1. Introduction
Candida albicans is the most common source of oral and systemic fungal infections [
1]. Despite improvements in treatment modalities, mortality rates for these infections remain relatively high, in part due to drug resistance [
2]. Formation of biofilms by fungal cells contributes to drug resistance due to the higher cell density preventing efficient drug trafficking into cells, the presence of a protective extracellular matrix (ECM), and alterations in gene expression of cells within the biofilm [
3]. Numerous in vitro studies have analyzed the growth and development of
C. albicans biofilms on many surfaces, including acrylic, silicone, plastic, glass, and catheters [
4,
5,
6,
7]. However, these studies have largely been carried out under static conditions in which the medium overlying the biofilm lacks flow.
C. albicans colonizes surfaces under dynamic flow in vivo, including the oral mucosa or denture surfaces in the presence of salivary flow. In the oral cavity, saliva is a nutrient poor, low flow, complex bodily fluid consisting of host proteins, microbiota, and various food remnants [
8]. The average shear stress generated by saliva across a tooth surface has been calculated to be 0.8 dynes/cm
2 [
9]. Despite this relatively low shear stress value; it is likely that salivary flow contributes to reduced formation of
C. albicans biofilms, as patients with xerostomia (reduced salivary flow) are at increased risk of developing oral candidiasis [
10,
11].
Few in vitro studies have examined the role that fluid flow may have on
C. albicans biofilm development [
12,
13,
14], even though colonization of
C. albicans occurs under flow in several human niches. Flow is important for three main reasons: first, flow generates a force parallel to the surface of the substrate (shear force) that works to remove cells from the biofilm and therefore reduces the mass of the biofilm; second, flow is responsible for dispersion of detached cells and subsequent delivery of these cells to new sites; and thirdly, flow allows bulk fluid movement to deliver fresh nutrients to cells within the biofilm. This last point is particularly important for biofilm development since environmental nutrient availability is a potent signal for
C. albicans hyphal formation [
15]. Earlier studies confirm that biofilms formed under flow have different morphologies than static biofilms [
16]. Under static conditions, biofilms develop through multiple temporal stages that are characterized as adherence, initiation, maturation, and dispersion. The initial founder yeast cells adhere to the substrate in the adherence phase (the first 2 h); followed by propagation of these yeast cells to form microcolonies (and in which some top-most cells form germ tubes perpendicular from the substrate) in the initiation phase (2–11 h). During the maturation phase, the biomass expands into confluent multilayers along with formation of the extracellular matrix (ECM) (12–30 h), and hyphae formation increases in the uppermost layers. The dispersion phase is characterized by release of yeast cells exclusively from the topmost region of biofilm to disperse to new sites [
16]. The growth of
C. albicans biofilms in in vivo catheter models subjected to transient (once or twice a day) low flow exhibited different structures than those grown under static conditions [
7,
17,
18,
19,
20]. Particularly striking was that the foundational layer of biofilms formed under flow consisted of yeast, hyphae and pseudohyphae rather than the uniform layer of yeast cells at the base of statically formed biofilms [
16]. In vivo biofilms of
Candida formed in cases of oral candidiasis show an additional unique characteristic, namely the development of patches of infection [
21].
Of the few in vitro biofilm studies performed with
C. albicans under flow, only end-points of growth were measured, therefore understanding of the crucial events needed for biofilm growth and development is lacking [
12,
14,
22]. In this study, we developed a novel system allowing real-time imaging of
C. albicans biofilms under flow. This system permits quantitative measurement of the dynamics of biofilm development that up to now have not been assessed. We analyzed the rates of cell attachment and detachment to the substrate, rates of cell detachment independent of available biomass (normalized detachment rate), cell–cell adhesion relative to cell-surface adhesion, and the total biofilm growth rate to understand critical components and developmental stages of
Candida albicans biofilms under flow. We found that biofilms formed in our flow system not only reproduced the mixed yeast, hyphae, and pseudohyphae layers of biofilms formed in catheter models, but also developed microcolonies from single
Candida albicans hyphal cells that are very similar phenotypically to the microcolonies formed upon invasion of epithelial monolayers [
23].
