugar-binding region containing DXD motif | Role in pathogenesis of plants by enabling hyphal growth |
| RT2_4975.p1 | Manganese lipoxygenase | Accelerates programmed spore germination |
| RT2_13809.p1 | Pollen proteins Ole e I like | Potentially implicated in pollen germination |
| RT2_18566.p1 | Spindle poison sensitivity protein | Role as spindle machinery for chromosome segregation and cytokinesis |
| RT2_10806.p1 | WD40 domain protein beta propeller | Regulates fungal cell differentiation |
| RT2_11269.p2 | Acid phosphatase, putative | Involved in fungal growth |
| RT2_4368.p1 | Class III aminotransferase, putative | Regulate the fungus-host interaction |
| Transport and signaling | ||
| RT2_12723.p2 | Bacterial extracellular solute-binding protein, family 3 | Act as chemoreceptors, recognition constituents of transport systems, and initiators of signal transduction pathways |
| RT2_18223.p1 | Eukaryotic porin | It is an important regulator of Ca2+ transport in and out of the mitochondria |
| RT2_1739.p1 | WSC domain-containing protein | Responses to stress cues and metal ions |
| Virulence | ||
| RT2_4851.p1 | Calcineurin-like phosphoesterase | Role in stress survival, sexual differentiation, and virulence |
| RT2_11269.p2 | Phosphoinositide phospholipase C, Ca2+-dependent | Roles in growth, stress tolerance, sexual development, and virulence |
| RT2_3804.p1 | Secretory lipase, putative | Potential virulence factors |
| RT2_5435.p1 | Versicolorin b synthase | Role in the biosynthesis of aflatoxins |
| RT2_15735.p1 | Zinc-binding dehydrogenase | Role in infection |
| Transcript ID | Protein Names | Possible Functions |
|---|---|---|
| Stress associated | ||
| RT3_6147_m.31990 | Acetyl-CoA acetyltransferase | Role in oxidative and cell wall stresses |
| RT3_8826_m.37297 | Calcineurin-like phosphoesterase | Involved in stress survival and fungal virulence |
| RT3_10144_m.39239 | Chitin synthesis regulation, resistance to Congo red | Role in cell wall stress, noxious chemicals, and osmotic pressure changes |
| RT3_12971_m.42487 | Cytochrome oxidase assembly protein | Cox11p is an integral protein of the inner mitochondrial membrane that is essential for cytochrome c oxidase assembly. Role in response to hydrogen peroxide exposure |
| RT3_12332_m.41840 | LipA and NB-ARC domain-containing protein | Function as key integrators of stress and nutrient availability signals |
| RT3_571_m.6736 | Pyridoxamine 5’-phosphate oxidase | Role in oxidative stress |
| RT3_15296_m.44520 | Spherulation-specific family 4 | Required for spherulation under starvation conditions |
| RT3_2386_m.18522 | SWIM zinc finger | Roles in the stress response and virulence |
| RT3_10404_m.39608 | WSC domain-containing protein | Responses to stress cues and metal ions |
| Cell wall, membrane transport, and signaling | ||
| RT3_17141_m.45950 | Ankyrin-3-like protein 3 | Roles in cell motility, activation, proliferation, contact, and the maintenance of specialized membrane domains |
| RT3_870_m.9203 | Aspartic endopeptidase | Role in cell-wall assembly, remodeling, and cell wall integrity |
| RT3_2825_m.20686 | Blastomyces yeast-phase-specific protein | Required for cell wall morphogenesis and pathogenesis |
| RT3_100_m.1747 | Cytochrome p450 | Role in redox signaling |
| RT3_6853_m.33623 | Endo-beta-1,6-glucanase | Cell wall morphogenesis and remodeling |
| RT3_3953_m.25310 | Gamma-glutamyltranspeptidase | Role in the vacuolar transport and metabolism of glutathione |
| RT3_2987_m.21396 | GPI biosynthesis protein family Pig-F | Required for Cell Wall Biogenesis and Normal Hyphal Growth |
