Modelling Bovine Granuloma Formation In Vitro upon Infection with Mycobacterium Avium Subspecies Paratuberculosis
Abstract
1. Introduction
2. Materials and Methods
2.1. Blood Sample Collection
2.2. Preparation and Separation of PBMC Culture
2.3. Culturing and Preparation of Map Strains
2.4. Setup and Maintenance of In Vitro Model
2.5. Monitoring Rate of Aggregation
2.6. RNA Isolation and Quantification
2.7. Measuring Viability of Host Cells
2.8. Map Enumeration and Viability
2.9. Host Cell Motility and Infection Rate
3. Results
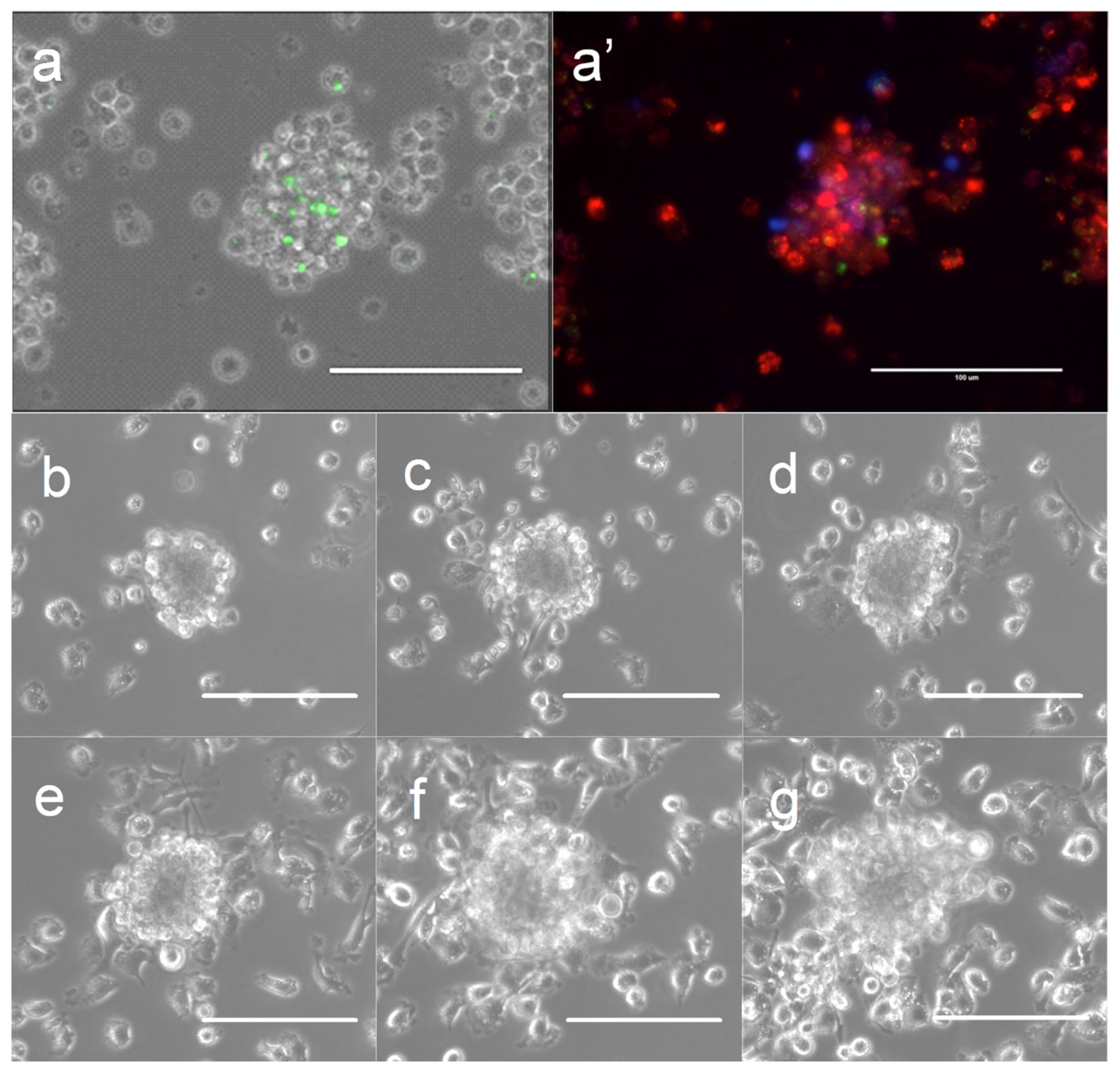
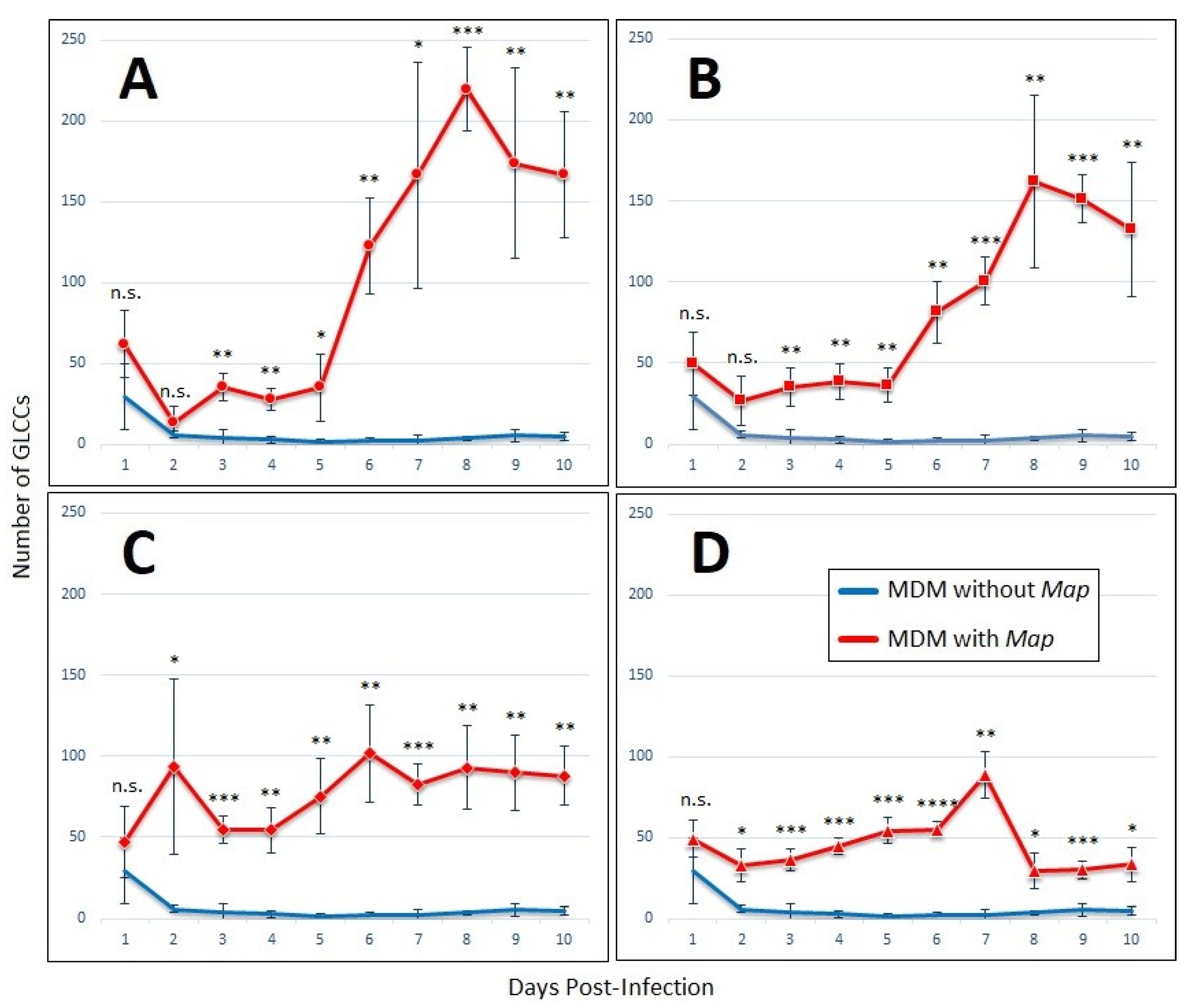
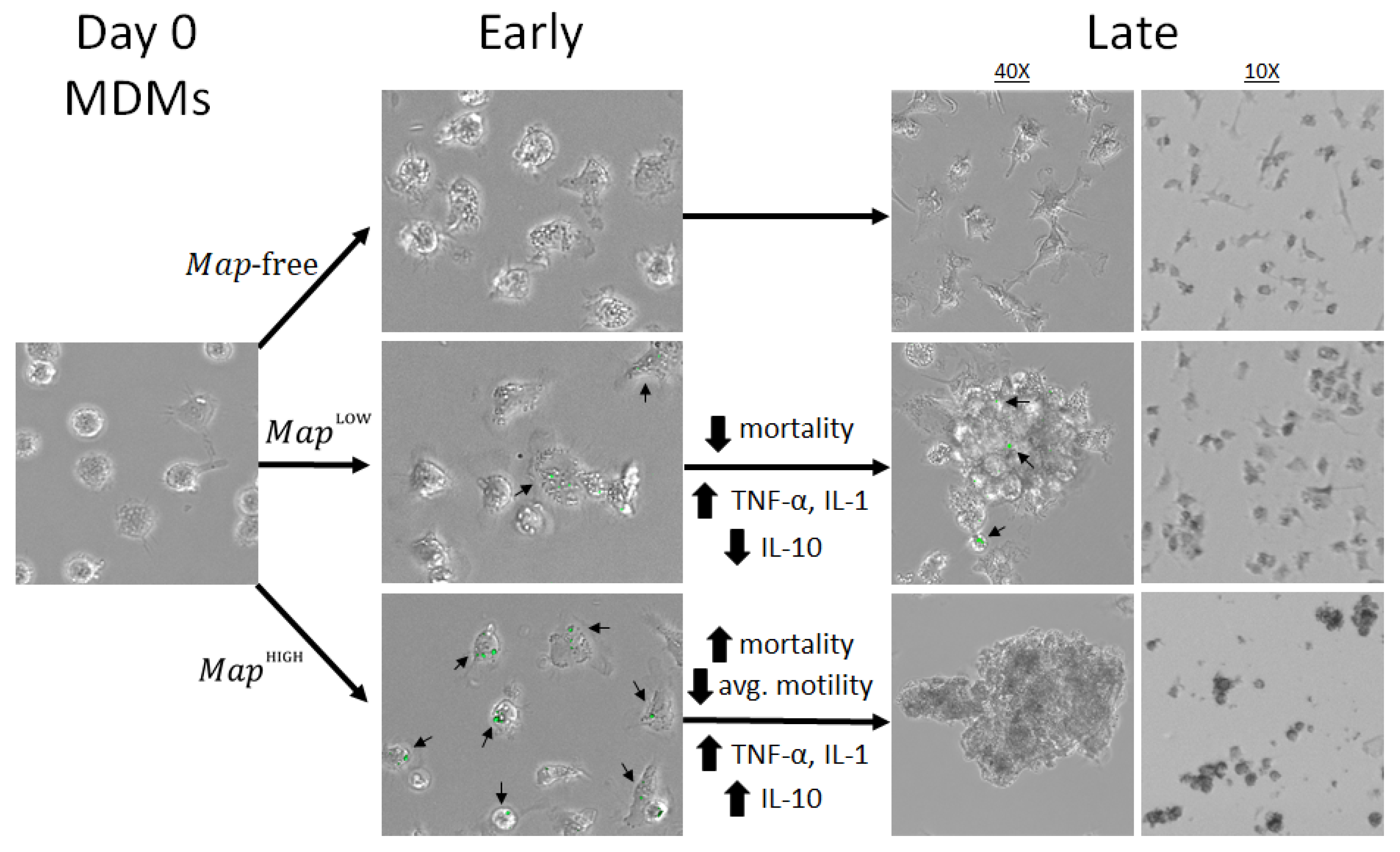
3.1. Host Cell Cluster Formation in the Presence of Lymphocyte-Specific Signaling Factors
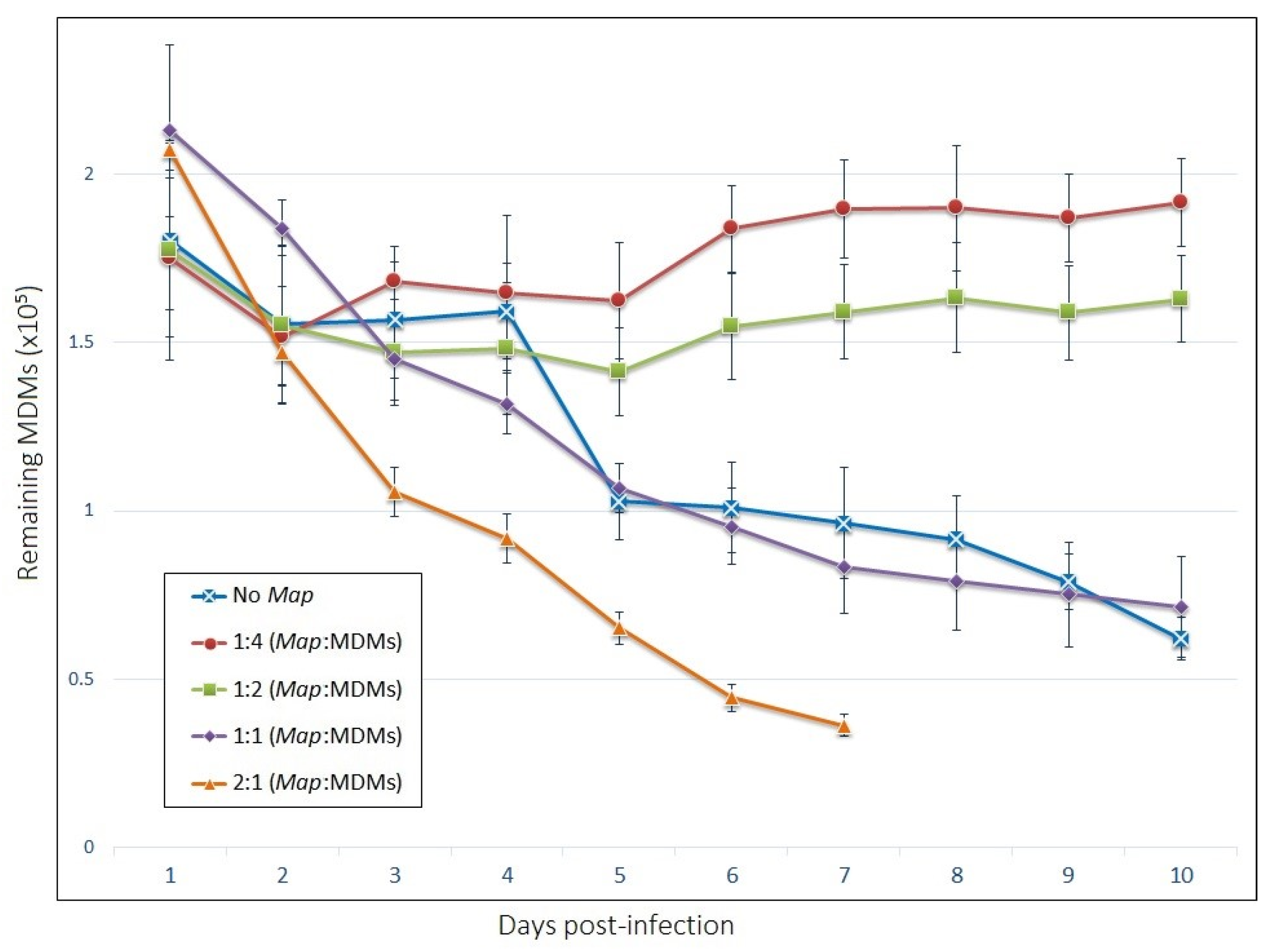
3.2. Viability of Host Mdms and Infection Rate
3.3. Host-Cell Behavior and Growth of Intracellular
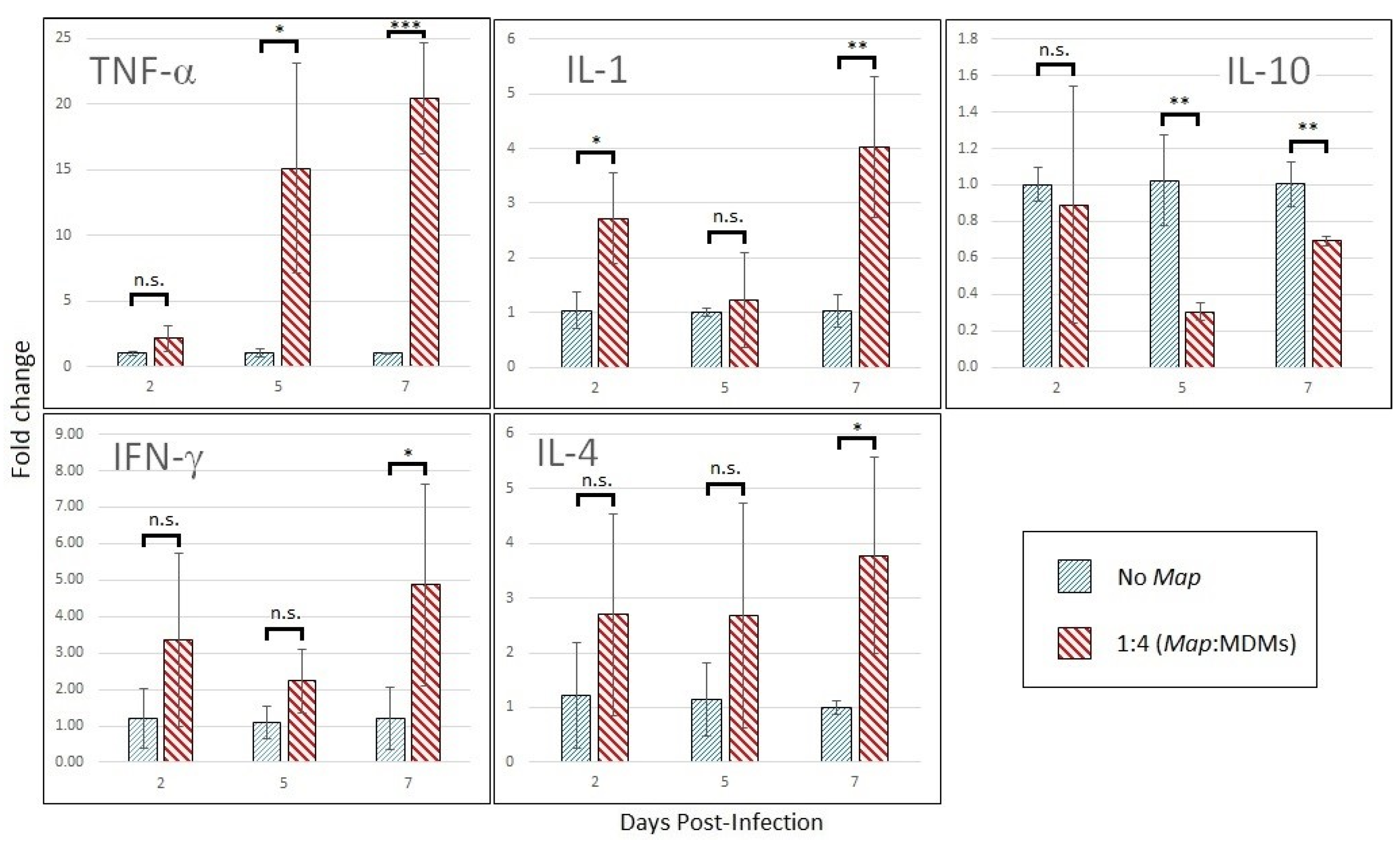
3.3.1. Cytokine Expression Profile Changes in Infected Mdms
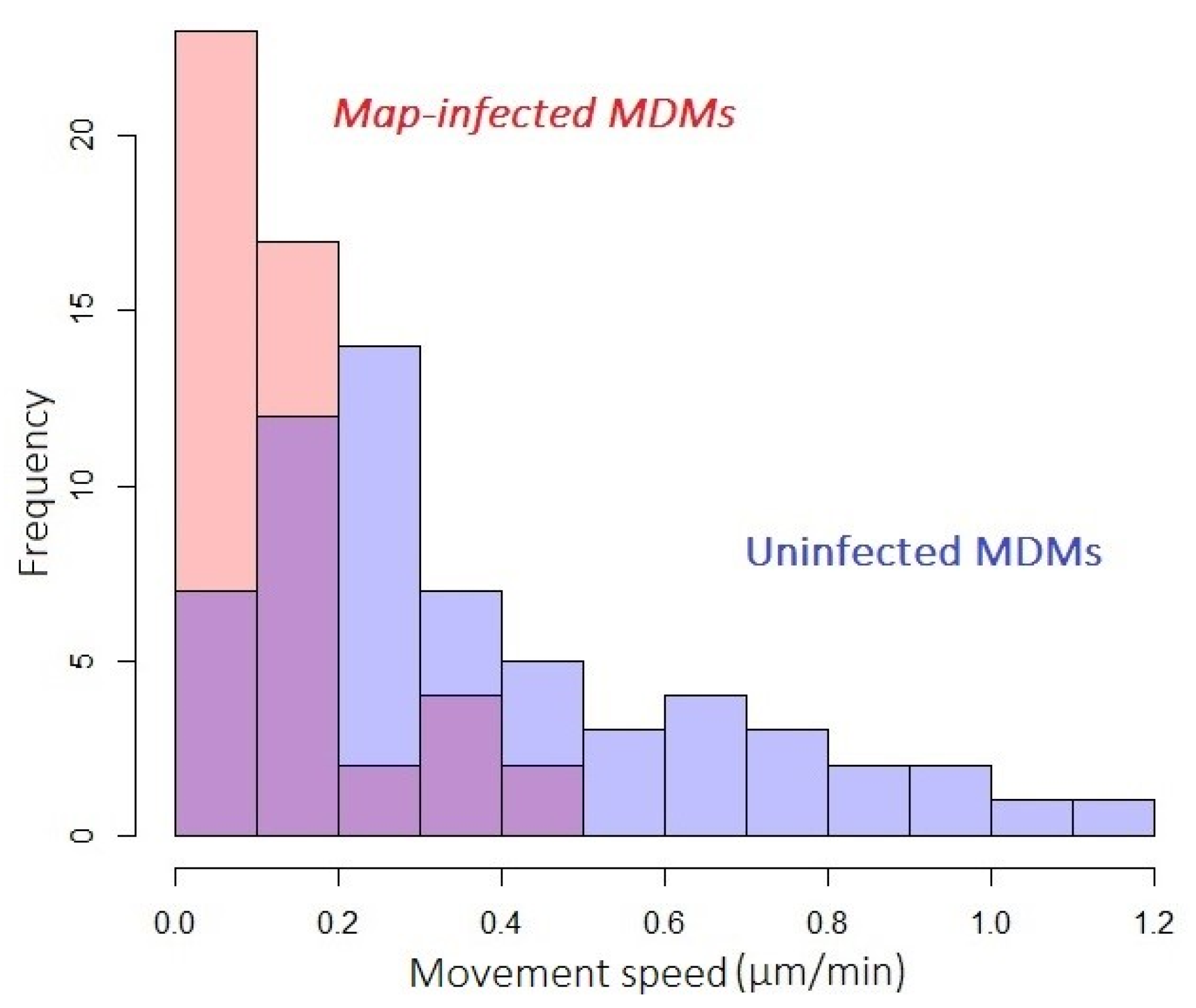
3.3.2. Host-Cell Motility
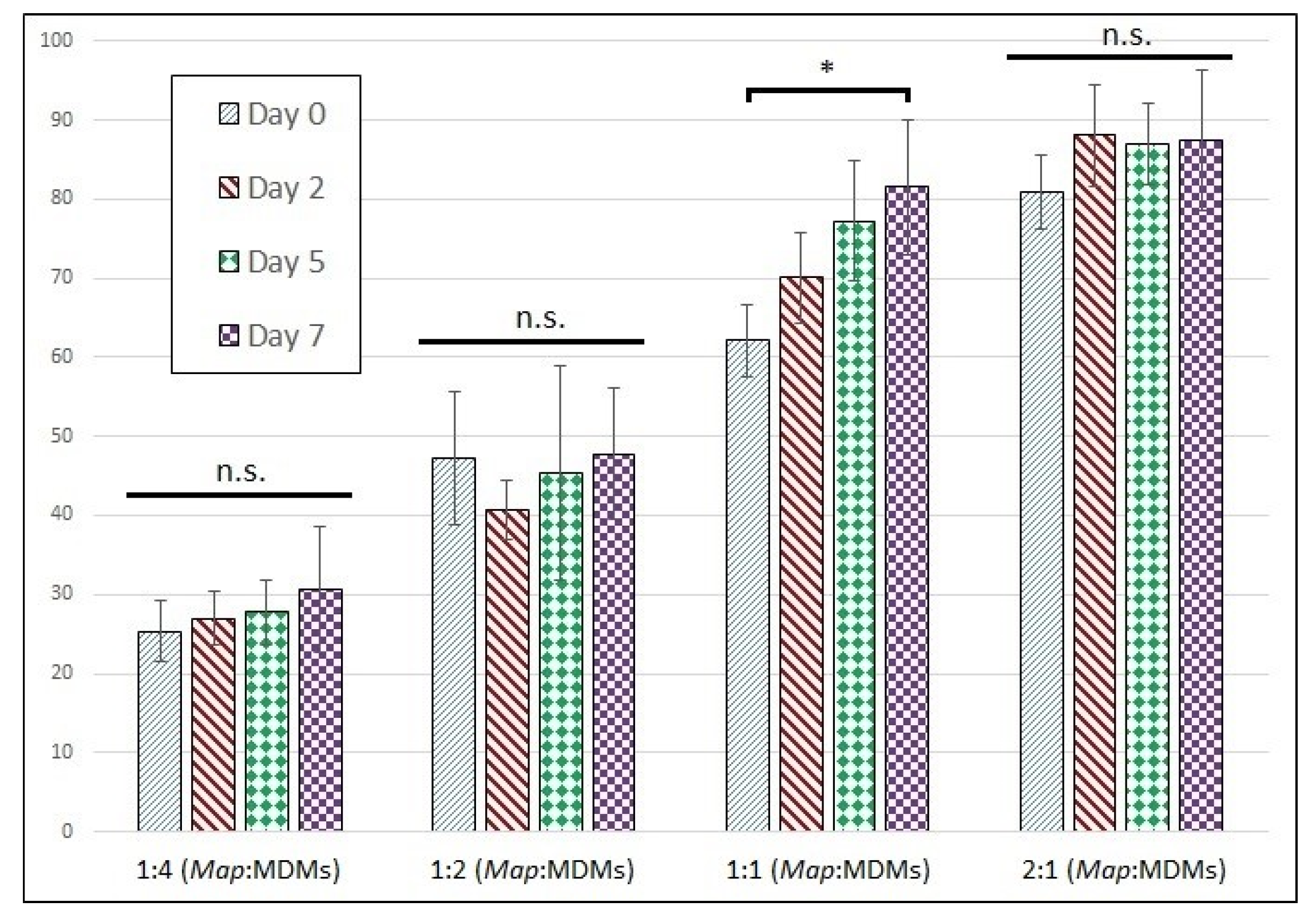
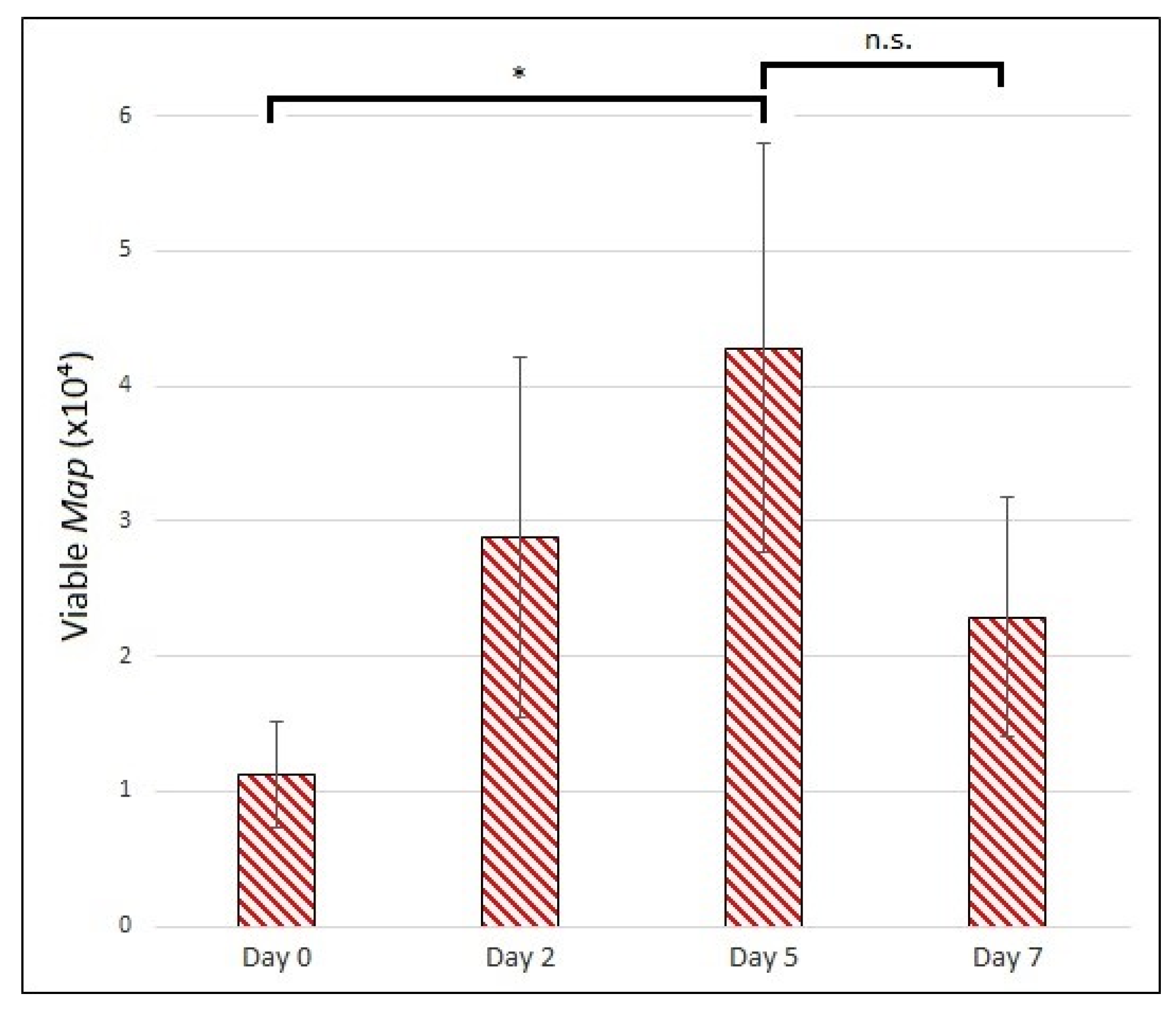
3.3.3. Map Viability
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Harris, N.B.; Barletta, R.G. Mycobacterium Avium Subsp. Paratuberculosis in Veterinary Medicine. Clin. Microbiol. Rev. 2001, 14, 489–512. [Google Scholar] [CrossRef] [PubMed]
- Ghazaei, C. Mycobacterium Tuberculosis and Lipids: Insights into Molecular Mechanisms from Persistence to Virulence. J. Res. Med. Sci. 2018, 23, 63. [Google Scholar] [CrossRef] [PubMed]
- Cook, G.M.; Berney, M.; Gebhard, S.; Heinemann, M.; Cox, R.A.; Danilchanka, O.; Niederweis, M. Physiology of Mycobacteria. Adv. Microb. Physiol. 2009, 55, 81–182, 318–319. [Google Scholar] [CrossRef] [PubMed]
- Whittington, R.J.; Begg, D.J.; De Silva, K.; Plain, K.M.; Purdie, A.C. Comparative Immunological and Microbiological Aspects of Paratuberculosis as a Model Mycobacterial Infection. Vet. Immunol. Immunopathol. 2012, 148, 29–47. [Google Scholar] [CrossRef]
- Kennedy, D.J.; Benedictus, G. Control of Mycobacterium Avium Subsp. Paratuberculosis Infection in Agricultural Species. Rev. Sci. Tech. 2001, 20, 151–179. [Google Scholar] [CrossRef]
- Linnabary, R.D.; Meerdink, G.L.; Collins, M.T.; Stabel, J.R.; Sweeney, R.W.; Washington, M.K.; Wells, S.J. Johne’s Disease in Cattle; Council for Agricultural Science and Technology: Ames, IA, USA, 2001; pp. 1–10. [Google Scholar]
- Lombard, J.E.; Gardner, I.A.; Jafarzadeh, S.R.; Fossler, C.P.; Harris, B.; Capsel, R.T.; Wagner, B.A.; Johnson, W.O. Herd-level prevalence of Mycobacterium avium subsp. paratuberculosis infection in United States dairy herds in 2007. Prev. Vet. Med. 2012, 108, 234–238. [Google Scholar] [CrossRef]
