Optimization of Fermentation Conditions for Enhanced Single Cell Protein Production by Rossellomorea marisflavi NDS and Nutritional Composition Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Strain and Culture Conditions
2.2. Determination of Protein Content
2.3. Optimization of Cultivation Condition
2.4. Optimization of Medium Composition
2.5. Fermentation Condition Optimization in a 10 L Bioreactor
2.6. Proximate Analysis
2.7. Amino Acid Analysis
2.8. Fatty Acid Analysis
2.9. Statistical Analysis
3. Results
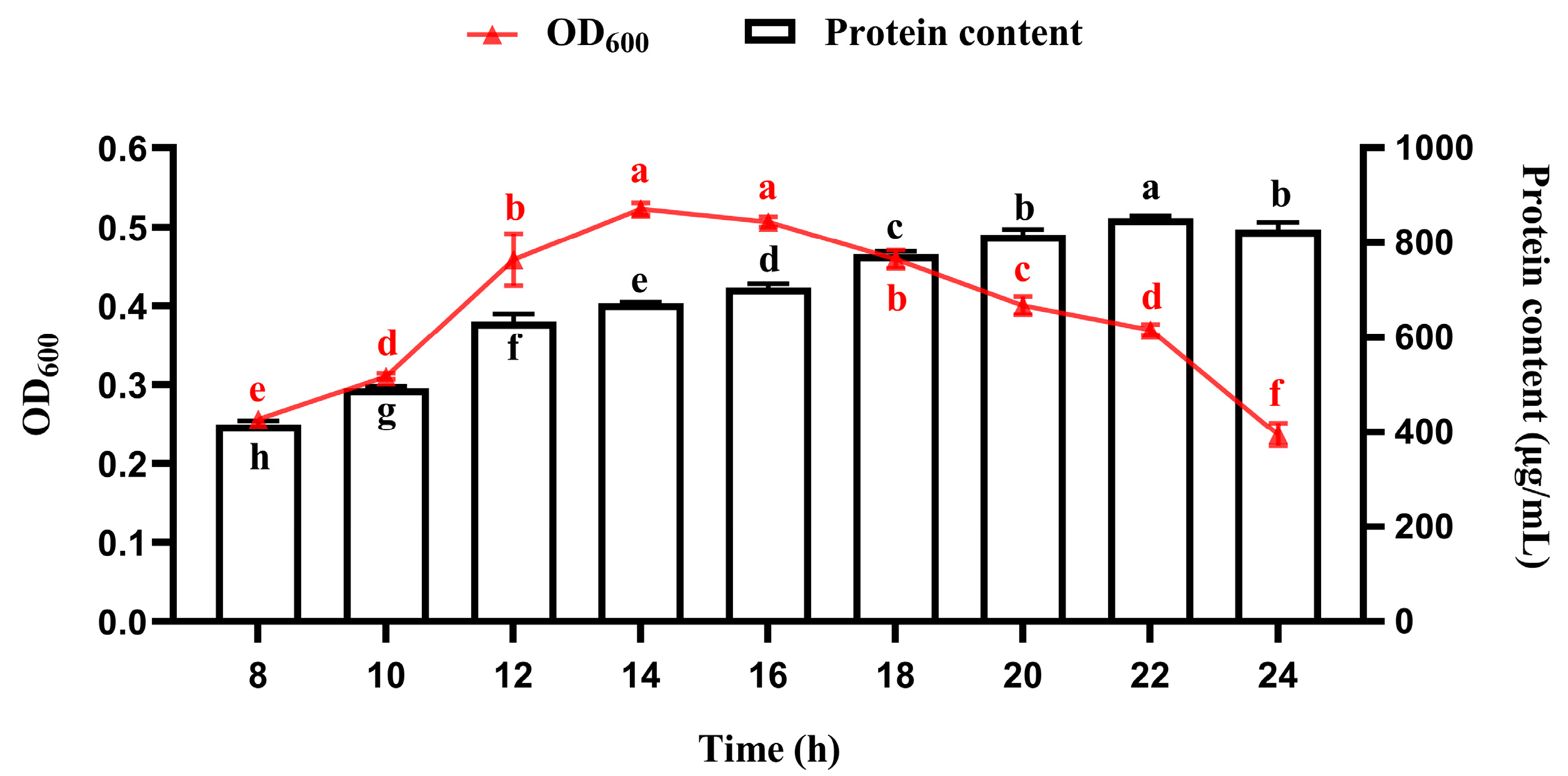
3.1. Temporal Profiles of Bacterial Growth and Protein Accumulation During Fermentation
