Identification of a Lactic Acid Bacteria to Degrade Biogenic Amines in Chinese Rice Wine and Its Enzymatic Mechanism
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Strains Screening and Identification
2.3. HPLC Determination of Biogenic Amines
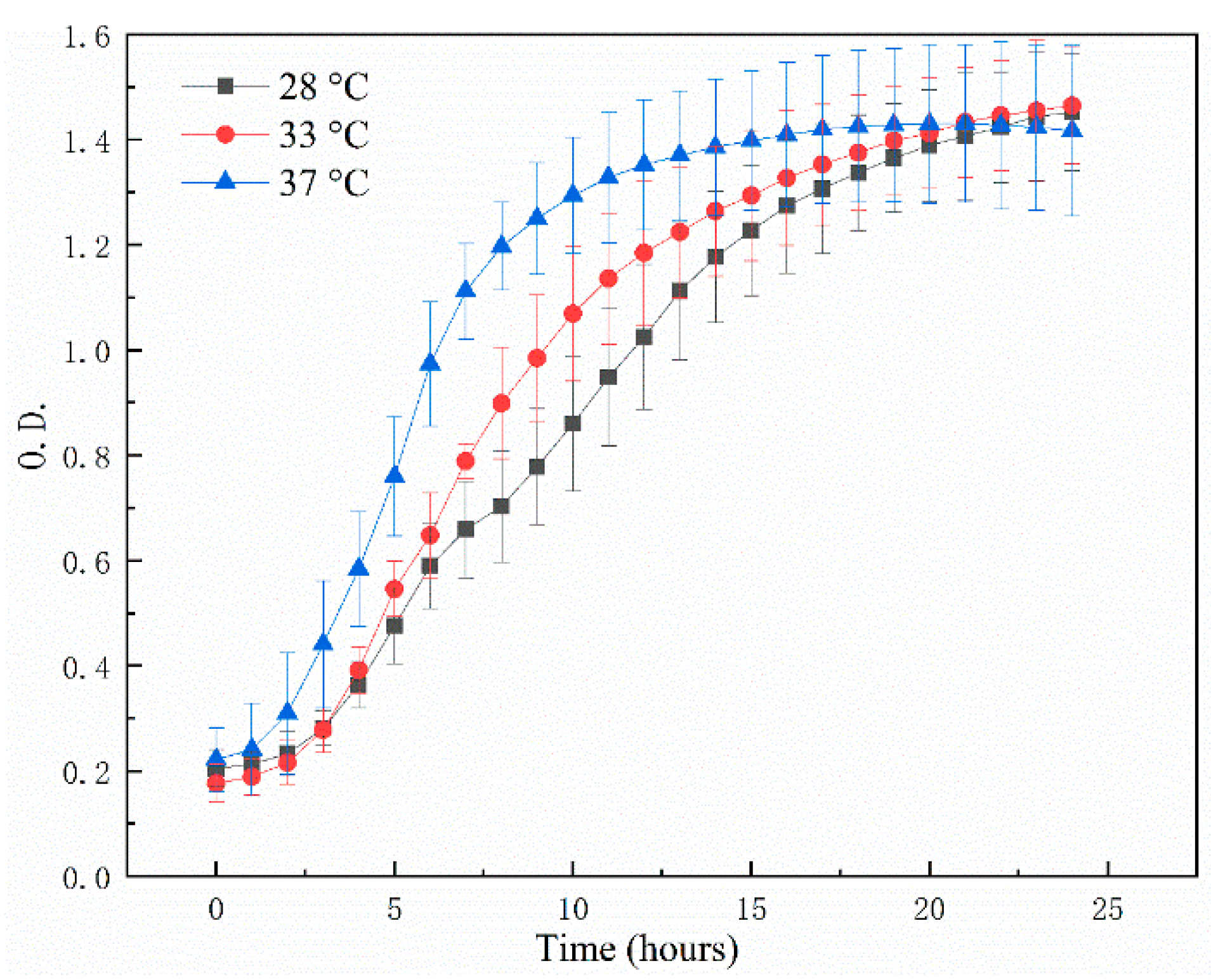
2.4. Bacterial Growth Analysis
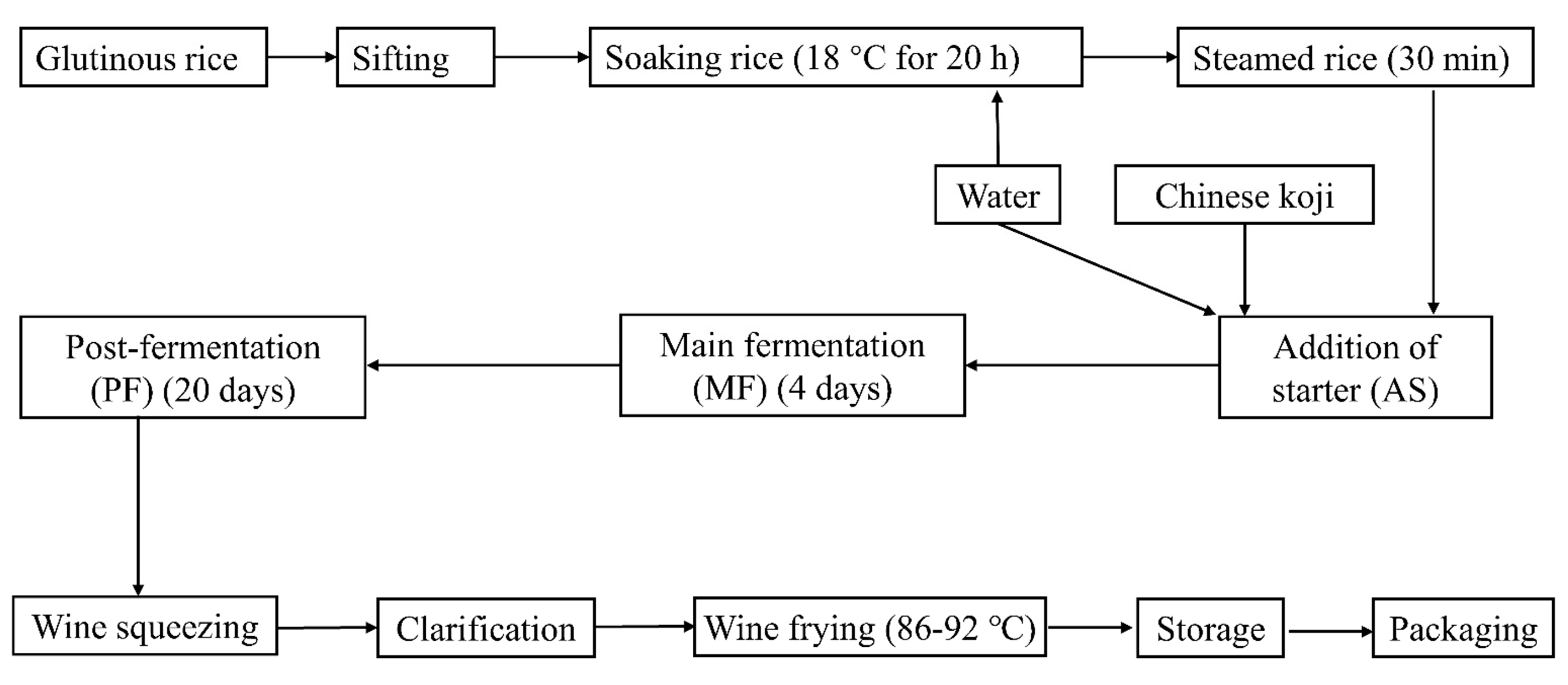
2.5. The Bacterial Starter Application in Pilot Scale Fermentation
2.6. Separation of the Amine Oxidases
2.7. Identification of the Amine Oxidases
2.8. Enzymatic Properties of the Amine Oxidases
2.9. Statistical Analysis
3. Result
3.1. Strains Screening and Identification
3.2. The Bacterial Growth Ability
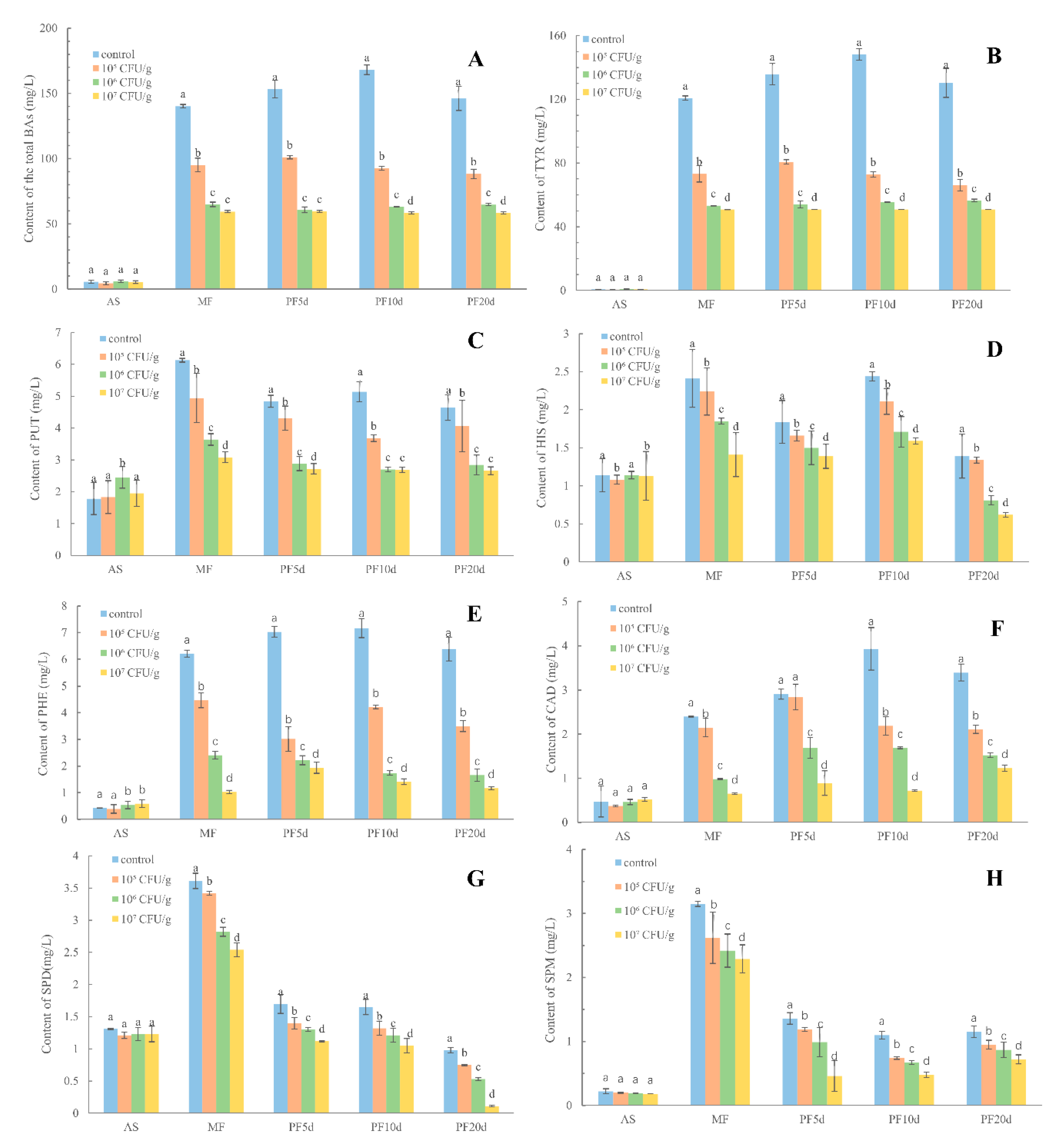
3.3. Changes in the BAs Induced by L. plantarum in Pilot Scale Fermentation
3.4. Total Acid and pH in Pilot Scale Fermentation
3.5. Alcohol Content, Total Sugar, Non-Sugar Solid and Amino Acid Nitrogen in Pilot Scale Fermentation
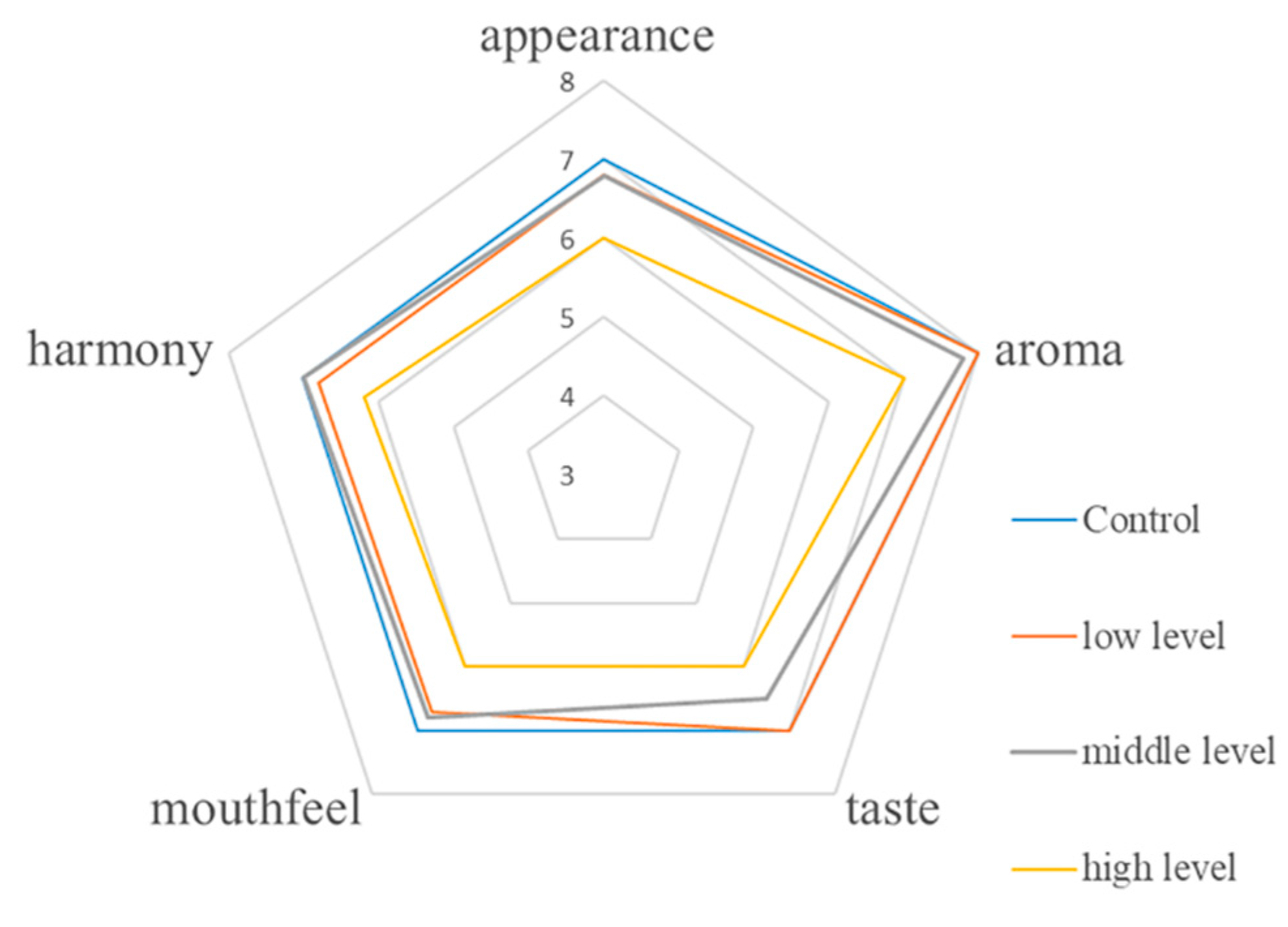
3.6. Sensory Evaluation
3.7. Purification and Identification of the Amine Oxidases
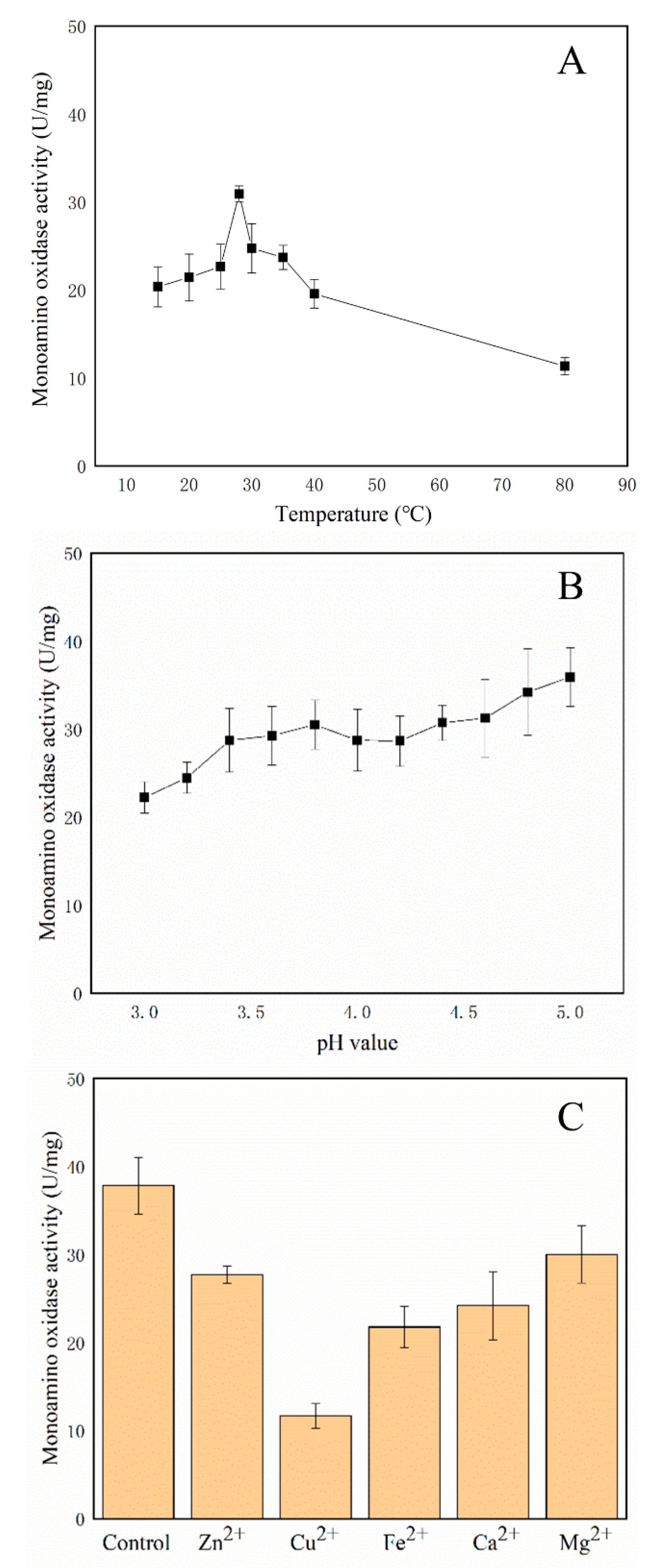
3.8. Amine Oxidases Assays
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kim, J.Y.; Kim, D.; Park, P.; Kang, H.-I.; Ryu, E.K.; Kim, S.M. Effects of storage temperature and time on the biogenic amine content and microflora in Korean turbid rice wine, Makgeolli. Food Chem. 2011, 128, 87–92. