Cryptococcal Traits Mediating Adherence to Biotic and Abiotic Surfaces
Abstract
1. Introduction
2. Cryptococcal Adhesion Tools
2.1. The Capsule Polysaccharide
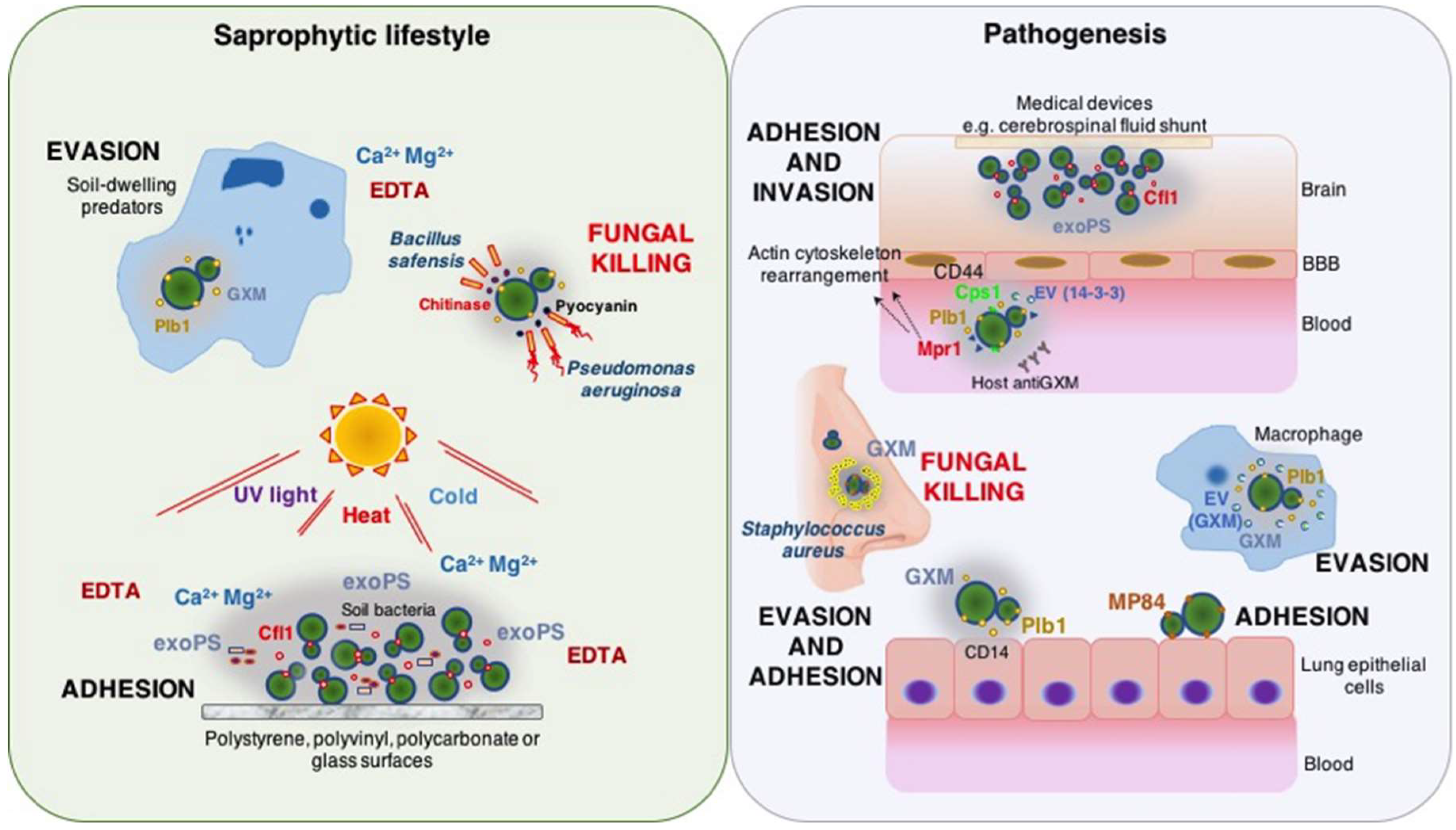
2.1.1. Interactions of Cryptococcus spp. in Soils: A Training Ground for Virulence
2.1.2. Intracellular Pathogenesis
2.2. Mannoprotein 84 (MP84)
2.3. Phospholipase B (Plb1)
2.4. Hyaluronic Acid Synthase
2.5. Metalloprotease (Mpr1)
2.6. 14-3-3 Adhesin
3. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- De Mattos-Shipley, K.M.; Ford, K.L.; Alberti, F.; Banks, A.M.; Bailey, A.M.; Foster, G.D. The good, the bad and the tasty: The many roles of mushrooms. Stud. Mycol. 2016, 85, 125–157. [Google Scholar] [CrossRef] [PubMed]
- Coelho, M.A.; Bakkeren, G.; Sun, S.; Hood, M.E.; Giraud, T. Fungal sex: The basidiomycota. Microbiol. Spectr. 2017, 5. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Bennett, J.E.; Wickes, B.L.; Meyer, W.; Cuomo, C.A.; Wollenburg, K.R.; Bicanic, T.A.; Castaneda, E.; Chang, Y.C.; Chen, J.; et al. The case for adopting the “species complex” nomenclature for the etiologic agents of cryptococcosis. mSphere 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Casadevall, A.; Perfect, J.R. Cryptococcus Neoformans; America Society Microbiology: Washington, DC, USA, 1998. [Google Scholar]
- Holmquist, L.; Russo, C.A.; Elixhauser, A. Meningitis-related hospitalizations in the United States, 2006: Statistical brief #57. In Healthcare Cost and Utilization Project (Hcup) Statistical Briefs; Agency for Healthcare Research and Quality (US): Rockville, MD, USA, 2006. [Google Scholar]
- Brizendine, K.D.; Baddley, J.W.; Pappas, P.G. Predictors of mortality and differences in clinical features among patients with cryptococcosis according to immune status. PLoS ONE 2013, 8, e60431. [Google Scholar] [CrossRef] [PubMed]
- Mirza, S.A.; Phelan, M.; Rimland, D.; Graviss, E.; Hamill, R.; Brandt, M.E.; Gardner, T.; Sattah, M.; de Leon, G.P.; Baughman, W.; et al. The changing epidemiology of cryptococcosis: An update from population-based active surveillance in 2 large metropolitan areas, 1992–2000. Clin. Infect. Dis. 2003, 36, 789–794. [Google Scholar] [CrossRef] [PubMed]
- McKenney, J.; Bauman, S.; Neary, B.; Detels, R.; French, A.; Margolick, J.; Doherty, B.; Klausner, J.D. Prevalence, correlates, and outcomes of cryptococcal antigen positivity among patients with aids, United States, 1986–2012. Clin. Infect. Dis. 2015, 60, 959–965. [Google Scholar] [CrossRef] [PubMed]
- Pyrgos, V.; Seitz, A.E.; Steiner, C.A.; Prevots, D.R.; Williamson, P.R. Epidemiology of cryptococcal meningitis in the US: 1997–2009. PLoS ONE 2013, 8, e56269. [Google Scholar] [CrossRef] [PubMed]
- O’Halloran, J.A.; Powderly, W.G.; Spec, A. Cryptococcosis today: It is not all about HIV infection. Curr. Clin. Microbiol. Rep. 2017, 4, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Spec, A.; Raval, K.; Powderly, W.G. End-stage liver disease is a strong predictor of early mortality in cryptococcosis. Open Forum Infect. Dis. 2016, 3. [Google Scholar] [CrossRef]
- Xu, J.; Vilgalys, R.; Mitchell, T.G. Multiple gene genealogies reveal recent dispersion and hybridization in the human pathogenic fungus Cryptococcus neoformans. Mol. Ecol. 2000, 9, 1471–1481. [Google Scholar] [CrossRef] [PubMed]
- Findley, K.; Rodriguez-Carres, M.; Metin, B.; Kroiss, J.; Fonseca, A.; Vilgalys, R.; Heitman, J. Phylogeny and phenotypic characterization of pathogenic cryptococcus species and closely related saprobic taxa in the tremellales. Eukaryot. Cell 2009, 8, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Casadevall, A.; Freij, J.B.; Hann-Soden, C.; Taylor, J. Continental drift and speciation of the Cryptococcus neoformans and Cryptococcus gattii species complexes. mSphere 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Emmons, C.W. Saprophytic sources of Cryptococcus neoformans associated with the pigeon (columba livia). Am. J. Hyg. 1955, 62, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Sorrell, T.C.; Ellis, D.H. Ecology of Cryptococcus neoformans. Rev. Iberoam. Micol. 1997, 14, 42–43. [Google Scholar] [PubMed]
- Harrison, T.S. Cryptococcus neoformans and cryptococcosis. J. Infect. 2000, 41, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.; De Obaldia, A.L.; Heitman, J. Cryptococcus neoformans mates on pigeon guano: Implications for the realized ecological niche and globalization. Eukaryot. Cell 2007, 6, 949–959. [Google Scholar] [CrossRef] [PubMed]
- Springer, D.J.; Billmyre, R.B.; Filler, E.E.; Voelz, K.; Pursall, R.; Mieczkowski, P.A.; Larsen, R.A.; Dietrich, F.S.; May, R.C.; Filler, S.G.; et al. Cryptococcus gattii vgiii isolates causing infections in HIV/AIDS patients in southern california: Identification of the local environmental source as arboreal. PLoS Pathog. 2014, 10, e1004285. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, T.G.; Perfect, J.R. Cryptococcosis in the era of AIDS—100 years after the discovery of Cryptococcus neoformans. Clin. Microbiol. Rev. 1995, 8, 515–548. [Google Scholar] [PubMed]
