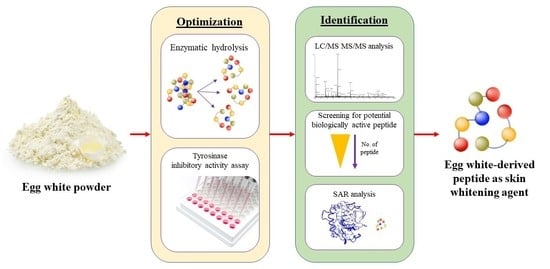
Chicken Egg White—Advancing from Food to Skin Health Therapy: Optimization of Hydrolysis Condition and Identification of Tyrosinase Inhibitor Peptides
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Enzymatic Hydrolysis of Egg White Proteins
2.2.1. Single-Factor Experiment
2.2.2. Optimization of Hydrolysis Conditions for Monophenolase and Diphenolase Inhibitory Activities
2.3. Determination of Tyrosinase Inhibitory Activities
2.3.1. Monophenolase Inhibitory Activity
2.3.2. Diphenolase Inhibitory Activity
2.4. Identification of Bioactive Peptides
2.5. Statistical Analysis
3. Results and Discussion
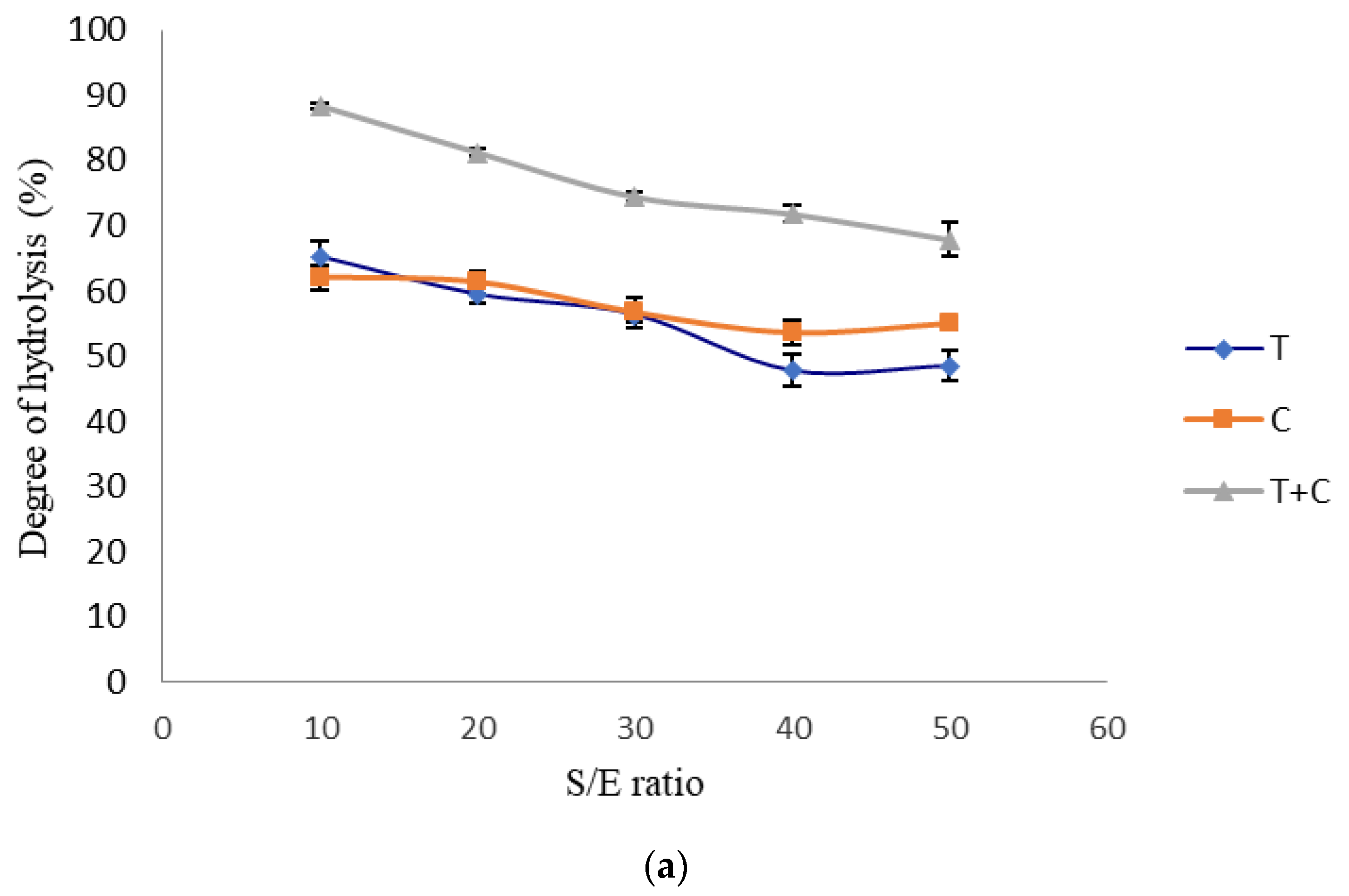
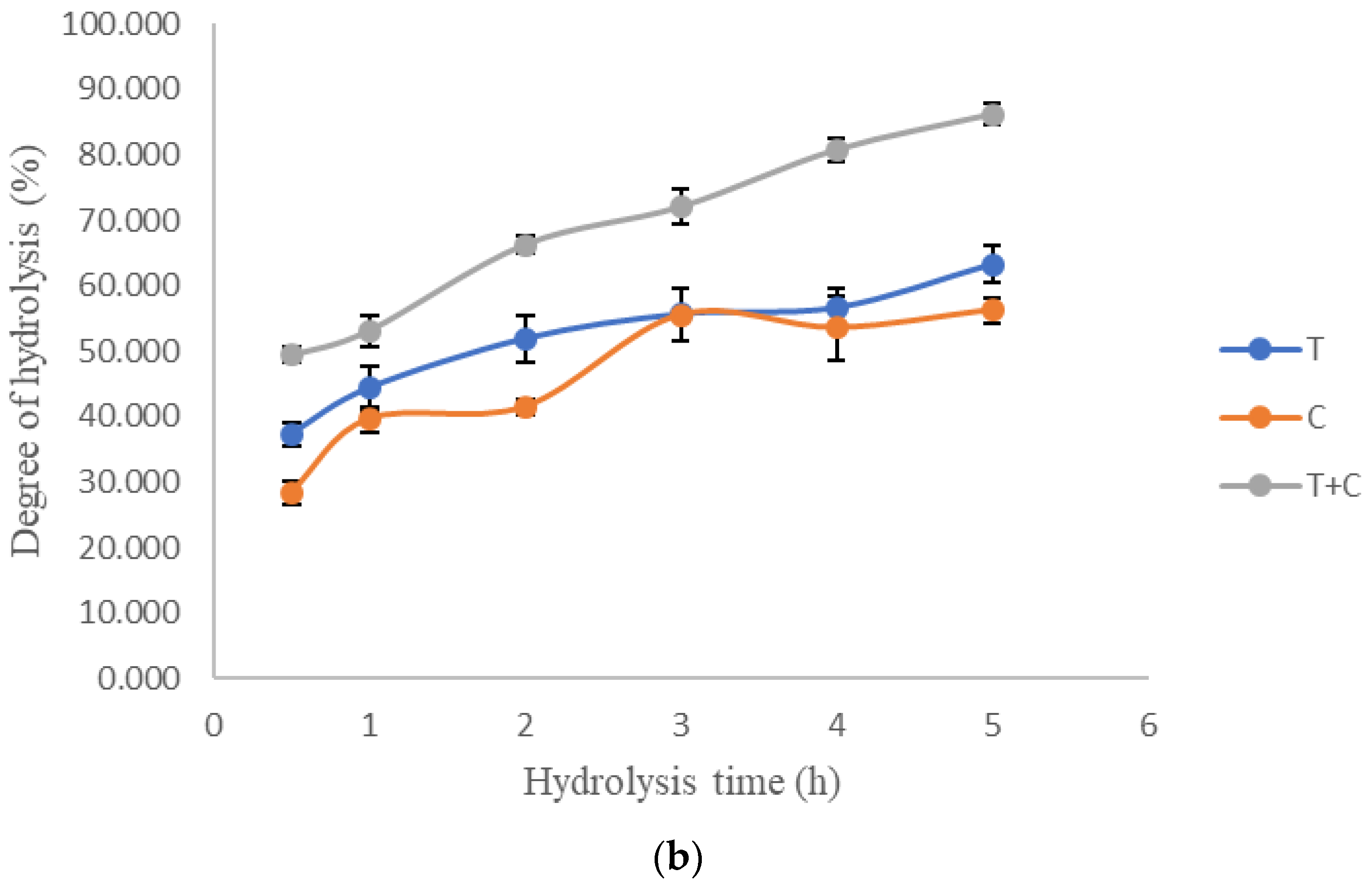
3.1. Single-Factor Experiment
3.2. Optimization of Tyrosinase Inhibitory Activities
3.3. Verification of Predictive Models
3.4. Identification of Bioactive Peptides
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A
| No. | Egg Protein Sequence Coverage | Peptide | Peptide Sequence Number | Score |
|---|---|---|---|---|
| (a) Monophenolase inhibitory activity | ||||
| 1 | Ovalbumin (77%) | ADHPF * | 361–365 | 0.8592 |
| 2 | FDKLPGFGD * | 60–68 | 0.6817 | |
| 3 | FDKLPGFGDSIEAQCGTSVN * | 60–79 | 0.5429 | |
| 4 | AFKDEDTKAMPF * | 188–199 | 0.5332 | |
| 5 | DKLPGFGD * | 61–68 | 0.5174 | |
| 6 | MSALAM | 36–41 | 0.4941 | |
| 7 | RGGLEPINF | 127–135 | 0.4842 | |
| 8 | YPILPEYL | 112–119 | 0.4732 | |
| 9 | AFKDEDTQAMPF | 188–199 | 0.4636 | |
| 10 | VLLPDEVSGLEKLESIINF | 244–262 | 0.4494 | |
| 11 | QVLLPDEVSGLEQLESIINF | 243–262 | 0.4335 | |
| 12 | KDEDTQAMPF | 190–199 | 0.4113 | |
| 13 | FDKLPGFGDSIEAQ | 60–73 | 0.4067 | |
| 14 | YPILPEYLQCVKELY | 112–126 | 0.3960 | |
| 15 | VLLPHEVSGLEQLESIINF | 244–262 | 0.3712 | |
| 16 | LVQLPDEVSGLEQLESIINF | 243–262 | 0.3593 | |
| 17 | LPDEVSGLEQLESIINF | 246–262 | 0.3544 | |
| 18 | MAMGITDVF | 299–307 | 0.3250 | |
| 19 | SSSANLSGISSAESLK | 308–323 | 0.3207 | |
| 20 | LVLLPDEVSGLEQLESIINF | 243–262 | 0.3189 | |
| 21 | VHHANENIFY | 21–30 | 0.3132 | |
| 22 | VTEQESKPVQMMYQIGLF | 210–218 | 0.3126 | |
| 23 | VHHANENIF | 21–29 | 0.2953 | |
| 24 | GGLEPINFQTAADQAR | 128–143 | 0.2810 | |
| 25 | VLLPDEVSGLEQLESIINFE | 244–263 | 0.2738 | |
| 26 | VASMASEKMK | 220–229 | 0.2728 | |
| 27 | AAHAEINEAGR | 330–340 | 0.2679 | |
| 28 | AEERYPILPEYL | 108–119 | 0.2648 | |
| 29 | YPILPEYLQ | 112–120 | 0.2638 | |
| 30 | ISQAVHAAHAEINEAGR | 324–340 | 0.2599 | |
| 31 | SANLSGISSAESLK | 310–323 | 0.2537 | |
| 32 | SWVESQTNGIIR | 148–159 | 0.2523 | |
| 33 | AFKDEDTQAMP | 188–198 | 0.2456 | |
| 34 | SGISSAESLK | 314–323 | 0.2242 | |
| 35 | MAMGITDVFSSSANLSGIS | 299–317 | 0.2239 | |
| 36 | INSWVESQTNGIIR | 146–159 | 0.2193 | |
| 37 | LYAEERYPILPEYL | 106–119 | 0.2015 | |
| 38 | SGISSAESL | 314–322 | 0.1982 | |
| 39 | VLLPDEVSGLEQLESIINFEK | 244–264 | 0.1885 | |
| 40 | VLQPSSVHSQTAM | 161–173 | 0.1836 | |
| 41 | VLLPDEVSGLEQL | 244–256 | 0.1835 | |
| 42 | LSGISSAESLK | 313–323 | 0.1793 | |
| 43 | HAEINEAGR | 332–340 | 0.1789 | |
| 44 | SAEAGVDAASVSEEF | 345–359 | 0.1784 | |
| 45 | VLLPDEVSGLEQLESIIN | 244–261 | 0.1723 | |
| 46 | EVVGSAEAGVDAASVSEEFR | 341–360 | 0.1703 | |
| 47 | NVLQPSSVHSQTAM | 160–173 | 0.1699 | |
| 48 | AMGITDVFSS | 300–309 | 0.1674 | |
| 49 | SSNVMEE | 270–276 | 0.1638 | |
| 50 | EVVGSAEAGVDAASVSEEF | 341–359 | 0.1629 | |
| 51 | SGISSAESLQ | 314–323 | 0.1544 | |
| 52 | ELINSWVESQTNGIIR | 144–159 | 0.1482 | |
| 53 | SGISSAESLE | 314–323 | 0.1469 | |
| 54 | VTEQESKPVQMM | 201–212 | 0.1453 | |
| 55 | AEINEAGR | 333–340 | 0.1437 | |
| 56 | VLKPSSVDSQTAM | 161–173 | 0.1431 | |
| 57 | VESQTNGIIR | 150–159 | 0.1401 | |
| 58 | NVLQPSSVDSQTAM | 160–173 | 0.1362 | |
| 59 | VLLPDEVSGLEQLESR | 244–259 | 0.1282 | |
| 60 | VLLPDEVSGLEKLESIINFEK | 244–264 | 0.1189 | |
| 61 | VASMASEK | 220–227 | 0.1086 | |
| 62 | ISQAVH | 324–329 | 0.1070 | |
| 63 | TSSNVMEER | 269–277 | 0.1050 | |
| 64 | VLLPHEVSGLEQLES | 244–258 | 0.0986 | |
| 65 | QITKPNDVY | 90–98 | 0.0968 | |
| 66 | DILNQITKPNDVY | 86–98 | 0.0951 | |
| 67 | VLLPDEVSGLEQLES | 244–258 | 0.0916 | |
| 68 | NQITKPNDVY | 89–98 | 0.0879 | |
| 69 | LTEWTSSNVMEER | 265–277 | 0.0807 | |
| 70 | VTEQESKPVQMK | 201–212 | 0.0805 | |
| 71 | RVTEQESKPVQM | 200–211 | 0.0787 | |
| 72 | NQITEPNDVY | 89–98 | 0.0742 | |
| 73 | VTEQESKPVQM | 201–211 | 0.0712 | |
| 74 | VTEQESKPV | 201–209 | 0.0395 | |
| 75 | NVLQPSSVDSQTAMVLVNAIVFK | 160–182 | 0.0365 | |