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Guan, X.; Wang, Y.; Li, L.; Wu, D.; Chen, Y.; Pei, H.; Xiao, D. Reduction of biogenic amines production by eliminating the PEP4 gene in Saccharomyces cerevisiae during fermentation of Chinese rice wine. Food Chem. 2015, 178, 208–211. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, M.A.; Moreno-Arribas, M.V. The problem of biogenic amines in fermented foods and the use of potential biogenic amine-degrading microorganisms as a solution. Trends Food Sci. Technol. 2014, 39, 146–155. [Google Scholar] [CrossRef]
- Xia, X.; Zhang, Q.; Zhang, B.; Zhang, W.; Wang, W. Insights into the Biogenic Amine Metabolic Landscape during Industrial Semidry Chinese Rice Wine Fermentation. J. Agric. Food Chem. 2016, 64, 7385–7393. [Google Scholar] [CrossRef] [PubMed]
- Ancín-Azpilicueta, C.; González-Marco, A.; Jiménez-Moreno, N. Current Knowledge about the Presence of Amines in Wine. Crit. Rev. Food Sci. Nutr. 2008, 48, 257–275. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.Y.; Yang, Y.P.; Peng, Q.; Han, Y. Biogenic amines in wine: A review. Int. J. Food Sci. Technol. 2015, 50, 1523–1532. [Google Scholar] [CrossRef]
- Liu, S.P.; Yu, J.X.; Wei, X.L.; Ji, Z.W.; Zhou, Z.L.; Meng, X.Y.; Mao, J. Sequencing-based screening of functional microorganism to decrease the formation of biogenic amines in Chinese rice wine. Food Control 2016, 64, 98–104. [Google Scholar] [CrossRef]
- Torlois, S.; Joyeux, A.; Moreno-Arribas, V.; Lonvaud-Funel, A.; Bertrand, A. Isolation, properties and behaviour of tyramine-producing lactic acid bacteria from wine. J. Appl. Microbiol. 2000, 88, 584–593. [Google Scholar]