| RT3_25_m.564 | Hkr1p | Regulate the filamentation and mitogen-activated protein kinase pathway |
| RT3_2892_m.20978 | Mid2 like cell wall stress sensor | Required for stress-induced nuclear to cytoplasmic translocation of Cyclin C and cell wall integrity signaling pathway |
| RT3_18063_m.46545 | MIP family channel protein | Involved in the transport of water and neutral solutes across the membranes |
| RT3_268_m.3768 | NCS1 nucleoside transporter | Responsible for uptake of purines, pyrimidines, or related metabolites |
| RT3_704_m.7811 | PAN domain | Essential for RasA-mediated morphogenetic signaling |
| RT3_5718_m.30849 | Pectin esterase | Facilitate plant cell wall modification and subsequent breakdown |
| RT3_18612_m.46914 | PQ loop repeat | Function as cargo receptors in vesicle transport |
| RT3_11944_m.41457 | Probable lipid transfer | Involved in lipid transfer |
| RT3_21269_m.48652 | Protein serine/threonine kinase | Roles in cellular function such as regulation of signaling pathways |
| RT3_3541_m.23752 | Protein YOP1 | Involved in membrane/vesicle trafficking |
| RT3_543_m.6478 | Proteolipid membrane potential modulator | Pmp3 is transcriptionally regulated by the Spc1 stress MAPK (mitogen-activated protein kinases) pathway |
| RT3_2326_m.18214 | SOCE-associated regulatory factor of calcium homoeostasis | Negative regulator of intracellular calcium signaling |
| RT3_2348_m.18321 | Stretch-activated Ca2+-permeable channel component | Required for Ca2+ influx induced by the mating pheromone, alpha-factor |
| Cytoskeleton, cell cycle, and fungal development | ||
| RT3_12912_m.42432 | Alkaline proteinase | Role in fungal physiology and development |
| RT3_20705_m.48270 | Coronin | Role in the organization and dynamics of actin, F-actin remodeling |
| RT3_13797_m.43265 | Frequency clock protein | Regulates various aspects of the circadian clock in Neurospora crassa |
| RT3_16749_m.45643 | Galactoside-binding lectin | Role in fruiting body development |
| RT3_15066_m.44317 | Nucleoside diphosphate kinase | Regulation of spore and sclerotia development |
| RT3_1_m.4 | WD40 domain-containing protein | Regulates fungal cell differentiation |
| RT3_3897_m.25109 | Wlm domain-containing protein (Fragment) | Related to the DNA damage response proteins WSS1 involved in sister chromatid separation and segregation |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mehta, N.; Patil, R.; Baghela, A. Differential Physiological Prerequisites and Gene Expression Profiles of Conidial Anastomosis Tube and Germ Tube Formation in Colletotrichum gloeosporioides. J. Fungi 2021, 7, 509. https://doi.org/10.3390/jof7070509
Mehta N, Patil R, Baghela A. Differential Physiological Prerequisites and Gene Expression Profiles of Conidial Anastomosis Tube and Germ Tube Formation in Colletotrichum gloeosporioides. Journal of Fungi. 2021; 7(7):509. https://doi.org/10.3390/jof7070509
Chicago/Turabian StyleMehta, Nikita, Ravindra Patil, and Abhishek Baghela. 2021. "Differential Physiological Prerequisites and Gene Expression Profiles of Conidial Anastomosis Tube and Germ Tube Formation in Colletotrichum gloeosporioides" Journal of Fungi 7, no. 7: 509. https://doi.org/10.3390/jof7070509
APA StyleMehta, N., Patil, R., & Baghela, A. (2021). Differential Physiological Prerequisites and Gene Expression Profiles of Conidial Anastomosis Tube and Germ Tube Formation in Colletotrichum gloeosporioides. Journal of Fungi, 7(7), 509. https://doi.org/10.3390/jof7070509