- Ott, S.L.; Wells, S.J.; Wagner, B.A. Herd-level economic losses associated with Johne’s disease on US dairy operations. Prev. Vet. Med. 1999, 40, 179–192. [Google Scholar] [CrossRef]
- Stabel, J.R. Johne’s Disease: A Hidden Threat. J. Dairy Sci. 1998, 81, 283–288. [Google Scholar] [CrossRef]
- Collins, M.T.; Gardner, I.A.; Garry, F.B.; Roussel, A.J.; Wells, S.J. Consensus Recommendations on Diagnostic Testing for the Detection of Paratuberculosis in Cattle in the United States. J. Am. Vet. Med. Assoc. 2006, 229, 1912–1919. [Google Scholar] [CrossRef]
- Sweeney, R.W.; Collins, M.T.; Koets, A.P.; McGuirk, S.M.; Roussel, A.J. Paratuberculosis (Johne’s Disease) in Cattle and Other Susceptible Species. J. Vet. Intern. Med. 2012, 26, 1239–1250. [Google Scholar] [CrossRef]
- Verteramo Chiu, L.J.; Tauer, L.W.; Al-Mamun, M.A.; Kaniyamattam, K.; Smith, R.L.; Grohn, Y.T. An Agent-Based Model Evaluation of Economic Control Strategies for Paratuberculosis in a Dairy Herd. J. Dairy Sci. 2018, 101, 6443–6454. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.L.; Al-Mamun, M.A.; Grohn, Y.T. Economic Consequences of Paratuberculosis Control in Dairy Cattle: A Stochastic Modeling Study. Prev. Vet. Med. 2017, 138, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Gardner, I.A.; Nielsen, S.S.; Whittington, R.J.; Collins, M.T.; Bakker, D.; Harris, B.; Sreevatsan, S.; Lombard, J.E.; Sweeney, R.; Smith, D.R.; et al. Consensus-Based Reporting Standards for Diagnostic Test Accuracy Studies for Paratuberculosis in Ruminants. Prev. Vet. Med. 2011, 101, 18–34. [Google Scholar] [CrossRef] [PubMed]
- Patton, E.A. Paratuberculosis Vaccination. Vet. Clin. N. Am. Food Anim. Pract. 2011, 27, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Bastida, F.; Juste, R.A. Paratuberculosis Control: A Review with a Focus on Vaccination. J. Immune Based Ther. Vaccines 2011, 9, 8. [Google Scholar] [CrossRef]
- Park, H.T.; Yoo, H.S. Development of Vaccines to Mycobacterium Avium Subsp. Paratuberculosis Infection. Clin. Exp. Vaccine Res. 2016, 5, 108–116. [Google Scholar] [CrossRef]
- Vrieling, M.; Santema, W.; Vordermeier, M.; Rutten, V.; Koets, A. Hsp70 Vaccination-Induced Primary Immune Responses in Efferent Lymph of the Draining Lymph Node. Vaccine 2013, 31, 4720–4727. [Google Scholar] [CrossRef]
- Sigurethardottir, O.G.; Valheim, M.; Press, C.M. Establishment of Mycobacterium Avium Subsp. Paratuberculosis Infection in the Intestine of Ruminants. Adv. Drug Deliv. Rev. 2004, 56, 819–834. [Google Scholar] [CrossRef]
- Sigur-Dardottir, O.G.; Press, C.M.; Evensen, O. Uptake of Mycobacterium Avium Subsp. Paratuberculosis Through the Distal Small Intestinal Mucosa in Goats: An Ultrastructural Study. Vet. Pathol. 2001, 38, 184–189. [Google Scholar] [CrossRef]