- Moreno-Arribas, M.; Polo, M.; Jorganes, F.; Muñoz, R. Screening of biogenic amine production by lactic acid bacteria isolated from grape must and wine. Int. J. Food Microbiol. 2003, 84, 117–123. [Google Scholar] [CrossRef]
- Zhang, F.; Xue, J.; Wang, D.; Wang, Y.; Zou, H.; Zhu, B. Dynamic changes of the content of biogenic amines in Chinese rice wine during the brewing process. J. Inst. Brew. 2013, 119, 294–302. [Google Scholar] [CrossRef]
- Chen, S.; Xu, Y. The Influence of Yeast Strains on the Volatile Flavour Compounds of Chinese Rice Wine. J. Inst. Brew. 2010, 116, 190–196. [Google Scholar] [CrossRef]
- Yongmei, L.; Xin, L.; Xiaohong, C.; Mei, J.; Chao, L.; Mingsheng, D. A survey of biogenic amines in Chinese rice wines. Food Chem. 2007, 100, 1424–1428. [Google Scholar] [CrossRef]
- Callejon, S.; Sendra, R.; Ferrer, S.; Pardo, I. Identification of a novel enzymatic activity from lactic acid bacteria able to degrade biogenic amines in wine. Appl. Microbiol. Biotechnol. 2014, 98, 185–198. [Google Scholar] [CrossRef] [PubMed]
- Capozzi, V.; Russo, P.; Ladero, V.; Fernandez, M.; Fiocco, D.; Alvarez, M.A.; Grieco, F.; Spano, G. Biogenic Amines Degradation by Lactobacillus plantarum: Toward a Potential Application in Wine. Front. Microbiol. 2012, 3, 122. [Google Scholar] [CrossRef] [PubMed]
- Leuschner, R.; Hammes, W. Tyramine degradation by micrococci during ripening of fermented sausage. Meat Sci. 1998, 49, 289–296. [Google Scholar] [CrossRef]
