Applying Supervised Machine Learning to Effusion Analysis for the Diagnosis of Feline Infectious Peritonitis
Abstract
1. Introduction
2. Methods
2.1. Dataset and Data Preparation
2.2. Exploratory Analysis
2.3. Feature Selection
2.4. Model Selection and Building
2.5. Model Evaluation
2.6. Software and Model Development Environments
3. Results
3.1. Case Summary
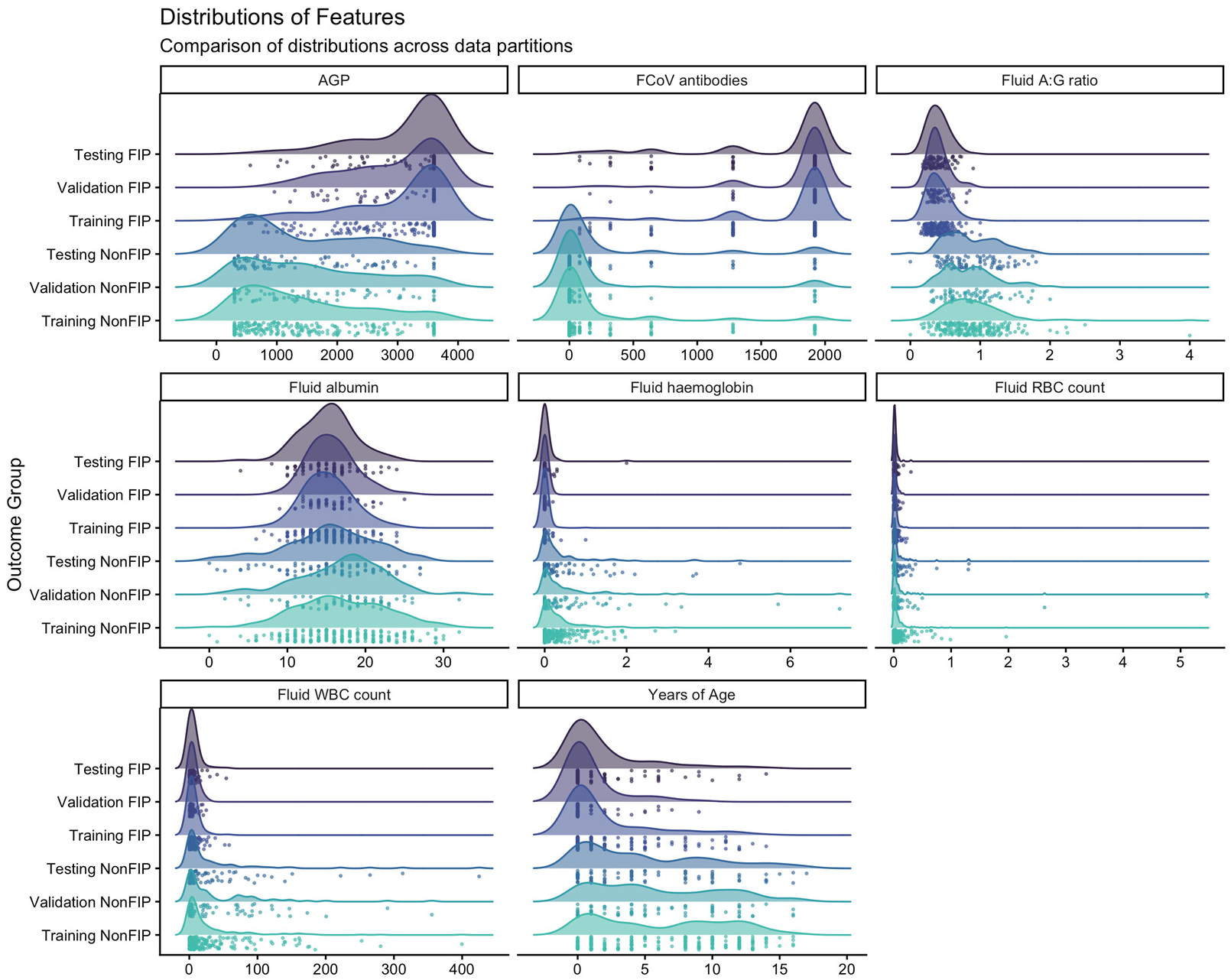
3.2. Exploratory Analysis
3.3. Data Partitions
3.4. Feature Selection
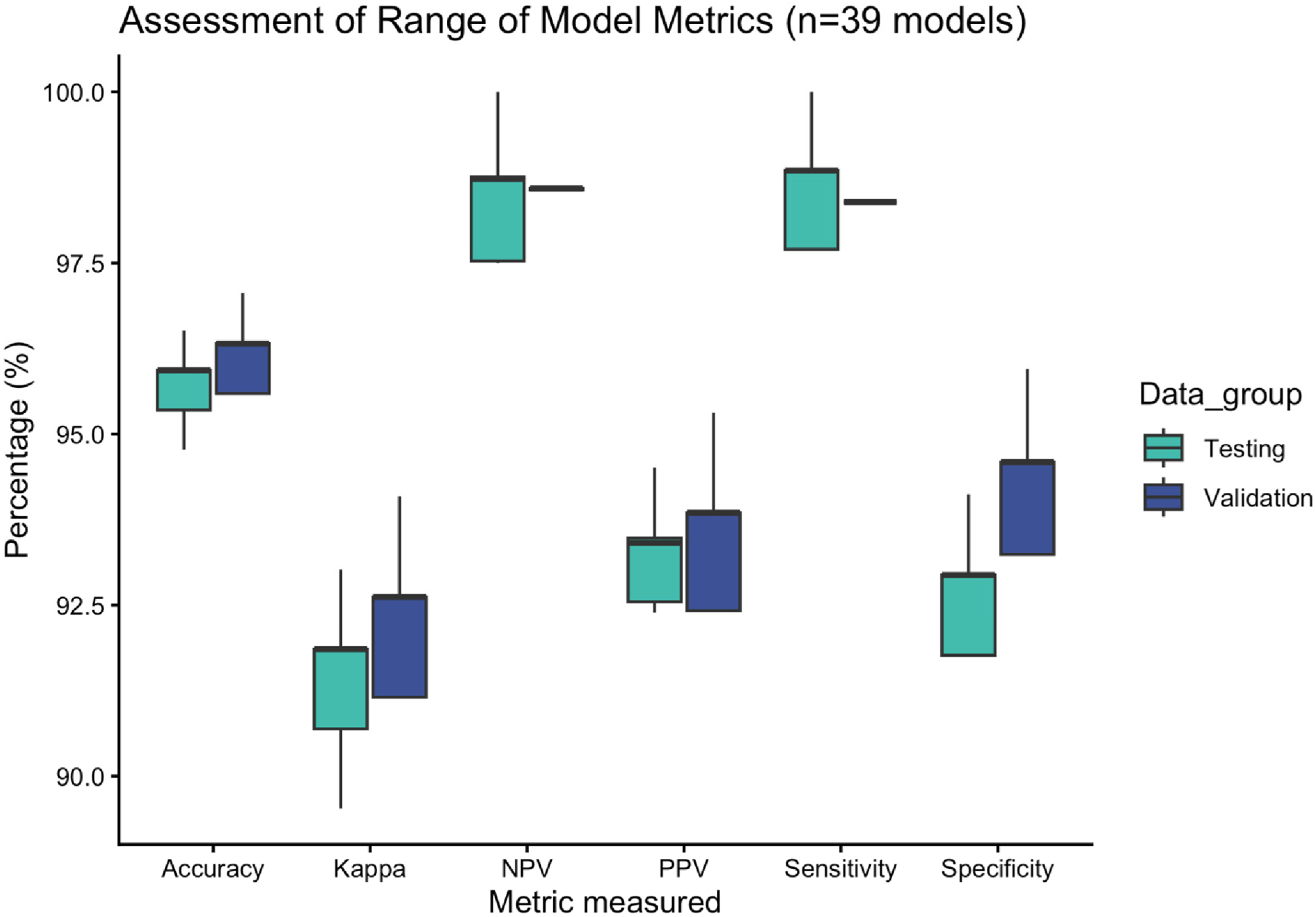
3.5. Model Performance and Statistical Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pedersen, N.C. An update on feline infectious peritonitis: Virology and immunopathogenesis. Vet. J. 2014, 201, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Weiss, R.C.; Scott, F.W. Pathogenesis of feline infectious peritonitis: Nature and development of viremia. Am. J. Vet. Res. 1981, 42, 382–390. [Google Scholar] [CrossRef] [PubMed]
- Kipar, A.; May, H.; Menger, S.; Weber, M.; Leukert, W.; Reinacher, M. Morphologic features and development of granulomatous vasculitis in feline infectious peritonitis. Vet. Pathol. 2005, 42, 321–330. [Google Scholar] [CrossRef]
- Felten, S.; Klein-Richers, U.; Unterer, S.; Bergmann, M.; Leutenegger, C.M.; Pantchev, N.; Balzer, J.; Zablotski, Y.; Hofmann-Lehmann, R.; Hartmann, K. Role of Feline Coronavirus as Contributor to Diarrhea in Cats from Breeding Catteries. Viruses 2022, 14, 858. [Google Scholar] [CrossRef]
- Felten, S.; Klein-Richers, U.; Hofmann-Lehmann, R.; Bergmann, M.; Unterer, S.; Leutenegger, C.M.; Hartmann, K. Correlation of Feline Coronavirus Shedding in Feces with Coronavirus Antibody Titer. Pathogens 2020, 9, 598. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, N.C.; Boyle, J.F.; Floyd, K.; Fudge, A.; Barker, J. An enteric coronavirus infection of cats and its relationship to feline infectious peritonitis. Am. J. Vet. Res. 1981, 42, 368–377. [Google Scholar] [CrossRef]
- Zehr, J.D.; Kosakovsky Pond, S.L.; Millet, J.K.; Olarte-Castillo, X.A.; Lucaci, A.G.; Shank, S.D.; Ceres, K.M.; Choi, A.; Whittaker, G.R.; Goodman, L.B. Natural selection differences detected in key protein domains between non-pathogenic and pathogenic feline coronavirus phenotypes. Virus Evol. 2023, 9, vead019. [Google Scholar] [CrossRef]
- Chang, H.W.; de Groot, R.J.; Egberink, H.F.; Rottier, P.J. Feline infectious peritonitis: Insights into feline coronavirus pathobiogenesis and epidemiology based on genetic analysis of the viral 3c gene. J. Gen. Virol. 2010, 91, 415–420. [Google Scholar] [CrossRef]
- Bank-Wolf, B.R.; Stallkamp, I.; Wiese, S.; Moritz, A.; Tekes, G.; Thiel, H.J. Mutations of 3c and spike protein genes correlate with the occurrence of feline infectious peritonitis. Vet. Microbiol. 2014, 173, 177–188. [Google Scholar] [CrossRef]
- Borschensky, C.M.; Reinacher, M. Mutations in the 3c and 7b genes of feline coronavirus in spontaneously affected FIP cats. Res. Vet. Sci. 2014, 97, 333–340. [Google Scholar] [CrossRef]
- Zhu, J.; Deng, S.; Mou, D.; Zhang, G.; Fu, Y.; Huang, W.; Zhang, Y.; Lyu, Y. Analysis of spike and accessory 3c genes mutations of less virulent and FIP-associated feline coronaviruses in Beijing, China. Virology 2024, 589, 109919. [Google Scholar] [CrossRef]
- Licitra, B.N.; Millet, J.K.; Regan, A.D.; Hamilton, B.S.; Rinaldi, V.D.; Duhamel, G.E.; Whittaker, G.R. Mutation in spike protein cleavage site and pathogenesis of feline coronavirus. Emerg. Infect. Dis. 2013, 19, 1066–1073. [Google Scholar] [CrossRef]
- Cave, T.A.; Golder, M.C.; Simpson, J.; Addie, D.D. Risk factors for feline coronavirus seropositivity in cats relinquished to a UK rescue charity. J. Feline Med. Surg. 2004, 6, 53–58. [Google Scholar] [CrossRef]
- Horzinek, M.C.; Osterhaus, A.D. Feline infectious peritonitis: A coronavirus disease of cats. J. Small Anim. Pract. 1978, 19, 623–630. [Google Scholar] [CrossRef]
- Pedersen, N.C. Virologic and immunologic aspects of feline infectious peritonitis virus infection. Adv. Exp. Med. Biol. 1987, 218, 529–550. [Google Scholar] [CrossRef] [PubMed]