- Kidd, S.E.; Hagen, F.; Tscharke, R.L.; Huynh, M.; Bartlett, K.H.; Fyfe, M.; Macdougall, L.; Boekhout, T.; Kwon-Chung, K.J.; Meyer, W. A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on vancouver island (British Columbia, Canada). Proc. Natl. Acad. Sci. USA 2004, 101, 17258–17263. [Google Scholar] [CrossRef] [PubMed]
- Kidd, S.E.; Bach, P.J.; Hingston, A.O.; Mak, S.; Chow, Y.; MacDougall, L.; Kronstad, J.W.; Bartlett, K.H. Cryptococcus gattii dispersal mechanisms, British Columbia, Canada. Emerg. Infect. Dis. 2007, 13, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Hagen, F.; Khayhan, K.; Theelen, B.; Kolecka, A.; Polacheck, I.; Sionov, E.; Falk, R.; Parnmen, S.; Lumbsch, H.T.; Boekhout, T. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal Genet. Biol. 2015, 78, 16–48. [Google Scholar] [CrossRef] [PubMed]
- Casadevall, A. Evolution of intracellular pathogens. Annu. Rev. Microbiol. 2008, 62, 19–33. [Google Scholar] [CrossRef] [PubMed]
- Simpson, G.G. The Major Features of Evolution; Columbia University Press: New York, NY, USA, 1953; p. 434. [Google Scholar]
- Casadevall, A.; Steenbergen, J.N.; Nosanchuk, J.D. ‘Ready made’ virulence and ‘dual use’ virulence factors in pathogenic environmental fungi—The Cryptococcus neoformans paradigm. Curr. Opin. Microbiol. 2003, 6, 332–337. [Google Scholar] [CrossRef]
- Casadevall, A.; Pirofski, L.A. Accidental virulence, cryptic pathogenesis, martians, lost hosts, and the pathogenicity of environmental microbes. Eukaryot. Cell 2007, 6, 2169–2174. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Ingber, D.E. The structural and mechanical complexity of cell-growth control. Nat. Cell Biol. 1999, 1, E131–E138. [Google Scholar] [CrossRef] [PubMed]
- De Groot, P.W.; Bader, O.; de Boer, A.D.; Weig, M.; Chauhan, N. Adhesins in human fungal pathogens: Glue with plenty of stick. Eukaryot. Cell 2013, 12, 470–481. [Google Scholar] [CrossRef] [PubMed]
- Ballal, S.; Inamdar, S.R. An overview of lectin-glycan interactions: A key event in initiating fungal infection and pathogenesis. Arch. Microbiol. 2018, 200, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Lipke, P.N.; Klotz, S.A.; Dufrene, Y.F.; Jackson, D.N.; Garcia-Sherman, M.C. Amyloid-like beta-aggregates as force-sensitive switches in fungal biofilms and infections. Microbiol. Mol. Biol. Rev. 2018, 82. [Google Scholar] [CrossRef]
- Casadevall, A. Cards of virulence and the global virulome for humans. Microbe 2006, 1, 359–364. [Google Scholar] [CrossRef]
- Casadevall, A.; Coelho, C.; Cordero, R.J.B.; Dragotakes, Q.; Jung, E.; Vij, R.; Wear, M.P. The capsule of Cryptococcus neoformans. Virulence 2018. [Google Scholar] [CrossRef] [PubMed]
- Pontes, B.; Frases, S. The Cryptococcus neoformans capsule: Lessons from the use of optical tweezers and other biophysical tools. Front. Microbiol. 2015, 6, 640. [Google Scholar] [CrossRef] [PubMed]
- Doering, T.L. How sweet it is! Cell wall biogenesis and polysaccharide capsule formation in Cryptococcus neoformans. Annu. Rev. Microbiol. 2009, 63, 223–247. [Google Scholar] [CrossRef] [PubMed]
- Cherniak, R.; Sundstrom, J.B. Polysaccharide antigens of the capsule of Cryptococcus neoformans. Infect. Immun. 1994, 62, 1507–1512. [Google Scholar] [PubMed]
- Cherniak, R.; Valafar, H.; Morris, L.C.; Valafar, F. Cryptococcus neoformans chemotyping by quantitative analysis of 1H nuclear magnetic resonance spectra of glucuronoxylomannans with a computer-simulated artificial neural network. Clin. Diagn. Lab. Immunol. 1998, 5, 146–159. [Google Scholar] [PubMed]
- Kozel, T.R.; Levitz, S.M.; Dromer, F.; Gates, M.A.; Thorkildson, P.; Janbon, G. Antigenic and biological characteristics of mutant strains of Cryptococcus neoformans lacking capsular O acetylation or xylosyl side chains. Infect. Immun 2003, 71, 2868–2875. [Google Scholar] [CrossRef] [PubMed]
- Urai, M.; Kaneko, Y.; Ueno, K.; Okubo, Y.; Aizawa, T.; Fukazawa, H.; Sugita, T.; Ohno, H.; Shibuya, K.; Kinjo, Y.; et al. Evasion of innate immune responses by the highly virulent Cryptococcus gattii by altering capsule glucuronoxylomannan structure. Front. Cell. Infect. Microbiol. 2015, 5, 101. [Google Scholar] [CrossRef] [PubMed]
- Cherniak, R.; Reiss, E.; Slodki, M.E.; Plattner, R.D.; Blumer, S.O. Structure and antigenic activity of the capsular polysaccharide of Cryptococcus neoformans serotype A. Mol. Immunol. 1980, 17, 1025–1032. [Google Scholar] [CrossRef]
- Turner, S.H.; Cherniak, R.; Reiss, E. Fractionation and characterization of galactoxylomannan from Cryptococcus neoformans. Carbohydr. Res. 1984, 125, 343–349. [Google Scholar] [CrossRef]
- De Jesus, M.; Nicola, A.M.; Rodrigues, M.L.; Janbon, G.; Casadevall, A. Capsular localization of the Cryptococcus neoformans polysaccharide component galactoxylomannan. Eukaryot. Cell 2009, 8, 96–103. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.C.; Casadevall, A. Polysaccharide antigen in brain tissue of aids patients with cryptococcal meningitis. Clin. Infect. Dis. 1996, 23, 194–195. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Casadevall, A. Specific antibody can prevent fungal biofilm formation and this effect correlates with protective efficacy. Infect. Immun. 2005, 73, 6350–6362. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.J.; Schlegel, R.; Moody, M.M.; Costerton, J.W.; Salcman, M. Ventriculoatrial shunt infection due to Cryptococcus neoformans: An ultrastructural and quantitative microbiological study. Neurosurgery 1986, 18, 373–375. [Google Scholar] [CrossRef] [PubMed]
