Identification of the Ricin-B-Lectin LdRBLk in the Colorado Potato Beetle and an Analysis of Its Expression in Response to Fungal Infections
Abstract
1. Introduction
2. Material and Methods
2.1. Identification and Bioinformatic Analysis of the LdRBlk Peptide
2.2. Phylogenetic Analysis
2.3. Fungi and Insects
2.4. Effect of B. bassiana and M. robertsii Infection
2.5. Effect of Different Susceptibility to Fungi
2.6. Effect of Cuticular Injuries
2.7. Sample Preparation and qPCR
2.8. Germination Assay
2.9. Statistics
3. Results
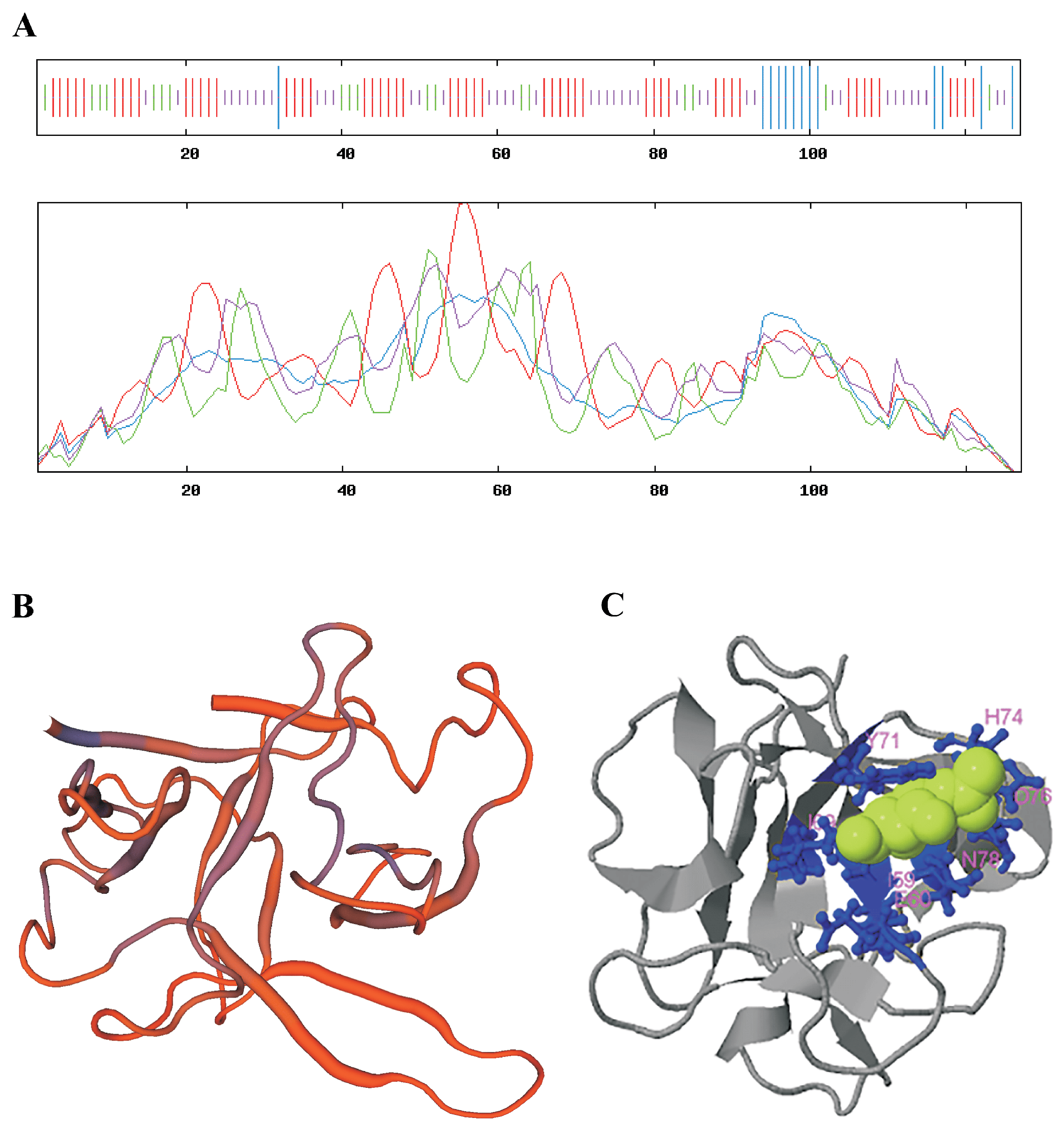
3.1. Identification and Bioinformatic Analysis of the LdRBlk Peptide
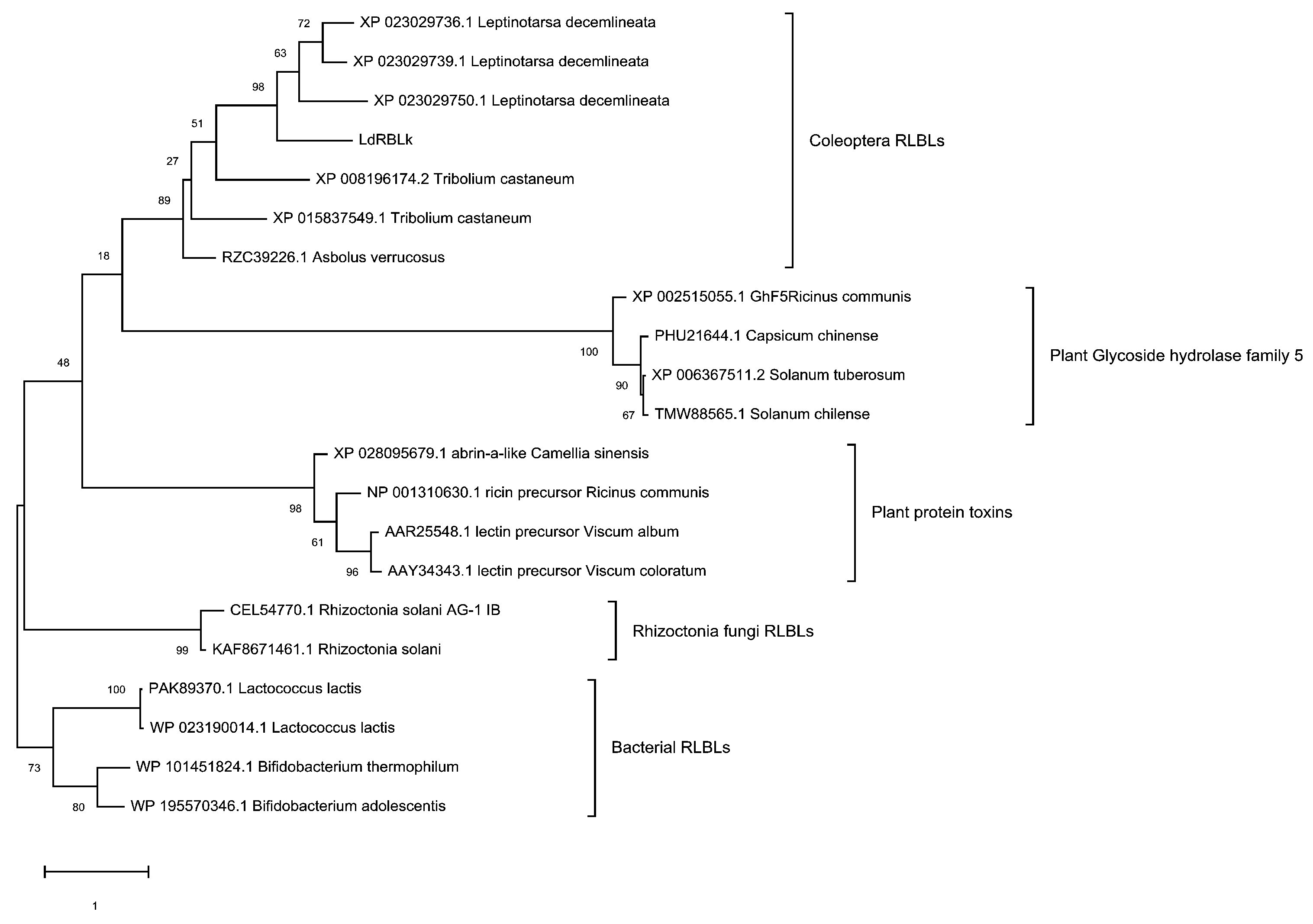
3.2. Phylogenetic Analysis
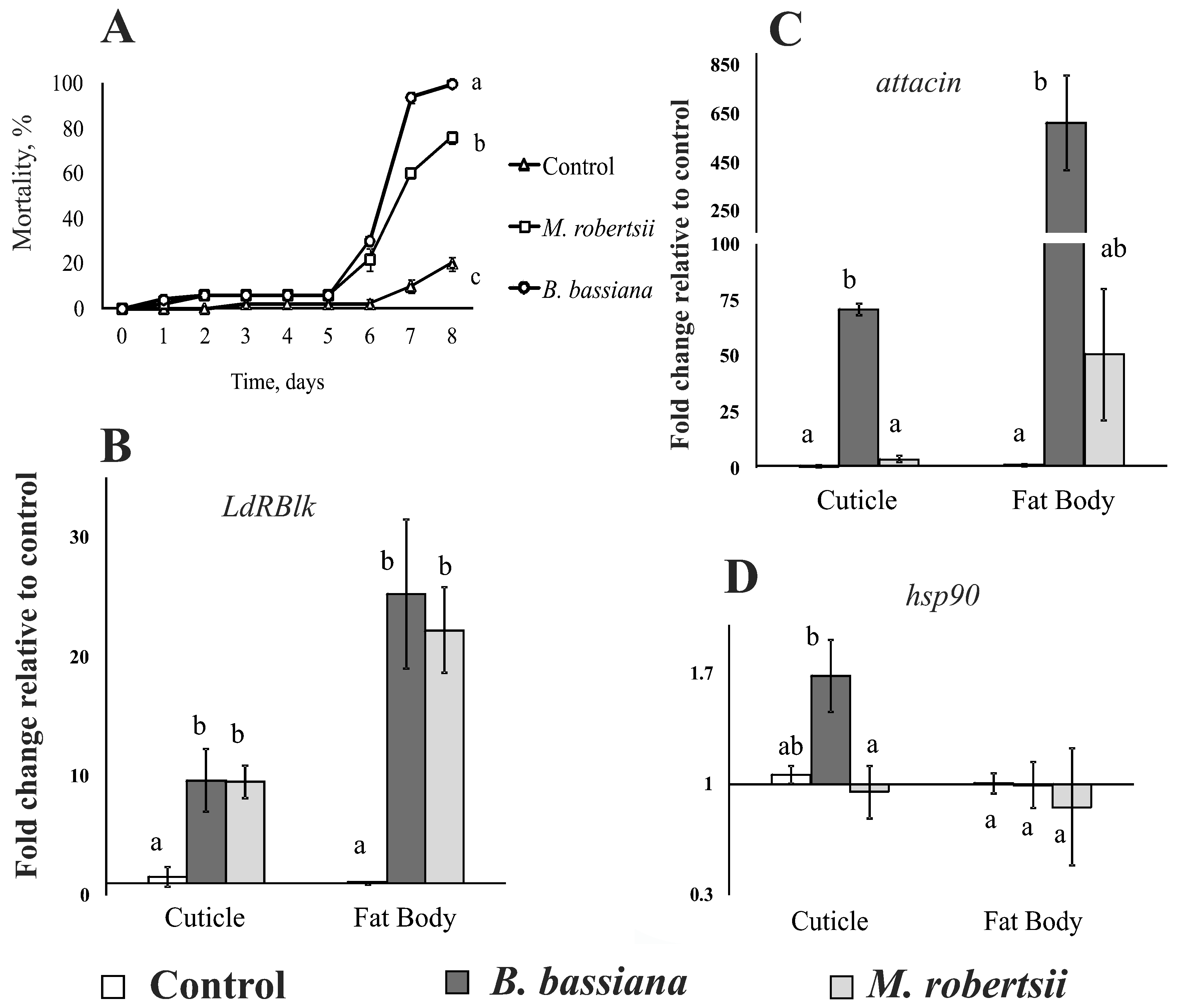
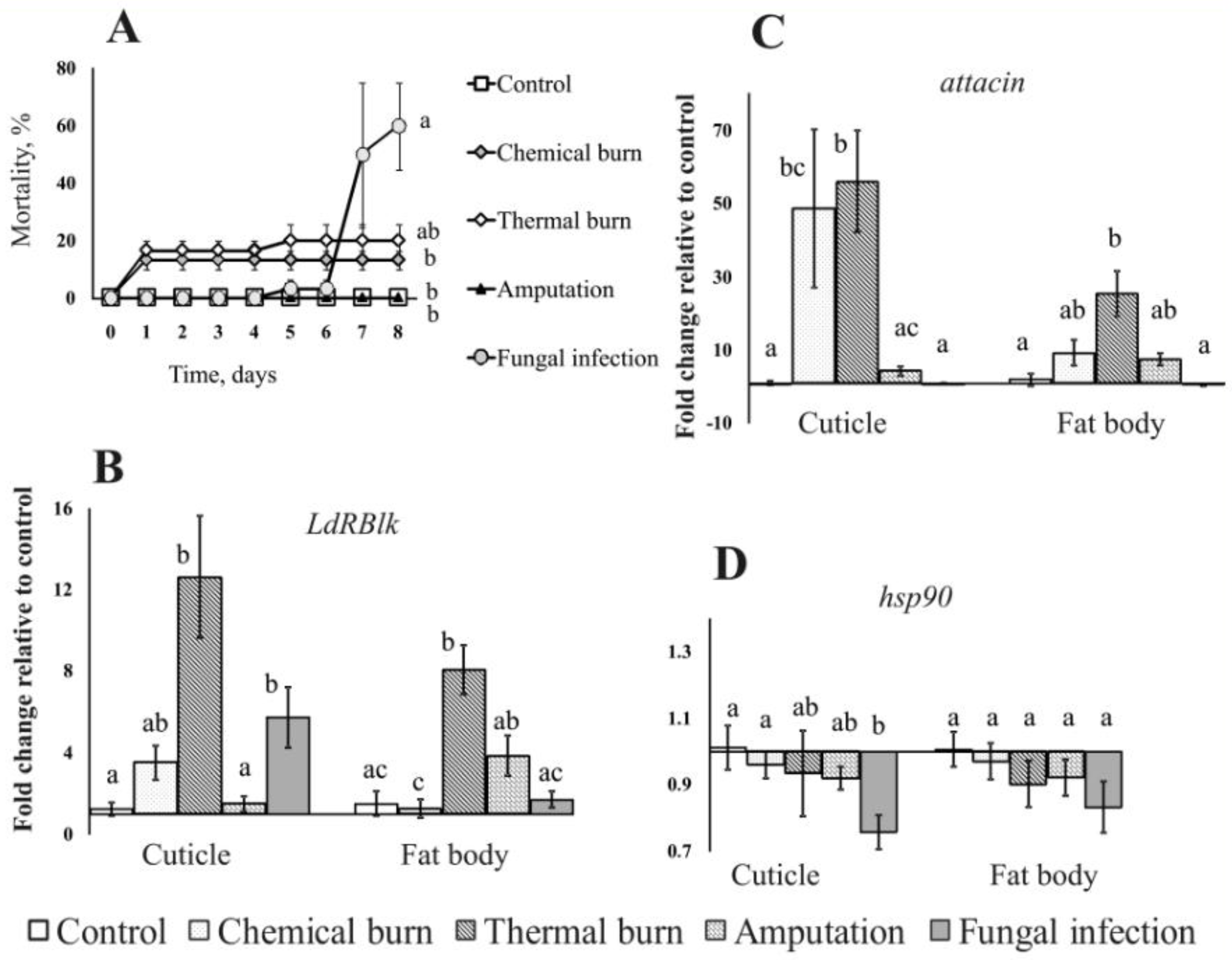
3.3. Effect of B. bassiana and M. robertsii on the Expression of the LdRBlk, Attacin and hsp90 Genes
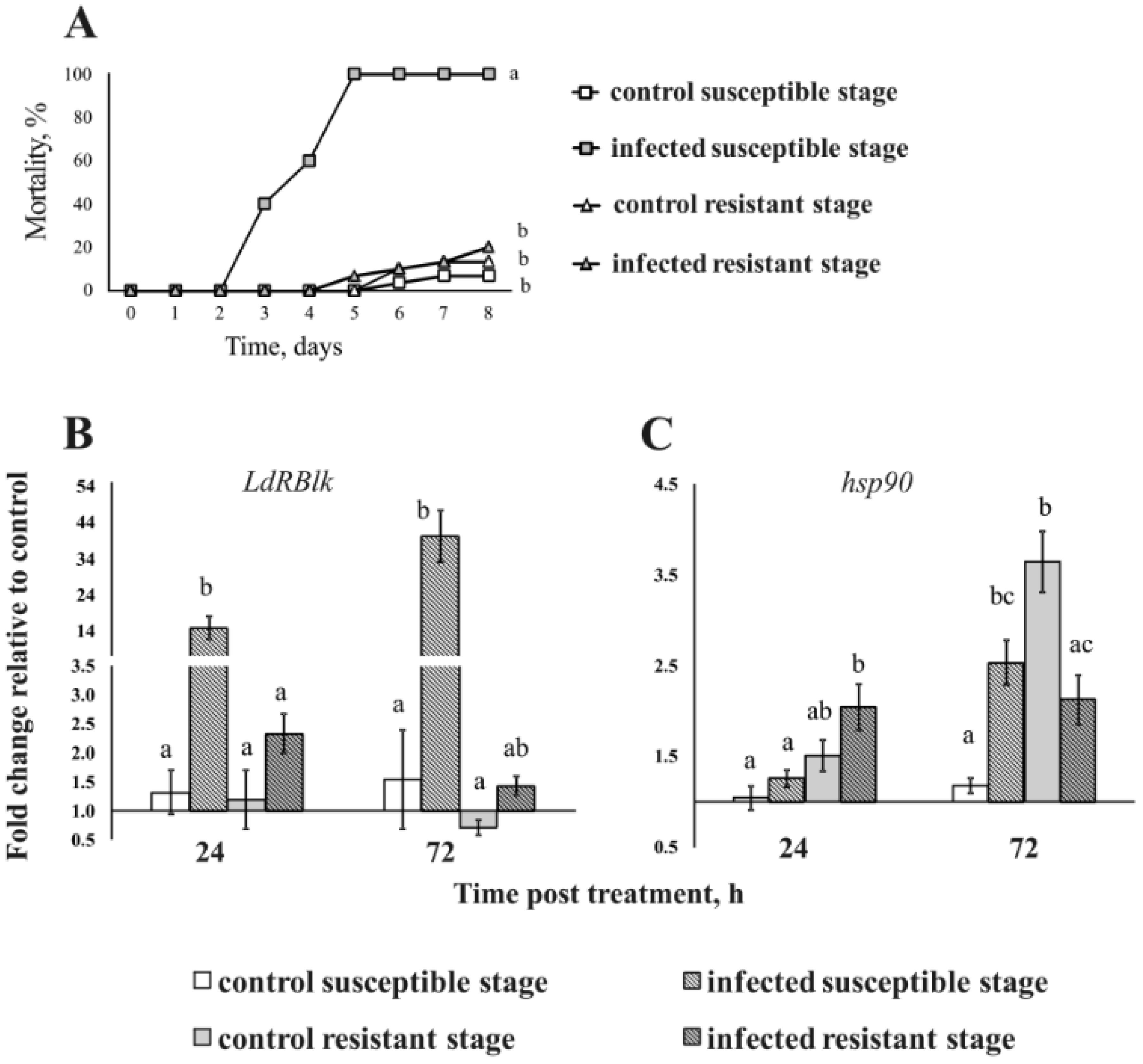
3.4. Expression of the LdRBlk and hsp90 Genes in Larvae with Different Susceptibility to Fungi
3.5. Gene Expression after Various Injuries to Integuments
3.6. Effect of β-Lectin of Ricin (RBL) from R. communis on the Germination of M. robertsii and B. bassiana Conidia
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Drickamer, K. C-type lectin-like domains. Curr. Opin. Struct. Biol. 1999, 5, 585–590. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Lin, G.; Langdon, W.Y.; Tao, L.; Zhang, J. Regulation of C-type lectin receptor-mediated antifungal immunity. Front. Immunol. 2018, 9, 123. [Google Scholar] [CrossRef] [PubMed]