| 76 | NVLQPSSVDSQTAMVLVNAIVF | 160–181 | 0.0218 | |
| 77 | Ovotransferrin (45%) | KDSNVNWNNLK | 458–468 | 0.4863 |
| 78 | GEADAVALHGGLVY | 406–419 | 0.4290 | |
| 79 | NLQMDDFELL | 579–588 | 0.4262 | |
| 80 | NLKMDDFELL | 579–588 | 0.3988 | |
| 81 | KDQLTPSPR | 352–360 | 0.3329 | |
| 82 | VPSLMHSQLY | 329–338 | 0.3155 | |
| 83 | GAIEWEGIESGSVEKAVAK | 155–173 | 0.2986 | |
| 84 | KGEADAVALDGGLVY | 405–419 | 0.2793 | |
| 85 | TAGVCGLVPVMAER | 420–433 | 0.2662 | |
| 86 | VPSLMDSQLY | 329–338 | 0.2612 | |
| 87 | AQSDFGVDTK | 289–298 | 0.2525 | |
| 88 | GAIEWEGIESGSVEQAVAK | 155–173 | 0.2308 | |
| 89 | RVPSLMDSQLY | 328–338 | 0.2249 | |
| 90 | VPVMAER | 427–433 | 0.1784 | |
| 91 | VAAHAVVAR | 266–274 | 0.1704 | |
| 92 | GAIEWEGIESGSVEQAVAE | 155–173 | 0.1683 | |
| 93 | AIEWEGIESGSVEQAVAK | 156–173 | 0.1624 | |
| 94 | LKPIAAEVY | 93–101 | 0.1209 | |
| 95 | KLKPIAAEVY | 92–101 | 0.1184 | |
| 96 | HAVVVRPEK | 611–619 | 0.1049 | |
| 97 | IQHSTVEENTGGK | 559–571 | 0.1005 | |
| 98 | TVNDLQGK | 124–131 | 0.0955 | |
| 99 | AVVVRPEK | 612–619 | 0.0953 | |
| 100 | TVNENAPDQKDEYELL | 231–246 | 0.0946 | |
| 101 | QGIESGSVEQAVAK | 160–173 | 0.0919 | |
| 102 | TDERPASY | 443–450 | 0.0891 | |
| 103 | IKHSTVEENTGGK | 559–571 | 0.0865 | |
| 104 | VAAHAVVARDDNQVEDIW | 266–283 | 0.0847 | |
| 105 | DLTQQER | 44–50 | 0.0846 | |
| 106 | VQHSTVEENTGGK | 559–571 | 0.0835 | |
| 107 | EGIESGSVEQAVAK | 160–173 | 0.0768 | |
| 108 | TVISSLK | 682–688 | 0.0710 | |
| 109 | HTTVNENAPDQKDEYELL | 229–246 | 0.0630 | |
| 110 | EGIESGSVEQAVAE | 160–173 | 0.0600 | |
| 111 | VVVRPEK | 613–619 | 0.0580 | |
| 112 | TVEENTGGK | 563–571 | 0.0491 | |
| 113 | Ovalbumin-related protein Y (32%) | ISDAVHGVF | 324–332 | 0.4692 |
| 114 | MISDAVHGVF | 323–332 | 0.4224 | |
| 115 | VLLPDEVSGLEHIEKTINF | 244–262 | 0.2846 | |
| 116 | HSLELEEFR | 354–362 | 0.2286 | |
| 117 | SLEIADKLY | 99–107 | 0.1898 | |
| 118 | VLLPDEVSGLER | 244–255 | 0.1871 | |
| 119 | VLLPDEVSGLERIEKTIN | 244–261 | 0.1605 | |
| 120 | VLLPDEVSGLERIEK | 244–258 | 0.1316 | |
| 121 | MEVNEEGTEATGSTGAIGNIK | 333–353 | 0.1211 | |
| 122 | TGGVEEVNFK | 127–136 | 0.1200 | |
| 123 | NVATLPAEK | 219–227 | 0.1152 | |
| 124 | Ovalbumin-related protein X (38%) | ILELPFASGDLLML * | 74–87 | 0.8643 |
| 125 | TGISSAESLK | 158–167 | 0.1223 | |
| 126 | NVATLPAEK | 63–71 | 0.1152 | |
| 127 | VLLPDEVSDLER | 88–99 | 0.1084 | |
| 128 | ISQAVH | 168–173 | 0.1070 | |
| 129 | AGSTGVIEDIK | 187–197 | 0.1025 | |
| 130 | VTKQESKPVQM | 45–55 | 0.0833 | |
| 131 | Ovomucoid (42%) | FPNATDKEGK | 32–41 | 0.3269 |
| 132 | DLRPICGTDGVTY | 49–61 | 0.2154 | |
| 133 | VEQGASVDKR | 137–146 | 0.0903 | |
| 134 | VEQGASVDER | 137–146 | 0.0734 | |
| 135 | Ovomucin (16%) | DGSGGCIPK * | 814–822 | 0.6103 |
| 136 | VTDSF | 1591–1595 | 0.2004 | |
| 137 | SNSLVILTQA | 1494–1503 | 0.1384 | |
| 138 | IQEIATDPGAEK | 941–952 | 0.1215 | |
| 139 | Cystatin (12%) | LLGAPVPVDENDEGLQR | 30–46 | 0.2450 |
| (b) Diphenolase inhibitory activity | ||||
| 1 | Ovalbumin (81%) | ADHPFLF * | 361–367 | 0.9699 |
| 2 | MYQIGLFR * | 212–219 | 0.8011 | |
| 3 | SMLVLLPDEVSGLEQLESIINFEK * | 241–264 | 0.6983 | |
| 4 | FDKLPGFGD * | 60–68 | 0.6817 | |
| 5 | HIATNAVLFFGR * | 371–382 | 0.5209 | |
| 6 | FKDEDTQAMPFR * | 189–200 | 0.5001 | |
| 7 | DEDTKAMPFR | 191–200 | 0.4978 | |
| 8 | ILELPFASGTMS | 230–241 | 0.4972 | |
| 9 | DEDTQAMPFR | 191–200 | 0.4842 | |
| 10 | MLVLLPDEVSGLEQLESIINFEK | 242–264 | 0.4842 | |
| 11 | YPILPEYLQCVK | 112–123 | 0.4769 | |
| 12 | AFKDEDTQAMPFR | 188–200 | 0.4702 | |
| 13 | SSSANLSGISSAESLK | 308–323 | 0.3207 | |
| 14 | SQAVHAAHAEINEAGR | 325–340 | 0.2965 | |
| 15 | VTEQESKPVQMMYQIGLFR | 201–219 | 0.2886 | |
| 16 | GGLEPINFQTAADQAR | 128–143 | 0.2810 | |
| 17 | SQTAMVLVNAIVFK | 169–182 | 0.2781 | |
| 18 | VASMASEKMK | 220–229 | 0.2728 | |
| 19 | AAHAEINEAGR | 330–340 | 0.2679 | |
| 20 | YPILPEYLQ | 112–120 | 0.2638 | |
| 21 | QAVHAAHAEINEAGR | 326–340 | 0.2606 | |
| 22 | ISQAVHAAHAEINEAGR | 324–340 | 0.2599 | |
| 23 | EAQCGTSVNVHSSLR | 71–85 | 0.2554 | |
| 24 | SANLSGISSAESLK | 310–323 | 0.2537 | |
| 25 | SSANLSGISSAESLK | 309–323 | 0.2497 | |
| 26 | EVCGSAEAGVDAASVSEEFR | 341–360 | 0.2424 | |
| 27 | GLEPINFQTAADQAR | 129–143 | 0.2420 | |
| 28 | VLLPDEVSGLEQLESIINFEQ | 244–264 | 0.2410 | |
| 29 | VLVNANVFK | 174–182 | 0.2362 | |
| 30 | NSQAVHAAHAEINEAGR | 324–340 | 0.2273 | |
| 31 | ISQAVHAAHAEIN | 324–336 | 0.2267 | |