- Glass, R.T.; Conrad, R.S.; Bullard, J.W.; Goodson, L.B.; Mehta, N.; Lech, S.J.; Loewy, Z.G. Evaluation of microbial flora found in previously worn prostheses from the northeast and southwest regions of the United States. J. Prosthet. Dent. 2010, 103, 384–389. [Google Scholar] [CrossRef]
- Shah, N.B.; Shoham, S.; Nayak, S. Cryptococcus neoformans prosthetic joint infection: Case report and review of the literature. Mycopathologia 2015, 179, 275–278. [Google Scholar] [CrossRef] [PubMed]
- Johannsson, B.; Callaghan, J.J. Prosthetic hip infection due to Cryptococcus neoformans: Case report. Diagn. Microbiol. Infect. Dis. 2009, 64, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Casadevall, A. Biofilm formation by Cryptococcus neoformans. Microbiol. Spectr. 2015, 3. [Google Scholar] [CrossRef]
- Martinez, L.R.; Moussai, D.; Casadevall, A. Antibody to Cryptococcus neoformans glucuronoxylomannan inhibits the release of capsular antigen. Infect. Immun. 2004, 72, 3674–3679. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Casadevall, A. Cryptococcus neoformans biofilm formation depends on surface support and carbon source and reduces fungal cell susceptibility to heat, cold, and UV light. Appl. Environ. Microbiol. 2007, 73, 4592–4601. [Google Scholar] [CrossRef] [PubMed]
- Robertson, E.J.; Casadevall, A. Antibody-mediated immobilization of Cryptococcus neoformans promotes biofilm formation. Appl. Environ. Microbiol. 2009, 75, 2528–2533. [Google Scholar] [CrossRef] [PubMed]
- Lopes, W.; Vainstein, M.H.; De Sousa Araujo, G.R.; Frases, S.; Staats, C.C.; de Almeida, R.M.C.; Schrank, A.; Kmetzsch, L.; Vainstein, M.H. Geometrical distribution of Cryptococcus neoformans mediates flower-like biofilm development. Front. Microbiol. 2017, 8, 2534. [Google Scholar] [CrossRef] [PubMed]
- Santi, L.; Beys-da-Silva, W.O.; Berger, M.; Calzolari, D.; Guimaraes, J.A.; Moresco, J.J.; Yates, J.R., 3rd. Proteomic profile of Cryptococcus neoformans biofilm reveals changes in metabolic processes. J. Proteome Res. 2014, 13, 1545–1559. [Google Scholar] [CrossRef] [PubMed]
- Frager, S.Z.; Chrisman, C.J.; Shakked, R.; Casadevall, A. Paramecium species ingest and kill the cells of the human pathogenic fungus Cryptococcus neoformans. Med. Mycol. 2010, 48, 775–779. [Google Scholar] [CrossRef] [PubMed]
- Steenbergen, J.N.; Nosanchuk, J.D.; Malliaris, S.D.; Casadevall, A. Cryptococcus neoformans virulence is enhanced after growth in the genetically malleable host dictyostelium discoideum. Infect. Immun. 2003, 71, 4862–4872. [Google Scholar] [CrossRef] [PubMed]
- Watkins, R.A.; Andrews, A.; Wynn, C.; Barisch, C.; King, J.S.; Johnston, S.A. Cryptococcus neoformans escape from dictyostelium amoeba by both wash-mediated constitutive exocytosis and vomocytosis. Front. Cell. Infect. Microbiol. 2018, 8, 108. [Google Scholar] [CrossRef] [PubMed]
- Steenbergen, J.N.; Shuman, H.A.; Casadevall, A. Cryptococcus neoformans interactions with amoebae suggest an explanation for its virulence and intracellular pathogenic strategy in macrophages. Proc. Natl. Acad. Sci. USA 2001, 98, 15245–15250. [Google Scholar] [CrossRef] [PubMed]
- Malliaris, S.D.; Steenbergen, J.N.; Casadevall, A. Cryptococcus neoformans var. Gattii can exploit Acanthamoeba castellanii for growth. Med. Mycol. 2004, 42, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Chrisman, C.J.; Alvarez, M.; Casadevall, A. Phagocytosis of Cryptococcus neoformans by, and nonlytic exocytosis from, Acanthamoeba castellanii. Appl. Environ. Microbiol. 2010, 76, 6056–6062. [Google Scholar] [CrossRef] [PubMed]
- Derengowski Lda, S.; Paes, H.C.; Albuquerque, P.; Tavares, A.H.; Fernandes, L.; Silva-Pereira, I.; Casadevall, A. The transcriptional response of Cryptococcus neoformans to ingestion by Acanthamoeba castellanii and macrophages provides insights into the evolutionary adaptation to the mammalian host. Eukaryot. Cell 2013, 12, 761–774. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, J.; Albuquerque, P.C.; Wolf, J.M.; Nascimento, R.; Pereira, M.D.; Nosanchuk, J.D.; Rodrigues, M.L. Analysis of multiple components involved in the interaction between Cryptococcus neoformans and Acanthamoeba castellanii. Fungal Biol. 2017, 121, 602–614. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, N.S.; Dos Santos, F.M.; Garcia, A.W.A.; Ferrareze, P.A.G.; Fabres, L.F.; Schrank, A.; Kmetzsch, L.; Rott, M.B.; Vainstein, M.H.; Staats, C.C. Modulation of zinc homeostasis in Acanthamoeba castellanii as a possible antifungal strategy against Cryptococcus gattii. Front. Microbiol. 2017, 8, 1626. [Google Scholar] [CrossRef] [PubMed]
- Fu, M.S.; Casadevall, A. Divalent metal cations potentiate the predatory capacity of amoeba for Cryptococcus neoformans. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef]
- Guimaraes, A.J.; Gomes, K.X.; Cortines, J.R.; Peralta, J.M.; Peralta, R.H. Acanthamoeba spp. As a universal host for pathogenic microorganisms: One bridge from environment to host virulence. Microbiol. Res. 2016, 193, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Moliner, C.; Fournier, P.E.; Raoult, D. Genome analysis of microorganisms living in amoebae reveals a melting pot of evolution. FEMS Microbiol. Rev. 2010, 34, 281–294. [Google Scholar] [CrossRef] [PubMed]