- Momotani, E.; Whipple, D.L.; Thiermann, A.B.; Cheville, N.F. Role of M Cells and Macrophages in the Entrance of Mycobacterium Paratuberculosis into Domes of Ileal Peyer’s Patches in Calves. Vet. Pathol. 1988, 25, 131–137. [Google Scholar] [CrossRef]
- Plattner, B.L.; Doyle, R.T.; Hostetter, J.M. Gamma-Delta T Cell Subsets are Differentially Associated with Granuloma Development and Organization in a Bovine Model of Mycobacterial Disease. Int. J. Exp. Pathol. 2009, 90, 587–597. [Google Scholar] [CrossRef] [PubMed]
- Plattner, B.L.; Huffman, E.L.; Hostetter, J.M. Gamma-Delta T-Cell Responses During Subcutaneous Mycobacterium Avium Subspecies Paratuberculosis Challenge in Sensitized or Naive Calves Using Matrix Biopolymers. Vet. Pathol. 2013, 50, 630–637. [Google Scholar] [CrossRef] [PubMed]
- Begg, D.J.; De Silva, K.; Carter, N.; Plain, K.M.; Purdie, A.; Whittington, R.J. Does a Th1 Over Th2 Dominancy Really Exist in the Early Stages of Mycobacterium Avium Subspecies Paratuberculosis Infections? Immunobiology 2011, 216, 840–846. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Hearn, M.; Magombedze, G.; Abendano, N.; Landin, M.; Juste, R.A. Deciphering the Virulence of Mycobacterium Avium Subsp. Paratuberculosis Isolates in Animal Macrophages Using Mathematical Models. J. Theor. Biol. 2019, 468, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Guirado, E.; Schlesinger, L.S. Modeling the Mycobacterium Tuberculosis Granuloma-The Critical Battlefield in Host Immunity and Disease. Front. Immunol. 2013, 4, 98. [Google Scholar] [CrossRef] [PubMed]
- Lenaerts, A.; Barry, C.E.; Dartois, V. Heterogeneity in Tuberculosis Pathology, Microenvironments and Therapeutic Responses. Immunol. Rev. 2015, 264, 288–307. [Google Scholar] [CrossRef] [PubMed]
- Seitzer, U.; Gerdes, J. Generation and Characterization of Multicellular Heterospheroids Formed by Human Peripheral Blood Mononuclear Cells. Cells Tissues Organs 2003, 174, 110–116. [Google Scholar] [CrossRef]
- Birkness, K.A.; Guarner, J.; Sable, S.B.; Tripp, R.A.; Kellar, K.L.; Bartlett, J.; Quinn, F.D. An In Vitro Model of the Leukocyte Interactions Associated with Granuloma Formation in Mycobacterium Tuberculosis Infection. Immunl. Cell Biol. 2007, 85, 160–168. [Google Scholar] [CrossRef]
- Puissegur, M.P.; Botanch, C.; Duteyrat, J.L.; Delsol, G.; Caratero, C.; Altare, F. An In Vitro Dual Model of Mycobacterial Granulomas to Investigate the Molecular Interactions between Mycobacteria and Human Host Cells. Cell Microbiol. 2004, 6, 423–433. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 Years of Image Analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Kralik, P.; Nocker, A.; Pavlik, I. Mycobacterium Avium Subsp. Paratuberculosis Viability Determination Using F57 Quantitative PCR in Combination with Propidium Monoazide Treatment. Int. J. Food Microbiol. 2010, 141, S80–S86. [Google Scholar] [CrossRef] [PubMed]