- Attipa, C.; Warr, A.S.; Epaminondas, D.; O’Shea, M.; Hanton, A.J.; Fletcher, S.; Malbon, A.; Lyraki, M.; Hammond, R.; Hardas, A. Feline infectious peritonitis epizootic caused by a recombinant coronavirus. Nature 2025, 645, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Shah, A.U.; Esparza, B.; Illanes, O.; Hemida, M.G. Comparative Genome Sequencing Analysis of Some Novel Feline Infectious Peritonitis Viruses Isolated from Some Feral Cats in Long Island. Viruses 2025, 17, 209. [Google Scholar] [CrossRef] [PubMed]
- Tasker, S. Diagnosis of feline infectious peritonitis: Update on evidence supporting available tests. J. Feline Med. Surg. 2018, 20, 228–243. [Google Scholar] [CrossRef]
- Pedersen, N.C. An update on feline infectious peritonitis: Diagnostics and therapeutics. Vet. J. 2014, 201, 133–141. [Google Scholar] [CrossRef]
- Fischer, Y.; Wess, G.; Hartmann, K. [Pericardial effusion in a cat with feline infectious peritonitis]. Schweiz. Arch. Tierheilkd. 2012, 154, 27–31. [Google Scholar] [CrossRef] [PubMed]
- Tucker, R.L.; Hodges, R.D. What is your diagnosis? Pericardial effusion in a cat. J. Am. Vet. Med. Assoc. 1994, 205, 825–826. [Google Scholar] [CrossRef] [PubMed]
- Savary, K.C.; Sellon, R.K.; Law, J.M. Chylous abdominal effusion in a cat with feline infectious peritonitis. J. Am. Anim. Hosp. Assoc. 2001, 37, 35–40. [Google Scholar] [CrossRef]
- Kopilovic, A.; Gvozdic, D.; Radakovic, M.; Spariosu, K.; Andric, N.; Francuski-Andric, J. Selected hematology ratios in cats with non-septic effusions highly suspected of feline infectious peritonitis. Vet. Glas. 2023, 77, 164–175. [Google Scholar] [CrossRef]
- Paltrinieri, S.; Parodi, M.C.; Cammarata, G. In vivo diagnosis of feline infectious peritonitis by comparison of protein content, cytology, and direct immunofluorescence test on peritoneal and pleural effusions. J. Vet. Diagn. Investig. 1999, 11, 358–361. [Google Scholar] [CrossRef] [PubMed]
- Tasker, S.; Addie, D.D.; Egberink, H.; Hofmann-Lehmann, R.; Hosie, M.J.; Truyen, U.; Belak, S.; Boucraut-Baralon, C.; Frymus, T.; Lloret, A.; et al. Feline Infectious Peritonitis: European Advisory Board on Cat Diseases Guidelines. Viruses 2023, 15, 1847. [Google Scholar] [CrossRef]
- Riemer, F.; Kuehner, K.A.; Ritz, S.; Sauter-Louis, C.; Hartmann, K. Clinical and laboratory features of cats with feline infectious peritonitis--a retrospective study of 231 confirmed cases (2000–2010). J. Feline Med. Surg. 2016, 18, 348–356. [Google Scholar] [CrossRef] [PubMed]
- Norris, J.M.; Bosward, K.L.; White, J.D.; Baral, R.M.; Catt, M.J.; Malik, R. Clinicopathological findings associated with feline infectious peritonitis in Sydney, Australia: 42 cases (1990–2002). Aust. Vet. J. 2005, 83, 666–673. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, D.; Kwok, W.; Graham, E.; Armitage, A.; Irvine, R.; Johnston, P.; McDonald, M.; Montgomery, D.; Nicolson, L.; Robertson, E.; et al. Diagnosis of non-effusive feline infectious peritonitis by reverse transcriptase quantitative PCR from mesenteric lymph node fine-needle aspirates. J. Feline Med. Surg. 2019, 21, 910–921. [Google Scholar] [CrossRef]
- Stranieri, A.; Scavone, D.; Paltrinieri, S.; Giordano, A.; Bonsembiante, F.; Ferro, S.; Gelain, M.E.; Meazzi, S.; Lauzi, S. Concordance between Histology, Immunohistochemistry, and RT-PCR in the Diagnosis of Feline Infectious Peritonitis. Pathogens 2020, 9, 852. [Google Scholar] [CrossRef]
- Stranieri, A.; Giordano, A.; Paltrinieri, S.; Giudice, C.; Cannito, V.; Lauzi, S. Comparison of the performance of laboratory tests in the diagnosis of feline infectious peritonitis. J. Vet. Diagn. Investig. 2018, 30, 459–463. [Google Scholar] [CrossRef]
- Herrewegh, A.A.; de Groot, R.J.; Cepica, A.; Egberink, H.F.; Horzinek, M.C.; Rottier, P.J. Detection of feline coronavirus RNA in feces, tissues, and body fluids of naturally infected cats by reverse transcriptase PCR. J. Clin. Microbiol. 1995, 33, 684–689. [Google Scholar] [CrossRef]
