Characterization of Blue Mold Penicillium Species Isolated from Stored Fruits Using Multiple Highly Conserved Loci
Abstract
:1. Introduction
2. Materials and Methods
2.1. Penicillium Isolation, Morphological Identification, and Mycotoxin Detection
2.2. DNA Extraction and PCR Amplification
2.3. Sequencing and Phylogenetic Analysis
3. Results
3.1. The Biological Characteristics of Penicillium spp.
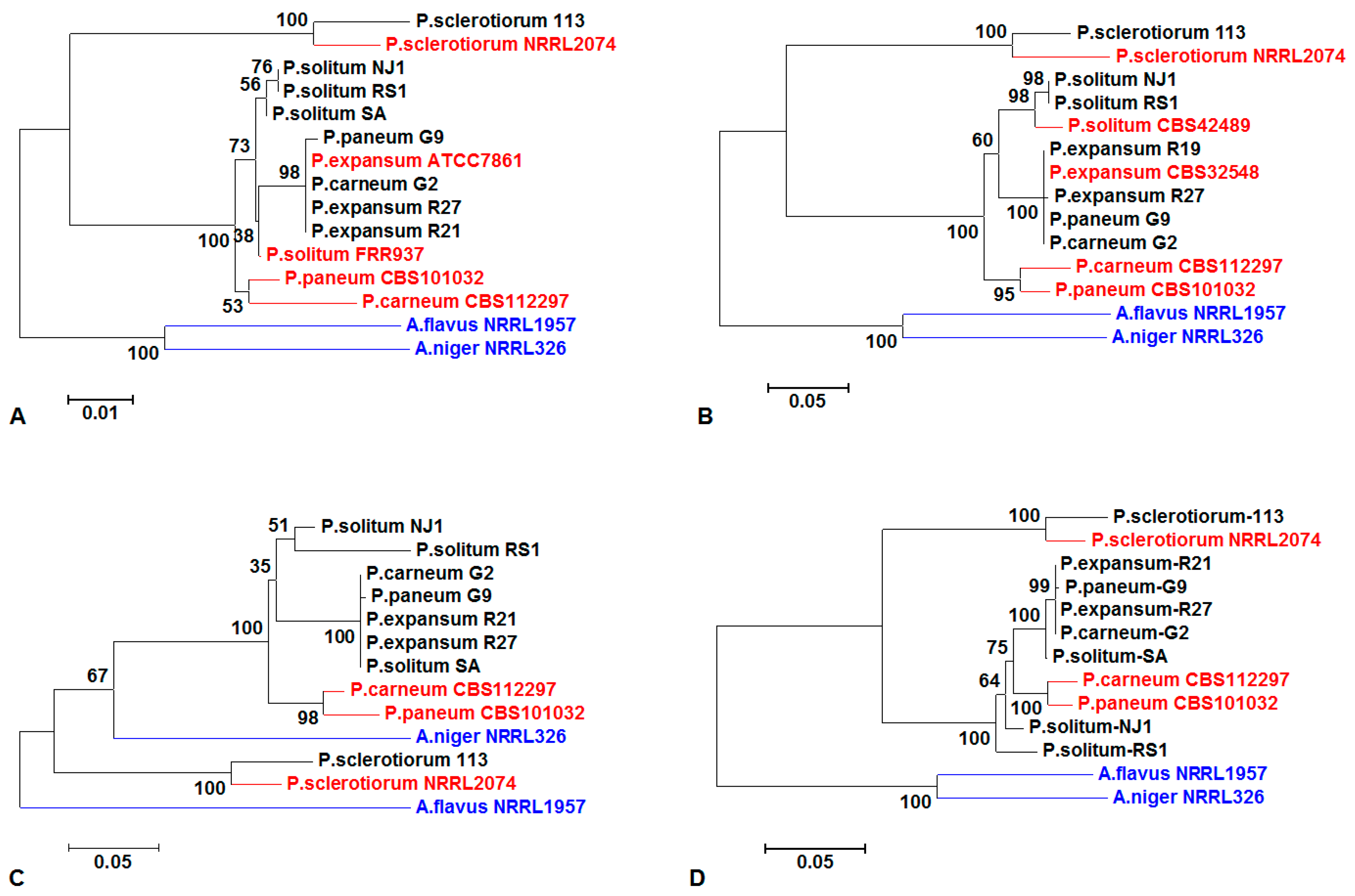
3.2. Sequencing Analysis of Penicillium spp. and Phylogenetic Analysis
3.3. Mycotoxin
4. Discussion and Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jurick, W.M., II; Yu, J.; Bennett, J.W. Blue Mould to Genomics and Beyond: Insights into the Biology and Virulence of Phytopathogenic Penicillium Species. In Aspergillus and Penicillium in the Post-Genmic Era; de Vries, R.P., Gelber, I.B., Anderson, M.R., Eds.; Caister Academic Press: Norfolk, UK, 2016; p. 210. [Google Scholar]
- Kim, W.K.; Sang, H.K.; Woo, S.K.; Park, M.S.; Paul, N.C.; Yu, S.H. Six species of Penicillium associated with blue mold of grape. Mycobiology 2007, 35, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Neri, F.; Mari, M.; Brigati, S. Control of Penicillium expansum by plant volatile compounds. Plant Pathol. 2006, 55, 100–105. [Google Scholar] [CrossRef]
- Jurick, W.M., II; Vico, I.; McEvoy, J.L.; Whitaker, B.D.; Janisiewicz, W.; Conway, W.S. Isolation, purification, and characterization of a polygalacturonase produced in Penicillium solitum-decayed "golden delicious" apple fruit. Phytopathology 2009, 99, 636–641. [Google Scholar] [CrossRef] [PubMed]
- Eckert, J.; Sievert, J.; Ratnayake, M. Reduction of imazalil effectiveness against citrus green mold in california packinghouses by resistant biotypes of Penicillium digitatum. Plant Dis. 1994, 78, 971–974. [Google Scholar] [CrossRef]
- Chalutz, E.; Wilson, C. Postharvest biocontrol of green and blue mold and sour rot of citrus fruit by Debaryomyces hansenii. Plant Dis. 1990, 74, 134–137. [Google Scholar] [CrossRef]
- Visagie, C.; Houbraken, J.; Frisvad, J.C.; Hong, S.-B.; Klaassen, C.; Perrone, G.; Seifert, K.; Varga, J.; Yaguchi, T.; Samson, R. Identification and nomenclature of the genus Penicillium. Stud. Mycol. 2014, 78, 343–371. [Google Scholar] [CrossRef] [PubMed]
- Hawksworth, D.L. Problems and prospects for improving the stability of names in Aspergillus and Penicillium. In Modern Concepts in Penicillium and Aspergillus Classification; Samson, R.A., Pitt, J.I., Eds.; Plenum Press: New York, NY, USA; London, UK, 1990; pp. 75–82. [Google Scholar]