2. Materials and Methods
2.1. Strains
Candida albicans CAI4
(URA+) (Δ
ura3::
imm434/Δ
ura3::
imm434 RPS1/Δ
rps1::
Clp10-
URA3) [
24] was used as WT control. The hyperfilamentous
C. albicans deletion mutant Δ
hog1 [
25] was used to compare morphologies of biofilm formation. Cultures were grown overnight in 1% (
w/
v) yeast extract, 2 % (
w/
v) bacto peptone, and 2% (
w/
v) glucose (YPD; Difco, Detroit, MI, USA). Cell densities of overnight cultures were determined using a cytometer, and values were used to determine volumes of overnight culture to add to the attachment flask (described below) to reach 1 × 10
6 cells/mL. After addition of culture, cells were allowed to acclimate for 15 m prior to initiation of flow.
2.2. Flow System
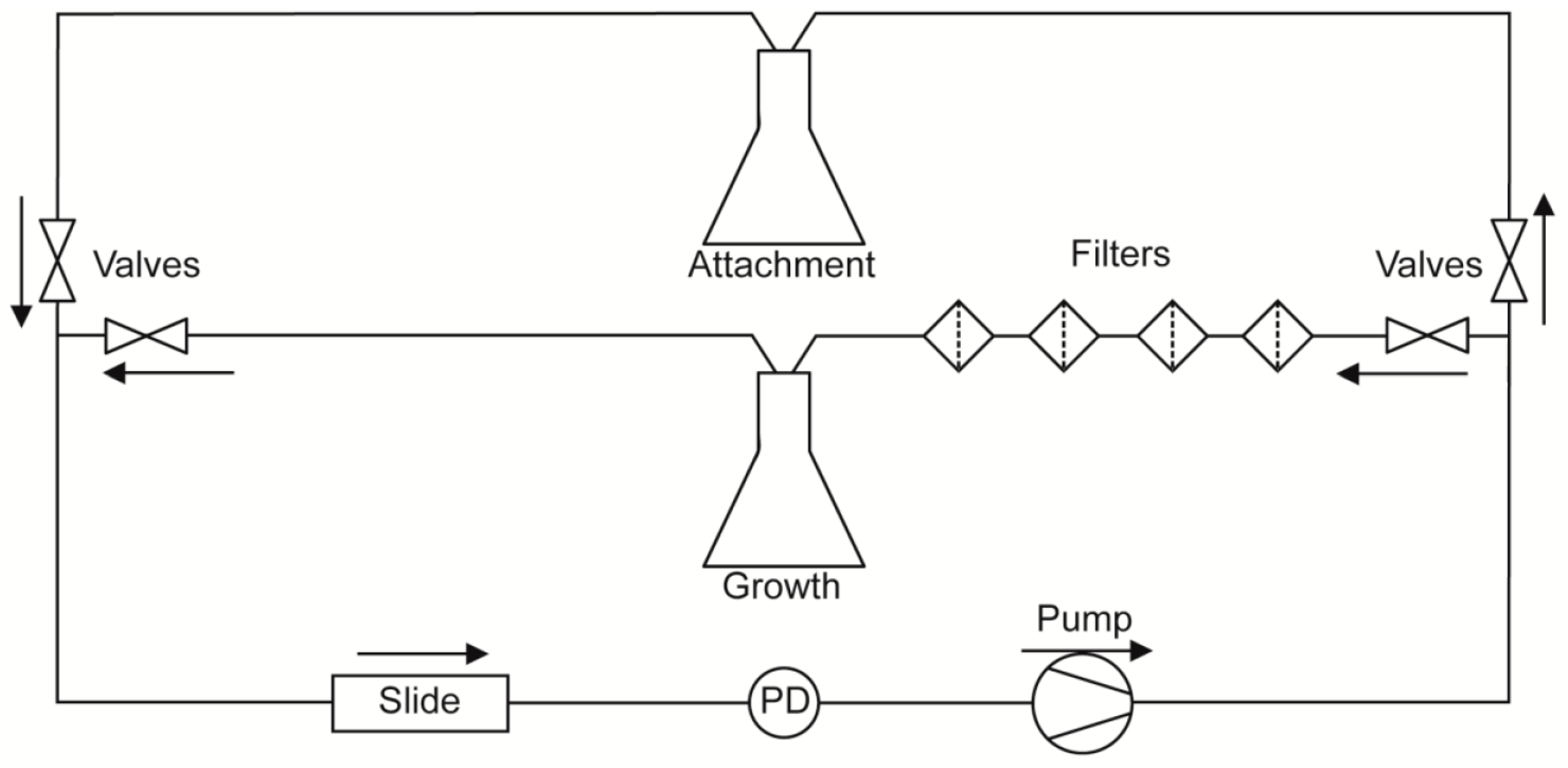
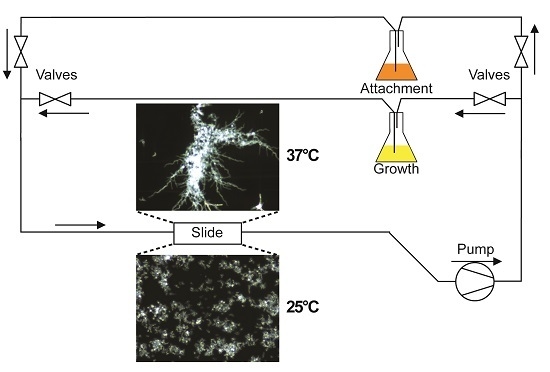
A diagram of the flow system used in this study is depicted in
Figure 1. To separate
C. albicans cell attachment from subsequent biofilm growth and development, and to allow evaluation of detachment independent of fresh cell seeding, flow experiments were split into two phases. During the first phase (attachment phase), fresh YPD seeded with
C. albicans cells (1 × 10
6 cells/mL) was circulated through a µ-Slide I 0.8 Luer family ibiTreat flow chamber (ibidi, Martinsried, Germany) using a Masterflex
® L/S
® variable speed pump (Cole-Parmer, Vernon Hills, IL, USA). This phase proceeded for 2 h, during which time cells were able to attach to the coverslip surface of the flow chamber. Afterwards, the source of media to the slide was switched to cell-free YPD for the remaining 16 h of the experiment (growth phase). The return flow during the growth phase was passed through four sequential cell filters: first two coarse filters (20 and 10 µm pore size; Analytical Scientific Instruments, Richmond, CA, USA), then a 2 µm pore size HPLC filter (Sigma