- Gunther, S.; Felten, S.; Wess, G.; Hartmann, K.; Weber, K. Detection of feline Coronavirus in effusions of cats with and without feline infectious peritonitis using loop-mediated isothermal amplification. J. Virol. Methods 2018, 256, 32–36. [Google Scholar] [CrossRef]
- Felten, S.; Leutenegger, C.M.; Balzer, H.J.; Pantchev, N.; Matiasek, K.; Wess, G.; Egberink, H.; Hartmann, K. Sensitivity and specificity of a real-time reverse transcriptase polymerase chain reaction detecting feline coronavirus mutations in effusion and serum/plasma of cats to diagnose feline infectious peritonitis. BMC Vet. Res. 2017, 13, 228. [Google Scholar] [CrossRef] [PubMed]
- Felten, S.; Weider, K.; Doenges, S.; Gruendl, S.; Matiasek, K.; Hermanns, W.; Mueller, E.; Matiasek, L.; Fischer, A.; Weber, K.; et al. Detection of feline coronavirus spike gene mutations as a tool to diagnose feline infectious peritonitis. J. Feline Med. Surg. 2017, 19, 321–335. [Google Scholar] [CrossRef]
- Stranieri, A.; Lauzi, S.; Paltrinieri, S. Clinicopathological and Molecular Analysis of Aqueous Humor for the Diagnosis of Feline Infectious Peritonitis. Vet. Sci. 2024, 11, 207. [Google Scholar] [CrossRef] [PubMed]
- Longstaff, L.; Porter, E.; Crossley, V.J.; Hayhow, S.E.; Helps, C.R.; Tasker, S. Feline coronavirus quantitative reverse transcriptase polymerase chain reaction on effusion samples in cats with and without feline infectious peritonitis. J. Feline Med. Surg. 2017, 19, 240–245. [Google Scholar] [CrossRef]
- Barua, S.; Sarkar, S.; Chenoweth, K.; Johnson, C.; Delmain, D.; Wang, C. Insights on feline infectious peritonitis risk factors and sampling strategies from polymerase chain reaction analysis of feline coronavirus in large-scale nationwide submissions. J. Am. Vet. Med. Assoc. 2025, 263, 82–89. [Google Scholar] [CrossRef]
- Pedersen, N.C.; Eckstrand, C.; Liu, H.; Leutenegger, C.; Murphy, B. Levels of feline infectious peritonitis virus in blood, effusions, and various tissues and the role of lymphopenia in disease outcome following experimental infection. Vet. Microbiol. 2015, 175, 157–166. [Google Scholar] [CrossRef]
- Felten, S.; Hartmann, K.; Doerfelt, S.; Sangl, L.; Hirschberger, J.; Matiasek, K. Immunocytochemistry of mesenteric lymph node fine-needle aspirates in the diagnosis of feline infectious peritonitis. J. Vet. Diagn. Investig. 2019, 31, 210–216. [Google Scholar] [CrossRef]
- Felten, S.; Matiasek, K.; Gruendl, S.; Sangl, L.; Hartmann, K. Utility of an immunocytochemical assay using aqueous humor in the diagnosis of feline infectious peritonitis. Vet. Ophthalmol. 2018, 21, 27–34. [Google Scholar] [CrossRef]
- Gruendl, S.; Matiasek, K.; Matiasek, L.; Fischer, A.; Felten, S.; Jurina, K.; Hartmann, K. Diagnostic utility of cerebrospinal fluid immunocytochemistry for diagnosis of feline infectious peritonitis manifesting in the central nervous system. J. Feline Med. Surg. 2017, 19, 576–585. [Google Scholar] [CrossRef]
- Felten, S.; Matiasek, K.; Gruendl, S.; Sangl, L.; Wess, G.; Hartmann, K. Investigation into the utility of an immunocytochemical assay in body cavity effusions for diagnosis of feline infectious peritonitis. J. Feline Med. Surg. 2017, 19, 410–418. [Google Scholar] [CrossRef]
- Howell, M.; Evans, S.J.M.; Cornwall, M.; Santangelo, K.S. Multiplex fluorescent immunocytochemistry for the diagnosis of feline infectious peritonitis: Determining optimal storage conditions. Vet. Clin. Pathol. 2020, 49, 640–645. [Google Scholar] [CrossRef]
- Ives, E.J.; Vanhaesebrouck, A.E.; Cian, F. Immunocytochemical demonstration of feline infectious peritonitis virus within cerebrospinal fluid macrophages. J. Feline Med. Surg. 2013, 15, 1149–1153. [Google Scholar] [CrossRef] [PubMed]