- Pitt, J.I. The Genus Penicillium and Its Teleomorphic States Eupenicillium and Talaromyces. In The Genus Penicillium and its Teleomorphic States Eupenicillium and Talaromyces; Academic Press Inc., Ltd.: London, UK, 1979. [Google Scholar]
- Seifert, K.; Louis-Seize, G. Phylogeny and Species Concepts in the Penicillium aurantiogriseum Complex as Inferred from Partial β-Tubulin gene DNA Sequences. In Integration of Modern Taxonomic Methods for Penicillium and Aspergillus Classification; Harwood Academic Publishers: Amsterdam, The Netherlands, 2000; pp. 189–198. [Google Scholar]
- Samson, R.A.; Hoekstra, E.S.; Frisvad, J.C. Introduction to Food-and Airborne Fungi; Centraalbureau voor Schimmelcultures (CBS): Utrecht, The Netherlands, 2004. [Google Scholar]
- Tiwari, K.; Jadhav, S.; Kumar, A. Morphological and molecular study of different Penicillium species. Middle-East J. Sci. Res. 2011, 7, 203–210. [Google Scholar]
- Pianzzola, M.; Moscatelli, M.; Vero, S. Characterization of Penicillium isolates associated with blue mold on apple in Uruguay. Plant Dis. 2004, 88, 23–28. [Google Scholar] [CrossRef]
- La Guerche, S.; Garcia, C.; Darriet, P.; Dubourdieu, D.; Labarère, J. Characterization of Penicillium species isolated from grape berries by their internal transcribed spacer (ITS1) sequences and by gas chromatography–mass spectrometry analysis of geosmin production. Curr. Microbiol. 2004, 48, 405–411. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W.; Bolchacova, E.; Voigt, K.; Crous, P.W. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [PubMed]
- Seifert, K.A.; Samson, R.A.; Houbraken, J.; Lévesque, C.A.; Moncalvo, J.M.; Louis-Seize, G.; Hebert, P.D. Prospects for fungus identification using CO1 DNA barcodes, with Penicillium as a test case. Proc. Natl. Acad. Sci. USA 2007, 104, 3901–3906. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Robbertse, B.; Robert, V.; Vu, D.; Cardinali, G.; Irinyi, L.; Meyer, W.; Nilsson, R.H.; Hughes, K.; Miller, A.N. Finding needles in haystacks: Linking scientific names, reference specimens and molecular data for fungi. Database 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- Houbraken, J.; Visagie, C.M.; Meijer, M.; Frisvad, J.C.; Busby, P.E.; Pitt, J.I.; Seifert, K.A.; Louis-Seize, G.; Demirel, R.; Yilmaz, N.; et al. A taxonomic and phylogenetic revision of Penicillium section Aspergilloides. Stud. Mycol. 2014, 78, 373–451. [Google Scholar] [CrossRef] [PubMed]
- Yin, G.; Zhang, Y.; Pennerman, K.K.; Hua, S.S.; Yu, J.; Guo, A.; Liu, Z.; Bennett, J.W. Draft genome sequence of the fungus Penicillium solitum NJ1. Genome Announc. 2016, 4, e01176-16. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Yin, G.; Inamda, A.A.; Luo, J.; Zhang, N.; Yang, I.; Buckley, B.; Bennett, J.W. Volatile organic compounds emitted by filamentous fungi isolated from flooded homes after hurricane Sandy show toxicity in a Drosophila bioassay. Indoor Air 2016. [Google Scholar] [CrossRef] [PubMed]