Aldrich, St. Louis, MO, USA) followed by a 0.22 µm polyvinylidene fluoride filter (Sterivex™; Millipore, Billerica, MA, USA), before being recycled so as to prevent contamination of the stock medium. Thus, during the attachment phase, cells are allowed to re-circulate across the surface of the slide, but during the growth phase all cells are removed prior to re-circulation, and media to the slide remains cell-free for the rest of the experiment.
In all experiments, the flow was set to generate a shear force of 0.8 dynes/cm
2 across the surface of the flow chamber. This value has been previously calculated as the approximate shear force that human saliva exerts on the tooth surface [
9]. A hotplate stirrer with an external temperature probe was used to warm the media to 37 °C. For those experiments indicated as 37 °C the microscope, including the slide being imaged, and several feet of preceding tubing, were warmed to 37 °C, maintaining biofilm growth at this temperature, otherwise these were kept at room temperature (RT).
2.3. Imaging
All images were taken using a Zeiss AxioScope A.1 transmitted light microscope (Zeiss, Göttingen, Germany) using darkfield illumination, and acquired using the µManger software [
26,
27]. All imaging conditions (exposure time, light intensity, magnification, N.A., condenser height, and approximate positioning of the slide) were maintained between experiments. For all experiments, images were acquired every two minutes during the attachment phase, and every 15 min during the growth phase.
2.4. Image Analyses
All image analyses were performed in the ImageJ software environment [
28] after conversion to an 8-bit grayscale file format. ImageJ macro scripts used for all analyses are functionally described below. Statistical analyses, including linear and non-linear regressions, were performed in Graphpad Prism
® version 5.03 software.