- Giordano, A.; Paltrinieri, S.; Bertazzolo, W.; Milesi, E.; Parodi, M. Sensitivity of Tru-cut and fine needle aspiration biopsies of liver and kidney for diagnosis of feline infectious peritonitis. Vet. Clin. Pathol. 2005, 34, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Sangl, L.; Matiasek, K.; Felten, S.; Grundl, S.; Bergmann, M.; Balzer, H.J.; Pantchev, N.; Leutenegger, C.M.; Hartmann, K. Detection of feline coronavirus mutations in paraffin-embedded tissues in cats with feline infectious peritonitis and controls. J. Feline Med. Surg. 2019, 21, 133–142. [Google Scholar] [CrossRef]
- Ziolkowska, N.; Pazdzior-Czapula, K.; Lewczuk, B.; Mikulska-Skupien, E.; Przybylska-Gornowicz, B.; Kwiecinska, K.; Ziolkowski, H. Feline Infectious Peritonitis: Immunohistochemical Features of Ocular Inflammation and the Distribution of Viral Antigens in Structures of the Eye. Vet. Pathol. 2017, 54, 933–944. [Google Scholar] [CrossRef] [PubMed]
- Paltrinieri, S.; Comazzi, S.; Spagnolo, V.; Giordano, A. Laboratory changes consistent with feline infectious peritonitis in cats from multicat environments. J. Vet. Med. A Physiol. Pathol. Clin. Med. 2002, 49, 503–510. [Google Scholar] [CrossRef]
- Paltrinieri, S.; Grieco, V.; Comazzi, S.; Cammarata Parodi, M. Laboratory profiles in cats with different pathological and immunohistochemical findings due to feline infectious peritonitis (FIP). J. Feline Med. Surg. 2001, 3, 149–159. [Google Scholar] [CrossRef]
- Giori, L.; Giordano, A.; Giudice, C.; Grieco, V.; Paltrinieri, S. Performances of different diagnostic tests for feline infectious peritonitis in challenging clinical cases. J. Small Anim. Pract. 2011, 52, 152–157. [Google Scholar] [CrossRef]
- Thayer, V.; Gogolski, S.; Felten, S.; Hartmann, K.; Kennedy, M.; Olah, G.A. 2022 AAFP/EveryCat Feline Infectious Peritonitis Diagnosis Guidelines. J. Feline Med. Surg. 2022, 24, 905–933. [Google Scholar] [CrossRef]
- Wu, J.; Zhang, P.; Zhang, L.; Meng, W.; Li, J.; Tong, C.; Li, Y.; Cai, J.; Yang, Z.; Zhu, J. Rapid and accurate identification of COVID-19 infection through machine learning based on clinical available blood test results. medRxiv 2020. [Google Scholar] [CrossRef]
- Hathaway, Q.A.; Roth, S.M.; Pinti, M.V.; Sprando, D.C.; Kunovac, A.; Durr, A.J.; Cook, C.C.; Fink, G.K.; Cheuvront, T.B.; Grossman, J.H.; et al. Machine-learning to stratify diabetic patients using novel cardiac biomarkers and integrative genomics. Cardiovasc. Diabetol. 2019, 18, 78. [Google Scholar] [CrossRef] [PubMed]
- Hornbrook, M.C.; Goshen, R.; Choman, E.; O’Keeffe-Rosetti, M.; Kinar, Y.; Liles, E.G.; Rust, K.C. Early colorectal cancer detected by machine learning model using gender, age, and complete blood count data. Dig. Dis. Sci. 2017, 62, 2719–2727. [Google Scholar] [CrossRef]
- Tseng, Y.-J.; Huang, C.-E.; Wen, C.-N.; Lai, P.-Y.; Wu, M.-H.; Sun, Y.-C.; Wang, H.-Y.; Lu, J.-J. Predicting breast cancer metastasis by using serum biomarkers and clinicopathological data with machine learning technologies. Int. J. Med. Inform. 2019, 128, 79–86. [Google Scholar] [CrossRef]
- Bradley, R.; Tagkopoulos, I.; Kim, M.; Kokkinos, Y.; Panagiotakos, T.; Kennedy, J.; De Meyer, G.; Watson, P.; Elliott, J. Predicting early risk of chronic kidney disease in cats using routine clinical laboratory tests and machine learning. J. Vet. Intern. Med. 2019, 33, 2644–2656. [Google Scholar] [CrossRef] [PubMed]
- Pfannschmidt, K.; Hüllermeier, E.; Held, S.; Neiger, R. Evaluating Tests in Medical Diagnosis: Combining Machine Learning with Game-Theoretical Concepts. In Proceedings of the Information Processing and Management of Uncertainty in Knowledge-Based Systems, Eindhoven, The Netherlands, 20–24 June 2016; pp. 450–461. [Google Scholar]
- Doguc, O.; Bilgi, S.B.; Cagdas, S.; Yilmazturk, N. A Decision Support System for Detecting FIP Disease in Cats Based on Machine Learning Methods. In Proceedings of the Emerging Trends and Applications in Artificial Intelligence, Istanbul, Turkey, 8–9 September 2023; pp. 176–186. [Google Scholar]
- Kuo, P.H.; Li, Y.H.; Yau, H.T. Development of feline infectious peritonitis diagnosis system by using CatBoost algorithm. Comput. Biol. Chem. 2024, 113, 108227. [Google Scholar] [CrossRef]