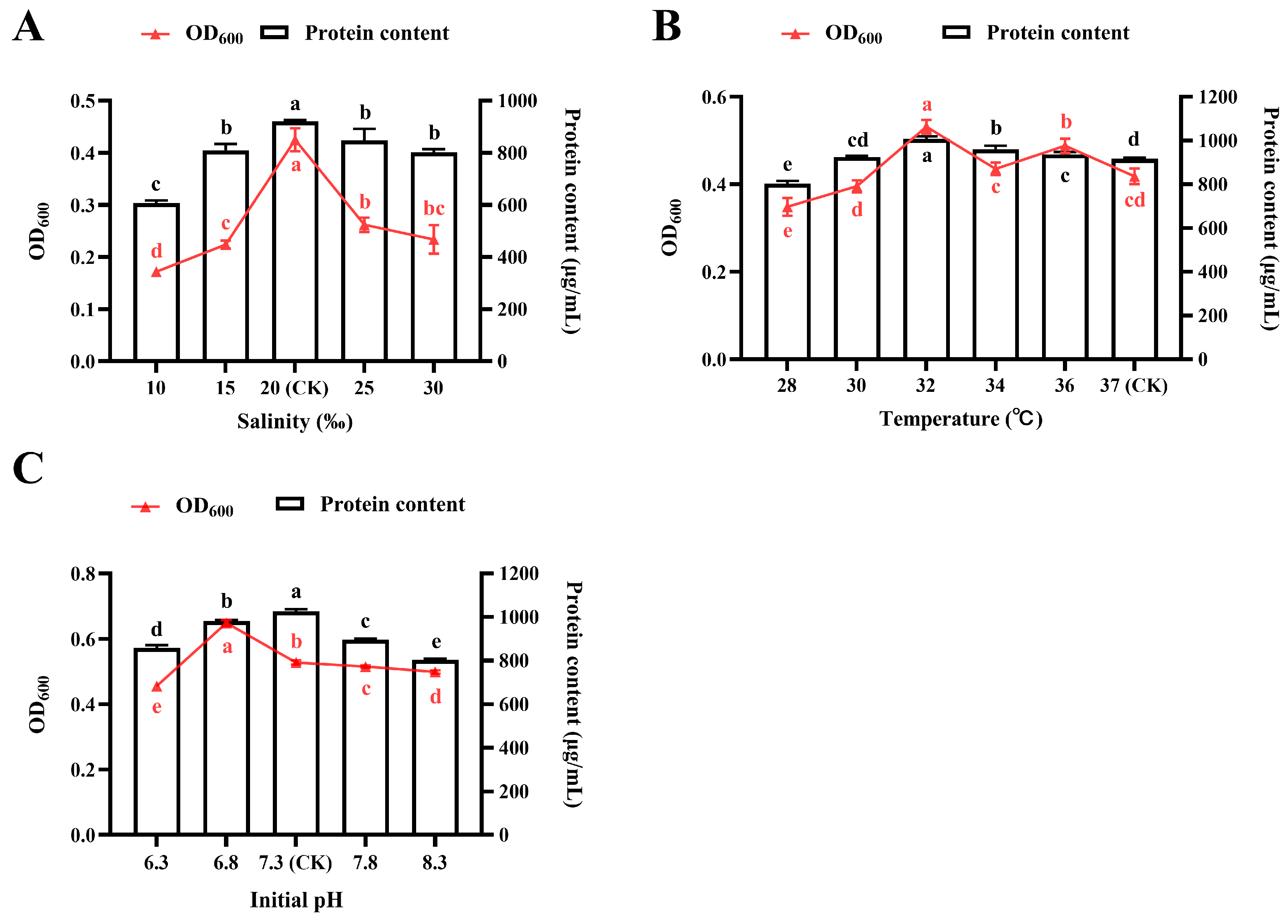
3.2. Effects of Cultivation Condition on Protein Accumulation and Bacterial Growth
3.2.1. Effect of NaCl Salinity
3.2.2. Effect of Temperature
3.2.3. Effect of Initial pH
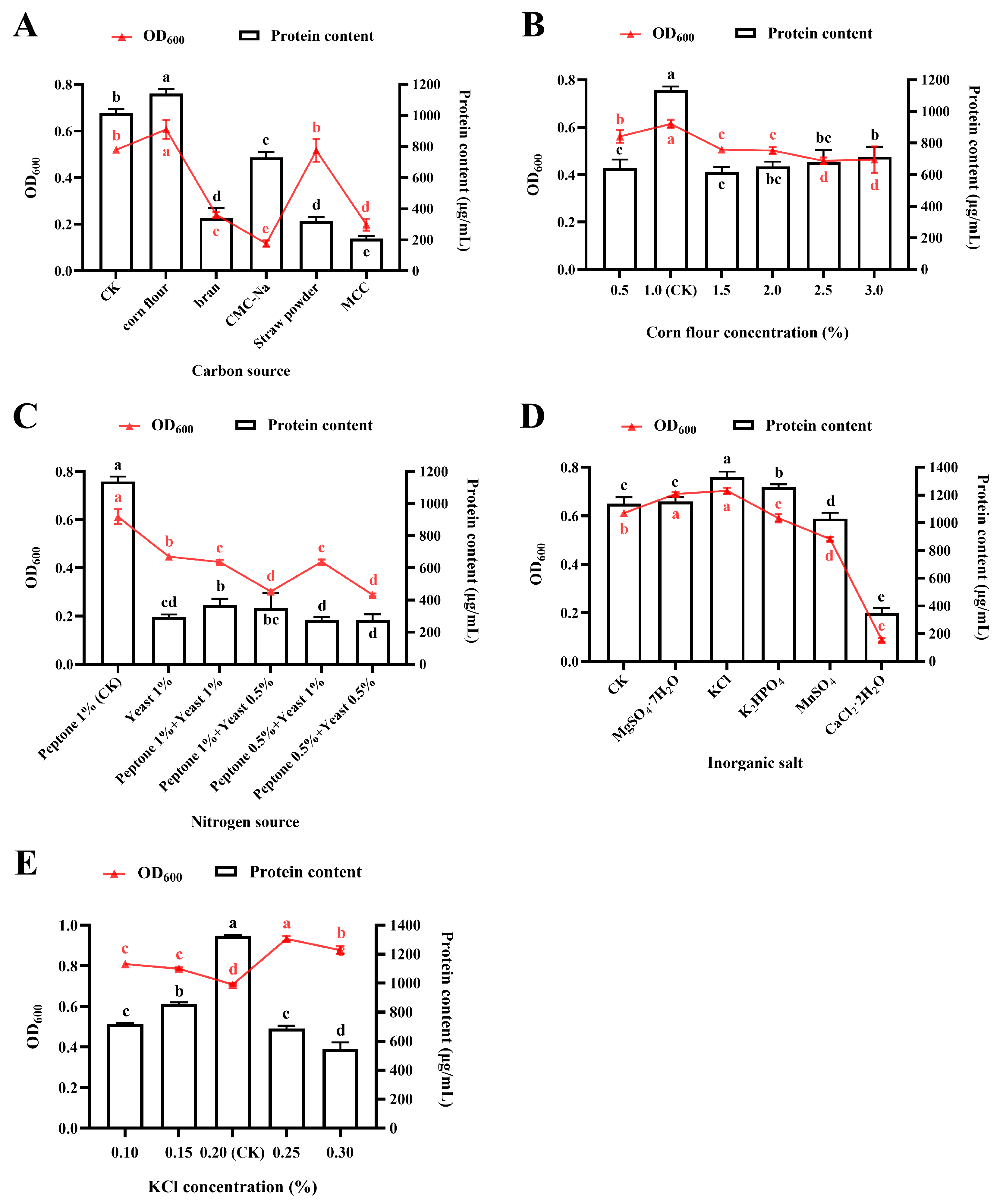
3.3. Effects of Medium Composition on Protein Accumulation and Bacterial Growth
3.3.1. Effect of Carbon Source
3.3.2. Effect of Nitrogen Source
3.3.3. Effect of Inorganic Ions
3.4. Effects of Fermentation Condition in a 10 L Bioreactor on Protein Accumulation and Bacterial Growth
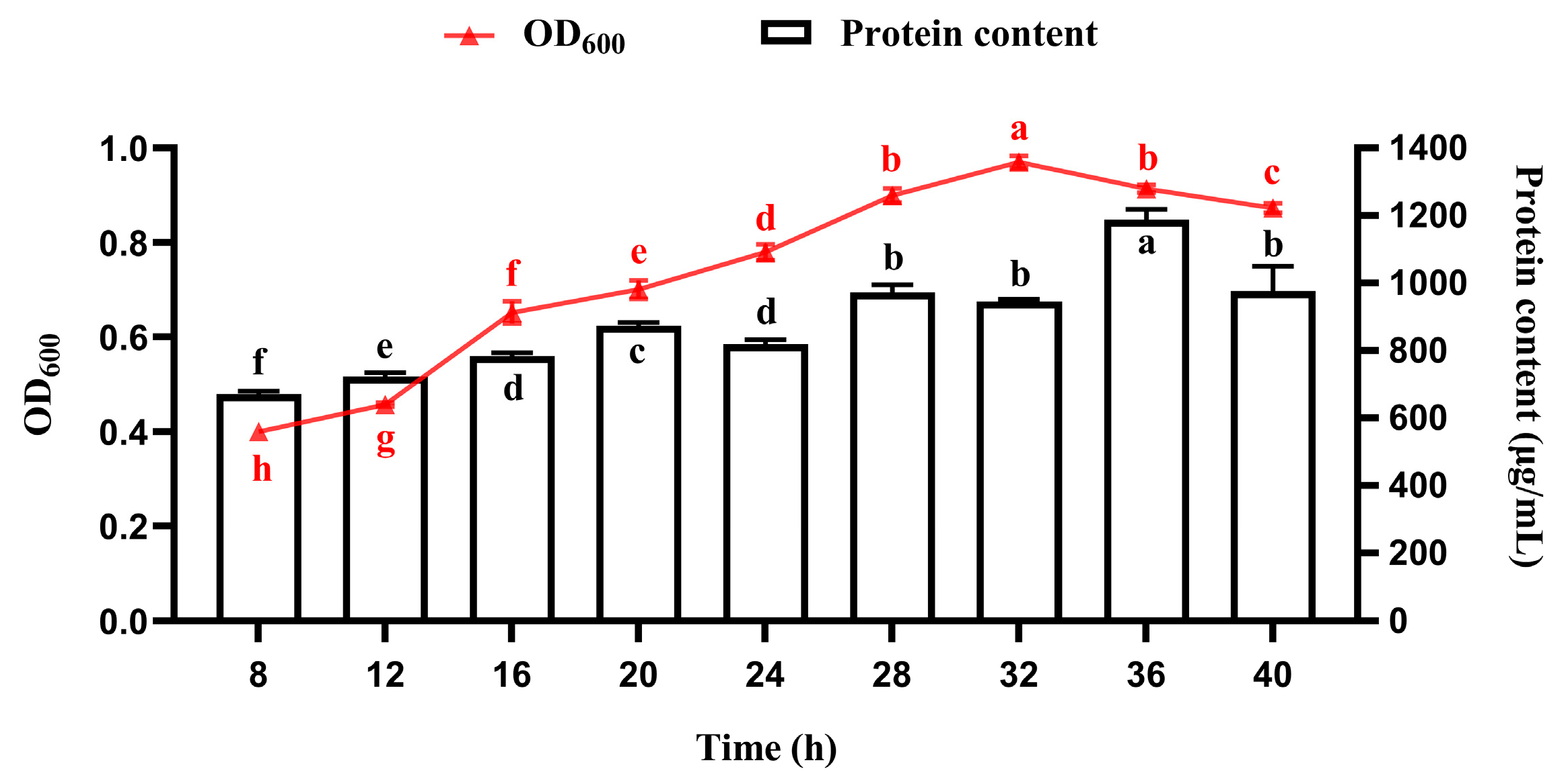
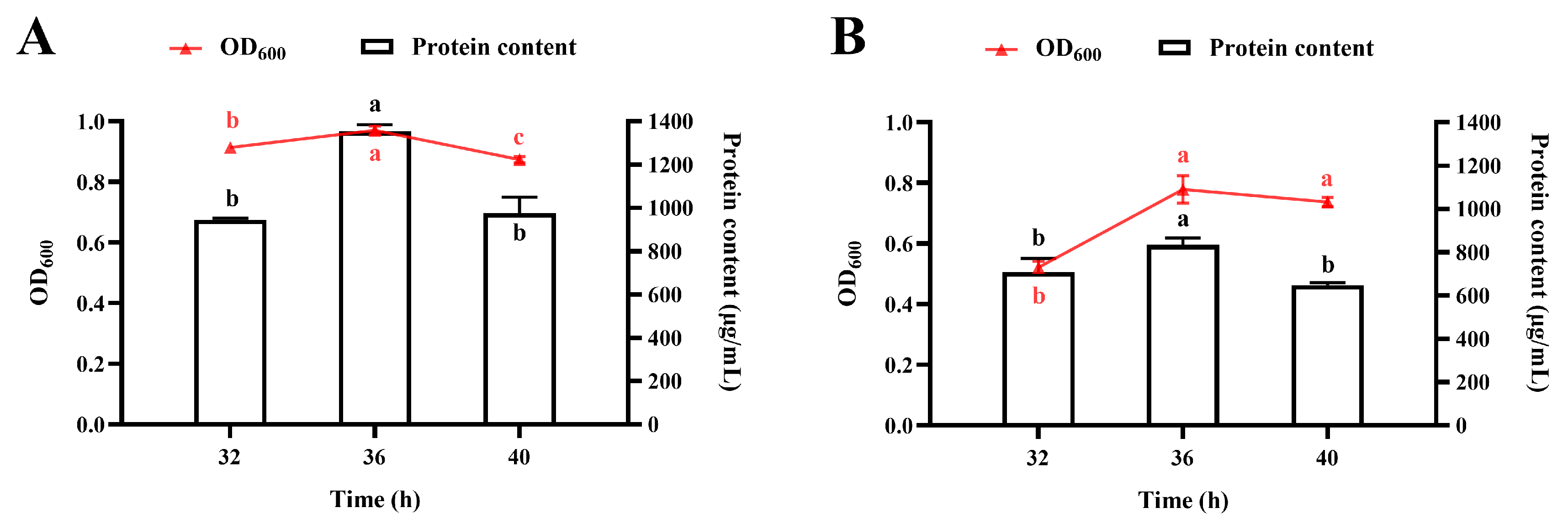
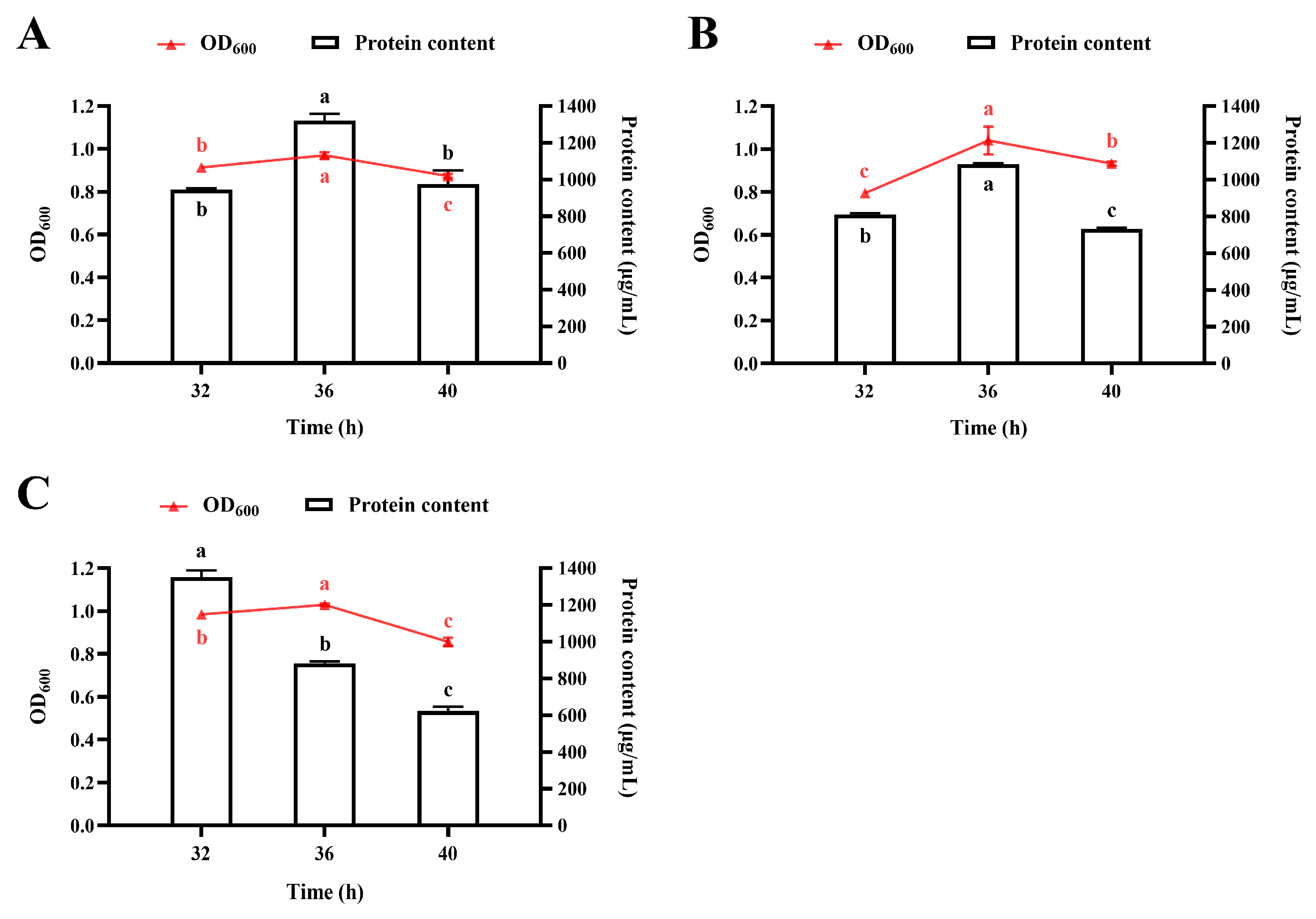