- Yin, G.; Zhang, Y.; Pennerman, K.K.; Hua, S.S.T.; Huang, Q.; Guo, A.; Liu, Z.; Bennett, J.W. Genome sequencing and analysis of the filamentous fungus Penicillium sclerotiorum 113, isolated after hurricane Sandy. Genome Announc. 2016, 4, e01153-16. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.-J.K.; Zabel, R.A. Identification Manual for Fungi from Utility Poles in the Eastern United States; American Type Culture Collection: Manassas, VA, USA, 1990. [Google Scholar]
- Samson, R.A.; Hong, S.; Peterson, S.; Frisvad, J.C.; Varga, J. Polyphasic taxonomy of Aspergillus section Fumigati and its teleomorph Neosartorya. Stud. Mycol. 2007, 59, 147–203. [Google Scholar] [CrossRef] [PubMed]
- Samson, R.A.; Noonim, P.; Meijer, M.; Houbraken, J.; Frisvad, J.C.; Varga, J. Diagnostic tools to identify black Aspergilli. Stud. Mycol. 2007, 59, 129–145. [Google Scholar] [CrossRef] [PubMed]
- Rivera, K.; Seifert, K. A taxonomic and phylogenetic revision of the Penicillium sclerotiorum complex. Stud. Mycol. 2011, 70, 139–158. [Google Scholar] [CrossRef] [PubMed]
- Gimeno, A.; Martins, M.L. Rapid thin layer chromatographic determination of patulin, citrinin, and aflatoxin in apples and pears, and their juices and jams. J. Assoc. Off. Anal. Chem. 1983, 66, 85–91. [Google Scholar] [PubMed]
- Frisvad, J.C.; Filtenborg, O.; Thrane, U. Analysis and screening for mycotoxins and other secondary metabolites in fungal cultures by thin-layer chromatography and high-performance liquid chromatography. Arch. Environ. Contam. Toxicol. 1989, 18, 331–335. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols Guide Methods Appl. 1990, 18, 315–322. [Google Scholar]
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [PubMed]
- Hong, S.B.; Cho, H.S.; Shin, H.D.; Frisvad, J.C.; Samson, R.A. Novel Neosartorya species isolated from soil in korea. Int. J. Syst. Evolut. Microbiol. 2006, 56, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Peterson, S.W.; Vega, F.E.; Posada, F.; Nagai, C. Penicillium coffeae, a new endophytic species isolated from a coffee plant and its phylogenetic relationship to P. fellutanum, P. thiersii and P. brocae based on parsimony analysis of multilocus DNA sequences. Mycologia 2005, 97, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef] [PubMed]
- Houbraken, J.; Spierenburg, H.; Frisvad, J.C. Rasamsonia, a new genus comprising thermotolerant and thermophilic Talaromyces and Geosmithia species. Antonie Leeuwenhoek 2012, 101, 403–421. [Google Scholar] [CrossRef] [PubMed]
