Genetically Related Mycobacterium bovis Strains Displayed Differential Intracellular Growth in Bovine Macrophages
Abstract
:1. Introduction
2. Methods
2.1. Ethic Statement
2.2. Macrophage Culture
2.3. Mycobacterium Bovis Strains
2.4. Spoligotyping
2.5. Bactericidal Assay
2.6. Statistical Analysis
3. Results
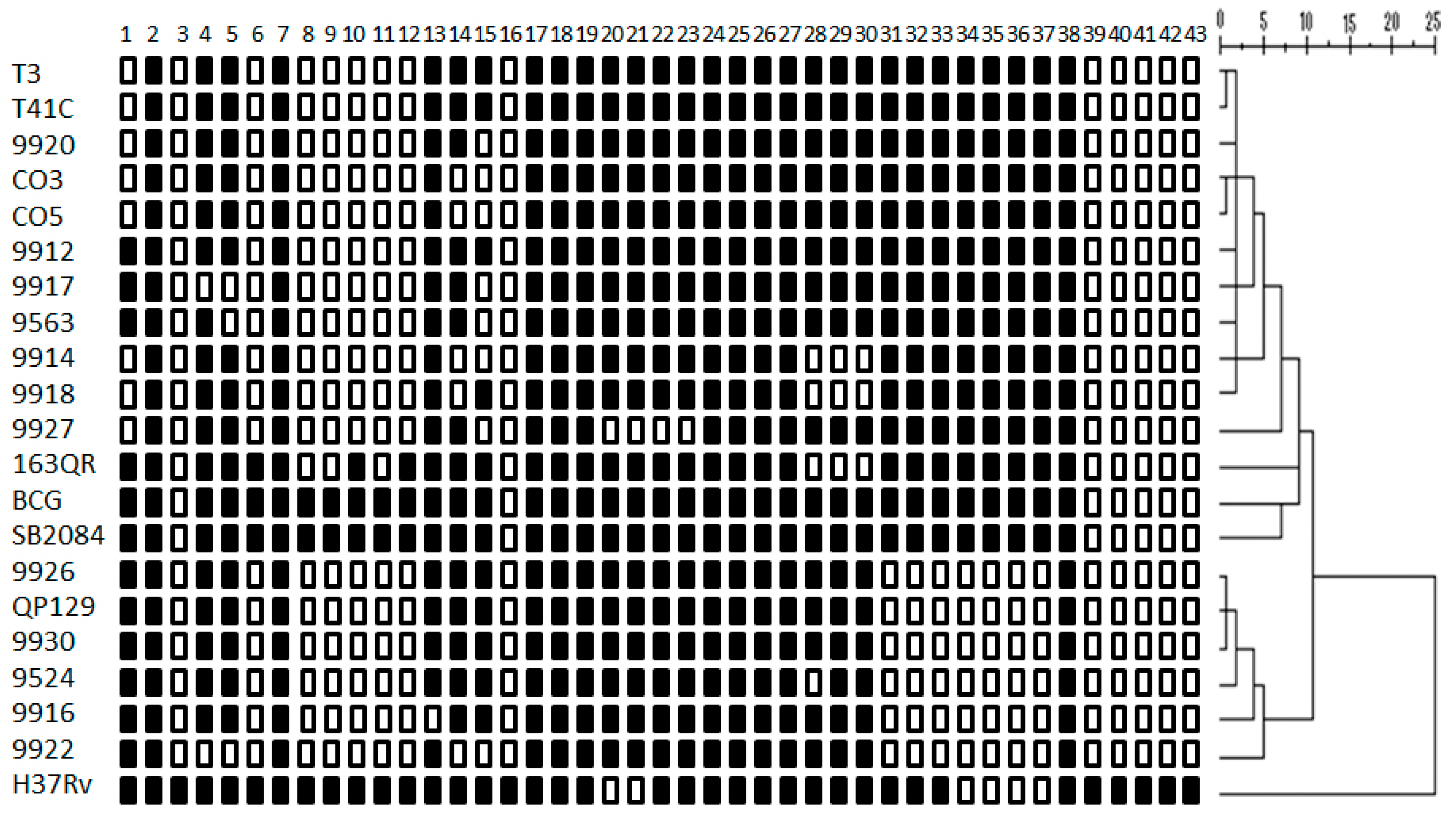
3.1. Spoligotyping Identified Strains Already Present in the Region and Three New Spoligotypes
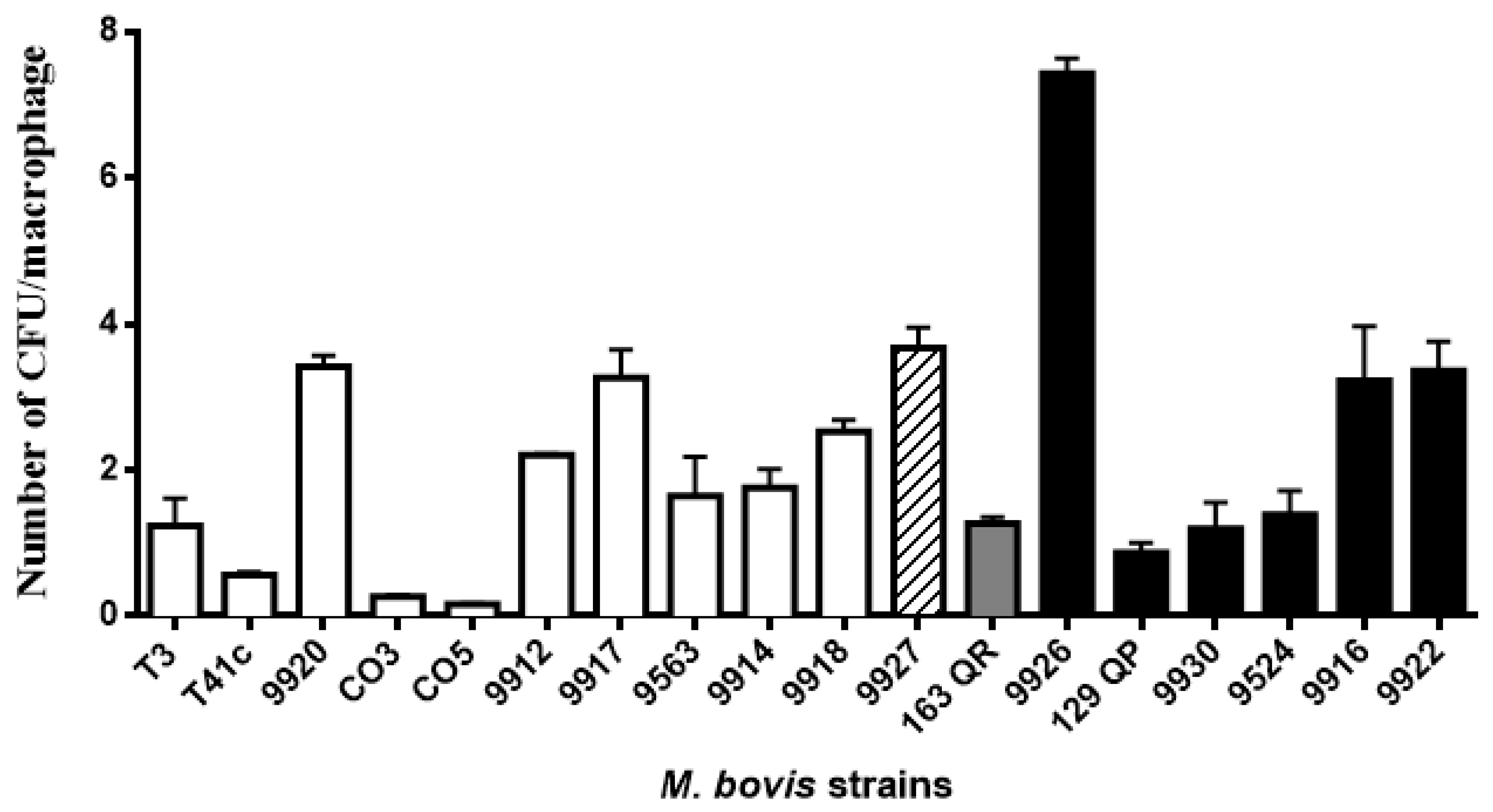
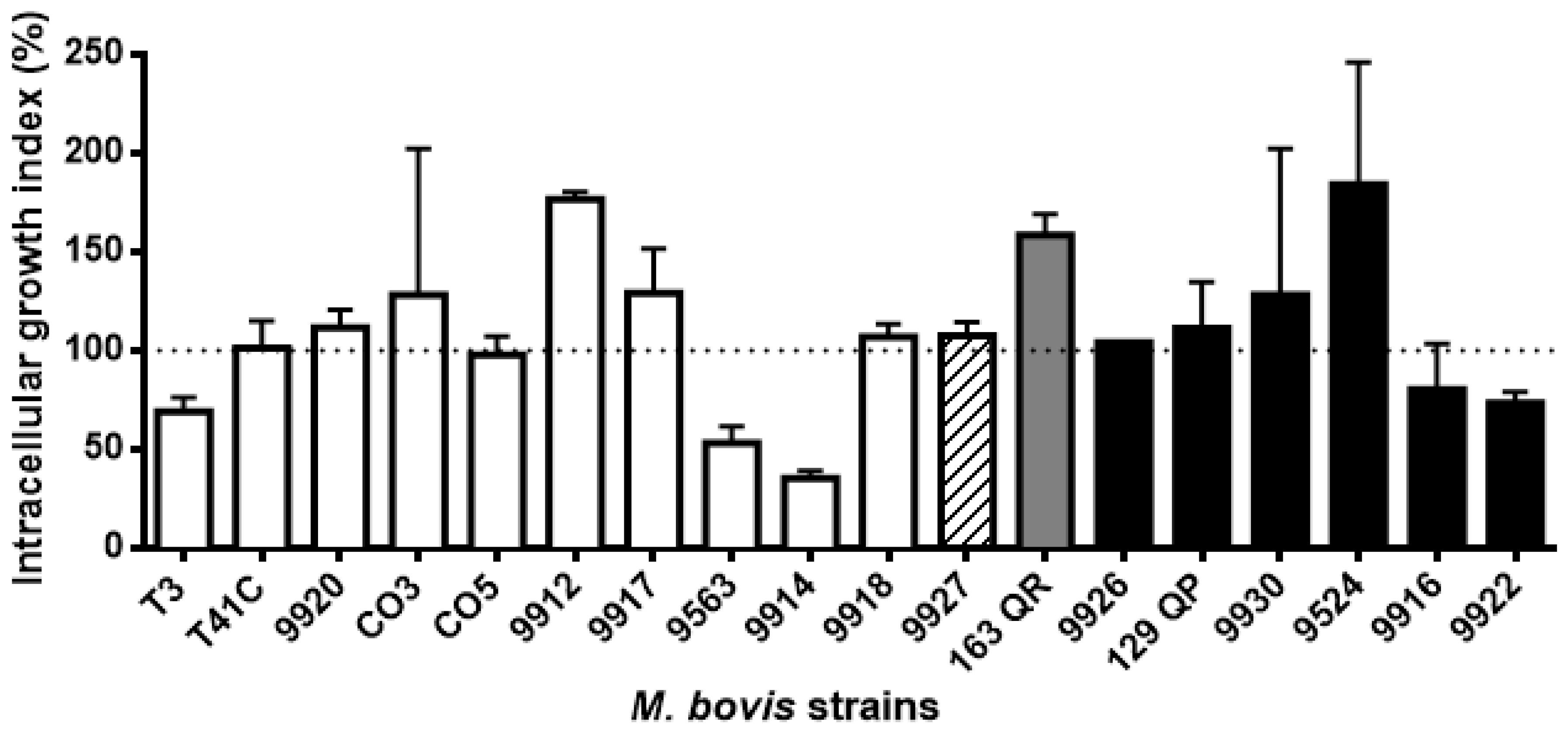
3.2. Mycobacterium Bovis Strains Showed Dissimilar Phagocytosis and Intracellular Survival/Replication Rates
3.3. Spacer Composition Does Not Influence M. bovis Intracellular Survival/Replication in Bovine Macrophages
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- White, P.C.L.; Böhm, M.; Marion, G.; Hutchings, M.R. Control of bovine tuberculosis in British livestock: There is no “silver bullet”. Trends Microbiol. 2008, 16, 420–427. [Google Scholar] [CrossRef] [PubMed]
- Smith, N.H. The global distribution and phylogeography of Mycobacterium bovis clonal complexes. Infect. Genet. Evol. 2012, 12, 857–865. [Google Scholar] [CrossRef] [PubMed]
- Boniotti, M.B.; Goria, M.; Loda, D.; Garrone, A.; Benedetto, A.; Mondo, A.; Tisato, E.; Zanoni, M.; Zoppi, S.; Dondo, A.; et al. Pacciarini, Molecular typing of Mycobacterium bovis strains isolated in Italy from 2000 to 2006 and evaluation of variable-number tandem repeats for geographically optimized genotyping. J. Clin. Microbiol. 2009, 47, 636–644. [Google Scholar] [CrossRef] [PubMed]
- Duarte, E.L.; Domingos, M.; Amado, A.; Cunha, M.V.; Botelho, A. MIRU-VNTR typing adds discriminatory value to groups of Mycobacterium bovis and Mycobacterium caprae strains defined by spoligotyping. Vet. Microbiol. 2010, 143, 299–306. [Google Scholar] [CrossRef]