- Bertelli, C.; Greub, G. Lateral gene exchanges shape the genomes of amoeba-resisting microorganisms. Front. Cell. Infect. Microbiol. 2012, 2, 110. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, M.; Casadevall, A. Phagosome extrusion and host-cell survival after Cryptococcus neoformans phagocytosis by macrophages. Curr. Biol. 2006, 16, 2161–2165. [Google Scholar] [CrossRef] [PubMed]
- Zaragoza, O.; Chrisman, C.J.; Castelli, M.V.; Frases, S.; Cuenca-Estrella, M.; Rodriguez-Tudela, J.L.; Casadevall, A. Capsule enlargement in Cryptococcus neoformans confers resistance to oxidative stress suggesting a mechanism for intracellular survival. Cell. Microbiol. 2008, 10, 2043–2057. [Google Scholar] [CrossRef] [PubMed]
- Chrisman, C.J.; Albuquerque, P.; Guimaraes, A.J.; Nieves, E.; Casadevall, A. Phospholipids trigger Cryptococcus neoformans capsular enlargement during interactions with amoebae and macrophages. PLoS Pathog. 2011, 7, e1002047. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Croudace, J.E.; Lammas, D.A.; May, R.C. Expulsion of live pathogenic yeast by macrophages. Curr. Biol. 2006, 16, 2156–2160. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.; Cox, G.M.; Litvintseva, A.P.; Mylonakis, E.; Malliaris, S.D.; Benjamin, D.K., Jr.; Giles, S.S.; Mitchell, T.G.; Casadevall, A.; Perfect, J.R.; et al. Cryptococcus neoformans α strains preferentially disseminate to the central nervous system during coinfection. Infect. Immun. 2005, 73, 4922–4933. [Google Scholar] [CrossRef] [PubMed]
- Fu, M.S.; Coelho, C.; DeLeon-Rodriguez, C.M.; Rossi, D.C.P.; Camacho, E.; Jung, E.H.; Kulkarni, M.; Casadevall, A. Cryptococcus neoformans urease affects the outcome of intracellular pathogenesis by modulating phagolysosomal pH. PLoS Pathog. 2018, in press. [Google Scholar] [CrossRef] [PubMed]
- Joubert, L.M.; Wolfaardt, G.M.; Botha, A. Microbial exopolymers link predator and prey in a model yeast biofilm system. Microb. Ecol. 2006, 52, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Abdulkareem, A.F.; Lee, H.H.; Ahmadi, M.; Martinez, L.R. Fungal serotype-specific differences in bacterial-yeast interactions. Virulence 2015, 6, 652–657. [Google Scholar] [CrossRef] [PubMed]
- Nowack, B.; VanBriesen, J.M. Chelating agents in the environment. In Biogeochemistry of Chelating Agents; American Chemical Society: Washington, DC, USA, 2005; pp. 1–18. [Google Scholar]
- Nimrichter, L.; Frases, S.; Cinelli, L.P.; Viana, N.B.; Nakouzi, A.; Travassos, L.R.; Casadevall, A.; Rodrigues, M.L. Self-aggregation of Cryptococcus neoformans capsular glucuronoxylomannan is dependent on divalent cations. Eukaryot. Cell 2007, 6, 1400–1410. [Google Scholar] [CrossRef] [PubMed]
- Henneken, L.; Nörtemann, B.; Hempel, D.C. Influence of physiological conditions on edta degradation. Appl. Microbiol. Biotechnol. 1995, 44, 190–197. [Google Scholar] [CrossRef]
- Rella, A.; Yang, M.W.; Gruber, J.; Montagna, M.T.; Luberto, C.; Zhang, Y.M.; Del Poeta, M. Pseudomonas aeruginosa inhibits the growth of cryptococcus species. Mycopathologia 2012, 173, 451–461. [Google Scholar] [CrossRef] [PubMed]
- Gish, S.R.; Maier, E.J.; Haynes, B.C.; Santiago-Tirado, F.H.; Srikanta, D.L.; Ma, C.Z.; Li, L.X.; Williams, M.; Crouch, E.C.; Khader, S.A.; et al. Computational analysis reveals a key regulator of cryptococcal virulence and determinant of host response. MBio 2016, 7, e00313–00316. [Google Scholar] [CrossRef] [PubMed]
- Zaragoza, O.; Rodrigues, M.L.; De Jesus, M.; Frases, S.; Dadachova, E.; Casadevall, A. The capsule of the fungal pathogen Cryptococcus neoformans. Adv. Appl. Microbiol. 2009, 68, 133–216. [Google Scholar] [PubMed]
- Ding, H.; Mayer, F.L.; Sanchez-Leon, E.; de, S.A.G.R.; Frases, S.; Kronstad, J.W. Networks of fibers and factors: Regulation of capsule formation in Cryptococcus neoformans. F1000Res 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Hommel, B.; Mukaremera, L.; Cordero, R.J.B.; Coelho, C.; Desjardins, C.A.; Sturny-Leclere, A.; Janbon, G.; Perfect, J.R.; Fraser, J.A.; Casadevall, A.; et al. Titan cells formation in Cryptococcus neoformans is finely tuned by environmental conditions and modulated by positive and negative genetic regulators. PLoS Pathog. 2018, 14, e1006982. [Google Scholar] [CrossRef] [PubMed]
- Zaragoza, O.; Casadevall, A. Experimental modulation of capsule size in Cryptococcus neoformans. Biol. Proced. Online 2004, 6, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F.L.; Kronstad, J.W. Disarming fungal pathogens: Bacillus safensis inhibits virulence factor production and biofilm formation by Cryptococcus neoformans and Candida albicans. MBio 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- de Beer, D.; Stoodley, P.; Lewandowski, Z. Liquid flow in heterogeneous biofilms. Biotechnol. Bioeng. 1994, 44, 636–641. [Google Scholar] [CrossRef] [PubMed]
- Berleman, J.E.; Zemla, M.; Remis, J.P.; Liu, H.; Davis, A.E.; Worth, A.N.; West, Z.; Zhang, A.; Park, H.; Bosneaga, E.; et al. Exopolysaccharide microchannels direct bacterial motility and organize multicellular behavior. ISME J. 2016, 10, 2620–2632. [Google Scholar] [CrossRef] [PubMed]