- Lindner, D.L.; Banik, M.T. Molecular phylogeny of Laetiporus and other brown rot polypore genera in North America. Mycologia 2008, 100, 417–430. [Google Scholar] [CrossRef] [PubMed]
- Goujon, M.; McWilliam, H.; Li, W.; Valentin, F.; Squizzato, S.; Paern, J.; Lopez, R. A new bioinformatics analysis tools framework at EMBL_EBI. Nucleic Acids Res. 2010, 38, W695–W699. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 783–791. [Google Scholar] [CrossRef]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. Mega6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Yu, L.; Kubatko, L.; Pearl, D.K.; Edwards, S.V. Coalescent methods for estimating phylogenetic trees. Mol. Phylogenet. Evol. 2009, 53, 320–328. [Google Scholar] [CrossRef] [PubMed]
- Gadagkar, S.R.; Rosenberg, M.S.; Kumar, S. Inferring species phylogenies from multiple genes: Concatenated sequence tree versus consensus gene tree. J. Exp. Zool. Part B Mol. Dev. Evol. 2005, 304, 64–74. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zong, Y.; Du, Z.; Chen, Y.; Zhang, Z.; Qin, G.; Zhao, W.; Tian, S. Genomic characterization reveals insights into patulin biosynthesis and pathogenicity in Penicillium species. Mol. Plant-Microbe Interact. 2015, 28, 635–647. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Jurick, W.M., II; Cao, H.; Yin, Y.; Gaskins, V.L.; Losada, L.; Zafar, N.; Kim, M.; Bennett, J.W.; Nierman, W.C. Draft genome sequence of Penicillium expansum strain R19, which causes postharvest decay of apple fruit. Genome Announc. 2014, 2, e00635-14. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Wu, G.; Jurick II, W.M.; Gaskins, V.L.; Yin, Y.; Yin, G.; Bennett, J.W.; Shelton, D.R. Genome sequence of Penicillium solitum RS1, which causes postharvest apple decay. Genome Announc. 2016, 4, e00363-16. [Google Scholar] [CrossRef] [PubMed]
- Pitt, J.; Spotts, R.; Holmes, R.; Cruickshank, R. Penicillium solitum revived, and its role as a pathogen of pomaceous fruit. Phytopathology 1991, 81, 1108–1112. [Google Scholar] [CrossRef]
- Gardes, M.; Bruns, T.D. Its primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef] [PubMed]
- Stielow, J.; Lévesque, C.; Seifert, K.; Meyer, W.; Iriny, L.; Smits, D.; Renfurm, R.; Verkley, G.; Groenewald, M.; Chaduli, D. One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Pers. Mol. Phylogeny Evol. Fungi 2015, 35, 242. [Google Scholar] [CrossRef] [PubMed]