- Dapkevicius, M.L.; Nout, M.; Rombouts, F.M.; Houben, J.H.; Wymenga, W. Biogenic amine formation and degradation by potential fish silage starter microorganisms. Int. J. Food Microbiol. 2000, 57, 107–114. [Google Scholar] [CrossRef]
- Landete, J.; Ferrer, S.; Pardo, I. Biogenic amine production by lactic acid bacteria, acetic bacteria and yeast isolated from wine. Food Control 2007, 18, 1569–1574. [Google Scholar] [CrossRef]
- García-Ruiz, A.; González-Rompinelli, E.M.; Bartolomé, B.; Moreno-Arribas, M.V. Potential of wine-associated lactic acid bacteria to degrade biogenic amines. Int. J. Food Microbiol. 2011, 148, 115–120. [Google Scholar] [CrossRef]
- Cui, Y.; Qu, X.; Li, H.; He, S.; Liang, H.; Zhang, H.; Ma, Y. Isolation of halophilic lactic acid bacteria from traditional Chinese fermented soybean paste and assessment of the isolates for industrial potential. Eur. Food Res. Technol. 2012, 234, 797–806. [Google Scholar] [CrossRef]
- Yu, H.; Ying, Y.; Fu, X.; Lu, H. Quality Determination of Chinese Rice Wine Based on Fourier Transform near Infrared Spectroscopy. J. Near Infrared Spectrosc. 2006, 14, 37–44. [Google Scholar] [CrossRef]
- Shen, F.; Ying, Y.; Li, B.; Zheng, Y.; Hu, J. Prediction of sugars and acids in Chinese rice wine by mid-infrared spectroscopy. Food Res. Int. 2011, 44, 1521–1527. [Google Scholar] [CrossRef]
- Callejon, S.; Sendra, R.; Ferrer, S.; Pardo, I. Ability of Kocuria varians LTH 1540 To Degrade Putrescine: Identification and Characterization of a Novel Amine Oxidase. J. Agric. Food Chem. 2015, 63, 4170–4178. [Google Scholar] [CrossRef] [PubMed]