- Kralik, P.; Babak, V.; Dziedzinska, R. Repeated Cycles of Chemical and Physical Disinfection and Their Influence on Mycobacterium Avium Subsp. Paratuberculosis Viability Measured by Propidium Monoazide F57 Quantitative real Time PCR. Vet. J. 2014, 201, 359–364. [Google Scholar] [CrossRef] [PubMed]
- Ricchi, M.; De Cicco, C.; Kralik, P.; Babak, V.; Boniotti, M.B.; Savi, R.; Cerutti, G.; Cammi, G.; Garbarino, C.; Arrigoni, N. Evaluation of Viable Mycobacterium Avium Subsp. Paratuberculosis in Milk Using Peptide-Mediated Separation and Propidium Monoazide qPCR. FEMS Microbiol. Lett. 2014, 356, 127–133. [Google Scholar] [CrossRef][Green Version]
- Odumeru, J.; Gao, A.; Chen, S.; Raymond, M.; Mutharia, L. Use of the Bead Beater for Preparation of Mycobacterium Paratuberculosis Template DNA in Milk. Can. J. Vet. Res. 2001, 65, 201–205. [Google Scholar]
- Nussbaum-Krammer, C.I.; Neto, M.F.; Brielmann, R.M.; Pedersen, J.S.; Morimoto, R.I. Investigating the Spreading and Toxicity of Prion-Like Proteins Using the Metazoan Model Organism C. Elegans. J. Vis. Exp. 2015. [Google Scholar] [CrossRef]
- Abendano, N.; Juste, R.A.; Alonso-Hearn, M. Anti-Inflammatory and Antiapoptotic Responses to Infection: A Common Denominator of Human and Bovine Macrophages Infected with Mycobacterium Avium Subsp. Paratuberculosis. Biomed. Res. Int. 2013, 2013, 908348. [Google Scholar] [CrossRef]
- Kabara, E.; Kloss, C.C.; Wilson, M.; Tempelman, R.J.; Sreevatsan, S.; Janagama, H.; Coussens, P.M. A large-Scale Study of Differential Gene Expression in Monocyte-Derived Macrophages Infected with Several Strains of Mycobacterium avium Subspecies Paratuberculosis. Brief. Funct. Genom. 2010, 9, 220–237. [Google Scholar] [CrossRef][Green Version]
- Machugh, D.E.; Taraktsoglou, M.; Killick, K.E.; Nalpas, N.C.; Browne, J.A.; DE Park, S.; Hokamp, K.; Gormley, E.; Magee, D.A. Pan-Genomic Analysis of Bovine Monocyte-Derived Macrophage Gene Expression in Response to In Vitro Infection with Mycobacterium Avium Subspecies Paratuberculosis. Vet. Res. 2012, 43, 25. [Google Scholar] [CrossRef]
- Abendano, N.; Sevilla, I.A.; Prieto, J.M.; Garrido, J.M.; Juste, R.A.; Alonso-Hearn, M. Mycobacterium Avium Subspecies Paratuberculosis Isolates from Sheep and Goats Show Reduced Persistence in Bovine Macrophages than Cattle, Bison, Deer and Wild Boar Strains Regardless of Genotype. Vet. Microbiol. 2013, 163, 325–334. [Google Scholar] [CrossRef]