| 32 | DILNQITKPNDVYSFSLASR | 86–105 | 0.2229 | |
| 33 | FQTAADQAR | 135–143 | 0.2202 | |
| 34 | VLVNAIVFK | 174–182 | 0.2049 | |
| 35 | MAMGITDVFSSSANLSGISSAESLK | 299–323 | 0.1963 | |
| 36 | AVHAAHAEINEAGR | 327–340 | 0.1935 | |
| 37 | SVNVHSSLR | 77–85 | 0.1930 | |
| 38 | LCEWTSSNVMEER | 265–277 | 0.1886 | |
| 39 | VLLPDEVSGLEQLESIINFEK | 244–264 | 0.1885 | |
| 40 | DLEPINFQTAADQAR | 129–143 | 0.1854 | |
| 41 | LSGISSAESLK | 313–323 | 0.1793 | |
| 42 | HAEINEAGR | 332–340 | 0.1789 | |
| 43 | LEPINFQTAADQAR | 130–143 | 0.1739 | |
| 44 | EVVGSAEAGVDAASVSEEFR | 341–360 | 0.1703 | |
| 45 | DILNQITKPNDVYSF | 86–100 | 0.1698 | |
| 46 | EAGVDAASVSEEFR | 437–360 | 0.1664 | |
| 47 | AEAGVDAASVSEEFR | 346–360 | 0.1641 | |
| 48 | VHAAHAEINEAGR | 328–340 | 0.1633 | |
| 49 | EVVGAAEAGVDAASVSEEFR | 341–360 | 0.1628 | |
| 50 | EPINFQTAADQAR | 131–143 | 0.1588 | |
| 51 | ISQAVHAAH | 324–332 | 0.1572 | |
| 52 | ELINSWVESQTNGIIR | 144–159 | 0.1482 | |
| 53 | VTEQESKPVQMM | 201–212 | 0.1453 | |
| 54 | VTEQESKPVQMMY | 201–213 | 0.1416 | |
| 55 | RVTEQESKPVQMMY | 200–213 | 0.1413 | |
| 56 | GITHVFSSSANLSGISSAESLK | 302–323 | 0.1401 | |
| 57 | VLVNAIVFE | 174–182 | 0.1401 | |
| 58 | AMGNTDVFSSSANLSGISSAESLK | 300–323 | 0.1395 | |
| 59 | NVLQPSSVDSQTAM | 160–173 | 0.1362 | |
| 60 | EWTSSNVMEER | 267–277 | 0.1280 | |
| 61 | GTSVNVHSSLR | 75–85 | 0.1274 | |
| 62 | LQPSSVDSQTAMVLVNAIVFK | 162–182 | 0.1259 | |
| 63 | AMGITDVFSSSANLSGISSAESLK | 300–323 | 0.1239 | |
| 64 | VTEQESKPVKM | 201–211 | 0.1172 | |
| 65 | GITDVFSSSANLSGISSAESLK | 302–323 | 0.1089 | |
| 66 | VASMASEK | 200–227 | 0.1086 | |
| 67 | TSSNVMEER | 269–277 | 0.1050 | |
| 68 | DILNQITKPNDVY | 86–98 | 0.0951 | |
| 69 | LTEWTSSNVMEER | 265–277 | 0.0807 | |
| 70 | ELINSWVESQTN | 144–155 | 0.0798 | |
| 71 | TEWTSSNVMEER | 266–277 | 0.0763 | |
| 72 | LYAEER | 106–111 | 0.0759 | |
| 73 | EAGVDAASVS | 347–356 | 0.0755 | |
| 74 | VASMASEE | 220–227 | 0.0721 | |
| 75 | VTEQESKPVQM | 201–211 | 0.0712 | |
| 76 | TQINK | 52–56 | 0.0607 | |
| 77 | NKVVR | 55–59 | 0.0591 | |
| 78 | VTEQESKPVQMMYQIGLFRVASMASEK | 201–227 | 0.0512 | |
| 79 | NVLKPSSVDSQTAMVLVNAIVFK | 160–182 | 0.0486 | |
| 80 | NVLQPSSVDSQTAMVLVNAIVFK | 160–182 | 0.0365 | |
| 81 | Ovotransferrin (63%) | SDFHLFGPPGK * | 299–309 | 0.8619 |
| 82 | SGYSGAFHCLK * | 208–218 | 0.8239 | |
| 83 | SGAFHCLK * | 211–218 | 0.8038 | |
| 84 | CQLCQGSGGIPPEK * | 518–531 | 0.6874 | |
| 85 | YFGYTGALRCLV * | 540–551 | 0.6541 | |
| 86 | DLLFKDSAIMLK * | 216–327 | 0.6400 | |
| 87 | SGALHCLK * | 211–218 | 0.5971 | |
| 88 | FMMFESQNKDLLFK * | 644–657 | 0.5892 | |
| 89 | YFGYTGALR * | 540–548 | 0.5801 | |
| 90 | KDSNVNWNNLK | 458–468 | 0.4863 | |
| 91 | DDNKVEDIWSFLSK | 275–288 | 0.4641 | |
| 92 | SGGIPPEK | 524–531 | 0.4622 | |
| 93 | NIPIGTLLHRG | 145–155 | 0.4328 | |
| 94 | DLLFKDSAIMLE | 316–327 | 0.4322 | |
| 95 | SAIQSMR | 345–351 | 0.4293 | |
| 96 | ANVMDYR | 595–601 | 0.4255 | |
| 97 | FFSASCVPGATIEQK | 174–188 | 0.4152 | |
| 98 | NIPIGTLLHR | 145–154 | 0.3756 | |
| 99 | TSCHTGLGR | 132–140 | 0.3624 | |
| 100 | GAIEWEGIESGSVEQAVAKFFSASCVPGA | 155–183 | 0.3503 | |
| 101 | FMMFESKNK | 644–652 | 0.3416 | |
| 102 | KDQLTPSPR | 352–360 | 0.3329 | |
| 103 | FMMFESQNK | 644–652 | 0.3265 | |
| 104 | GDVAFVK | 222–228 | 0.3236 | |
| 105 | EAGLAPYK | 85–92 | 0.3173 | |
| 106 | GLIHNR | 488–493 | 0.3124 | |
| 107 | VEDIWSFLSE | 278–288 | 0.3066 | |
| 108 | GAIEWEGIESGSVEKAVAK | 155–173 | 0.2986 | |
| 109 | DGKGDVAFVK | 219–228 | 0.2939 | |
| 110 | SDFGVDTK | 291–298 | 0.2580 | |
| 111 | AQSDFGVDTK | 289–298 | 0.2525 | |
| 112 | GDVAFIKHSTVEENTGGK | 554–571 | 0.2429 | |
| 113 | TDERPASYF | 443–451 | 0.2356 | |
| 114 | GAIEWEGIESGSVEQAVAK | 155–173 | 0.2308 | |
| 115 | AAHAVVAR | 267–274 | 0.2295 | |
| 116 | GANEWEGIESGSVEQAVAK | 155–173 | 0.2076 | |
| 117 | GAIEWEGNESGSVEQAVAK | 155–173 | 0.2067 | |
| 118 | AQSDFGVDTE | 289–298 | 0.1852 | |
| 119 | FYTVISSLKTCNPS | 680–693 | 0.1792 | |
| 120 | VAAHAVVAR | 266–274 | 0.1704 | |
| 121 | GAIEWEGIESGSVEQAVAE | 155–173 | 0.1683 | |
| 122 | FGVHGSEK | 634–641 | 0.1601 | |
| 123 | FGVNGSEKSK | 634–643 | 0.1364 | |
| 124 | RFGVNGSEK | 633–641 | 0.1351 | |
| 125 | GDVAFVQHSTVEENTGGK | 554–571 | 0.1323 | |
| 126 | KGTEFTVNDLQGK | 119–131 | 0.1190 | |
| 127 | GTEFTVNDLQGK | 120–131 | 0.1166 | |
| 128 | DVAFIQHSTVEENTGGK | 555–571 | 0.1158 | |
| 129 | KCVAS | 531–535 | 0.1136 | |