- Speakman, E.A.; Dambuza, I.M.; Salazar, F.; Brown, G.D. T cell antifungal immunity and the role of C-type lectin receptors. Trends. Immunol. 2020, 41, 61–76. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.; Wu, R.; Yang, T.; Shen, H.; Hu, Z. Effect of pathogenic bacteria on a novel C-type lectin, hemocyte and superoxide dismutase/alkaline phosphatase activity in Onchidium reevesii. Fish Shellfish. Immunol. 2020, 102, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Hatakeyama, T.; Unno, H.; Kouzuma, Y.; Uchida, T.; Eto, S.; Hidemura, H.; Kato, N.; Yonekura, M.; Kusunoki, M. C-type lectin-like carbohydrate recognition of the hemolytic lectin CEL-III containing ricin-type β-trefoil folds. J. Biol. Chem. 2007, 282, 37826–37835. [Google Scholar] [CrossRef] [PubMed]
- Rao, X.J.; Shahzad, T.; Liu, S.; Wu, P.; He, Y.T.; Sun, W.J.; Fan, X.Y.; Yang, Y.F.; Shi, Q.; Yu, X.Q. Identification of C-type lectin-domain proteins (CTLDPs) in silkworm Bombyx mori. Dev. Comp. Immunol. 2015, 53, 328–338. [Google Scholar] [CrossRef]
- Bi, J.; Ning, M.; Li, J.; Zhang, P.; Wang, L.; Xu, S.; Zhong, Y.; Wang, Z.; Song, Q.; Li, B. A C-type lectin with dual-CRD from Tribolium castaneum is induced in response to bacterial challenge. Pest. Manag. Sci. 2020, 76, 3965–3974. [Google Scholar] [CrossRef] [PubMed]
- Lua, Y.; Sua, F.; Zhua, K.; Zhua, M.; Lia, Q.; Hua, Q.; Zhanga, J.; Zhanga, R.; Yu, X.-Q. Comparative genomic analysis of C-type lectin-domain genes in seven holometabolous insect species. Insect Biochem. Mol. Biol. 2020, 126, 103451. [Google Scholar] [CrossRef]