- Weiss, D.J.; Evanson, O.A.; Moritz, A.; Deng, M.Q.; Abrahamsen, M.S. Differential Responses of Bovine Macrophages to Mycobacterium Avium Subsp. Paratuberculosis and Mycobacterium Avium Subsp. Avium. Infect. Immun. 2002, 70, 5556–5561. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Janagama, H.K.; Jeong, K.; Kapur, V.; Coussens, P.; Sreevatsan, S. Cytokine Responses of Bovine Macrophages to Diverse Clinical Mycobacterium Avium Subspecies Paratuberculosis Strains. BMC Microbiol. 2006, 6, 10. [Google Scholar] [CrossRef] [PubMed]
- Abendano, N.; Tyukalova, L.; Barandika, J.F.; Balseiro, A.; Sevilla, I.A.; Garrido, J.M.; Juste, R.A.; Alonso-Hearn, M. Mycobacterium Avium Subsp. Paratuberculosis Isolates Induce In Vitro Granuloma Formation and Show Successful Survival Phenotype, Common Anti-Inflammatory and Antiapoptotic Responses within Ovine Macrophages Regardless of Genotype or Host of Origin. PLoS ONE 2014, 9, e104238. [Google Scholar] [CrossRef]
- Weiss, D.J.; Evanson, O.A.; Deng, M.; Abrahamsen, M.S. Gene Expression and Antimicrobial Activity of Bovine Macrophages in Response to Mycobacterium Avium Subsp. Paratuberculosis. Vet. Pathol. 2004, 41, 326–337. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.T.; Sommer, S.; Kabara, E.A.; Verman, N.; Kuelbs, M.A.; Saama, P.; Halgren, R.; Coussens, P.M. Gene Expression Profiling of Monocyte-Derived Macrophages Following Infection with Mycobacterium Avium Subspecies Avium and Mycobacterium Avium Subspecies Paratuberculosis. Physiol. Genom. 2006, 28, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Periasamy, S.; Tripathi, B.N.; Singh, N. Mechanisms of Mycobacterium Avium Subsp. Paratuberculosis Induced Apoptosis and Necrosis in Bovine Macrophages. Vet. Microbiol. 2013, 165, 392–401. [Google Scholar] [CrossRef] [PubMed]
- Le Gros, G.; Ben-Sasson, S.Z.; Seder, R.; Finkelman, F.D.; Paul, W.E. Generation of Interleukin 4 (IL-4)-Producing Cells In Vivo and In Vitro: IL-2 and IL-4 are Required for In Vitro Generation of IL-4-Producing Cells. J. Exp. Med. 1990, 172, 921–929. [Google Scholar] [CrossRef]
- Choi, P.; Reiser, H. IL-4: Role in Disease and Regulation of Production. Clin. Exp. Immunol. 1998, 113, 317–319. [Google Scholar] [CrossRef]
- Raphael, I.; Nalawade, S.; Eagar, T.N.; Forsthuber, T.G. T cell Subsets and Their Signature Cytokines in Autoimmune and Inflammatory Diseases. Cytokine 2015, 74, 5–17. [Google Scholar] [CrossRef]
- Cose, S. T-cell Migration: A Naive Paradigm? Immunology 2007, 120, 1–7. [Google Scholar] [CrossRef]
- Karlsson, M.; Linton, L.; Lampinen, M.; Karlen, P.; Glise, H.; Befrits, R.; Janczewska, I.; Carlson, M.; Winqvist, O.; Eberhardson, M. Naive T cells Correlate with Mucosal Healing in Patients with Inflammatory Bowel Disease. Scand. J. Gastroenterol. 2014, 49, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Carragher, D.M.; Rangel-Moreno, J.; Randall, T.D. Ectopic Lymphoid Tissues and Local Immunity. Semin. Immunol. 2008, 20, 26–42. [Google Scholar] [CrossRef] [PubMed]