- Mbovis.org. Mycobacterium bovis Molecular Typing Database. 2002. Available online: http://www.mbovis.org/.
- Caws, M.; Thwaites, G.; Dunstan, S.; Hawn, T.R.; Lan, N.T.N.; Thuong, N.T.T.; Stepniewska, K.; Huyen, M.N.T.; Bang, N.D.; Loc, T.H.; et al. The influence of host and bacterial genotype on the development of disseminated disease with Mycobacterium tuberculosis. PLoS Pathog. 2008, 4, e1000034. [Google Scholar] [CrossRef]
- Zhang, M.; Gong, J.; Yang, Z.; Samten, B.; Cave, M.D.; Barnes, P.F. Enhanced capacity of a widespread strain of Mycobacterium tuberculosis to grow in human macrophages. J. Infect. Dis. 1999, 179, 1213–1217. [Google Scholar] [CrossRef]
- Theus, S.; Eisenach, K.; Fomukong, N.; Silver, R.F.; Cave, M.D. Beijing family Mycobacterium tuberculosis strains differ in their intracellular growth in THP-1 macrophages SUMMARY. Int. J. 2007, 11, 1087–1093. [Google Scholar]
- Qureshi, T.; Templeton, J.W.; Adams, L.G. Intracellular survival of Brucella abortus, Mycobacterium bovis BCG, Salmonella dublin, and Salmonella typhimurium in macrophages from cattle genetically resistant to Brucella abortus. Vet. Immunol. Immunopathol. 1996, 50, 55–65. [Google Scholar] [CrossRef]
- Gutiérrez-pabello, J.A.; Adams, L.G. Survival of Mycobacterium bovis in macrophages from cattle naturally resistant and susceptible to intracellular pathogens. Vet. México 2003, 34, 277–281. [Google Scholar]
- Esquivel-Solís, H.; Vallecillo, A.J.; Benítez-Guzmán, A.; Adams, L.G.; López-Vidal, Y.; Gutiérrez-Pabello, J.A. Nitric oxide not apoptosis mediates differential killing of Mycobacterium bovis in bovine macrophages. PLoS ONE 2013, 8, e63464. [Google Scholar] [CrossRef]