| 130 | TDERPASY | 443–450 | 0.0891 | |
| 131 | LKPIAAEVYEHTEGSTTSYY | 93–112 | 0.0753 | |
| 132 | YTVISSLK | 681–688 | 0.0742 | |
| 133 | CTVVDETK | 390–397 | 0.0495 | |
| 134 | HTTVNENAPDQK | 229–240 | 0.0390 | |
| 135 | Ovomucoid (38%) | FPNATDKEGK | 32–41 | 0.3269 |
| 136 | VMVLCNR | 107–113 | 0.2887 | |
| 137 | GASVDKR | 140–146 | 0.2203 | |
| 138 | SIEFGTNISK | 71–80 | 0.1899 | |
| 139 | AVVESNGTLTLSHFGK | 194–209 | 0.1597 | |
| 140 | FCNAVVES | 191–198 | 0.1383 | |
| 141 | VEQGASVDKR | 137–146 | 0.0903 | |
| 142 | CAHKVEQ | 133–139 | 0.0902 | |
| 143 | VEQGASVDER | 137–146 | 0.0734 | |
| 144 | CNFCNAVVESNGTLTLSHFGK | 189–209 | 0.0623 | |
| 145 | VEQGASVDK | 137–145 | 0.0585 | |
| 146 | VEQGASVDE | 137–145 | 0.0467 | |
| 147 | Ovomucin (26%) | SGGQFSLTSTVKVC * | 1973–1986 | 0.5560 |
| 148 | SSCEDCVCT * | 1888–1896 | 0.5459 | |
| 149 | FNCSSAGPGAIGSEC * | 771–785 | 0.5429 | |
| 150 | SSCICS * | 270–275 | 0.5397 | |
| 151 | FDGRSR * | 1000–1005 | 0.5330 | |
| 152 | PAQEQLM | 1255–1261 | 0.3931 | |
| 153 | KSLSICSLK | 916–924 | 0.3501 | |
| 154 | VTSDGCCK | 2001–2008 | 0.3200 | |
| 155 | LEGCYPECS | 1163–1171 | 0.3088 | |
| 156 | ECGNSC | 312–317 | 0.2923 | |
| 157 | EPSELCK | 1629–1635 | 0.2680 | |
| 158 | TCTCNKR | 846–852 | 0.2315 | |
| 159 | DTCADPE | 319–325 | 0.2167 | |
| 160 | VTDSF | 1591–1595 | 0.2004 | |
| 161 | TATGAVEDSAAAFGNSWE | 547–564 | 0.1661 | |
| 162 | GTCSTYS | 2044–2050 | 0.1278 | |
| 163 | IQEIATDPGAEK | 941–952 | 0.1215 | |
| 164 | EVIVDTLLSR | 1722–1731 | 0.1101 | |
| 165 | VQVSTVGR | 28–35 | 0.0990 | |
| 166 | AVTGTN | 1901–1906 | 0.0685 | |
| 167 | Lysozyme (54%) | GYSLGNWVCAAK * | 40–51 | 0.6563 |
| 168 | HGLDNYRG | 33–40 | 0.4763 | |
| 169 | HGLDNYR | 33–39 | 0.3753 | |
| 170 | GTDVQAWIR | 135–143 | 0.3365 | |
| 171 | GILQINSR | 72–79 | 0.2008 | |
| 172 | FESNFNTQATNR | 52–63 | 0.1820 | |
| 173 | SNFNTQATNR | 54–63 | 0.1487 | |
| 174 | ILQINSR | 73–79 | 0.1212 | |
| 175 | NTDGSTDYGILQINSR | 64–79 | 0.0900 | |
| 176 | Ovalbumin-related protein Y (28%) | VHNLFK | 80–85 | 0.3580 |
| 177 | HSLELEEFR | 354–362 | 0.2286 | |
| 178 | VATLPAEKMK | 220–229 | 0.1937 | |
| 179 | KFYTGGVEEVNFK | 124–136 | 0.1917 | |
| 180 | VLLPDEVSGLER | 244–255 | 0.1871 | |
| 181 | FYTGGVEEVNFK | 125–136 | 0.1570 | |
| 182 | ATGSTGAI | 342–349 | 0.1262 | |
| 183 | TESQMK | 50–55 | 0.0899 | |
| 184 | Ovostatin (2%) | EKMAPALRLLV | 538–548 | 0.4863 |
| 185 | LVDKDNSPISNK | 379–390 | 0.1410 | |
| 186 | Ovalbumin-related protein X (29%) | HNPTNTIVYFGR | 217–228 | 0.2810 |
| 187 | ILELPFASGDLSMLVLLPDEVSDLER | 74–99 | 0.1682 | |
| 188 | ILELPFASGDLSMLVLLPDEVSHLER | 74–99 | 0.1607 | |
| 189 | ISQAVHGAFMELSEDGIEMAGSTGVIEDIK | 168–197 | 0.0118 | |
References
- Sánchez-Ferrer, Á.; Rodríguez-López, J.N.; García-Cánovas, F.; García-Carmona, F. Tyrosinase: A comprehensive review of its mechanism. BBA Protein Struct. Mol. Enzymol. 1995, 1247, 1–11. [Google Scholar] [CrossRef]
- Michalek, I.M.; Benn, E.K.; dos Santos, F.L.C.; Gordon, S.; Wen, C.; Liu, B. A systematic review of global legal regulations on the permissible level of heavy metals in cosmetics with particular emphasis on skin lightening products. Environ. Res. 2019, 170, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Westerhof, W.; Kooyers, T.J. Hydroquinone and its analogues in dermatology—A potential health risk. J. Cosmet Dermatol. 2005, 4, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.C. Tyrosinase Inhibitor Extract. U.S. Patent 7,125,572, 24 October 2006. [Google Scholar]
- Ubeid, A.A.; Zhao, L.; Wang, Y.; Hantash, B.M. Short-sequence oligopeptides with inhibitory activity against mushroom and human tyrosinase. J. Investig. Dermatol. 2009, 129, 2242–2249. [Google Scholar] [CrossRef]
- Kubglomsong, S.; Theerakulkait, C.; Reed, R.L.; Yang, L.; Maier, C.S.; Stevens, J.F. Isolation and identification of tyrosinase-inhibitory and copper-chelating peptides from hydrolyzed rice-bran-derived albumin. J. Agric. Food Chem. 2018, 66, 8346–8354. [Google Scholar] [CrossRef] [PubMed]
- Marini, A.; Farwick, M.; Grether-Beck, S.; Brenden, H.; Felsner, I.; Jaenicke, T.; Weber, M.; Schild, J.; Maczkiewitz, U.; Köhler, T.; et al. Modulation of skin pigmentation by the tetrapeptide PKEK: In vitro and in vivo evidence for skin whitening effects. Exp. Dermatol. 2012, 21, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Albericio, F.; Kruger, H.G. Therapeutic peptides. Future Med. Chem. 2012, 4, 1527–1531. [Google Scholar] [CrossRef]