- Dunbar, D.; Babayan, S.A.; Krumrie, S.; Haining, H.; Hosie, M.J.; Weir, W. Assessing the feasibility of applying machine learning to diagnosing non-effusive feline infectious peritonitis. Sci. Rep. 2024, 14, 2517. [Google Scholar] [CrossRef]
- Sanders, N.A. Pleural transudates and modified transudates. In Textbook of Respiratory Disease in Dogs and Cats; W.B. Saunders: Philadelphia, PA, USA, 2004; pp. 587–597. [Google Scholar]
- Stockham, S.L.; Scott, M.A. Cavitary effusion. In Fundamentals of Veterinary Clinical Pathology, 2nd ed.; Wiley-Blackwell: Ames, IA, USA, 2008. [Google Scholar]
- Beatty, J.; Barrs, V. Pleural Effusion in the Cat: A Practical Approach to Determining Aetiology. J. Feline Med. Surg. 2010, 12, 693–707. [Google Scholar] [CrossRef]
- Tasker, S.; Gunn-Moore, D. Differential diagnosis of ascites in cats. Practice 2000, 22, 472–479. [Google Scholar] [CrossRef]
- Raab, O. Abdominocentsis and Diagnostic Peritoneal Lavage. In Textbook of Veterinary Internal Medicine: Diseases of the Dog and the Cat, 8th ed.; Ettinger, S.J., Feldman, E.C., Côté, E., Eds.; Elsevier: St. Louis, MI, USA, 2017; Volume 1. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2022. [Google Scholar]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. Xgboost: A scalable tree boosting system. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge, Discovery and Data Mining, San Fransisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]
- Kuhn, M. Caret: Classification and Regression Training; Boston, MA, USA, 2020. [Google Scholar]
- Boser, B.E.; Guyon, I.M.; Vapnik, V.N. A training algorithm for optimal margin classifiers. In Proceedings of the Fifth Annual Workshop on Computational Learning Theory, Pittsburgh, PA, USA, 1 July 1992; pp. 144–152. [Google Scholar]
- de Oliveira, A.; Engelmann, A.M.; Jaguezeski, A.M.; da Silva, C.B.; Barbosa, N.V.; de Andrade, C.M. Retrospective study of the aetiopathological diagnosis of pleural or peritoneal effusion exams of dogs and cats. Comp. Clin. Pathol. 2021, 30, 811–820. [Google Scholar] [CrossRef]
- Wright, K.N.; Gompf, R.E.; DeNovo, R.C., Jr. Peritoneal effusion in cats: 65 cases (1981–1997). J. Am. Vet. Med. Assoc. 1999, 214, 375–381. [Google Scholar] [CrossRef]
- Davies, C.; Forrester, S. Pleural effusion in cats: 82 cases (1987 to 1995). J. Small Anim. Pract. 1996, 37, 217–224. [Google Scholar] [CrossRef]
- König, A.; Hartmann, K.; Mueller, R.S.; Wess, G.; Schulz, B.S. Retrospective analysis of pleural effusion in cats. J. Feline Med. Surg. 2019, 21, 1102–1110. [Google Scholar] [CrossRef]
- Hall, D.J.; Shofer, F.; Meier, C.K.; Sleeper, M.M. Pericardial effusion in cats: A retrospective study of clinical findings and outcome in 146 cats. J. Vet. Intern. Med. 2007, 21, 1002–1007. [Google Scholar] [CrossRef] [PubMed]
- Hardwick, J.J.; Ioannides-Hoey, C.S.; Finch, N.; Black, V. Bicavitary effusion in cats: Retrospective analysis of signalment, clinical investigations, diagnosis and outcome. J. Feline Med. Surg. 2024, 26, 1098612X241227122. [Google Scholar] [CrossRef] [PubMed]
- Helfer-Hungerbuehler, A.K.; Spiri, A.M.; Meili, T.; Riond, B.; Krentz, D.; Zwicklbauer, K.; Buchta, K.; Zuzzi-Krebitz, A.M.; Hartmann, K.; Hofmann-Lehmann, R.; et al. Alpha-1-Acid Glycoprotein Quantification via Spatial Proximity Analyte Reagent Capture Luminescence Assay: Application as Diagnostic and Prognostic Marker in Serum and Effusions of Cats with Feline Infectious Peritonitis Undergoing GS-441524 Therapy. Viruses 2024, 16, 791. [Google Scholar] [CrossRef] [PubMed]
- Zwicklbauer, K.; Grassl, P.; Alberer, M.; Kolberg, L.; Schweintzger, N.A.; Härtle, S.; Matiasek, K.; Hofmann-Lehmann, R.; Hartmann, K.; Friedel, C.C.; et al. Whole blood RNA profiling in cats dissects the host immunological response during recovery from feline infectious peritonitis. PLoS ONE 2025, 20, e0332248. [Google Scholar] [CrossRef]
- Tršar, L.; Štrljič, M.; Svete, A.N.; Koprivec, S.; Tozon, N.; Žel, M.K.; Pavlin, D. Evaluation of selected inflammatory markers in cats with feline infectious peritonitis before and after therapy. BMC Vet. Res. 2025, 21, 330. [Google Scholar] [CrossRef]