- Stone, W.; Kroukamp, O.; Korber, D.R.; McKelvie, J.; Wolfaardt, G.M. Microbes at surface-air interfaces: The metabolic harnessing of relative humidity, surface hygroscopicity, and oligotrophy for resilience. Front. Microbiol. 2016, 7, 1563. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Mihu, M.R.; Han, G.; Frases, S.; Cordero, R.J.; Casadevall, A.; Friedman, A.J.; Friedman, J.M.; Nosanchuk, J.D. The use of chitosan to damage Cryptococcus neoformans biofilms. Biomaterials 2010, 31, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Miyake, Y.; Tsunoda, T.; Minagi, S.; Akagawa, Y.; Tsuru, H.; Suginaka, H. Antifungal drugs affect adherence of Candida albicans to acrylic surfaces by changing the zeta-potential of fungal cells. FEMS Microbiol. Lett. 1990, 57, 211–214. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhai, B.; Lin, X. The link between morphotype transition and virulence in Cryptococcus neoformans. PLoS Pathog. 2012, 8, e1002765. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Tian, X.; Gyawali, R.; Lin, X. Fungal adhesion protein guides community behaviors and autoinduction in a paracrine manner. Proc. Natl. Acad. Sci. USA 2013, 110, 11571–11576. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, P.; Nicola, A.M.; Nieves, E.; Paes, H.C.; Williamson, P.R.; Silva-Pereira, I.; Casadevall, A. Quorum sensing-mediated, cell density-dependent regulation of growth and virulence in Cryptococcus neoformans. MBio 2013, 5, e00986. [Google Scholar] [CrossRef] [PubMed]
- Taylor-Smith, L.M. Cryptococcus-epithelial interactions. J. Fungi (Basel) 2017, 3, 53. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Rodas, R.; Zaragoza, O. Catch me if you can: Phagocytosis and killing avoidance by Cryptococcus neoformans. FEMS Immunol. Med. Microbiol. 2012, 64, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Kozel, T.R.; Gotschlich, E.C. The capsule of Cryptococcus neoformans passively inhibits phagocytosis of the yeast by macrophages. J. Immunol. 1982, 129, 1675–1680. [Google Scholar] [PubMed]
- Shoham, S.; Huang, C.; Chen, J.M.; Golenbock, D.T.; Levitz, S.M. Toll-like receptor 4 mediates intracellular signaling without TNF-alpha release in response to Cryptococcus neoformans polysaccharide capsule. J. Immunol. 2001, 166, 4620–4626. [Google Scholar] [CrossRef] [PubMed]
- Monari, C.; Bistoni, F.; Casadevall, A.; Pericolini, E.; Pietrella, D.; Kozel, T.R.; Vecchiarelli, A. Glucuronoxylomannan, a microbial compound, regulates expression of costimulatory molecules and production of cytokines in macrophages. J. Infect. Dis. 2005, 191, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Feldmesser, M.; Kress, Y.; Novikoff, P.; Casadevall, A. Cryptococcus neoformans is a facultative intracellular pathogen in murine pulmonary infection. Infect. Immun. 2000, 68, 4225–4237. [Google Scholar] [CrossRef] [PubMed]
- Voelz, K.; May, R.C. Cryptococcal interactions with the host immune system. Eukaryot. Cell 2010, 9, 835–846. [Google Scholar] [CrossRef] [PubMed]
- Tucker, S.C.; Casadevall, A. Replication of Cryptococcus neoformans in macrophages is accompanied by phagosomal permeabilization and accumulation of vesicles containing polysaccharide in the cytoplasm. Proc. Natl. Acad. Sci. USA 2002, 99, 3165–3170. [Google Scholar] [CrossRef] [PubMed]
- Abeyratne-Perera, H.K.; Chandran, P.L. Mannose surfaces exhibit self-latching, water structuring, and resilience to chaotropes: Implications for pathogen virulence. Langmuir 2017, 33, 9178–9189. [Google Scholar] [CrossRef] [PubMed]
- Vartivarian, S.E.; Anaissie, E.J.; Cowart, R.E.; Sprigg, H.A.; Tingler, M.J.; Jacobson, E.S. Regulation of cryptococcal capsular polysaccharide by iron. J. Infect. Dis. 1993, 167, 186–190. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Bakker, T.; Harris, J.; Tsang, C.; Brown, G.D.; Wormald, M.R.; Gordon, S.; Dwek, R.A.; Rudd, P.M.; Martinez-Pomares, L. Glycosylation influences the lectin activities of the macrophage mannose receptor. J. Biol. Chem. 2005, 280, 32811–32820. [Google Scholar] [CrossRef] [PubMed]
- Cordero, R.J.; Pontes, B.; Guimaraes, A.J.; Martinez, L.R.; Rivera, J.; Fries, B.C.; Nimrichter, L.; Rodrigues, M.L.; Viana, N.B.; Casadevall, A. Chronological aging is associated with biophysical and chemical changes in the capsule of Cryptococcus neoformans. Infect. Immun. 2011, 79, 4990–5000. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, F.L.; Nimrichter, L.; Cordero, R.J.; Frases, S.; Rodrigues, J.; Goldman, D.L.; Andruszkiewicz, R.; Milewski, S.; Travassos, L.R.; Casadevall, A.; et al. Role for chitin and chitooligomers in the capsular architecture of Cryptococcus neoformans. Eukaryot. Cell 2009, 8, 1543–1553. [Google Scholar] [CrossRef] [PubMed]
- Saito, F.; Ikeda, R. Killing of Cryptococcus neoformans by Staphylococcus aureus: The role of cryptococcal capsular polysaccharide in the fungal-bacteria interaction. Med. Mycol. 2005, 43, 603–612. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, R.; Saito, F.; Matsuo, M.; Kurokawa, K.; Sekimizu, K.; Yamaguchi, M.; Kawamoto, S. Contribution of the mannan backbone of cryptococcal glucuronoxylomannan and a glycolytic enzyme of Staphylococcus aureus to contact-mediated killing of Cryptococcus neoformans. J. Bacteriol. 2007, 189, 4815–4826. [Google Scholar] [CrossRef] [PubMed]