- Caston, J.; Eaton, C.; Gheorghiu, B.; Ware, L. Tyramine induced hypertensive episodes and panic attacks in hereditary deficient monoamine oxidase patients. J. S. C. Med. Assoc. 1975 2002, 98, 187. [Google Scholar]
- Beneduce, L.; Romano, A.; Capozzi, V.; Lucas, P.; Barnavon, L.; Bach, B.; Vuchot, P.; Grieco, F.; Spano, G. Biogenic amine in wines. Ann. Microbiol. 2010, 60, 573–578. [Google Scholar] [CrossRef]
- Ouyang, Q.; Zhao, J.; Chen, Q. Measurement of non-sugar solids content in Chinese rice wine using near infrared spectroscopy combined with an efficient characteristic variables selection algorithm. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2015, 151, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Levering, P.R.; Van Dijken, J.P.; Veenhuis, M.; Harder, W.; Dijken, J.P. Arthrobacter P1, a fast growing versatile methylotroph with amine oxidase as a key enzyme in the metabolism of methylated amines. Arch. Microbiol. 1981, 129, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Yagodina, O.V.; Nikol’Skaya, E.B.; Khovanskikh, A.E.; Kormilitsyn, B.N. Amine Oxidases of Microorganisms. Zhurnal Evoliutsionnoĭ Biokhimii I Fiziol. 2002, 29, 864–869. [Google Scholar]
- Sekiguchi, Y.; Makita, H.; Yamamura, A.; Matsumoto, K. A thermostable histamine oxidase from Arthrobacter crystallopoietes KAIT-B-007. J. Biosci. Bioeng. 2004, 97, 104–110. [Google Scholar] [CrossRef]
- Tipton, K.F.; Boyce, S.; O’Sullivan, J.; Davey, G.P.; Healy, J. Monoamine Oxidases: Certainties and Uncertainties. Curr. Med. Chem. 2004, 11, 1965–1982. [Google Scholar] [CrossRef]