- Jiang, H.; Vilcinskas, A.; Kanost, M.R. Immunity in Lepidopteran Insects. In Invertebrate Immunity, Advances in Experimental Medicine and Biology; Söderhäll, K., Ed.; Springer: Boston, MA, USA, 2010; pp. 181–204. [Google Scholar] [CrossRef]
- Koizumi, N.; Imai, Y.; Morozumi, A.; Imamura, M.; Kadotani, T.; Yaoi, K.; Iwahana, H.; Sato, R. Lipopolysaccharide-binding protein of Bombyx mori participates in a hemocyte-mediated defense reaction against Gram-negative bacteria. J. Insect Physiol. 1999, 45, 853–859. [Google Scholar] [CrossRef]
- Shin, S.W.; Park, D.S.; Kim, S.C.; Park, H.Y. Two carbohydrate recognition domains of Hyphantria cunea lectin bind to bacterial lipopolysaccharides through O-specific chain. FEBS Lett. 2000, 467, 70–74. [Google Scholar] [CrossRef]
- Yu, X.Q.; Ling, E.; Tracy, M.E.; Zhu, Y. Immulectin-4 from the tobacco hornworm Manduca sexta binds to lipopolysaccharide and lipoteichoic acid. Insect Mol. Biol. 2006, 15, 119–128. [Google Scholar] [CrossRef]
- Yu, X.Q.; Tracy, M.E.; Ling, E.; Scholz, F.R.; Trenczek, T. A novel C-type immulectin-3 from Manduca sexta is translocated from hemolymph into the cytoplasm of hemocytes. Insect Biochem. Mol. Biol. 2005, 35, 285–295. [Google Scholar] [CrossRef]