- Ochiai, A.; Tanaka, S.; Tanaka, T.; Taniguchi, M. Rice bran protein as a potent source of antimelanogenic peptides with tyrosinase inhibitory activity. J. Nat. Prod. 2016, 79, 2545–2551. [Google Scholar] [CrossRef]
- Oh, G.W.; Ko, S.C.; Heo, S.Y.; Nguyen, V.T.; Kim, G.; Jang, C.H.; Park, W.S.; Choi, I.-W.; Qian, Z.-J.; Jung, W.K. A novel peptide purified from the fermented microalga Pavlova lutheri attenuates oxidative stress and melanogenesis in B16F10 melanoma cells. Process. Biochem. 2015, 50, 1318–1326. [Google Scholar] [CrossRef]
- Eriksson, L.; Johansson, E.; Kettaneh-Wold, N.; Wikström, C.; Wold, S. D-optimal design. In Design of Experiments: Principles and Applications; UMETRICS: Stockholm, Sweden, 2000. [Google Scholar]
- Alkali, L.M.T.B.M. D-optimal design optimization of Jatropha curcas L. seed oil hydrolysis via alkali-catalyzed reactions. Sains Malays. 2012, 41, 731–738. [Google Scholar]
- Bahadi, M.; Yusoff, M.F.; Derawi, J.S.D. Optimization of response surface methodology by d-optimal design for alkaline hydrolysis of crude palm kernel oil. Sains Malays. 2020, 49, 29–41. [Google Scholar] [CrossRef]
- Olsen, J.V.; Ong, S.E.; Mann, M. Trypsin cleaves exclusively C-terminal to arginine and lysine residues. Mol. Cell. Proteom. 2004, 3, 608–614. [Google Scholar] [CrossRef] [PubMed]
- Blow, D.M. The Structure of Chymotrypsin. In The Enzymes, 3rd ed.; Boyer, P.D., Ed.; Academic Press: New York, NY, USA, 1971; pp. 185–212. [Google Scholar]
- Hsiao, N.W.; Tseng, T.S.; Lee, Y.C.; Chen, W.C.; Lin, H.H.; Chen, Y.R.; Wang, Y.-T.; Hsu, H.-J.; Tsai, K.C. Serendipitous discovery of short peptides from natural products as tyrosinase inhibitors. J. Chem. Inf. Model. 2014, 54, 3099–3111. [Google Scholar] [CrossRef] [PubMed]
- Schurink, M.; van Berkel, W.J.; Wichers, H.J.; Boeriu, C.G. Novel peptides with tyrosinase inhibitory activity. Peptides 2007, 28, 485–495. [Google Scholar] [CrossRef]
- Miguel, M.; Recio, I.; Gomez-Ruiz, J.A.; Ramos, M.; Lopez-Fandino, R. Angiotensin I–converting enzyme inhibitory activity of peptides derived from egg white proteins by enzymatic hydrolysis. J. Food Protect. 2004, 67, 1914–1920. [Google Scholar] [CrossRef]
- Baharuddin, N.A.; Halim NR, A.; Sarbon, N.M. Effect of degree of hydrolysis (DH) on the functional properties and angiotensin I-converting enzyme (ACE) inhibitory activity of eel (Monopterus sp.) protein hydrolysate. Int. Food Res. J. 2016, 23, 1424–1431. [Google Scholar]
- Takahashi, M.; Takara, K.; Toyozato, T.; Wada, K. A novel bioactive chalcone of Morus australis inhibits tyrosinase activity and melanin biosynthesis in B16 melanoma cells. J. Oleo Sci. 2012, 61, 585–592. [Google Scholar] [CrossRef]
- Siow, H.L.; Gan, C.Y. Extraction of antioxidative and antihypertensive bioactive peptides from Parkia speciosa seeds. Food Chem. 2013, 141, 3435–3442. [Google Scholar] [CrossRef]
- Ma, B.; Zhang, K.; Hendrie, C.; Liang, C.; Li, M.; Doherty-Kirby, A.; Lajoie, G. PEAKS: Powerful software for peptide de novo sequencing by tandem mass spectrometry. Rapid Commun. Mass Spectrom. 2003, 17, 2337–2342. [Google Scholar] [CrossRef]
- Mooney, C.; Haslam, N.J.; Pollastri, G.; Shields, D.C. Towards the improved discovery and design of functional peptides: Common features of diverse classes permit generalized prediction of bioactivity. PLoS ONE 2012, 7, e45012. [Google Scholar] [CrossRef]
- Trabuco, L.G.; Lise, S.; Petsalaki, E.; Russell, R.B. PepSite: Prediction of peptide-binding sites from protein surfaces. Nucleic Acids Res. 2012, 40, W423–W427. [Google Scholar] [CrossRef] [PubMed]
- Abdou, A.M.; Kim, M.; Sato, K. Functional proteins and peptides of hen’s egg origin. In Bioactive Food Peptides in Health and Disease; Blanca, H.L., Hsieh, C.C., Eds.; InTech: Rijeka, Croatia, 2013; pp. 115–144. [Google Scholar]
- Akazawa, T.; Ogawa, M.; Hayakawa, S. Migration of chicken egg-white protein ovalbumin-related protein X and its alteration in heparin-binding affinity during embryogenesis of fertilized egg. Poult. Sci. 2019, 98, 5100–5108. [Google Scholar] [CrossRef] [PubMed]
- Hirose, J.; Doi, Y.; Kitabatake, N.; Narita, H. Ovalbumin-related gene Y protein bears carbohydrate chains of the ovomucoid type. Biosci. Biotechnol. Biochem. 2006, 70, 144–151. [Google Scholar] [CrossRef][Green Version]