- Cona, A.; Rea, G.; Angelini, R.; Federico, R.; Tavladoraki, P. Functions of amine oxidases in plant development and defence. Trends Plant Sci. 2006, 11, 80–88. [Google Scholar] [CrossRef]
- Grimsby, J.; Chen, K.; Wang, L.J.; Lan, N.C.; Shih, J.C. Human monoamine oxidase A and B genes exhibit identical exon-intron organization. Proc. Natl. Acad. Sci. USA 1991, 88, 3637–3641. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.Y.; Neff, N.H. Beta-phenylethylamine: A specific substrate for type B monoamine oxidase of brain. J. Pharmacol. Exp. Ther. 1973, 187, 365–371. [Google Scholar]
- Li, M.; Binda, C.; Mattevi, A.; Edmondson, D.E. Functional Role of the “Aromatic Cage” in Human Monoamine Oxidase B: Structures and Catalytic Properties of Tyr435 Mutant Proteins. Biochemistry 2006, 45, 4775–4784. [Google Scholar] [CrossRef] [PubMed]
- Van Hellemond, E.W.; Van Dijk, M.; Heuts, D.P.H.M.; Janssen, D.B.; Fraaije, M.W. Discovery and characterization of a putrescine oxidase from Rhodococcus erythropolis NCIMB 11540. Appl. Microbiol. Biotechnol. 2008, 78, 455–463. [Google Scholar] [CrossRef] [PubMed]
| Strains | Tryptamine | Phenylethylamine | Putrescine | Cadaverine | Tyramine | HISTAMINE | Spermidine | Spermine |
|---|---|---|---|---|---|---|---|---|
| Lactobacillus plantarum CAU 3823 | 55.95 ± 6.59 | 40.85 ± 9.87 | 41.82 ± 7.97 | 42.79 ± 7.76 | 40.12 ± 8.09 | 44.72 ± 7.56 | 43.51 ± 8.39 | 42.56 ± 8.41 |