- Watanabe, A.; Miyazawa, S.; Kitami, M.; Tabunoki, H.; Ueda, K.; Sato, R. Characterization of a novel C-type lectin, Bombyx mori multibinding protein, from the B. mori hemolymph. J. Immunol. 2006, 177, 4594–4604. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.Q.; Gan, H.; Kanost, M.R. Immulectin, an inducible C-type lectin from an insect, Manduca sexta, stimulates activation of plasma prophenol oxidase. Insect Biochem. Mol. Biol. 1999, 29, 585–597. [Google Scholar] [CrossRef]
- Ling, E.; Yu, X.Q. Cellular encapsulation and melanization are enhanced by immulectins, pattern recognition receptors from the tobacco hornworm Manduca sexta. Dev. Comp. Immunol. 2006, 30, 289–299. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, M.; Ohta, M.; Suzuki, A.; Takase, H.; Yoshizawa, Y.; Kitami, M.; Sato, R. Localization of the serine protease homolog BmSPH-1 in nodules of E. coli-injected Bombyx mori larvae and functional analysis of its role in nodule melanization. Dev. Comp. Immunol. 2011, 35, 611–619. [Google Scholar] [CrossRef] [PubMed]
- Komarov, D.A.; Slepneva, I.A.; Dubovskii, I.M.; Grizanova, E.V.; Khramtsov, V.V.; Glupov, V.V. Generation of superoxide radical and hydrogen peroxide in insect hemolymph in the course of immune response. Dokl. Biol. Sci. 2006, 411, 482–485. [Google Scholar] [CrossRef]
- Olsnes, S.; Pihl, A. Different biological properties of the two constituent peptide chains of ricin a toxic protein inhibiting protein synthesis. Biochemistry 1973, 12, 3121–3126. [Google Scholar] [CrossRef]
- Broom, A.; Doxey, A.C.; Lobsanov, Y.D.; Berthin, L.G.; Rose, D.R.; Howell, P.L.; McConkey, B.J.; Meiering, E.M. Modular evolution and the origins of symmetry: Reconstruction of a three-fold symmetric globular protein. Structure 2012, 20, 161–171. [Google Scholar] [CrossRef]