To determine the coverage area of the biofilm, thresholds were applied to every image at a gray value minimum of 15, and percent surface area was measured. To evaluate the biomass of the attached cells (biofilm biomass), a densitometry analysis was performed. Specifically, the cumulative gray values of all pixels above 15 were evaluated for every frame of the darkfield time-lapse videos. The growth rate of each biofilm was then evaluated by linear regression of all biomass data collected.
To evaluate the rate of cell attachment during the attachment phase, a given frame (n) was subtracted from its next frame (n + 1) for every image of the attachment phase ((n + 1) – n). This subtraction resulted in an image where any cell that attached to the imaging region between the frames remained bright (at their original intensity), while cells that remained constant between frames were removed from the image. A threshold was then applied to these calculated images, highlighting newly attached cells, and subsequently processed using the ImageJ binary erosion filter to limit background noise and minor shifts in cell position. The area of newly attached cells (µm2) was then determined on each of these images using the analyze particles tool in ImageJ. To increase specificity towards cells, particles had to be a minimum of 20 µm2. Rates of cell attachment were then determined by fitting the cumulative attachment area for the first 2 h with linear regressions.
Rate of cell detachment was determined in a similar manner to cell attachment, but the image subtraction was reversed (n – (n + 1)), resulting in an image that highlighted cells that detached between frames. Detachment rates were evaluated over the entire duration of the experiment (attachment and growth phases) in a manner similar to attachment rates. The rate of total cell detachment was found to be dependent on the biomass of the biofilm, but this simply arises due to the increased number of cells available to detach. Thus, values obtained for this variable did not reflect the relative ease with which cells are removed from the biofilm or substrate surface, which is the parameter we were interested in. Thus, we normalized the area of detachment obtained between each frame to the biomass of the biofilm prior to these detachments ((n – (n + 1))area/nbiomass), resulting in a value that effectively represents the proportion of cells that detached (referred to as normalized detachment). The rates of normalized detachment were also evaluated using linear regressions, however data during the attachment phase was excluded from the analyses, as these values were often not stable, likely due to low biomass values.
To estimate relative cell–cell to cell-surface binding strengths, we performed image subtraction ((n + 1) – n) to determine newly attached cells at each frame, and applied a threshold to these images (as described above). These images were then processed using the ImageJ binary erosion filter, and particles at least 20 µm2 and with a circularity value of at least 0.4 were counted as cells. These particles were then compared to images of the biofilm coverage area of the preceding frame (n), to determine regions of overlap (completed using the “AND” operator in the ImageJ image calculator). Regions of overlap were counted as cell–cell adhesion events if they were at least 2.5 µm2. The number of cell–cell adhesions was then normalized to the total number of adhesion events, giving the relative cell–cell adhesion.
2.5. Full Slide Scans
All slide scans were conducted with a Bio-Rad GS-700 Imaging Densitometer (Bio-Rad, Hercules, CA, USA), and analyzed by densitometry analysis using ImageJ with no lower threshold applied.
2.6. Statistical Comparisons
For all rates determined through regression analyses, non-overlapping 95% confidence intervals were considered statistically significant at p < 0.05. For cell–cell adhesion, means ± S.D. of relative cell–cell adhesion between 1 and 2 h were determined and statistical significance was evaluated using a one-way ANOVA followed by a post-hoc Tukey’s t-test (significance at p < 0.05).
4. Discussion
In this study, we were able to develop a novel system to analyze in real-time the attachment and development of C. albicans biofilms under flow. With this system, we were able to analyze the first 18 h of biofilm development under several different conditions for unique quantitative measures of biofilm development, including the cell attachment rate, detachment rate, relative cell–cell adhesion and biofilm biomass over time. To the best of our knowledge, this is the first time that cell attachment and detachment of biofilm forming microbes has been measured in this manner.