- Weninger, W.; Carlsen, H.S.; Goodarzi, M.; Moazed, F.; Crowley, M.A.; Baekkevold, E.S.; Cavanagh, L.L.; Von Andrian, U.H. Naive T cell Recruitment to Nonlymphoid Tissues: A Role for Endothelium-Expressed CC Chemokine Ligand 21 in Autoimmune Disease and Lymphoid Neogenesis. J. Immunol. 2003, 170, 4638–4648. [Google Scholar] [CrossRef] [PubMed]
- Pozzi, L.A.; Maciaszek, J.W.; Rock, K.L. Both deNdritic Cells and Macrophages Can Stimulate Naive CD8 T Cells In Vivo to Proliferate, Develop Effector Function, and Differentiate into Memory Cells. J. Immunol. 2005, 175, 2071–2081. [Google Scholar] [CrossRef] [PubMed]
- Hume, D.A. Macrophages as APC and the Dendritic Cell Myth. J. Immunol. 2008, 181, 5829–5835. [Google Scholar] [CrossRef] [PubMed]
- Wilson, R.A.; Zolnai, A.; Rudas, P.; Frenyo, L.V. T-Cell Subsets in Blood and Lymphoid Tissues Obtained from Fetal Calves, Maturing Calves, and Adult Bovine. Vet. Immunol. Immunopathol. 1996, 53, 49–60. [Google Scholar] [CrossRef]
- Fournie, J.J.; Bonneville, M. Stimulation of Gamma Delta T Cells by Phosphoantigens. Res. Immunol. 1996, 147, 338–347. [Google Scholar] [CrossRef]
- Paul, S.; Lal, G. Regulatory and Effector Functions of Gamma-Delta (Gammadelta) T Cells and Their Therapeutic Potential in Adoptive Cellular Therapy for Cancer. Int. J. Cancer 2016, 139, 976–985. [Google Scholar] [CrossRef]
- Albarrak, S.M.; Waters, W.R.; Stabel, J.R.; Hostetter, J.M. WC1(+) Gammadelta T Cells from Cattle Naturally Infected with Mycobacterium Avium Subsp. Paratuberculosis Respond Differentially to Stimulation with PPD-J. Vet. Immunol. Immunopathol. 2017, 190, 57–64. [Google Scholar] [CrossRef]
- Guzman, E.; Hope, J.; Taylor, G.; Smith, A.L.; Cubillos-Zapata, C.; Charleston, B. Bovine Gammadelta T Cells are a Major Regulatory T Cell Subset. J. Immunol. 2014, 193, 208–222. [Google Scholar] [CrossRef]
- Silva, D.; Silva, M.V.D.; Barros, C.C.O.; Alexandre, P.B.D.; Timoteo, R.P.; Catarino, J.S.; Sales-Campos, H.; Machado, J.R.; Rodrigues, D.B.R.; Oliveira, C.J.; et al. TNF-Alpha Blockade Impairs In Vitro Tuberculous Granuloma Formation and Down Modulate Th1, Th17 and Treg Cytokines. PLoS ONE 2018, 13, e0194430. [Google Scholar] [CrossRef]
- Gerber, D.J.; Azuara, V.; Levraud, J.P.; Huang, S.Y.; Lembezat, M.P.; Pereira, P. IL-4-Producing Gamma Delta T Cells that Express a Very Restricted TCR Repertoire are Preferentially Localized in Liver and Spleen. J. Immunol. 1999, 163, 3076–3082. [Google Scholar] [PubMed]
- Magombedze, G.; Eda, S.; Stabel, J. Predicting the Role of IL-10 in the Regulation of the Adaptive Immune Responses in Mycobacterium Avium Subsp. Paratuberculosis Infections Using Mathematical Models. PLoS ONE 2015, 10, e0141539. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Singh, V.K.; Hunter, R.L.; Jagannath, C. Macrophage Heterogeneity and Plasticity in Tuberculosis. J. Leukoc. Biol. 2019. [Google Scholar] [CrossRef]