| Addition the Selected Strain (cfu/mL) | The Addition of Starter | Main Fermentation | Post-Fermentation 5d | Post-Fermentation 10d | Post-Fermentation 20d | |
|---|---|---|---|---|---|---|
| Total acid (g/L) | Control | 6.03 ± 0.22 a | 3.81 ± 0.15 a | 4.91 ± 0.07 a | 5.55 ± 0.33 a | 5.94 ± 0.20 a |
| 105 (low level) | 6.17 ± 0.13 a | 4.93 ± 0.13 b | 5.41 ± 0.19 b | 5.92 ± 0.19 a | 6.53 ± 0.13 b | |
| 106 (middle level) | 5.92 ± 0.19 a | 6.01 ± 0.14 c | 6.51 ± 0.14 c | 6.74 ± 0.44 b | 6.86 ± 0.13 d | |
| 107 (high level) | 6.03 ± 0.15 a | 6.04 ± 0.10 c | 7.06 ± 0.09 c | 8.01 ± 0.23 c | 9.14 ± 0.45 c | |
| pH | Control | 6.33 ± 0.19 a | 4.04 ± 0.12 a | 4.19 ± 0.05 a | 4.21 ± 0.03 b | 4.14 ± 0.12 a |
| 105 (low level) | 6.37 ± 0.28 a | 4.00 ± 0.14 a | 4.36 ± 0.12 a | 4.12 ± 0.07 a | 3.99 ± 0.16 a | |
| 106 (middle level) | 6.45 ± 0.22 a | 3.84 ± 0.16 a | 4.34 ± 0.08 a | 4.45 ± 0.13 b | 3.87 ± 0.12 a | |
| 107 (high level) | 6.43 ± 0.23 a | 3.71 ± 0.04 b | 4.24 ± 0.12 a | 4.32 ± 0.12 b | 3.63 ± 0.03 b |
| Addition the Selected Strain (cfu/mL) | Alcohol Content (% vol) | Amino Acid Nitrogen (g/L) | Total Sugar (g/L) | Non-Sugar Solid (g/L) |
|---|---|---|---|---|
| Control | 11.52 ± 0.23 a | 1.44 ± 0.11 a | 31.98 ± 1.37 a | 39.81 ± 0.33 a |
| 105 (low level) | 11.49 ± 0.35 a | 1.28 ± 0.35 a | 15.35 ± 2.34 b | 62.34 ± 0.32 c |
| 106 (middle level) | 10.33 ± 0.41 b | 0.82 ± 0.13 b | 11.98 ± 3.25 b | 71.52 ± 0.18 d |
| 107 (high level) | 9.29 ± 0.25 c | 0.59 ± 0.02 c | 10.97 ± 2.23 b | 51.16 ± 0.25 b |