- Castillo-Velázquez, U.; Aranday-Cortés, E.; Gutiérrez-Pabello, J.A. Alternative activation modifies macrophage resistance to Mycobacterium bovis. Vet. Microbiol. 2011, 151, 51–59. [Google Scholar] [CrossRef]
- Harmon, B.G.; Adams, L.G.; Templeton, J.W.; Smith, R. Macrophage function in mammary glands of Brucella abortus-infected cows and cows that resisted infection after inoculation of Brucella abortus. Am. J. Vet. Res. 1989, 50, 459–465. [Google Scholar] [PubMed]
- León, D.A.; Zumárraga, M.J.; Oropeza, R.J.; Gioffré, A.K.; Bernardelli, A.; Estévez, H.O.; Cataldi, A.A.; Pando, R.H. Mycobacterium bovis with different genotypes and from different hosts induce dissimilar immunopathological lesions in a mouse model of tuberculosis. Clin. Exp. Immunol. 2009, 157, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Alfonseca-Silva, E.; Hernández-Pando, R.; Gutiérrez-Pabello, J.A. Mycobacterium bovis-infected macrophages from resistant and susceptible cattle exhibited a differential pro-inflammatory gene expression profile depending on strain virulence. Vet. Immunol. Immunopathol. 2015, 176, 34–43. [Google Scholar] [CrossRef]
- Theus, S.A.; Cave, M.D.; Eisenach, K.; Walrath, J.; Lee, H.; Mackay, W.; Whalen, C.; Silver, R.F. Differences in the growth of paired Ugandan isolates of Mycobacterium tuberculosis within human mononuclear phagocytes correlate with epidemiological evidence of strain virulence. Infect. Immun. 2006, 74, 6865–6876. [Google Scholar] [CrossRef]