- Woudenberg, J.; Seidl, M.; Groenewald, J.; de Vries, M.; Stielow, J.; Thomma, B.; Crous, P. Alternaria section Alternaria: Species, formae speciales or pathotypes? Stud. Mycol. 2015, 82, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; An, C.; Xu, S.; Lee, S.; Yamamoto, N. High-throughput sequencing reveals unprecedented diversities of Aspergillus species in outdoor air. Lett. Appl. Microbiol. 2016, 63, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Yamamoto, N. Accuracy of the high-throughput amplicon sequencing to identify species within the genus Aspergillus. Fungal Biol. 2015, 119, 1311–1321. [Google Scholar] [CrossRef] [PubMed]
- Diepeningen, A.D.; Feng, P.; Ahmed, S.; Sudhadham, M.; Bunyaratavej, S.; Hoog, G.S. Spectrum of Fusarium infections in tropical dermatology evidenced by multilocus sequencing typing diagnostics. Mycoses 2015, 58, 48–57. [Google Scholar] [CrossRef] [PubMed]
- Jančič, S.; Nguyen, H.D.; Frisvad, J.C.; Zalar, P.; Schroers, H.-J.; Seifert, K.A.; Gunde-Cimerman, N. A taxonomic revision of the Wallemia sebi species complex. PLoS ONE 2015, 10, e0125933. [Google Scholar] [CrossRef] [PubMed]
- Sholberg, P.L.; Harlton, C.; Haag, P.; Le´vesque, C.A.; O’Gorman, D.; Seifert, K. Benzimidazole and diphenylamine sensitivity and identity of Penicillium spp. that cause postharvest blue mould of apples using β-tubulin gene sequences. Postharv. Biol. Technol. 2005, 36, 41–49. [Google Scholar] [CrossRef]
- Pitt, J. The current role of Aspergillus and Penicillium in human and animal health. J. Med. Vet. Mycol. 1994, 32, 17–32. [Google Scholar] [CrossRef] [PubMed]
- Pitt, J. Toxigenic fungi and mycotoxins. Br. Med. Bull. 2000, 56, 184–192. [Google Scholar] [CrossRef] [PubMed]
- Julca, I.; Droby, S.; Sela, N.; Marcet-Houben, M.; Gabaldón, T. Contrasting genomic diversity in two closely related postharvest pathogens: Penicillium digitatum and Penicillium expansum. Genome Biol. Evol. 2016, 8, 218–227. [Google Scholar] [CrossRef] [PubMed]
| Gene | Primer | Sequence (5′→3′) | Length (bp) | Reference |
|---|---|---|---|---|
| Internal transcribed spacer (ITS) | ITS1F | CTTGGTCATTTAGAGGAAGTAA | ~600 | [28] |
| ITS4 | TCCTCCGCTTATTGATATGC | |||
| β-tubulin (benA) | Bt2a | GGTAACCAAATCGGTGCTGCTTTC | ~550 | [29] |
| Bt2b | ACCCTCAGTGTAGTGACCCTTGGC | |||
| Calmodulin (CaM) | CMD5 | CCGAGTACAAGGARGCCTTC | ~580 | [30] |
| CMD6 | CCGATRGAGGTCATRACGTGG | |||
| CF1 | GCCGACTCTTTGACYGARGAR | ~750 | [31] | |
| CF4 | TTTYTGCATCATRAGYTGGAC | |||
| RNA polymerase II second largest subunit (RPB2-1) | 5F | GAYGAYMGWGATCAYTTYGG | ~1000 | [32] |
| 7CR | CCCATRGCTTGYTTRCCCAT | |||
| RNA polymerase II second largest subunit (RPB2-2) | 5Feur | GAYGAYCGKGAYCAYTTCGG | ~1000 | [33] |
| 7CReur | CCCATRGCYTGYTTRCCCAT |
| Species | Host | Source (Location) | Year | Patulin 1 | Citrinin | Back Color |
|---|---|---|---|---|---|---|