- Golovko, L.; Lyons, L.A.; Liu, H.; Sorensen, A.; Wehnert, S.; Pedersen, N.C. Genetic susceptibility to feline infectious peritonitis in Birman cats. Virus Res. 2013, 175, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Pesteanu-Somogyi, L.D.; Radzai, C.; Pressler, B.M. Prevalence of feline infectious peritonitis in specific cat breeds. J. Feline Med. Surg. 2006, 8, 1–5. [Google Scholar] [CrossRef] [PubMed]



| Training | Validation | Testing | Overall | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Feature Group | Feature Name | Method | Non-FIP (n = 223) | FIP (n = 187) | Non-FIP (n = 74) | FIP (n = 62) | Non-FIP (n = 85) | FIP (n = 87) | Non-FIP (n = 382) | FIP (n = 336) |
| Serology | FCoV titre | Immunofluorescence ◊ | ||||||||
| Median [Min, Max] | 0 [0, 1920] | 1920 [80.0, 1920] | 0 [0, 1920] | 1920 [160, 1920] | 0 [0, 1920] | 1920 [80.0, 1920] | 0 [0, 1920] | 1920 [80.0, 1920] | ||
| Cytology | Fluid RBC count (×1012/L) | Siemens Advia 120 analyser † | ||||||||
| Mean [Min, Max] | 0.0894 [0, 1.96] | 0.0282 [0, 0.230] | 0.17 [0, 5.45] | 0.026 [0, 0.160] | 0.097 [0, 1.31] | 0.0252 [0, 0.300] | 0.107 [0, 5.45] | 0.027 [0, 0.300] | ||
| Fluid haemoglobin (g/dL) | Siemens Advia 120 analyser † | |||||||||
| Mean [Min, Max] | 0.347 [0, 3.19] | 0.0134 [0, 1.00] | 0.62 [0, 7.20] | 0.008 [0, 0.200] | 0.459 [0, 4.77] | 0.0393 [0, 2.00] | 0.425 [0, 7.20] | 0.019 [0, 2.00] | ||
| Fluid WBC count (×109/L) | Siemens Advia 120 analyser †/manual count | |||||||||
| Mean [Min, Max] | 30.1 [0.0300, 400] | 5.47 [0.110, 58.5] | 45.3 [0.170, 356] | 5.09 [0.320, 24.9] | 36.8 [0.0800, 425] | 5.65 [0.200, 54.2] | 34.6 [0.030, 425] | 5.45 [0.110, 58.5] | ||
| Lymphocyte count (×109/L) | Manual cell count | |||||||||
| Mean [Min, Max] | 1.39 [0, 52.2] | 0.243 [0, 5.17] | 0.916 [0, 18.2] | 0.25 [0, 2.25] | 1.16 [0, 20.5] | 0.295 [0, 3.25] | 1.25 [0, 52.2] | 0.258 [0, 5.17] | ||
| Neutrophil count (×109/L) | Manual cell count | |||||||||
| Mean [Min, Max] | 12.1 [0, 376] | 3.66 [0, 49.1] | 24.9 [0, 338] | 3.39 [0, 18.5] | 14.2 [0, 209] | 3.83 [0, 41.2] | 15.1 [0, 376] | 3.65 [0, 49.1] | ||
| Macrophage count (×109/L) | Manual cell count | |||||||||
| Mean [Min, Max] | 1.18 [0, 51.9] | 0.709 [0, 8.78] | 2.11 [0, 58.1] | 0.604 [0, 5.52] | 1.44 [0, 34.8] | 0.833 [0, 9.76] | 1.42 [0, 58.1] | 0.722 0, 9.76] | ||
| Eosinophil count (×109/L) | Manual cell count | |||||||||
| Mean [Min, Max] | 0.0477 [0, 2.83] | 0.0043 [0, 0.283] | 0.0129 [0, 0.270] | 0.0052 [0, 0.251] | 3.56 [0, 297] | 0.002 [0, 0.0988] | 0.822 [0, 297] | 0.004 [0, 0.283] | ||
| Plasma cell count | Manual cell count | |||||||||
| Mean [Min, Max] | 0.0016 [0, 0.357] | 0.0016 [0, 0.283] | 0 [0, 0] | 0.00087 [0, 0.0364] | 0.0002 [0, 0.0174] | 0 [0, 0] | 0.001 [0, 0.357] | 0.001 [0, 0.283] | ||
| Mast cell count (×109/L) | Manual cell count | |||||||||
| Mean [Min, Max] | 0.0056 [0, 1.23] | 0.0026 [0, 0.283] | 0.0108 [0, 0.786] | 0.0021 [0, 0.120] | 0.0031 [0, 0.226] | 0.0714 [0, 0.301] | 0.006 [0, 1.23] | 0.004 [0, 0.301] | ||
| Mesothelial cells (×109/L) | Manual cell count | |||||||||
| Mean [Min, Max] | 0.0005 [0, 0.0712] | 0 [0, 0] | 0.0031 [0, 0.222] | 0.0001 [0, 0.0044] | 0.0069 [0, 0.590] | 0 [0, 0] | 0.002 [0, 0.590] | 0.00001 [0, 0.004] | ||
| Biochemistry | Fluid total protein (g/L) | Siemens Dimension Xpand Plus analyser | ||||||||
| Mean [Min, Max] | 39.6 [1.00, 110] | 56.5 [28.0, 109] | 40.6 [4.00, 109] | 57.6 [26.0, 88.0] | 34.9 [1.00, 69.0] | 56.7 [18.0, 94.0] | 38.8 [1.00, 110] | 56.8 [18.0, 109] | ||
| Fluid albumin (g/L ) | Siemens Dimension Xpand Plus analyser | |||||||||
| Mean [Min, Max] | 16.9 [0, 32.0] | 15.1 [6.00, 24.0] | 17.0 [2.00, 32.0] | 15.6 [9.00, 25.0] | 15.4 [0, 27.0] | 15.2 [4.00, 24.0] | 16.6 [0, 32.0] | 15.2 [4.00, 25.0] | ||
| Fluid globulins (g/L) | Siemens Dimension Xpand Plus analyser | |||||||||
| Mean [Min, Max] | 22.7 [1.00, 87.0] | 41.4 [16.0, 97.0] | 23.6 [2.00, 87.0] | 42.0 [17.0, 65.0] | 19.5 [1.00, 48.0] | 41.6 [14.0, 81.0] | 22.1 1.00, 87.0] | 41.5 [14.0, 97.0] | ||