- Wong, K.C.; Leong, W.M.; Law, H.K.W.; Ip, K.F.; Lam, J.T.H.; Yuen, K.Y.; Ho, P.L.; Tse, W.S.; Weng, X.H.; Zhang, W.H.; et al. Molecular characterization of clinical isolates of Mycobacterium tuberculosis and their association with phenotypic virulence in human macrophages. Clin. Vaccine Immunol. 2007, 14, 1279–1284. [Google Scholar] [CrossRef]
- Garbaccio, S.; Macias, A.; Shimizu, E.; Paolicchi, F.; Pezzone, N.; Magnano, G.; Zapata, L.; Abdala, A.; Tarabla, H.; Peyru, M.; et al. Association between spoligotype-VNTR types and virulence of Mycobacterium bovis in cattle. Virulence 2014, 5, 297–302. [Google Scholar] [CrossRef]
- Fang, Z.; Doig, C.; Kenna, D.T.; Smittipat, N.; Palittapongarnpim, P.; Watt, B.; Forbes, K.J. IS6110-mediated deletions of wild-type chromosomes of Mycobacterium tuberculosis. J. Bacteriol. 1999, 181, 1014–1020. [Google Scholar]
- Groenen, P.M.; Bunschoten, A.E.; Soolingen, D.V.; Errtbden, J.D.V. Nature of DNA polymorphism in the direct repeat cluster of Mycobacterium tuberculosis; application for strain differentiation by a novel typing method. Mol. Microbiol. 1993, 10, 1057–1065. [Google Scholar] [CrossRef]
- Van Embden, J.D.A.; van Gorkom, T.; Kremer, K.; Jansen, R.; van der Zeijst, B.A.M.; Schouls, L.M. Genetic variation and evolutionary origin of the direct repeat locus of Mycobacterium tuberculosis complex bacteria. J. Bacteriol. 2000, 182, 2393–2401. [Google Scholar] [CrossRef]
- Wright, D.M.; Allen, A.R.; Mallon, T.R.; McDowell, S.W.J.; Bishop, S.C.; Glass, E.J.; Bermingham, M.L.; Woolliams, J.A.; Skuce, R.A. Field-isolated genotypes of Mycobacterium bovis vary in virulence and influence case pathology but do not affect outbreak size. PLoS ONE 2013, 8, e74503. [Google Scholar] [CrossRef] [PubMed]