- Kim, S. Ricin B-like lectin orthologues from two mushrooms, Hericium erinaceus and Stereum hirsutum, enable recognition of highly fucosylated N-glycans. Int. J. Biol. Macromol. 2020, 147, 560–568. [Google Scholar] [CrossRef]
- Pohleven, J.; Obermajer, N.; Sabotic, J.; Anzlovar, S.; Sepcić, K.; Kos, J.; Kralj, B.; Strukelj, B.; Brzin, J. Purification, characterization and cloning of a ricin B-like lectin from mushroom Clitocybe nebularis with antiproliferative activity against human leukemic T cells. Biochim. Biophys. Acta 2009, 1790, 173–181. [Google Scholar] [CrossRef] [PubMed]
- Dumas, P.; Morin, M.D.; Boquel, S.; Moffat, C.E.; Morin, P.J. Expression status of heat shock proteins in response to cold, heat, or insecticide exposure in the Colorado potato beetle Leptinotarsa decemlineata. Cell Stress Chaperon 2019, 24, 539–547. [Google Scholar] [CrossRef]
- Wu, Q.; Patočka, J.; Kuča, K. Insect antimicrobial peptides, a mini review. Toxins 2018, 10, 461. [Google Scholar] [CrossRef]
- Clements, J.; Schoville, S.; Peterson, N.; Lan, Q.; Groves, R.L. Characterizing molecular mechanisms of imidacloprid resistance in select populations of Leptinotarsa decemlineata in the Central Sands region of Wisconsin. PLoS ONE 2016, 28. [Google Scholar] [CrossRef]
- ExPASy Bioinformatics Resource Portal. Translate Tool. Available online: https://web.expasy.org/translate/ (accessed on 31 March 2021).