| A:G ratio | Siemens Dimension Xpand Plus analyser | |||||||||
| Median [Min, Max] | 0.820 [0, 4.00] | 0.380 [0.120, 1.00] | 0.840 [0.300, 2.00] | 0.360 [0.180, 0.850] | 0.800 [0, 1.80] | 0.390 [0.180, 0.900] | 0.820 [0, 4.00] | 0.380 [0.120, 1.00] | ||
| AGP (µg/mL) | RID */ELISA | |||||||||
| Mean [Min, Max] | 1440 [299, 3600] | 2970 [299, 3600] | 1540 [299, 3600] | 2940 [960, 3600] | 1440 [299, 3600] | 3100 [560, 3600] | 1460 [299, 3600] | 3000 [299, 3600] | ||
| Signalment | Age (years) | |||||||||
| Mean [Min, Max] | 6.26 [0, 16.0] | 2.07 [0, 14.0] | 5.96 [0, 16.0] | 0.968 [0, 9.00] | 5.06 [0, 17.0] | 1.97 [0, 14.0] | 5.93 [0, 17.0] | 1.84 [0, 14.0] | ||
| Sex n (%) | ||||||||||
| Male | 114 (51%) | 119 (64%) | 43 (58%) | 43 (69%) | 51 (60%) | 59 (68%) | 208 (54%) | 221 (66%) | ||
| Female | 109 (49%) | 68 (36%) | 31 (42%) | 19 (31%) | 34 (40%) | 28 (32%) | 174 (46%) | 115 (34%) | ||
| Pedigree n (%) | ||||||||||
| Pedigree | 44 (20%) | 106 (57%) | 18 (24%) | 41 (66%) | 24 (28%) | 51 (59%) | 86 (23%) | 198 (59%) | ||
| Not Pedigree | 179 (80%) | 81 (43%) | 56 (76%) | 21 (34%) | 61 (72%) | 36 (41%) | 296 (77%) | 138 (41%) | ||
| Effusion Information | Bicavitary effusion n (%) | |||||||||
| Present | 7 (3%) | 1 (1%) | 3 (4%) | 1 (2%) | 1 (1%) | 2 (2%) | 11 (3%) | 4 (1%) | ||
| Absent | 216 (97%) | 186 (99%) | 71 (96%) | 61 (98%) | 84 (99%) | 85 (98%) | 371 (97%) | 332 (99%) | ||
| Effusion type n (%) | ||||||||||
| Transudate | 10 (4%) | 1 (1%) | 4 (5%) | 0 (0%) | 11 (13%) | 0 (0%) | 25 (7%) | 1 (0%) | ||
| Modified Transudate | 87 (39%) | 114 (61%) | 28 (38%) | 39 (63%) | 29 (34%) | 56 (64%) | 144 (38%) | 209 (62%) | ||
| Exudate | 126 (57%) | 72 (39%) | 42 (57%) | 23 (37%) | 45 (53%) | 31 (36%) | 213 (56%) | 126 (38%) | ||
| Effusion site n (%) | ||||||||||
| Ascites | 85 (38%) | 116 (62%) | 36 (49%) | 42 (68%) | 38 (45%) | 57 (66%) | 159 (42%) | 215 (64%) | ||
| Pleural | 114 (51%) | 41 (22%) | 35 (47%) | 8 (13%) | 36 (42%) | 11 (13%) | 185 (48%) | 60 (18%) | ||
| Peritoneal | 22 (10%) | 29 (16%) | 3 (4%) | 12 (19%) | 8 (9%) | 19 (22%) | 33 (9%) | 60 (18%) | ||
| Thoracic | 1 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 1 (1%) | 0 (0%) | 2 (1%) | 0 (0%) | ||
| Pericardial | 1 (0%) | 1 (1%) | 0 (0%) | 0 (0%) | 2 (2%) | 0 (0%) | 3 (1%) | 1 (0%) | ||
| Clinical signs | Lethargy n (%) | |||||||||
| Present | 31 (14%) | 38 (20%) | 11 (15%) | 8 (13%) | 8 (9%) | 19 (22%) | 50 (13%) | 65 (19%) | ||
| Absent | 192 (86%) | 149 (80%) | 63 (85%) | 54 (87%) | 77 (91%) | 68 (78%) | 332 (87%) | 271 (81%) | ||
| Icterus n (%) | ||||||||||
| Present | 7 (3%) | 9 (5%) | 2 (3%) | 4 (6%) | 1 1%) | 6 (7%) | 10 (3%) | 19 (6%) | ||
| Absent | 216 (97%) | 178 (95%) | 72 (97%) | 58 (94%) | 84 (99%) | 81 (93%) | 372 (97%) | 317 (94%) | ||
| Pyrexia n (%) | ||||||||||
| Present | 28 (13%) | 55 (29%) | 11 (15%) | 12 (19%) | 12 (14%) | 23 (26%) | 51 (13%) | 90 (27%) | ||
| Absent | 195 (87%) | 132 (71%) | 63 (85%) | 50 (81%) | 73 (86%) | 64 (74%) | 331 (87%) | 246 (73%) | ||
| Anorexia n (%) | ||||||||||
| Present | 37 (17%) | 46 (25%) | 14 (19%) | 11 (18%) | 13 (15%) | 15 (17%) | 64 (17%) | 72 (21%) | ||
| Absent | 186 (83%) | 141 (75%) | 60 (81%) | 51 (82%) | 72 (85%) | 72 (83%) | 318 (83%) | 264 (79%) | ||
| Inappetence n (%) | ||||||||||
| Present | 31 (14%) | 33 (18%) | 9 (12%) | 10 (16%) | 8 (9%) | 17 (20%) | 48 (13%) | 60 (18%) | ||
| Absent | 192 (86%) | 154 (82%) | 65 (88%) | 52 (84%) | 77 (91%) | 70 (80%) | 334 (87%) | 276 (82%) | ||
| Dyspnoea n (%) | ||||||||||
| Present | 25 (11%) | 10 (5%) | 8 (11%) | 1 (2%) | 10 (12%) | 2 (2%) | 43 (11%) | 13 (4%) | ||
| Absent | 198 (89%) | 177 (95%) | 66 (89%) | 61 (98%) | 75 (88%) | 85 (98%) | 339 (89%) | 323 (96%) | ||
| Diarrhoea n (%) | ||||||||||
| Present | 6 (3%) | 14 (7%) | 1 (1%) | 3 (5%) | 3 (4%) | 3 (3%) | 10 (3%) | 20 (6%) | ||
| Absent | 217 (97%) | 173 (93%) | 73 (99%) | 59 (95%) | 82 (96%) | 84 (97%) | 372 (97%) | 316 (94%) | ||