- Abeyrathne, E.D.N.S.; Lee, H.Y.; Jo, C.; Nam, K.C.; Ahn, D.U. Enzymatic hydrolysis of ovalbumin and the functional properties of the hydrolysates. Poult. Sci. 2014, 93, 2678–2686. [Google Scholar] [CrossRef] [PubMed]
- Stevens, L. Egg white proteins. Comp. Biochem. Phys. B Comp. Biochem. 1991, 100, 1–9. [Google Scholar] [CrossRef]
- Custódio, M.F.; Goulart, A.J.; Marques, D.P.; Giordano, R.C.; Giordano, R.D.L.C.; Monti, R. Hydrolysis of cheese whey proteins with trypsin, chymotrypsin and carboxypeptidase A. Aliment. Nutr. Araraquara 2009, 16, 105–109. [Google Scholar]
- Noh, D.O.; Suh, H.J. Preparation of egg white liquid hydrolysate (ELH) and its radical-scavenging activity. Prev. Nutr. Food Sci. 2015, 20, 183. [Google Scholar] [CrossRef]
- De Castro, R.J.S.; Sato, H.H. A response surface approach on optimization of hydrolysis parameters for the production of egg white protein hydrolysates with antioxidant activities. Biocatal. Agric. Biotechnol. 2015, 4, 55–62. [Google Scholar] [CrossRef]
- Chen, C.; Chi, Y.J.; Zhao, M.Y.; Xu, W. Influence of degree of hydrolysis on functional properties, antioxidant and ACE inhibitory activities of egg white protein hydrolysate. Food Sci. Biotechnol. 2012, 21, 27–34. [Google Scholar] [CrossRef]
- Garg, S.; Apostolopoulos, V.; Nurgali, K.; Mishra, V.K. Evaluation of in silico approach for prediction of presence of opioid peptides in wheat. J. Funct. Foods 2018, 41, 34–40. [Google Scholar] [CrossRef]
- Mudgil, P.; Kamal, H.; Yuen, G.C.; Maqsood, S. Characterization and identification of novel antidiabetic and anti-obesity peptides from camel milk protein hydrolysates. Food Chem. 2018, 259, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Salim MA, S.M.; Gan, C.Y. Dual-function peptides derived from egg white ovalbumin: Bioinformatics identification with validation using in vitro assay. J. Funct. Foods 2020, 64, 103618. [Google Scholar]
- Strothkamp, K.G.; Jolley, R.L.; Mason, H.S. Quaternary structure of mushroom tyrosinase. Biochem. Bioph. Res. Co. 1976, 70, 519–524. [Google Scholar]
- Ismaya, W.T.; Rozeboom, H.J.; Weijn, A.; Mes, J.J.; Fusetti, F.; Wichers, H.J.; Dijkstra, B.W. Crystal structure of Agaricus bisporus mushroom tyrosinase: Identity of the tetramer subunits and interaction with tropolone. Biochemistry 2011, 50, 5477–5486. [Google Scholar]
- Drickamer, K.; Taylor, M.E. Recent insights into structures and functions of C-type lectins in the immune system. Curr. Opin. Struct. Biol. 2015, 34, 26–34. [Google Scholar] [PubMed]
- Weijn, A.; Bastiaan-Net, S.; Wichers, H.J.; Mes, J.J. Melanin biosynthesis pathway in Agaricus bisporus mushrooms. Fungal Genet. Biol. 2013, 55, 42–53. [Google Scholar]
- Flurkey, W.H.; Inlow, J.K. Proteolytic processing of polyphenol oxidase from plants and fungi. J. Inorg. Biochem. 2008, 102, 2160–2170. [Google Scholar] [CrossRef]
- Kanteev, M.; Goldfeder, M.; Fishman, A. Structure–function correlations in tyrosinases. Protein Sci. 2015, 24, 1360–1369. [Google Scholar]
- Valley, C.C.; Cembran, A.; Perlmutter, J.D.; Lewis, A.K.; Labello, N.P.; Gao, J.; Sachs, J.N. The methionine-aromatic motif plays a unique role in stabilizing protein structure. J. Biol. Chem. 2012, 287, 34979–34991. [Google Scholar]
- Kanteev, M.; Goldfeder, M.; Chojnacki, M.; Adir, N.; Fishman, A. The mechanism of copper uptake by tyrosinase from Bacillus megaterium. JBIC J. Biol. Inorg. Chem. 2013, 18, 895–903. [Google Scholar]
- Hassani, S.; Haghbeen, K.; Fazli, M. Non-specific binding sites help to explain mixed inhibition in mushroom tyrosinase activities. Eur. J. Med. Chem. 2016, 122, 138–148. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.J.; Noh, S.G.; Park, Y.; Kang, D.; Chun, P.; Chung, H.Y.; Moon, H.R. In vitro and in silico insights into tyrosinase inhibitors with (E)-benzylidene-1-indanone derivatives. Comput. Struct. Biotechnol. J. 2019, 17, 1255–1264. [Google Scholar] [CrossRef] [PubMed]
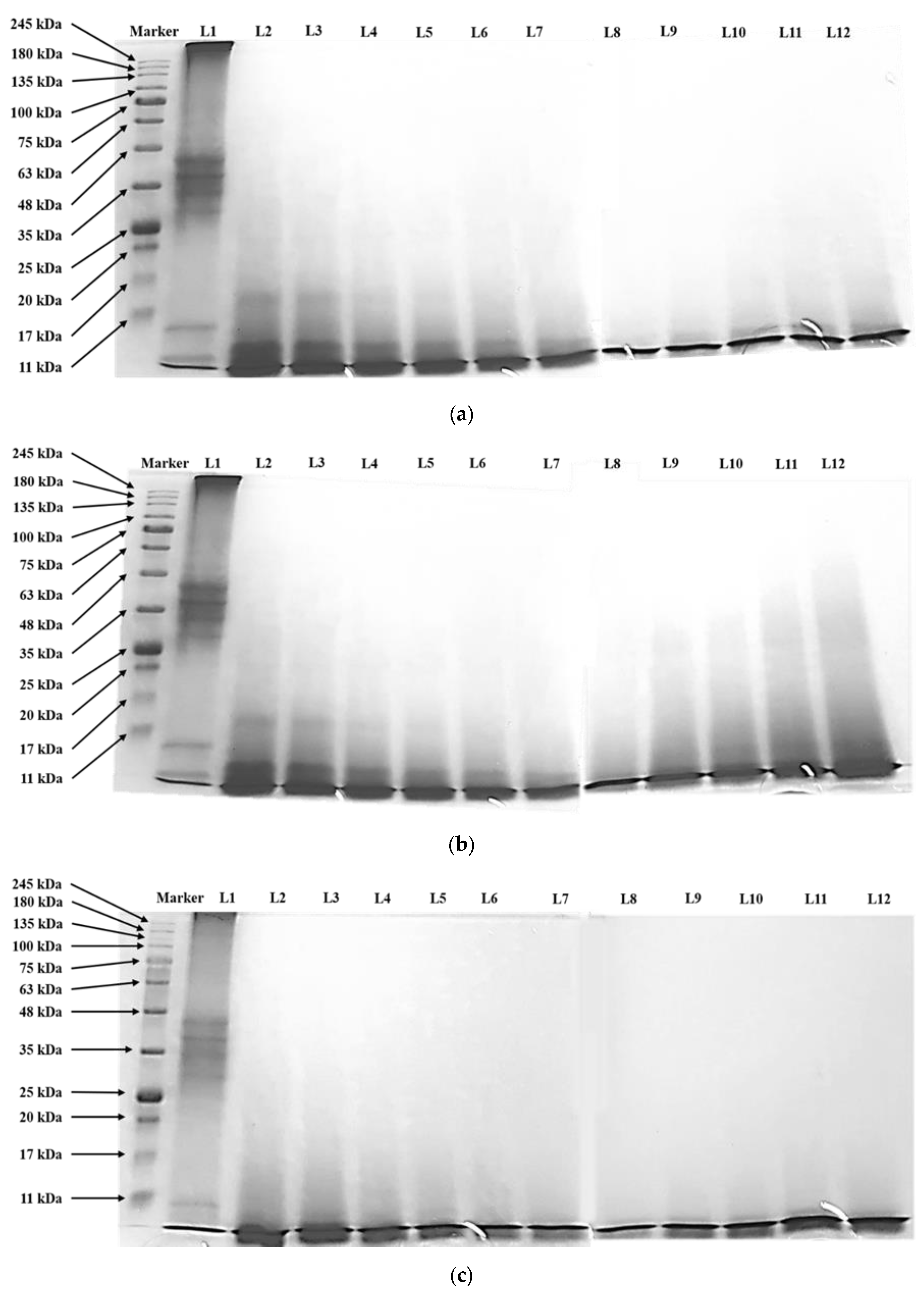

| Variable | Coded Variable | Coded Variable Level | ||||
|---|---|---|---|---|---|---|
| −1 | −0.5 | 0 | 0.5 | 1 | ||
| Trypsin composition (%) | X1 | 0 | 25 | 50 | 75 | 100 |
| Chymotrypsin composition (%) | X2 | 0 | 25 | 50 | 75 | 100 |
| S/E ratio (w/w) | X3 | 10 | 15 | 20 | 25 | 30 |
| Hydrolysis time (h) | X4 | 2 | 2.75 | 3.5 | 4.25 | 5 |
| S/E Ratio, X3 | Hydrolysis Time, X4 (h) | (a) | (b) | |||||
|---|---|---|---|---|---|---|---|---|