3.4.1. Temporal Profiles of Bacterial Growth and Protein Accumulation During Fermentation in a 10 L Bioreactor
3.4.2. Effect of pH
3.4.3. Effect of Agitator Speed
3.5. Analysis of Nutritional Composition of R. marisflavi NDS
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| WW | Wet weight |
| SCFAs | Short-chain fatty acids |
| BPM | Beef extract peptone liquid medium |
| PBS | Phosphate-buffered solution |
| CMC-Na | Carboxymethylcellulose sodium |
| MCC | Microcrystalline cellulose |
| HCl | Hydrochloric acid |
| NaOH | Sodium hydroxide |
| H2SO4 | Sulfuric acid |
| NH3 | Ammonia |
| HPLC | High performance liquid chromatography |
| GC | Gas chromatography |
| FAMEs | Fatty acid methyl esters |
| ANOVA | One-way analysis of variance |
| SD | Standard deviation |
| CK | Control check |
References
- Kim, S.W.; Less, J.F.; Wang, L.; Yan, T.; Kiron, V.; Kaushik, S.J.; Lei, X.G. Meeting global feed protein demand: Challenge, opportunity, and strategy. Annu. Rev. Anim. Biosci. 2019, 7, 221–243. [Google Scholar] [CrossRef]
- Amara, A.A.; Abd, E.N. Fungi as a source of edible proteins and animal feed. J. Fungi 2023, 9, 73. [Google Scholar] [CrossRef] [PubMed]
- Makkar, H.P.S. Review: Feed demand landscape and implications of food-not feed strategy for food security and climate change. Animal 2017, 12, 1744–1754. [Google Scholar] [CrossRef] [PubMed]