- Geourjon, C.; Deléage, G. SOPMA: Significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Comput. Appl. Biosci. 1995, 11, 681–684. [Google Scholar] [CrossRef] [PubMed]
- Prabi-Gerland Rhone-Alpes Bioinformatic Pole Garland Site. Sopma Secondary Structure Prediction Method. Available online: https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_sopma.html (accessed on 31 March 2021).
- Biozentrum. SWISS-MODEL. Available online: https://www.swissmodel.expasy.org/ (accessed on 31 March 2021).
- Almagro Armenteros, J.J.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsenet, H. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 2019, 37, 420–423. [Google Scholar] [CrossRef]
- Center of Biological Sequence Analysis. SignalP-5.0 Server. Available online: http://www.cbs.dtu.dk/services/SignalP (accessed on 31 March 2021).
- Ikeda, M.; Arai, M.; Okuno, T.; Shimizu, T. TMPDB: A database of experimentally-characterized transmembrane topologies. Nucleic Acids Res. 2003, 31, 406–409. [Google Scholar] [CrossRef] [PubMed]
- ExPASy Bioinformatics Resource Portal. Prediction of Transmembrane Regions and Orientation. Available online: http://www.ch.embnet.org/software/TMPRED_form.html (accessed on 31 March 2021).
- Roy, A.; Kucukural, A.; Zhang, Y. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc. 2010, 5, 725–738. [Google Scholar] [CrossRef]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 2015, 43, 174–181. [Google Scholar] [CrossRef]
- Zhang Lab. I-TASSER Protein Structure and Function Predictions. Available online: https://zhanglab.ccmb.med.umich.edu/I-TASSER/ (accessed on 31 March 2021).
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook, 1st ed.; Editor Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- ExPASy Bioinformatics Resource Portal. ProtScale Tool. Available online: https://web.expasy.org/protscale/ (accessed on 31 March 2021).
- Nachman, M.W.; Crowell, S.L. Estimate of the mutation rate per nucleotide in humans. Genetics 2000, 156, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Kryukov, V.Y.; Kabilov, M.R.; Smirnova, N.; Tomilova, O.G.; Tyurin, M.V.; Akhanaev, Y.B.; Polenogova, O.V.; Danilov, V.P.; Zhangissina, S.K.; Alikina, T.; et al. Bacterial decomposition of insects post-Metarhizium infection: Possible influence on plant growth. Fungal Biol. 2019, 123, 927–935. [Google Scholar] [CrossRef]
- National Center for Biotechnology Information. BLAST. Standard Protein BLASTp. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 31 March 2021).
- National Center for Biotechnology Information. SPARCLE. Architecture Viewer. Available online: https://www.ncbi.nlm.nih.gov/Structure/sparcle/archview.html?archid=10624646#top (accessed on 31 March 2021).
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.; Clamp, M.; Barton, G.J. Jalview Version 2--a multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [PubMed]
- EMBL-EBI. MAFFT. Available online: https://www.ebi.ac.uk/Tools/msa/mafft/ (accessed on 31 March 2021).
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 1, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Whelan, S.; Goldman, N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 2001, 18, 691–699. [Google Scholar] [CrossRef] [PubMed]
- Tomilova, O.G.; Yaroslavtseva, O.N.; Ganina, M.D.; Tyurin, M.V.; Chernyak, E.I.; Senderskiy, I.V.; Noskov, Y.A.; Polenogova, O.V.; Akhanaev, Y.B.; Kryukov, V.Y.; et al. Changes in antifungal defence systems during the intermoult period in the Colorado potato beetle. J. Insect Physiol. 2019, 116, 106–117. [Google Scholar] [CrossRef] [PubMed]
- Integrated DNA Technology, IDT. Oligo Analyser Tool. Available online: http://eu.idtdna.com/calc/analyzer (accessed on 31 March 2021).