| Pallor n (%) | ||||||||||
| Present | 9 (4%) | 6 (3%) | 2 (3%) | 1 (2%) | 2 (2%) | 1 (1%) | 13 (3%) | 8 (2%) | ||
| Absent | 214 (96%) | 181 (97%) | 72 (97%) | 61 (98%) | 83 (98%) | 86 (99%) | 369 (97%) | 328 (98%) | ||
| Vomiting n (%) | ||||||||||
| Present | 5 (2%) | 4 (2%) | 2 (3%) | 2 (3%) | 4 (5%) | 2 (2%) | 11 (3%) | 8 (2%) | ||
| Absent | 218 (98%) | 183 (98%) | 72 (97%) | 60 (97%) | 81 (95%) | 85 (98%) | 371 (97%) | 328 (98%) | ||
| Mass n (%) | ||||||||||
| Present | 5 (2%) | 7 (4%) | 0 (0%) | 0 (0%) | 2 (2%) | 2 (2%) | 7 (2%) | 9 (3%) | ||
| Absent | 218 (98%) | 180 (96%) | 74 (100%) | 62 (100%) | 83 (98%) | 85 (98%) | 375 (98%) | 327 (97%) | ||
| Dataset Assessed | Features Included | No. of Features (n) | Accuracy (95% CI) | Cohen’s Kappa (%) | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | Ensemble RF Mean Resample Accuracy [Min, Max] (%) |
|---|---|---|---|---|---|---|---|---|---|
| Validation dataset | LM (-rbc, -tp, -glob) + CS + effusion info + PlasC + MstC + MesC | 25 | 96.32 (91.63–98.8) | 92.62 | 98.39 | 94.59 | 93.85 | 98.59 | |
| LM (-rbc, -tp, -glob) + CS + effusion site and type | 21 | 96.32 (91.63–98.8) | 92.62 | 98.39 | 94.59 | 93.85 | 98.59 | ||
| LM (-rbc, -tp, -glob) + CS + effusion site, type and bicavitary | 22 | 96.32 (91.63–98.8) | 92.62 | 98.39 | 94.59 | 93.85 | 98.59 | ||
| LM (-rbc, -tp, -glob) + CS + effusion site + bicavitary | 23 | 96.32 (91.63–98.8) | 92.62 | 98.39 | 94.59 | 93.85 | 98.59 | ||
| * | LM (-rbc, -tp, -glob) + CS + effusion site | 20 | 96.32 (91.63–98.8) | 92.62 | 98.39 | 94.59 | 93.85 | 98.59 | |
| Testing dataset | LM (-rbc, -tp, -glob) + CS, + effusion info + PlasC + MstC + MesC | 25 | 96.51 (92.56–98.71) | 93.02 | 98.85 | 94.12 | 94.51 | 98.77 | 94.02 [89.8, 96.52] |
| LM (-rbc, -tp, -glob) + CS + effusion site and type | 21 | 96.51 (92.56–98.71) | 93.02 | 98.85 | 94.12 | 94.51 | 98.77 | 94.64 [91.2, 96.26] | |
| LM (-rbc, -tp, -glob) + CS + effusion site, type and bicavitary | 22 | 96.51 (92.56–98.71) | 93.02 | 98.85 | 94.12 | 94.51 | 98.77 | 94.69 [90.4, 96.78] | |
| LM (-rbc, -tp, -glob) + CS + effusion site + bicavitary | 23 | 96.51 (92.56–98.71) | 93.02 | 98.85 | 94.12 | 94.51 | 98.77 | 94.81 [92.51, 96.52] | |
| * | LM (-rbc, -tp, -glob) + CS + effusion site | 20 | 96.51 (92.56–98.71) | 93.02 | 98.85 | 94.12 | 94.51 | 98.77 | 94.82 [92.53, 96.28] |
| Reference | ||||
|---|---|---|---|---|
| FIP | Non-FIP | |||
| Prediction | FIP | 61 | 4 | Sensitivity 98.39% |
| Non-FIP | 1 | 70 | Specificity 94.59% | |
| PPV 93.85% | NPV 98.59% | Accuracy 96.32% (91.63–98.8%) | ||
| Reference | ||||
|---|---|---|---|---|
| FIP | Non-FIP | |||
| Prediction | FIP | 86 | 5 | Sensitivity 98.85% |
| Non-FIP | 1 | 80 | Specificity 94.12% | |
| PPV 94.51% | NPV 98.77% | Accuracy 96.51% (92.56–98.71%) | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Dunbar, D.E.; Babayan, S.A.; Krumrie, S.; Rennie, S.; Waugh, E.M.; Hosie, M.J.; Weir, W. Applying Supervised Machine Learning to Effusion Analysis for the Diagnosis of Feline Infectious Peritonitis. Bioengineering 2026, 13, 127. https://doi.org/10.3390/bioengineering13020127
Dunbar DE, Babayan SA, Krumrie S, Rennie S, Waugh EM, Hosie MJ, Weir W. Applying Supervised Machine Learning to Effusion Analysis for the Diagnosis of Feline Infectious Peritonitis. Bioengineering. 2026; 13(2):127. https://doi.org/10.3390/bioengineering13020127
Chicago/Turabian StyleDunbar, Dawn E., Simon A. Babayan, Sarah Krumrie, Sharmila Rennie, Elspeth M. Waugh, Margaret J. Hosie, and William Weir. 2026. "Applying Supervised Machine Learning to Effusion Analysis for the Diagnosis of Feline Infectious Peritonitis" Bioengineering 13, no. 2: 127. https://doi.org/10.3390/bioengineering13020127
APA StyleDunbar, D. E., Babayan, S. A., Krumrie, S., Rennie, S., Waugh, E. M., Hosie, M. J., & Weir, W. (2026). Applying Supervised Machine Learning to Effusion Analysis for the Diagnosis of Feline Infectious Peritonitis. Bioengineering, 13(2), 127. https://doi.org/10.3390/bioengineering13020127