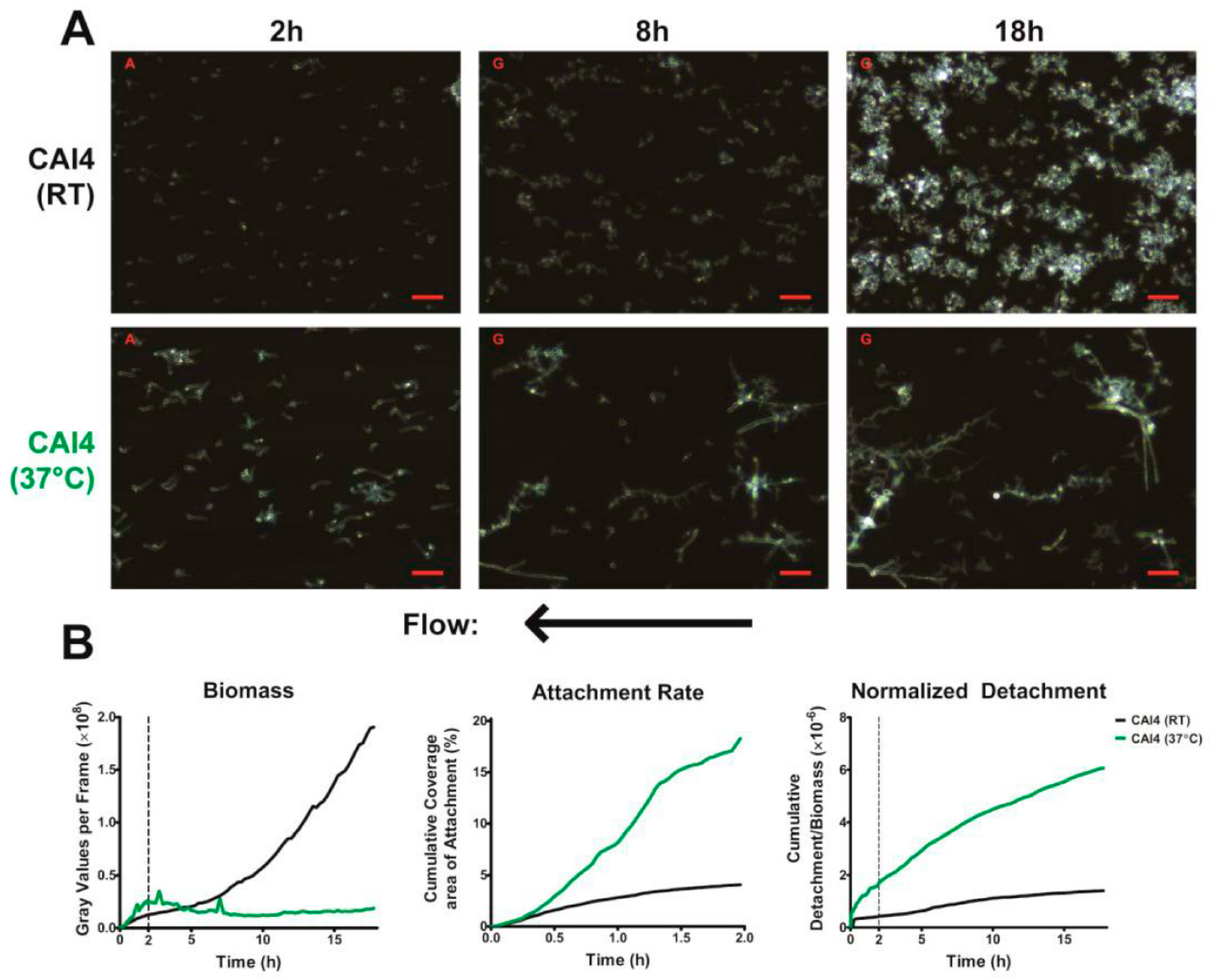
Our results support a previous study that showed that the dispersion of
C. albicans is a continuous process (
Figure 2B—right) [
14]. Cells detached from our biofilm throughout their growth and development, with the number of detaching cells increasing with increasing biofilm biomass. This is in contrast to the process of dispersion in bacterial cells, where they exhibit discontinuous large-scale dispersals by forming pillar and mushroom structures that ultimately release numerous bacteria upon maturation. As the process of dispersion was constant, and was found to be dependent on the available biomass, we normalized the detachment to the total biofilm biomass at each frame, permitting measurement of how easily cells were being removed from the biofilm (normalized detachment rate). We also found that the normalized detachment rate remains relatively constant for the first 18 h of biofilm development under flow. This was a surprising finding, since dispersion is considered a property of mature biofilms. However, our results show that cells under constant laminar flow undergo continuous detachment and seeding, and that these conditions may be more representative of the development of in vivo biofilms.