- Abdelaal, H.F.M.; Spalink, D.; Amer, A.; Steinberg, H.; Hashish, E.A.; Nasr, E.A.; Talaat, A.M. Genomic polymorphism associated with the emergence of virulent isolates of Mycobacterium bovis in the Nile Delta. Sci. Rep. 2019, 9, 1–15. [Google Scholar] [CrossRef] [PubMed]
- JReyes, A.G.; Casanova, L.G.; Torres, C.R.; Gallegos, S.L.S.; Alarcón, G.J.C.; Pezzat, M.M.; Martínez, O.P.; Chávez, C.E.; Suazo, F.M. Population structure of Mycobacterium bovis isolates from cattle in Mexico. Prev. Vet. Med. 2012, 106, 1–8. [Google Scholar] [CrossRef]
- Rodwell, T.C.; Kapasi, A.J.; Moore, M.; Milian-Suazo, F.; Harris, B.; Guerrero, L.P.; Moser, K.; Strathdee, S.A.; Garfein, R.S. Tracing the origins of Mycobacterium bovis tuberculosis in humans in the USA to cattle in Mexico using spoligotyping. Int. J. Infect. Dis. 2010, 14, e129–e135. [Google Scholar] [CrossRef] [PubMed]
- MZumárraga, J.; Arriaga, C.; Barandiaran, S.; Cobos-Marín, L.; de Waard, J.; Estrada-Garcia, I.; Figueiredo, T.; Figueroa, A.; Giménez, F.; Gomes, H.M.; et al. Understanding the relationship between Mycobacterium bovis spoligotypes from cattle in Latin American Countries. Res. Vet. Sci. 2013, 94, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Milian-Suazo, F.; Harris, B.; Díaz, C.A.; Torres, C.R.; Stuber, T.; Ojeda, G.A.; Loredo, A.M.; Soria, M.P.; Payeur, J.B. Molecular epidemiology of Mycobacterium bovis: Usefulness in international trade. Prev. Vet. Med. 2008, 87, 261–271. [Google Scholar] [CrossRef]
- Cobos-Marín, L.; Montes-Vargas, J.; Zumarraga, M.; Cataldi, A.; Romano, M.I.; Estrada-Garcia, I.; Gonzalez-y-Merchand, J.A. Spoligotype analysis of Mycobacterium bovis isolates from Northern México. Can. J. Microbiol. 2005, 51, 996–1000. [Google Scholar] [CrossRef]