| Run | Enzyme Composition (%) | Monophenolase Inhibitory Activity (%) | Diphenolase Inhibitory Activity (%) | |||||
| Trypsin, X1 | Chymotrypsin, X2 | Experimental (y1) | Predicted * (y0) | Experimental (z1) | Predicted # (z0) | |||
| 1 | 100 | 0 | 10 | 5 | 32.5 ± 2.8 | 35.6 | 30.7 ± 1.7 | 33.6 |
| 2 | 50 | 50 | 30 | 2 | 20.8 ± 1.8 | 20.2 | 44.3 ± 4.5 | 44.1 |
| 3 | 0 | 100 | 20 | 3.5 | 15.2 ± 1.4 | 12.7 | 31.4 ± 3.5 | 32.1 |
| 4 | 0 | 100 | 10 | 2 | 31.6 ± 2.4 | 30.5 | 34.2 ± 4.0 | 33.8 |
| 5 | 100 | 0 | 30 | 2 | 24.7 ± 1.3 | 24.3 | 35.7 ± 2.1 | 36.1 |
| 6 | 0 | 100 | 30 | 5 | 39.1 ± 3.4 | 38.5 | 40.6 ± 1.8 | 40.9 |
| 7 | 0 | 100 | 30 | 2 | 18.9 ± 1.8 | 19.6 | 30.2 ± 2.3 | 31.1 |
| 8 | 100 | 0 | 20 | 2 | 29.5 ± 4.7 | 30.7 | 45.5 ± 3.6 | 48.1 |
| 9 | 0 | 100 | 30 | 2 | 20.0 ± 3.8 | 19.6 | 32.0 ± 0.5 | 31.1 |
| 10 | 50 | 50 | 10 | 5 | 37.2 ± 2.5 | 37.5 | 32.8 ± 0.9 | 31.7 |
| 11 | 0 | 100 | 10 | 2 | 30.2 ± 5.0 | 30.5 | 33.7 ± 1.7 | 33.8 |
| 12 | 50 | 50 | 30 | 5 | 26.1 ± 3.7 | 25.3 | 28.8 ± 2.7 | 30.0 |
| 13 | 100 | 0 | 30 | 3.5 | 26.3 ± 3.5 | 27.3 | 38.1 ± 2.4 | 37.3 |
| 14 | 100 | 0 | 10 | 2 | 30.3 ± 4.5 | 32.8 | 35.8 ± 3.9 | 35.6 |
| 15 | 50 | 50 | 20 | 2 | 15.8 ± 3.8 | 16.9 | 44.7 ± 3.9 | 42.5 |
| 16 | 50 | 50 | 30 | 3.5 | 19.2 ± 2.8 | 21.0 | 43.4 ± 3.8 | 44.3 |
| 17 | 75 | 25 | 25 | 2.75 | 26.1 ± 3.5 | 23.5 | 49.6 ± 3.2 | 47.0 |
| 18 | 0 | 100 | 20 | 5 | 21.8 ± 3.6 | 23.9 | 36.2 ± 3.4 | 35.1 |
| 19 | 25 | 75 | 15 | 2.75 | 19.0 ± 2.1 | 20.9 | 31.8 ± 2.5 | 35.0 |
| 20 | 0 | 100 | 30 | 5 | 38.4 ± 0.6 | 38.5 | 41.6 ± 2.2 | 40.9 |
| 21 | 0 | 100 | 10 | 5 | 43.4 ± 4.0 | 41.5 | 46.6 ± 2.0 | 48.7 |
| 22 | 0 | 100 | 10 | 3.5 | 37.4 ± 4.5 | 39.7 | 41.7 ± 2.7 | 39.3 |
| 23 | 75 | 25 | 20 | 5 | 48.0 ± 4.1 | 48.1 | 35.9 ± 2.8 | 34.5 |
| 24 | 100 | 0 | 30 | 5 | 44.7 ± 0.2 | 44.4 | 25.0 ± 2.3 | 25.2 |
| 25 | 100 | 0 | 10 | 5 | 38.7 ± 0.7 | 35.6 | 35.7 ± 4.2 | 33.6 |
| 26 | 100 | 0 | 10 | 2 | 35.2 ± 1.3 | 32.8 | 36.9 ± 1.4 | 35.6 |
| 27 | 50 | 50 | 10 | 2 | 47.2 ± 2.2 | 45.9 | 37.7 ± 2.5 | 38.6 |
| 28 | 75 | 25 | 15 | 3.5 | 36.1 ± 1.4 | 34.6 | 39.1 ± 1.7 | 39.7 |
| Source | Sum of Squares | DF | Mean Square | F Value | Prob > F |
|---|---|---|---|---|---|
| (a) Monophenolase inhibitory activity (Y) | |||||
| Model | 20.668 | 16 | 1.292 | 20.447 | <0.0001 |
| X2 | 1.678 | 1 | 1.678 | 26.563 | 0.0003 |
| X3 | 0.009 | 1 | 0.009 | 0.139 | 0.7167 |
| X4 | 4.155 | 1 | 4.155 | 65.766 | <0.0001 |
| X22 | 0.075 | 1 | 0.075 | 1.189 | 0.2988 |
| X32 | 1.469 | 1 | 1.469 | 23.260 | 0.0005 |
| X42 | 0.258 | 1 | 0.258 | 4.079 | 0.0685 |
| X2X3 | 1.728 | 1 | 1.728 | 27.346 | 0.0003 |
| X2X4 | 0.973 | 1 | 0.973 | 15.396 | 0.0024 |
| X3X4 | 0.873 | 1 | 0.873 | 13.812 | 0.0034 |
| X22X3 | 1.489 | 1 | 1.489 | 23.572 | 0.0005 |
| X22X4 | 1.222 | 1 | 1.222 | 19.342 | 0.0011 |
| X2X32 | 5.107 | 1 | 5.107 | 80.846 | <0.0001 |
| X2X42 | 0.217 | 1 | 0.217 | 3.442 | 0.0905 |
| X32X4 | 2.024 | 1 | 2.024 | 32.038 | 0.0001 |
| X3X42 | 0.005 | 1 | 0.005 | 0.071 | 0.7943 |
| X2X3X4 | 0.073 | 1 | 0.073 | 1.148 | 0.3069 |
| Residual | 0.695 | 11 | 0.063 | ||
| Lack of Fit | 0.447 | 6 | 0.075 | 1.507 | 0.3348 |
| Pure Error | 0.247 | 5 | 0.049 | ||
| Cor Total | 21.362 | 27 | |||
| R2 | 0.967 | ||||
| Adjusted R2 | 0.920 | ||||
| C.V. | 4.611 | ||||
| (b) Diphenolase inhibitory activity (Z) | |||||
| Model | 6.078 | 14 | 0.434 | 13.106 | <0.0001 |
| X2 | 0.185 | 1 | 0.185 | 5.573 | 0.0345 |
| X3 | 0.071 | 1 | 0.071 | 2.143 | 0.1669 |
| X4 | 0.519 | 1 | 0.519 | 15.658 | 0.0016 |
| X22 | 0.010 | 1 | 0.010 | 0.316 | 0.5833 |
| X32 | 1.283 | 1 | 1.283 | 38.730 | <0.0001 |
| X42 | 0.214 | 1 | 0.214 | 6.454 | 0.0246 |
| X2X3 | 0.235 | 1 | 0.235 | 7.104 | 0.0194 |
| X2X4 | 0.385 | 1 | 0.385 | 11.632 | 0.0046 |
| X3X4 | 0.258 | 1 | 0.258 | 7.799 | 0.0152 |
| X22X3 | 0.263 | 1 | 0.263 | 7.939 | 0.0145 |
| X22X4 | 1.060 | 1 | 1.060 | 31.990 | <0.0001 |
| X2X32 | 1.771 | 1 | 1.771 | 53.473 | <0.0001 |
| X3X42 | 0.314 | 1 | 0.314 | 9.481 | 0.0088 |
| X2X3X4 | 0.040 | 1 | 0.040 | 1.198 | 0.2936 |
| Residual | 0.431 | 13 | 0.033 | ||
| Lack of Fit | 0.315 | 8 | 0.039 | 1.696 | 0.2906 |
| Pure Error | 0.116 | 5 | 0.023 | ||
| Cor Total | 6.508 | 27 | |||
| R2 | 0.934 | ||||
| Adjusted R2 | 0.863 | ||||
| C.V. | 2.996 | ||||
| No. | Peptide Sequence | Egg Protein Fragment | Peptide Length | Potential Binding Sites of Mushroom Tyrosinase (PDB ID: 2Y9X) | PepSite 2 p-Value |