- Fernandez, M.; Benavides, J.; Castano, P.; Elguezabal, N.; Fuertes, M.; Munoz, M.; Royo, M.; Ferreras, M.C.; Perez, V. Macrophage Subsets Within Granulomatous Intestinal Lesions in Bovine Paratuberculosis. Vet. Pathol. 2017, 54, 82–93. [Google Scholar] [CrossRef]
- Coussens, P.M.; Colvin, C.J.; Rosa, G.J.; Perez Laspiur, J.; Elftman, M.D. Evidence for a Novel Gene Expression Program in Peripheral Blood Mononuclear Cells from Mycobacterium Avium Subsp. Paratuberculosis-Infected Cattle. Infect. Immun. 2003, 71, 6487–6498. [Google Scholar] [CrossRef]
- Puissegur, M.P.; Lay, G.; Gilleron, M.; Botella, L.; Nigou, J.; Marrakchi, H.; Mari, B.; Duteyrat, J.L.; Guerardel, Y.; Kremer, L.; et al. Mycobacterial Lipomannan Induces Granuloma Macrophage Fusion Via a TLR2-Dependent, ADAM9-And Beta1 Integrin-Mediated Pathway. J. Immunol. 2007, 178, 3161–3169. [Google Scholar] [CrossRef]
- Kapoor, N.; Pawar, S.; Sirakova, T.D.; Deb, C.; Warren, W.L.; Kolattukudy, P.E. Human Granuloma In Vitro Model, for TB Dormancy and Resuscitation. PLoS ONE 2013, 8, e53657. [Google Scholar] [CrossRef]
- Parasa, V.R.; Rahman, M.J.; Ngyuen Hoang, A.T.; Svensson, M.; Brighenti, S.; Lerm, M. Modeling Mycobacterium Tuberculosis early Granuloma Formation in Experimental Human Lung Tissue. Dis. Model. Mech. 2014, 7, 281–288. [Google Scholar] [CrossRef]
- Huang, Z.; Luo, Q.; Guo, Y.; Chen, J.; Xiong, G.; Peng, Y.; Ye, J.; Li, J. Mycobacterium Tuberculosis-Induced Polarization of Human Macrophage Orchestrates the Formation and Development of Tuberculous Granulomas In Vitro. PLoS ONE 2015, 10, e0129744. [Google Scholar] [CrossRef]
- Guirado, E.; Mbawuike, U.; Keiser, T.L.; Arcos, J.; Azad, A.K.; Wang, S.H.; Schlesinger, L.S. Characterization of Host and Microbial Determinants in Individuals with Latent Tuberculosis Infection Using a Human Granuloma Model. mBio 2015, 6, e02537-14. [Google Scholar] [CrossRef] [PubMed]
- Reyes, N.; Bettin, A.; Reyes, I.; Geliebter, J. Microarray Analysis of the In Vitro Granulomatous Response to Mycobacterium Tuberculosis H37Ra. Colomb. Med. 2015, 46, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, N.; Bhattacharyya, C.; Mukherjee, A.; Ullah, U.; Pandit, B.; Rao, K.V.S.; Majumder, P.P. Dissecting Host Factors that Regulate the Early Stages of Tuberculosis Infection. Tuberculosis 2016, 100, 102–113. [Google Scholar] [CrossRef] [PubMed]
- Tezera, L.B.; Bielecka, M.K.; Chancellor, A.; Reichmann, M.T.; Shammari, B.A.; Brace, P.; Batty, A.; Tocheva, A.; Jogai, S.; Marshall, B.G.; et al. Dissection of the Host-Pathogen Interaction in Human Tuberculosis Using a Bioengineered 3-Dimensional Model. eLife 2017, 6. [Google Scholar] [CrossRef]