- Santillan-Flores, M.A.; Flores, J.; Arriaga-Diaz, C.; Romero-Torres, C.; Suárez-Güemes, F.; Espitia, C. Polymorphism of the PE domain of PE/PE_PGRS sequences in clinical isolates of Mycobacterium bovis in Mexico. Vet. Microbiol. 2006, 115, 364–369. [Google Scholar] [CrossRef]
- Milián-Suazo, F.; Banda-Ruíz, V.; Ramírez-Casillas, C.; Arriaga-Díaz, C. Genotyping of Mycobacterium bovis by geographic location within Mexico. Prev. Vet. Med. 2002, 55, 255–264. [Google Scholar] [CrossRef]

| Strain ID | Spoligotype ID | Place of Isolation | Strain Origin * |
|---|---|---|---|
| T3 | SB0673 | Torreón | United Kingdom |
| T41C | SB0673 | Torreón | United Kingdom |
| 9920 | SB0669 | Jalisco | United Kingdom |
| CO3 | SB1112 | Edo de Mex | Mexico |
| CO5 | SB1112 | Edo de Mex | Mexico |
| 9912 | SB0140 | Nayarit | Argentina/United Kingdom |
| 9917 | SB1816 | Jalisco | Mexico |
| 9563 | SB1498 | Jalisco | Mexico |
| 9914 | SB1811 | Jalisco | Mexico |
| 9918 | SB2351 (Nil) | Jalisco | Mexico |
| 9927 | SB1115 | Jalisco | Mexico |
| 163QR | SB1503 | Querétaro | Mexico |
| BCG | SB0120 | ||
| Control | SB2084 | Mexico | |
| 9926 | SB2352 (Nil) | Jalisco | Mexico |
| QP129 | SB2352 (Nil) | Querétaro | Mexico |
| 9930 | SB2352 (Nil) | Jalisco | Mexico |
| 9524 | SB2353 (Nil) | Edo de Mex | Mexico |
| 9916 | SB1819 | Jalisco | Mexico |
| 9922 | SB1815 | Mexico | Mexico |
| Strain ID | Spoligotype ID | Phagocytosis % | Statistical Group | Intracellular Growth % | Statistical Group |
|---|---|---|---|---|---|
| T3 | SB0673 | 1.24 | 2 | 69.1 | 2 |
| T41c | SB0673 | 0.57 | 1 | 101.3 | 2 |
| 9920 | SB0669 | 3.42 | 4 | 111.6 | 2 |
| CO3 | SB1112 | 0.27 | 1 | 128.2 | 2 |
| CO5 | SB1112 | 0.17 | 1 | 98.1 | 2 |
| 9912 | SB0140 | 2.21 | 3 | 176.5 | 3 |
| 9917 | SB1816 | 3.27 | 4 | 129.3 | 2 |