- Kernien, J.F.; Snarr, B.D.; Sheppard, D.C.; Nett, J.E. The interface between fungal biofilms and innate immunity. Front. Immunol. 2017, 8, 1968. [Google Scholar] [CrossRef] [PubMed]
- Bach, M.C.; Tally, P.W.; Godofsky, E.W. Use of cerebrospinal fluid shunts in patients having acquired immunodeficiency syndrome with cryptococcal meningitis and uncontrollable intracranial hypertension. Neurosurgery 1997, 41, 1280–1282; discussion 1282–1283. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Fries, B.C. Fungal biofilms: Relevance in the setting of human disease. Curr. Fungal Infect. Rep. 2010, 4, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Casadevall, A. Susceptibility of Cryptococcus neoformans biofilms to antifungal agents in vitro. Antimicrob. Agents Chemother. 2006, 50, 1021–1033. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Casadevall, A. Cryptococcus neoformans cells in biofilms are less susceptible than planktonic cells to antimicrobial molecules produced by the innate immune system. Infect. Immun. 2006, 74, 6118–6123. [Google Scholar] [CrossRef] [PubMed]
- Nosanchuk, J.D.; Casadevall, A. Cellular charge of Cryptococcus neoformans: Contributions from the capsular polysaccharide, melanin, and monoclonal antibody binding. Infect. Immun. 1997, 65, 1836–1841. [Google Scholar] [PubMed]
- Alvarez, M.; Saylor, C.; Casadevall, A. Antibody action after phagocytosis promotes Cryptococcus neoformans and Cryptococcus gattii macrophage exocytosis with biofilm-like microcolony formation. Cell. Microbiol. 2008, 10, 1622–1633. [Google Scholar] [CrossRef] [PubMed]
- Jain, N.; Guerrero, A.; Fries, B.C. Phenotypic switching and its implications for the pathogenesis of Cryptococcus neoformans. FEMS Yeast Res. 2006, 6, 480–488. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Ibom, D.C.; Casadevall, A.; Fries, B.C. Characterization of phenotypic switching in Cryptococcus neoformans biofilms. Mycopathologia 2008, 166, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Fries, B.C.; Goldman, D.L.; Cherniak, R.; Ju, R.; Casadevall, A. Phenotypic switching in Cryptococcus neoformans results in changes in cellular morphology and glucuronoxylomannan structure. Infect. Immun. 1999, 67, 6076–6083. [Google Scholar] [PubMed]
- Fries, B.C.; Taborda, C.P.; Serfass, E.; Casadevall, A. Phenotypic switching of Cryptococcus neoformans occurs in vivo and influences the outcome of infection. J. Clin. Investig. 2001, 108, 1639–1648. [Google Scholar] [CrossRef] [PubMed]
- Aslanyan, L.; Sanchez, D.A.; Valdebenito, S.; Eugenin, E.A.; Ramos, R.L.; Martinez, L.R. The crucial role of biofilms in Cryptococcus neoformans survival within macrophages and colonization of the central nervous system. J. Fungi (Basel) 2017, 3, 10. [Google Scholar] [CrossRef] [PubMed]
- Moranova, Z.; Virtudazo, E.; Hricova, K.; Ohkusu, M.; Kawamoto, S.; Husickova, V.; Raclavsky, V. The CRZ1/SP1-like gene links survival under limited aeration, cell integrity and biofilm formation in the pathogenic yeast Cryptococcus neoformans. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc Czech Repub. 2014, 158, 212–220. [Google Scholar] [CrossRef] [PubMed]
- Robertson, E.J.; Wolf, J.M.; Casadevall, A. Edta inhibits biofilm formation, extracellular vesicular secretion, and shedding of the capsular polysaccharide glucuronoxylomannan by Cryptococcus neoformans. Appl. Environ. Microbiol. 2012, 78, 7977–7984. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L.R.; Christaki, E.; Casadevall, A. Specific antibody to Cryptococcus neoformans glucurunoxylomannan antagonizes antifungal drug action against cryptococcal biofilms in vitro. J. Infect. Dis. 2006, 194, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Levitz, S.M.; Specht, C.A. The molecular basis for the immunogenicity of Cryptococcus neoformans mannoproteins. FEMS Yeast Res. 2006, 6, 513–524. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, M.L.; Nimrichter, L. In good company: Association between fungal glycans generates molecular complexes with unique functions. Front. Microbiol. 2012, 3, 249. [Google Scholar] [CrossRef] [PubMed]
- Jesus, M.D.; Nicola, A.M.; Chow, S.K.; Lee, I.R.; Nong, S.; Specht, C.A.; Levitz, S.M.; Casadevall, A. Glucuronoxylomannan, galactoxylomannan, and mannoprotein occupy spatially separate and discrete regions in the capsule of Cryptococcus neoformans. Virulence 2010, 1, 500–508. [Google Scholar] [CrossRef] [PubMed]
- Vartivarian, S.E.; Reyes, G.H.; Jacobson, E.S.; James, P.G.; Cherniak, R.; Mumaw, V.R.; Tingler, M.J. Localization of mannoprotein in Cryptococcus neoformans. J. Bacteriol. 1989, 171, 6850–6852. [Google Scholar] [CrossRef] [PubMed]
- Gow, N.A.R.; Latge, J.P.; Munro, C.A. The fungal cell wall: Structure, biosynthesis, and function. Microbiol. Spectr. 2017, 5. [Google Scholar] [CrossRef]
- de Nobel, H.; Lipke, P.N. Is there a role for gpis in yeast cell-wall assembly? Trends Cell Biol. 1994, 4, 42–45. [Google Scholar] [CrossRef]