- Peumans, W.J.; Van Damme, E.J.M. Lectins as plant defense proteins. Plant Physiol. 1995, 109, 347–352. [Google Scholar] [CrossRef]
- Tsaneva, M.; Van Damme, E.J.M. 130 years of plant lectin research. Glycoconj. J. 2020, 37, 533–551. [Google Scholar] [CrossRef] [PubMed]
- Notova, S.; Bonnardel, F.; Lisacek, F.; Varrot, A.; Imberty, A. Structure and engineering of tandem repeat lectins. Curr. Opin. Struct. Biol. 2019, 62, 39–47. [Google Scholar] [CrossRef]
- Hazes, B. The (QxW)3 domain: A flexible lectin scaffold. Protein Sci. 1996, 5, 1490–1501. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Lu, Y.; Chen, H.; Wu, C.; Zhang, H.; Chen, L.; Chen, X. Antifungal peptides produced by actinomycetes and their biological activities against plant diseases. J. Antibiot. (Tokyo) 2020, 73, 265–282. [Google Scholar] [CrossRef] [PubMed]
- De Lucca, A.J.; Walsh, T.J. Antifungal peptides: Novel therapeutic compounds against emerging pathogens. Antimicrob. Agents Chemother. 1999, 43, 1–11. [Google Scholar] [CrossRef] [PubMed]
- La Rocca, P.; Biggin, P.C.; Tieleman, D.P.; Sansom, M.S. Simulation studies of the interaction of antimicrobial peptides and lipid bilayers. Biochim. Biophys. Acta 1999, 1462, 185–200. [Google Scholar] [CrossRef] [PubMed]
- Diaz, M.; Casali, P. Somatic immunoglobulin hypermutation. Curr. Opin. Immunol. 2002, 14, 235–240. [Google Scholar] [CrossRef]
- Drake, J.W.; Charlesworth, B.; Charlesworth, D.; Crow, J.F. Rates of spontaneous mutation. Genetics 1998, 148, 1667–1686, PMC1460098. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, T.; Tonozuka, T.; Ide, A.; Yuzawa, T.; Oguma, K.; Nishikawa, A. Sugar-binding sites of the HA1 subcomponent of Clostridium botulinum type C progenitor toxin. J. Mol. Biol. 2008, 376, 854–867. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, T.; Tonozuka, T.; Ito, S.; Takeda, Y.; Sato, R.; Matsuo, I.; Ito, Y.; Oguma, K.; Nishikawa, A. Molecular diversity of the two sugar-binding sites of the β-trefoil lectin HA33/C (HA1) from Clostridium botulinum type C neurotoxin. Arch. Biochem. Biophys. 2011, 512, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Nakano, M.; Tabata, S.; Sugihara, K.; Kouzuma, Y.; Kimura, M.; Yamasaki, N. Primary structure of hemolytic lectin CEL-III from marine invertebrate Cucumaria echinata and its cDNA: Structural similarity to the B-chain from plant lectin, ricin. Biochim. Biophys. Acta 1999, 1435, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Uchida, T.; Yamasaki, T.; Eto, S.; Sugawara, H.; Kurisu, G.; Nakagawa, A.; Kusunoki, M.; Hatakeyama, T. Crystal structure of the hemolytic lectin CEL-III isolated from the marine invertebrate Cucumaria echinata: Implications of domain structure for its membrane pore-formation mechanism. J. Biol. Chem. 2004, 279, 37133–37141. [Google Scholar] [CrossRef] [PubMed]
- Hatakeyama, T.; Furukawa, M.; Nagatomo, H.; Yamasaki, N.; Mori, T. Oligomerization of the hemolytic lectin CEL-III from the marine invertebrate Cucumaria echinate induced by the binding of carbohydrate ligands. J. Biol. Chem. 1996, 28, 16915–16920. [Google Scholar] [CrossRef]
- Van Holle, S.; De Schutter, K.; Eggermont, L.; Tsaneva, M.; Dang, L.; Van Damme, E.J.M. Comparative study of lectin domains in model species: New insights into evolutionary dynamics. Int. J. Mol. Sci. 2017, 18, 1136. [Google Scholar] [CrossRef] [PubMed]
- Van Holle, S.; Van Damme, E.J.M. Messages from the past: New insights in plant lectin evolution. Front. Plant Sci. 2019, 10, 36. [Google Scholar] [CrossRef] [PubMed]
- Van Damme, E.J.M.; Peumans, W.J.; Barre, A.; Rougé, P. Plant lectins: A composite of several distinct families of structurally and evolutionary related proteins with diverse biological roles. Crit. Rev. Plant Sci. 1998, 17, 575–692. [Google Scholar] [CrossRef]
- Vandenbussche, F.; Peumans, W.J.; Desmyter, S.; Proost, P.; Ciani, M.; Van Damme, E.J.M. The type-1 and type-2 ribosome-inactivating proteins from Iris confer transgenic tobacco plants local but not systemic protection against viruses. Planta 2004, 220, 211–221. [Google Scholar] [CrossRef]
- Shahidi-Noghabi, S.; Van Damme, E.J.M.; Smagghe, G. Expression of Sambucus nigra agglutinin (SNA-I’) from elderberry bark in transgenic tobacco plants results in enhanced resistance to different insect species. Transgenic Res. 2009, 18, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.Q.; Liu, R.S.; Wang, Q.; Liu, W.Y. Toxicity of two type II ribosome-inactivating proteins (cinnamomin and ricin) to domestic silkworm larvae. Arch. Insect Biochem. Physiol. 2004, 57, 160–165. [Google Scholar] [CrossRef] [PubMed]
- Babu, M.; Griffiths, J.S.; Huang, T.S.; Wang, A. Altered gene expression changes in Arabidopsis leaf tissues and protoplasts in response to Plum pox virus infection. BMC Genom. 2008, 9, 325. [Google Scholar] [CrossRef] [PubMed]