Using our imaging system at 37 °C, we were successfully able to observe the development of two major phenotypic characteristics of biofilms grown under flow. The first being that these biofilms formed mixed yeast, hyphae, and pseudohyphae basal layers (
Figure 2A), which has been previously observed in flow catheter models of biofilm formation [
7,
17,
18,
19,
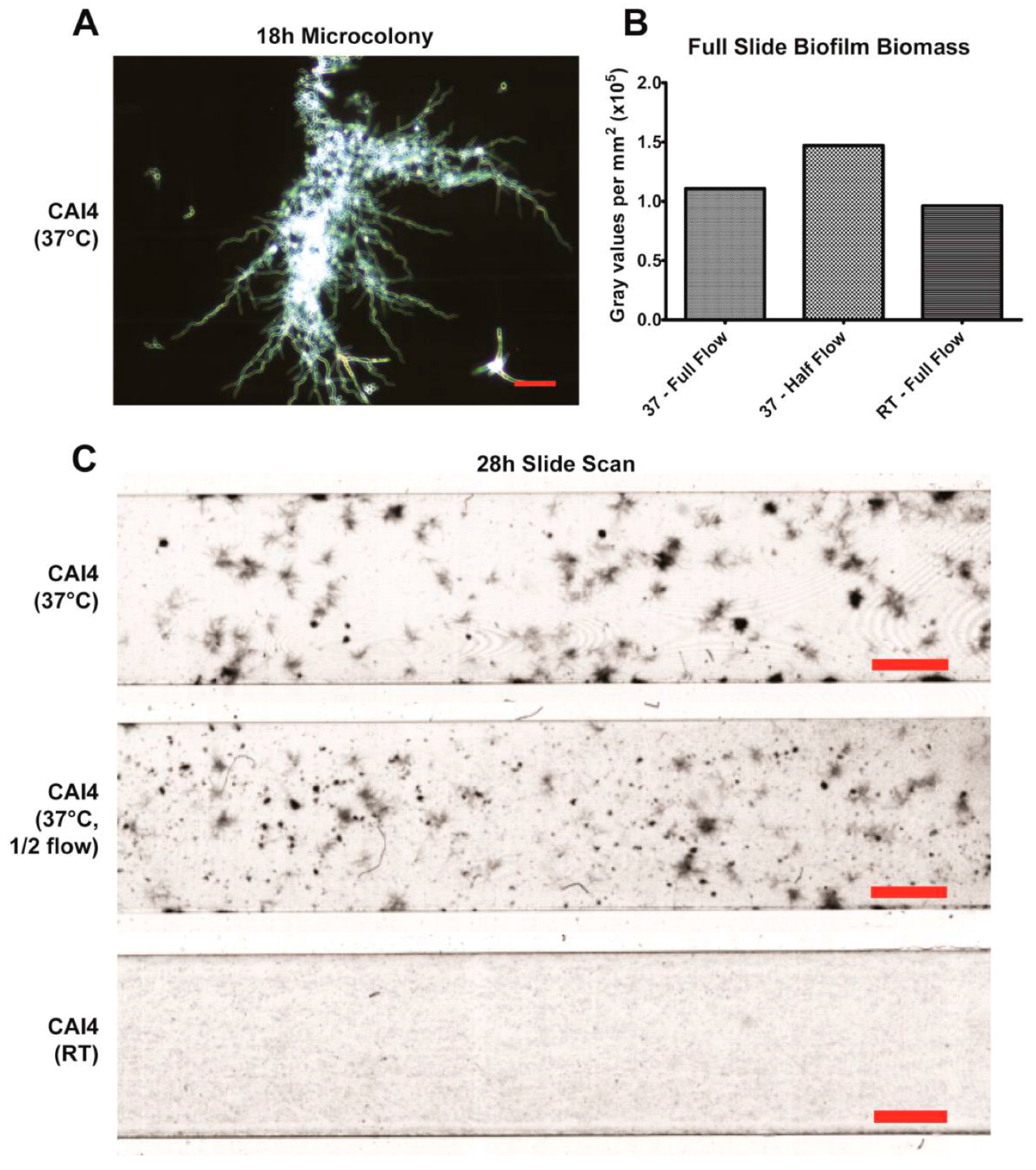
20]. This mixture seems to largely be the result of hyphae growing along the substrate surface, branching and forming laterally budded yeast and pseudohyphae. Additionally, flow-generated biofilms formed distinct microcolonies (
Figure 3A,C), similar to those found within tongue plaques in in vivo infection [
30]. These microcolonies evolved from single hyphal cells that branched extensively, forming vast tree-like structures.