- Biondo, C.; Messina, L.; Bombaci, M.; Mancuso, G.; Midiri, A.; Beninati, C.; Cusumano, V.; Gerace, E.; Papasergi, S.; Teti, G. Characterization of two novel cryptococcal mannoproteins recognized by immune sera. Infect. Immun. 2005, 73, 7348–7355. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, P.A.; Penha, L.L.; Mendonca-Previato, L.; Previato, J.O. Mannoprotein MP84 mediates the adhesion of Cryptococcus neoformans to epithelial lung cells. Front. Cell. Infect. Microbiol. 2014, 4, 106. [Google Scholar] [CrossRef] [PubMed]
- Siafakas, A.R.; Sorrell, T.C.; Wright, L.C.; Wilson, C.; Larsen, M.; Boadle, R.; Williamson, P.R.; Djordjevic, J.T. Cell wall-linked cryptococcal phospholipase B1 is a source of secreted enzyme and a determinant of cell wall integrity. J. Biol. Chem. 2007, 282, 37508–37514. [Google Scholar] [CrossRef] [PubMed]
- Djordjevic, J.T.; Del Poeta, M.; Sorrell, T.C.; Turner, K.M.; Wright, L.C. Secretion of cryptococcal phospholipase B1 (PLB1) is regulated by a glycosylphosphatidylinositol (GPI) anchor. Biochem. J. 2005, 389, 803–812. [Google Scholar] [CrossRef] [PubMed]
- Siafakas, A.R.; Wright, L.C.; Sorrell, T.C.; Djordjevic, J.T. Lipid rafts in Cryptococcus neoformans concentrate the virulence determinants phospholipase B1 and Cu/Zn superoxide dismutase. Eukaryot. Cell 2006, 5, 488–498. [Google Scholar] [CrossRef] [PubMed]
- Ganendren, R.; Carter, E.; Sorrell, T.; Widmer, F.; Wright, L. Phospholipase B activity enhances adhesion of Cryptococcus neoformans to a human lung epithelial cell line. Microbes Infect. 2006, 8, 1006–1015. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.C.; Wright, L.C.; Golding, J.C.; Sorrell, T.C. Purification and characterization of secretory phospholipase b, lysophospholipase and lysophospholipase/transacylase from a virulent strain of the pathogenic fungus Cryptococcus neoformans. Biochem. J. 2000, 347, 431–439. [Google Scholar] [CrossRef] [PubMed]
- Evans, R.J.; Li, Z.; Hughes, W.S.; Djordjevic, J.T.; Nielsen, K.; May, R.C. Cryptococcal phospholipase b1 is required for intracellular proliferation and control of titan cell morphology during macrophage infection. Infect. Immun. 2015, 83, 1296–1304. [Google Scholar] [CrossRef] [PubMed]
- De Leon-Rodriguez, C.M.; Rossi, D.C.P.; Fu, M.S.; Dragotakes, Q.; Coelho, C.; Guerrero Ros, I.; Caballero, B.; Nolan, S.J.; Casadevall, A. The outcome of the Cryptococcus neoformans-macrophage interaction depends on phagolysosomal membrane integrity. J. Immunol. 2018, 201, 583–603. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.C.; Crary, B.; Chang, Y.C.; Kwon-Chung, K.J.; Kim, K.J. Cryptococcus neoformans activates rhogtpase proteins followed by protein kinase C, focal adhesion kinase, and ezrin to promote traversal across the blood-brain barrier. J. Biol. Chem. 2012, 287, 36147–36157. [Google Scholar] [CrossRef] [PubMed]
- Maruvada, R.; Zhu, L.; Pearce, D.; Zheng, Y.; Perfect, J.; Kwon-Chung, K.J.; Kim, K.S. Cryptococcus neoformans phospholipase B1 activates host cell RAC1 for traversal across the blood-brain barrier. Cell. Microbiol. 2012, 14, 1544–1553. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.C.; Jong, A.; Huang, S.; Zerfas, P.; Kwon-Chung, K.J. Cps1, a homolog of the streptococcus pneumoniae type 3 polysaccharide synthase gene, is important for the pathobiology of Cryptococcus neoformans. Infect. Immun. 2006, 74, 3930–3938. [Google Scholar] [CrossRef] [PubMed]
- Jong, A.; Wu, C.H.; Chen, H.M.; Luo, F.; Kwon-Chung, K.J.; Chang, Y.C.; Lamunyon, C.W.; Plaas, A.; Huang, S.H. Identification and characterization of CPS1 as a hyaluronic acid synthase contributing to the pathogenesis of Cryptococcus neoformans infection. Eukaryot. Cell 2007, 6, 1486–1496. [Google Scholar] [CrossRef] [PubMed]
- Jong, A.; Wu, C.H.; Shackleford, G.M.; Kwon-Chung, K.J.; Chang, Y.C.; Chen, H.M.; Ouyang, Y.; Huang, S.H. Involvement of human CD44 during Cryptococcus neoformans infection of brain microvascular endothelial cells. Cell. Microbiol. 2008, 10, 1313–1326. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.H.; Long, M.; Wu, C.H.; Kwon-Chung, K.J.; Chang, Y.C.; Chi, F.; Lee, S.; Jong, A. Invasion of Cryptococcus neoformans into human brain microvascular endothelial cells is mediated through the lipid rafts-endocytic pathway via the dual specificity tyrosine phosphorylation-regulated kinase 3 (DYRK3). J. Biol. Chem. 2011, 286, 34761–34769. [Google Scholar] [CrossRef] [PubMed]
- Vu, K.; Eigenheer, R.A.; Phinney, B.S.; Gelli, A. Cryptococcus neoformans promotes its transmigration into the central nervous system by inducing molecular and cellular changes in brain endothelial cells. Infect. Immun. 2013, 81, 3139–3147. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhang, K.Q. Independent expansion of zincin metalloproteinases in onygenales fungi may be associated with their pathogenicity. PLoS ONE 2014, 9, e90225. [Google Scholar] [CrossRef] [PubMed]