| P. carneum G2 | Golden delicious | Pennsylvania | 2011 | +/− | +/− | tan |
| P. paneum G9 | Golden delicious | Pennsylvania | 2011 | +/− | +/− | yellow |
| P. expansum R19 | Red delicious | Pennsylvania | 2011 | + | + | tan |
| P. expansum R21 | Red delicious | Pennsylvania | 2011 | + | − | green |
| P. expansum R27 | Red delicious | Pennsylvania | 2011 | + | + | green |
| P. solitum RS1 | Apple | Oregon | 2011 | − | − | tan |
| P. solitum SA | Peach seed | West Virginia | 2011 | − | − | tan |
| P. solitum NJ1 | - | NRRL, Illinois | 2012 | − | − | tan |
| P. sclerotiorum 113 | Obtained from home living room | New Jersey | 2013 | +/− | +/− | orange |
| Penicillium spp. | ITS | benA | CaM a | CaM b | RPB2-1 | RPB2-2 |
|---|---|---|---|---|---|---|
| P. carneum G2 | KX243324 | KX243333 | KX243341 | - | KX243353 | KX243356 |
| P. paneum G9 | KX243325 | KX243334 | KX243342 | - | KX243354 | KX243357 |
| P. expansum R19 | - | KX243337 | - | - | - | - |
| P. expansum R21 | KX243328 | - | KX243345 | KX243351 | KX243355 | KX243358 |
| P. expansum R27 | KX243329 | KX243338 | KX243346 | - | - | - |
| P. solitum SA | KX243330 | - | KX243347 | - | - | - |
| P. solitum RS1 | KX243331 | KX243339 | KX243348 | - | - | - |
| P. solitum NJ1 | KX243323 | KX243332 | KX243340 | - | KX243352 | - |
| P. sclerotiorum 113 | KX365203 | KX365204 | KX365205 | - | KX365206 | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yin, G.; Zhang, Y.; Pennerman, K.K.; Wu, G.; Hua, S.S.T.; Yu, J.; Jurick, W.M., II; Guo, A.; Bennett, J.W. Characterization of Blue Mold Penicillium Species Isolated from Stored Fruits Using Multiple Highly Conserved Loci. J. Fungi 2017, 3, 12. https://doi.org/10.3390/jof3010012
Yin G, Zhang Y, Pennerman KK, Wu G, Hua SST, Yu J, Jurick WM II, Guo A, Bennett JW. Characterization of Blue Mold Penicillium Species Isolated from Stored Fruits Using Multiple Highly Conserved Loci. Journal of Fungi. 2017; 3(1):12. https://doi.org/10.3390/jof3010012
Chicago/Turabian StyleYin, Guohua, Yuliang Zhang, Kayla K. Pennerman, Guangxi Wu, Sui Sheng T. Hua, Jiujiang Yu, Wayne M. Jurick, II, Anping Guo, and Joan W. Bennett. 2017. "Characterization of Blue Mold Penicillium Species Isolated from Stored Fruits Using Multiple Highly Conserved Loci" Journal of Fungi 3, no. 1: 12. https://doi.org/10.3390/jof3010012
APA StyleYin, G., Zhang, Y., Pennerman, K. K., Wu, G., Hua, S. S. T., Yu, J., Jurick, W. M., II, Guo, A., & Bennett, J. W. (2017). Characterization of Blue Mold Penicillium Species Isolated from Stored Fruits Using Multiple Highly Conserved Loci. Journal of Fungi, 3(1), 12. https://doi.org/10.3390/jof3010012