|---|---|---|---|---|---|
| (a) Monophenolase inhibitory activity | |||||
| 1 | ADHPF | Ovalbumin | 5 | Y140, K389, H390 | 0.002658 |
| 2 | AFKDEDTKAMPF | Ovalbumin | 12 | N22, F135, D137, Y140. R301, P366, D367, W386, H390, Y391 | 0.02053 |
| 3 | ILELPFASGDLLML | Ovalbumin-related protein X | 14 | Y36, L40, F54, G58, H61, H85, F90, H94, W101, Q133, H259, H263, M280, H285, A286, A287, F288, D289, P290, F292, W293, H296 | 0.03464 |
| 4 | DKLPGFGD | Ovalbumin | 8 | Y140, P370, Y382, W386, K389, H390 | 0.05277 |
| 5 | FDKLPGFGD | Ovalbumin | 9 | Y140, P370, Y382, W386, K389, H390 | 0.0722 |
| 6 | FDKLPGFGDSIEAQCGTSVN | Ovalbumin | 20 | Y140, T233, R301, M309, D367, P370, Y382, N384, W386, H388, K389, H390 | 0.08107 |
| 7 | DGSGGCIPK | Ovomucin | 9 | N22, F135, D137, Y140, R301, D367, Y382, W386, K389, H390 | 0.1274 |
| (b) Diphenolase inhibitory activity | |||||
| 1 | SDFHLFGPPGK | Ovotransferrin | 11 | Y140, R301, D367, Y382, W386, H390 | 0.009412 |
| 2 | FDGRSR | Ovomucin | 6 | D137, R301, P366, D367, W386, H390, Y391 | 0.01312 |
| 3 | FNCSSAGPGAIGSEC | Ovomucin | 15 | N22, F135, D137, Y140, R301, P366, D367, W386, H390, Y391 | 0.01614 |
| 4 | MYQIGLFR | Ovalbumin | 8 | D137, Y140, R301, D367, P370, Y382, W386, H390, Y391 | 0.01832 |
| 5 | GYSLGNWVCAAK | Lysozyme | 12 | H61, N81, Y82, C83, T84, H85, F90, W93, H94, R95, Y97, E98, E256, H259, H263, M280, V283, A286, A287, F292, W293, H296 | 0.01891 |
| 6 | DLLFKDSAIMLK | Ovotransferrin | 12 | D137, Y140, R301, D367, Y382, W386, K389, H390, Y391 | 0.02538 |
| 7 | CQLCQGSGGIPPEK | Ovotransferrin | 14 | D137, Y140, R301, P366, D367, Y382, W386, H390, Y391 | 0.02578 |
| 8 | ADHPFLF | Ovalbumin | 7 | Y140, R301, P366, D367, F368, P370, W386, H390 | 0.03051 |
| 9 | SGAFHCLK | Ovotransferrin | 8 | Y140, Y382, W386, H390 | 0.03994 |
| 10 | YFGYTGALRCLV | Ovotransferrin | 12 | H61, H85, H94, Y97, Y140, H259, H263, M280, V283, A287, F292, W293, H295, H296, V299, R301, D367, Y382, W386, H390 | 0.04488 |
| 11 | HIATNAVLFFGR | Ovalbumin | 12 | G58, H61, C83, H85, F90, Typ93, H94, Y97, D137, Y140, H259, H263, M280, H285, A286, A287, F288, D289, F292, W293, H296, R301, D367, W386, H390, Y391 | 0.04707 |
| 12 | FKDEDTQAMPFR | Ovalbumin | 12 | D137, Y140, R301, P366, D367, W386, H390, Y391 | 0.05594 |
| 13 | FMMFESQNKDLLFK | Ovotransferrin | 14 | H61, N81, Y82, C83, T84, H85, F90, W93, H94, Y97, D137, Y140, H259, H263, M280, A286, A287, F292, W293, H296, R301, P366, D367, W386, H390, Y391 | 0.06287 |
| 14 | FDKLPGFGD | Ovalbumin | 9 | Y140, P370, Y382, W386, K389, H390 | 0.0722 |
| 15 | SMLVLLPDEVSGLEQLESIINFEK | Ovalbumin | 24 | D137, Y140, R301, D367, P370, Y382, W386, K389, H390, Y391 | 0.08195 |
| 16 | SGYSGAFHCLK | Ovotransferrin | 11 | Y140, R301, P366, D367, Y382, W386, H390 | 0.0849 |
| 17 | SGGQFSLTSTVKVC | Ovomucin | 14 | D137, Y140, R301, D367, W386, H390, Y391 | 0.08667 |
| 18 | SGALHCLK | Ovotransferrin | 8 | H61, N81, Y82, C83, T84, H85, W93, H94, R95, Y97, E98, H259, H263, A286, A287, F292, W293, H296 | 0.1247 |
| 19 | SSCICS | Ovomucin | 6 | N22, F135, D137, R301, P366, D367, W386, H390 | 0.1938 |
| 20 | SSCEDCVCT | Ovomucin | 9 | D137, Y140, R301, D367, W386, H390, Y391 | 0.2086 |
| 21 | YFGYTGALR | Ovotransferrin | 9 | H61, H85, H94, Y97, H259, H263, M280, V283, A286, Als287, F292, W293, H295, H296, V299 | 0.2364 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yap, P.-G.; Gan, C.-Y. Chicken Egg White—Advancing from Food to Skin Health Therapy: Optimization of Hydrolysis Condition and Identification of Tyrosinase Inhibitor Peptides. Foods 2020, 9, 1312. https://doi.org/10.3390/foods9091312
Yap P-G, Gan C-Y. Chicken Egg White—Advancing from Food to Skin Health Therapy: Optimization of Hydrolysis Condition and Identification of Tyrosinase Inhibitor Peptides. Foods. 2020; 9(9):1312. https://doi.org/10.3390/foods9091312
Chicago/Turabian StyleYap, Pei-Gee, and Chee-Yuen Gan. 2020. "Chicken Egg White—Advancing from Food to Skin Health Therapy: Optimization of Hydrolysis Condition and Identification of Tyrosinase Inhibitor Peptides" Foods 9, no. 9: 1312. https://doi.org/10.3390/foods9091312
APA StyleYap, P.-G., & Gan, C.-Y. (2020). Chicken Egg White—Advancing from Food to Skin Health Therapy: Optimization of Hydrolysis Condition and Identification of Tyrosinase Inhibitor Peptides. Foods, 9(9), 1312. https://doi.org/10.3390/foods9091312