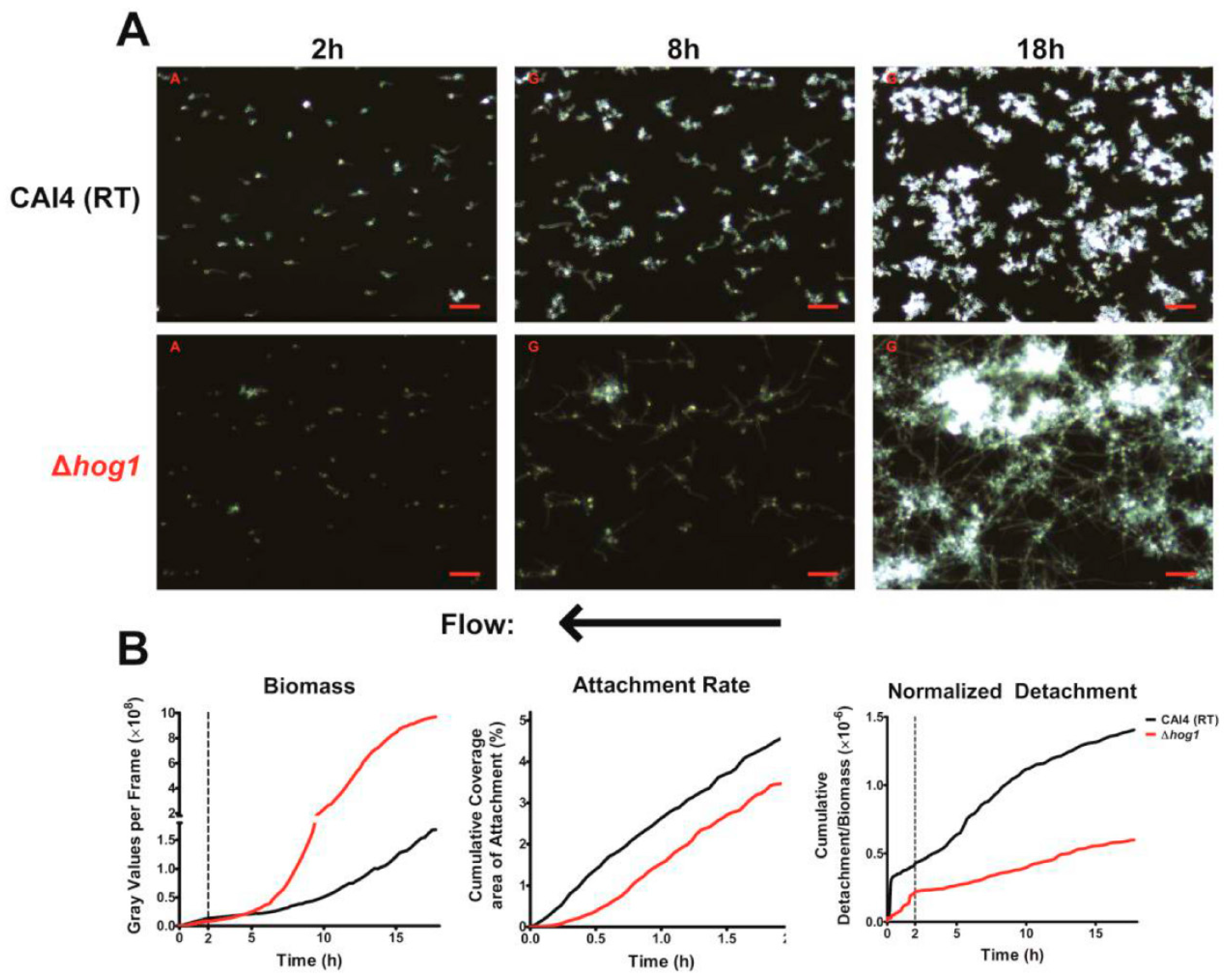
In addition to these two major characteristics, we also found that most hyphae (from both WT cells grown at 37 °C or Δ
hog1 cells grown at RT) grew in a sinusoidal morphology under flow conditions. This morphology has been observed previously, most notably when
C. albicans cells were grown on cellophane overlying a 2% agar base [
31,
32]. Previous studies identified these structures as 3-dimensional helices; however, upon closer examination, we found that the hyphae in our experiments were predominantly lying flat along the surface while growing in a sinusoidal fashion. It is likely that this morphology may be a result of laminar flow, but it is unclear if this hyphal morphology is also found in vivo. Interestingly, helical hyphae were found to be more resistant to antifungals than their straight hyphae counterparts [
31].
Flow rates can affect biofilm formation by both altering the shear force exerted on the biofilm and by altering the rate at which fresh nutrients are delivered to the cells. The latter is determined by the bulk flow velocity (cm/min) of the fluid, which can be calculated by dividing the flow rate (mL/min) by the cross-sectional surface area of the channel (cm2). Alterations to the flow rate will also alter both the shear force and the flow velocity, and can be used to model changes in salivary flow or to model other settings of fungal biofilm development such as catheters. It is possible to isolate the effects of variations in shear force or flow velocity independently from one another by adjusting both the flow rate and the cross-sectional surface area simultaneously. The slides used in our study are available in multiple channel heights, thus one parameter can be set as a constant (e.g., shear force = 0.8 dynes/cm2) across multiple channel heights by adjusting the flow rate. Thus, factors limiting biofilm development at a particular flow rate (either acquiring adequate nutrients in low flow velocity vs. maintaining adhesion in high shear force) can be determined.