- Agrawal, N.; Streata, I.; Pei, G.; Weiner, J.; Kotze, L.; Bandermann, S.; Lozza, L.; Walzl, G.; Du Plessis, N.; Ioana, M.; et al. Human Monocytic Suppressive Cells Promote Replication of Mycobacterium Tuberculosis and Alter Stability of In Vitro Generated Granulomas. Front. Immunol. 2018, 9, 2417. [Google Scholar] [CrossRef]
- Wang, H.; Maeda, Y.; Fukutomi, Y.; Makino, M. An In Vitro Model of Mycobacterium Leprae Induced Granuloma Formation. BMC Infect. Dis. 2013, 13, 279. [Google Scholar] [CrossRef]
- Garza-Cuartero, L.; McCarthy, E.; Brady, J.; Cassidy, J.; Hamilton, C.; Sekiya, M.; NcNair, J.; Mulcahy, G. Development of an In Vitro Model of the Early-Stage Bovine Tuberculous Granuloma Using Mycobacterium Bovis-BCG. Vet. Immunol. Immunopathol. 2015, 168, 249–257. [Google Scholar] [CrossRef]
- Je, S.; Quan, H.; Na, Y.; Cho, S.N.; Kim, B.J.; Seok, S.H. An In Vitro Model of Granuloma-Like Cell Aggregates Substantiates Early Host Immune Responses Against Mycobacterium Massiliense Infection. Biol. Open 2016, 5, 1118–1127. [Google Scholar] [CrossRef]
- Hussen, J.; Duvel, A.; Sandra, O.; Smith, D.; Sheldon, I.M.; Zieger, P.; Schuberth, H.J. Phenotypic and Functional Heterogeneity of Bovine Blood Monocytes. PLoS ONE 2013, 8, e71502. [Google Scholar] [CrossRef]
- Comas, I.; Chakravartti, J.; Small, P.M.; Galagan, J.; Niemann, S.; Kremer, K.; Ernst, J.D.; Gagneux, S. Human T Cell Epitopes of Mycobacterium Tuberculosis are Evolutionarily Hyperconserved. Nat. Genet. 2010, 42, 498–503. [Google Scholar] [CrossRef]
- Verrall, A.J.; Netea, M.G.; Alisjahbana, B.; Hill, P.C.; Van Crevel, R. Early Clearance of Mycobacterium Tuberculosis: A New Frontier in Prevention. Immunology 2014, 141, 506–513. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rice, J.H.; McDaniel, M.M.; Holland, A.; Eda, S. Modelling Bovine Granuloma Formation In Vitro upon Infection with Mycobacterium Avium Subspecies Paratuberculosis. Vet. Sci. 2019, 6, 80. https://doi.org/10.3390/vetsci6040080
Rice JH, McDaniel MM, Holland A, Eda S. Modelling Bovine Granuloma Formation In Vitro upon Infection with Mycobacterium Avium Subspecies Paratuberculosis. Veterinary Sciences. 2019; 6(4):80. https://doi.org/10.3390/vetsci6040080
Chicago/Turabian StyleRice, J. Hunter, Margaret M. McDaniel, Alyson Holland, and Shigetoshi Eda. 2019. "Modelling Bovine Granuloma Formation In Vitro upon Infection with Mycobacterium Avium Subspecies Paratuberculosis" Veterinary Sciences 6, no. 4: 80. https://doi.org/10.3390/vetsci6040080
APA StyleRice, J. H., McDaniel, M. M., Holland, A., & Eda, S. (2019). Modelling Bovine Granuloma Formation In Vitro upon Infection with Mycobacterium Avium Subspecies Paratuberculosis. Veterinary Sciences, 6(4), 80. https://doi.org/10.3390/vetsci6040080