| Protein Concentration (mg/mL) | Monoamine Oxidase Activity (U/mg) | Diamine Oxidase Activity (×10−4 U/mg) | |
|---|---|---|---|
| Cell-free extracts | 5.5 | 36.9 ± 5.2 | 1.3 ± 0.1 |
| Fraction 1 | 3.1 | 19.8 ± 2.6 | ND |
| Fraction 2 | 1.6 | 2.4 ± 1.2 | ND |
| Fraction 3 | 0.5 | ND | ND |
| Tryptamine | Phenylethylamine | Putrescine | Cadaverine | Histamine | Tyramine | Spermidine | Spermine | |
|---|---|---|---|---|---|---|---|---|
| Fraction 1 | 47.9 | 44.3 | 40.3 | 41.1 | 41.1 | 41.9 | 41 | 43.5 |
| Fraction 2 | ND | 0.3 | 0.7 | ND | 1.2 | 3.8 | ND | ND |
| Fraction 3 | ND | ND | ND | ND | ND | ND | ND | ND |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niu, T.; Li, X.; Guo, Y.; Ma, Y. Identification of a Lactic Acid Bacteria to Degrade Biogenic Amines in Chinese Rice Wine and Its Enzymatic Mechanism. Foods 2019, 8, 312. https://doi.org/10.3390/foods8080312
Niu T, Li X, Guo Y, Ma Y. Identification of a Lactic Acid Bacteria to Degrade Biogenic Amines in Chinese Rice Wine and Its Enzymatic Mechanism. Foods. 2019; 8(8):312. https://doi.org/10.3390/foods8080312
Chicago/Turabian StyleNiu, Tianjiao, Xing Li, Yongjie Guo, and Ying Ma. 2019. "Identification of a Lactic Acid Bacteria to Degrade Biogenic Amines in Chinese Rice Wine and Its Enzymatic Mechanism" Foods 8, no. 8: 312. https://doi.org/10.3390/foods8080312
APA StyleNiu, T., Li, X., Guo, Y., & Ma, Y. (2019). Identification of a Lactic Acid Bacteria to Degrade Biogenic Amines in Chinese Rice Wine and Its Enzymatic Mechanism. Foods, 8(8), 312. https://doi.org/10.3390/foods8080312