| 9563 | SB1498 | 1.65 | 2 | 53.2 | 2 |
| 9914 | SB1811 | 1.76 | 2 | 35.6 | 1 |
| 9918 | SB2351 | 2.53 | 3 | 107.0 | 2 |
| 9927 | SB1115 | 3.68 | 4 | 107.7 | 2 |
| 163QR | SB1503 | 1.27 | 2 | 158.6 | 3 |
| 9926 | SB2352 | 7.43 | 5 | 104.0 | 2 |
| 129QP | SB2352 | 0.87 | 1 | 111.4 | 2 |
| 9930 | SB2352 | 1.20 | 2 | 128.2 | 2 |
| 9524 | SB2353 | 1.39 | 2 | 184.0 | 3 |
| 9916 | SB1819 | 3.22 | 4 | 80.6 | 2 |
| 9922 | SB1815 | 3.36 | 4 | 73.7 | 2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Benítez-Guzmán, A.; Esquivel-Solís, H.; Romero-Torres, C.; Arriaga-Díaz, C.; Gutiérrez-Pabello, J.A. Genetically Related Mycobacterium bovis Strains Displayed Differential Intracellular Growth in Bovine Macrophages. Vet. Sci. 2019, 6, 81. https://doi.org/10.3390/vetsci6040081
Benítez-Guzmán A, Esquivel-Solís H, Romero-Torres C, Arriaga-Díaz C, Gutiérrez-Pabello JA. Genetically Related Mycobacterium bovis Strains Displayed Differential Intracellular Growth in Bovine Macrophages. Veterinary Sciences. 2019; 6(4):81. https://doi.org/10.3390/vetsci6040081
Chicago/Turabian StyleBenítez-Guzmán, Alejandro, Hugo Esquivel-Solís, Cecilia Romero-Torres, Camila Arriaga-Díaz, and José A. Gutiérrez-Pabello. 2019. "Genetically Related Mycobacterium bovis Strains Displayed Differential Intracellular Growth in Bovine Macrophages" Veterinary Sciences 6, no. 4: 81. https://doi.org/10.3390/vetsci6040081
APA StyleBenítez-Guzmán, A., Esquivel-Solís, H., Romero-Torres, C., Arriaga-Díaz, C., & Gutiérrez-Pabello, J. A. (2019). Genetically Related Mycobacterium bovis Strains Displayed Differential Intracellular Growth in Bovine Macrophages. Veterinary Sciences, 6(4), 81. https://doi.org/10.3390/vetsci6040081