Nutrient availability in our study is likely slightly affected by our use of a recirculating media system, which would also retain and recirculate cell signaling molecules. However, given that we saw continued cell division, and hyphal growth at >30 h (data not shown), we do not believe this is a major factor in our experiments. For prolonged, multi-day experiments, it would be possible to modify the system to continuously supply fresh nutrients, as described previously [
33].
The use of darkfield microscopy with our system is advantageous over the use of fluorescently labeled cells since there is no photo-bleaching. Additionally, fluorescence microscopy typically requires longer exposure times and higher intensity lighting that can cause phototoxicity [
34]. While fluorescent microscopy does have many tools available to assist in image analysis, particularly COMSTAT [
35], we were able to develop our own algorithms that took advantage of several features of darkfield microscopy. Traditional microscopy (no optical sectioning), as was used here, is power conservative, meaning that objects which are out of focus contribute nearly the same number of photons to an image as they would if they were in focus [
36]. In this system, cells that are further from the surface are still contributing to the analyses, therefore the contribution of all cells in the 3D biofilm are calculated despite capturing only a single 2D image at each time point.
Furthermore, this flow system can be easily adapted to study the effects of antimicrobial agents on biofilms by placement of an upstream in-line injection port for drug delivery. Also, epithelial or endothelial cells can be grown inside the slide channel prior to introduction of microbial cells to study fungal cell attachment, growth, and cell invasion in real time. This flow system can be assembled without use of custom manufactured parts and is reasonably inexpensive. The use of darkfield microscopy also allows images to be acquired with relatively simple microscopes, and allows seamless analysis of multiple microbial organisms. The versatility and adaptability of our flow system enable it to be used to study many different potential phenomena related to biofilm development.