- Vu, K.; Tham, R.; Uhrig, J.P.; Thompson, G.R., 3rd; Na Pombejra, S.; Jamklang, M.; Bautos, J.M.; Gelli, A. Invasion of the central nervous system by Cryptococcus neoformans requires a secreted fungal metalloprotease. MBio 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Fang, W.; Fa, Z.Z.; Xie, Q.; Wang, G.Z.; Yi, J.; Zhang, C.; Meng, G.X.; Gu, J.L.; Liao, W.Q. Complex roles of annexin a2 in host blood-brain barrier invasion by Cryptococcus neoformans. CNS Neurosci. Ther. 2017, 23, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Na Pombejra, S.; Salemi, M.; Phinney, B.S.; Gelli, A. The metalloprotease, mpr1, engages annexina2 to promote the transcytosis of fungal cells across the blood-brain barrier. Front. Cell. Infect. Microbiol. 2017, 7, 296. [Google Scholar] [CrossRef] [PubMed]
- Stukes, S.; Coelho, C.; Rivera, J.; Jedlicka, A.E.; Hajjar, K.A.; Casadevall, A. The membrane phospholipid binding protein annexin A2 promotes phagocytosis and nonlytic exocytosis of Cryptococcus neoformans and impacts survival in fungal infection. J. Immunol. 2016, 197, 1252–1261. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R. An account of fungal 14-3-3 proteins. Eur. J. Cell Biol. 2017, 96, 206–217. [Google Scholar] [CrossRef] [PubMed]
- Kelly, M.N.; Johnston, D.A.; Peel, B.A.; Morgan, T.W.; Palmer, G.E.; Sturtevant, J.E. Bmh1p (14-3-3) mediates pathways associated with virulence in Candida albicans. Microbiology 2009, 155, 1536–1546. [Google Scholar] [CrossRef] [PubMed]
- Pham, C.D.; Yu, Z.; Sandrock, B.; Bolker, M.; Gold, S.E.; Perlin, M.H. Ustilago maydis Rho1 and 14-3-3 homologues participate in pathways controlling cell separation and cell polarity. Eukaryot. Cell 2009, 8, 977–989. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, M.L.; Nakayasu, E.S.; Oliveira, D.L.; Nimrichter, L.; Nosanchuk, J.D.; Almeida, I.C.; Casadevall, A. Extracellular vesicles produced by Cryptococcus neoformans contain protein components associated with virulence. Eukaryot. Cell 2008, 7, 58–67. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Chang, Y.C.; Wu, C.H.; Liu, J.; Kwon-Chung, K.J.; Huang, S.H.; Shimada, H.; Fante, R.; Fu, X.; Jong, A. The 14-3-3 gene function of Cryptococcus neoformans is required for its growth and virulence. J. Microbiol. Biotechnol. 2016, 26, 918–927. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.H.; Wu, C.H.; Chang, Y.C.; Kwon-Chung, K.J.; Brown, R.J.; Jong, A. Cryptococcus neoformans-derived microvesicles enhance the pathogenesis of fungal brain infection. PLoS ONE 2012, 7, e48570. [Google Scholar] [CrossRef] [PubMed]
- Lipke, P.N. What we do not know about fungal cell adhesion molecules. J. Fungi (Basel) 2018, 4, 59. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.B.; Gulati, M.; Johnson, A.D.; Nobile, C.J. Development and regulation of single- and multi-species Candida albicans biofilms. Nat. Rev. Microbiol. 2018, 16, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Freij, J.B.; Fu, M.S.; De Leon Rodriguez, C.M.; Dziedzic, A.; Jedlicka, A.E.; Dragotakes, Q.; Rossi, D.C.P.; Jung, E.H.; Coelho, C.; Casadevall, A. Conservation of intracellular pathogenic strategy among distantly related cryptococcal species. Infect. Immun. 2018, 86. [Google Scholar] [CrossRef] [PubMed]
| Cryptococcal Attributes | Adherence-Related Roles | |
|---|---|---|
| Saprophytic Lifestyle | Pathogenesis | |
| GXM capsular polysaccharide | Binding to respiratory epithelia through interaction with CD14 and TLR2, and TLR 4 receptors [94,97]. Biofilm formation favoring: adherence to intracranial medical devices [110], increased resistance to antifungal drugs [111,112], less susceptibility to host oxidative stress molecules [113], and colonization of the host CNS [120]. | |
| Exo-polysaccharides | Biofilm formation promoting attachment to abiotic surfaces [51]. | |
| Mannoprotein 84 (MP84) | Adhesion to lung epithelial cells in a capsule-independent manner [131]. | |
| Phospholipase B (Plb1) | Adhesion to lung epithelial cells via fatty acid released from host substrates [135]. Binding to HBMEC monolayers mediated by alteration of the host actin cytoskeleton, which leads to transmigration across BBB, and colonization of the brain [139,140]. | |
| Hyaluronic acid synthase (CPS1) | Interaction with CD44-containing lipid rafts expressed by HBMEC that promotes rearrangement of the host cytoskeleton, migration across BBB, and invasion of brain endothelial cells [143,144,145]. | |
| Metalloprotease (Mpr1) | Selective attachment and internalization to BBB via disruption of surface proteins of the HBMEC [147]. | |
| 14-3-3 adhesin | Vesicle-secreted protein involved in adherence to HBMEC monolayer possible promoted by other vesicle transported fungal components such as urease [155,156]. | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Camacho, E.; Casadevall, A. Cryptococcal Traits Mediating Adherence to Biotic and Abiotic Surfaces. J. Fungi 2018, 4, 88. https://doi.org/10.3390/jof4030088
Camacho E, Casadevall A. Cryptococcal Traits Mediating Adherence to Biotic and Abiotic Surfaces. Journal of Fungi. 2018; 4(3):88. https://doi.org/10.3390/jof4030088
Chicago/Turabian StyleCamacho, Emma, and Arturo Casadevall. 2018. "Cryptococcal Traits Mediating Adherence to Biotic and Abiotic Surfaces" Journal of Fungi 4, no. 3: 88. https://doi.org/10.3390/jof4030088
APA StyleCamacho, E., & Casadevall, A. (2018). Cryptococcal Traits Mediating Adherence to Biotic and Abiotic Surfaces. Journal of Fungi, 4(3), 88. https://doi.org/10.3390/jof4030088