- Wanchoo, A.; Lewis, M.W.; Keyhani, N.O. Lectin mapping reveals stage-specific display of surface carbohydrates in in vitro and haemolymph-derived cells of the entomopathogenic fungus Beauveria bassiana. Microbiology 2009, 9, 3121–3133. [Google Scholar] [CrossRef] [PubMed]
- Faruck, M.O.; Yusof, F.; Chowdhury, S. An overview of antifungal peptides derived from insect. Peptides 2016, 80, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Schubert, M.; Bleuler-Martinez, S.; Butschi, A.; Wälti, M.A.; Egloff, P.; Stutz, K.; Yan, S.; Collot, M.; Mallet, J.M.; Wilson, I.B.H.; et al. Plasticity of the β-trefoil protein fold in the recognition and control of invertebrate predators and parasites by a fungal defence system. PLoS Pathog. 2012, 8, e1002706. [Google Scholar] [CrossRef]
- Žurga, S.; Pohleven, J.; Renko, M.; Bleuler-Martinez, S.; Sosnowski, P.; Turk, D.; Künzler, M.; Kos, J.; Sabotič, J. A novel β-trefoil lectin from the parasol mushroom (Macrolepiota procera) is nematotoxic. FEBS J. 2014, 281, 3489–3506. [Google Scholar] [CrossRef] [PubMed]
- Mowlds, P.; Kavanagh, K. Effect of pre-incubation temperature on susceptibility of Galleria mellonella larvae to infection by Candida albicans. Mycopathologia 2008, 165, 5–12. [Google Scholar] [CrossRef] [PubMed]
- Kryukov, V.Y.; Yaroslavtseva, O.N.; Whitten, M.M.A.; Tyurin, M.V.; Ficken, K.J.; Greig, C.; Melo, N.R.; Glupov, V.V.; Dubovskiy, I.M.; Butt, T.M. Fungal infection dynamics in response to temperature in the lepidopteran insect Galleria mellonella. Insect Sci. 2018, 25, 454–466. [Google Scholar] [CrossRef] [PubMed]
- Wojda, I.; Taszłow, P. Heat shock affects host-pathogen interaction in Galleria mellonella infected with Bacillus thuringiensis. J. Insect Physiol. 2013, 59, 894–905. [Google Scholar] [CrossRef] [PubMed]
- Martinson, E.O.; Wheeler, D.; Wright, J.; Mrinalini; Siebert, A.L.; Werren, J.H. Nasonia vitripennis venom causes targeted gene expression changes in its fly host. Mol. Ecol. 2014, 23, 5918–5930. [Google Scholar] [CrossRef] [PubMed]
- Dushay, M.S.; Roethele, J.B.; Chaverri, J.M.; Dulek, D.E.; Syed, S.K.; Kitami, T.; Eldon, E.D. Two attacin antibacterial genes of Drosophila melanogaster. Gene 2000, 246, 49–57. [Google Scholar] [CrossRef]
- Vey, A.; Fargues, J. Histological and ultrastructural studies of Beauveria bassiana infection in Leptinotarsa decemlineta larvae during ecdysis. J. Invertebr. Pathol. 1977, 30, 207–215. [Google Scholar] [CrossRef]
- Kryukov, V.Y.; Polenogova, O.V.; Kosman, E.S.; Krivopalov, A.; Kabilov, M.; Alikina, T.; Slepneva, I.A.; Vorontsova, Y.; Akhanaev, Y.; Glupov, V.V.; et al. Fungi-bacteria interactions in Colorado potato beetle larvae during mycoses development. Pest. Manag. Sci. 2021. Manuscript in preparation. [Google Scholar]
- Jaronski, S. Soil ecology of the entomopathogenic Ascomycetes: A critical examination of what we (think) we know. In Use of Entomopathogenic Fungi in Biological Pest Management; Ekesi, S., Maniania, N.K., Eds.; Research Signpost, Fort, P.O., Trivandrum-695 023: Kerala, India, 2008; pp. 91–144. ISBN 978-81-308-0192-6. [Google Scholar]
- Kryukov, V.Y.; Rotskaya, U.; Yaroslavtseva, O.; Polenogova, O.; Kryukova, N.; Akhanaev, Y.; Krivopalov, A.; Alikina, T.; Vorontsova, Y.L.; Slepneva, I.; et al. Fungus Metarhizium robertsii and neurotoxic insecticide affect gut immunity and microbiota in Colorado potato beetles. Sci. Rep. 2021, 11, 1299. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rotskaya, U.N.; Kryukov, V.Y.; Kosman, E.; Tyurin, M.; Glupov, V.V. Identification of the Ricin-B-Lectin LdRBLk in the Colorado Potato Beetle and an Analysis of Its Expression in Response to Fungal Infections. J. Fungi 2021, 7, 364. https://doi.org/10.3390/jof7050364
Rotskaya UN, Kryukov VY, Kosman E, Tyurin M, Glupov VV. Identification of the Ricin-B-Lectin LdRBLk in the Colorado Potato Beetle and an Analysis of Its Expression in Response to Fungal Infections. Journal of Fungi. 2021; 7(5):364. https://doi.org/10.3390/jof7050364
Chicago/Turabian StyleRotskaya, Ulyana N., Vadim Yu. Kryukov, Elena Kosman, Maksim Tyurin, and Viktor V. Glupov. 2021. "Identification of the Ricin-B-Lectin LdRBLk in the Colorado Potato Beetle and an Analysis of Its Expression in Response to Fungal Infections" Journal of Fungi 7, no. 5: 364. https://doi.org/10.3390/jof7050364
APA StyleRotskaya, U. N., Kryukov, V. Y., Kosman, E., Tyurin, M., & Glupov, V. V. (2021). Identification of the Ricin-B-Lectin LdRBLk in the Colorado Potato Beetle and an Analysis of Its Expression in Response to Fungal Infections. Journal of Fungi, 7(5), 364. https://doi.org/10.3390/jof7050364