- Giuliana, P.; Francesca, T.; Riccardo, F.; Rosaria, M.; Paolo, B.; Antonella, D.Z.; Anna, D.A.; Giovanni, P.; Luciano, P.; Achille, S. Protein hunger of the feed sector: The alternatives offered by the plant world. Ital. J. Anim. Sci. 2020, 19, 1204–1225. [Google Scholar] [CrossRef]
- He, J.; Tang, M.; Zhong, F.; Deng, J.; Li, W.; Zhang, L.; Lin, Q.; Xia, X.; Li, J.; Guo, T. Current trends and possibilities of typical microbial protein production approaches: A review. Crit. Rev. Biotechnol. 2024, 44, 1515–1532. [Google Scholar] [CrossRef] [PubMed]
- Tara, P.C.; Dmitry, A.R.; Andrei, L.O.; Scott, N.P. B Vitamins and their role in immune regulation and cancer. Nutrients 2020, 12, 3380. [Google Scholar] [CrossRef]
- Bogale, T.T. Microbial protein production from agro-industrial wastes as food and feed. Am. J. Life Sci. 2020, 8, 121–126. [Google Scholar] [CrossRef]
- Kashif, R.; Sabir, H.; Asif, S.; Waheed, M.; Mahmoud, K.A.; Nisar, A.; Fares, A. Comprehensive insights into sustainable conversion of agricultural and food waste into microbial protein for animal feed production. Rev. Environ. Sci. Bio. 2023, 22, 527–562. [Google Scholar] [CrossRef]
- Samara, C.A.; Erick, D.-R.; Bruna, L.; Minaxi, S.; Solange, I.M.; Kumar, T.V.; Kalaskar, D.M.; Kumar, G.V.; Anuj, K.C. Microbial meat: A sustainable vegan protein source produced from agri-waste to feed the world. Food Res. Int. 2023, 166, 112596. [Google Scholar] [CrossRef]
- Bojana, B.; Damjan, V.; Đurđina, V.; Rada, J.; Siniša, D. Biotechnological production of sustainable microbial proteins from agro-industrial residues and by-products. Foods 2022, 12, 107. [Google Scholar] [CrossRef]
- Heringer, H.C.E.; Marchioro, M.L.K.; Meneguzzi, D.; Barbosa-Dekker, A.M.; Dekker, R.F.H.; da Cunha, M.A.A. Valorization of spent Brewers yeast in the integrated production of the fungal exopolysaccharide (1→6)-β-D-glucan (lasiodiplodan) and single-cell protein. Biocatal. Agric. Biotechnol. 2023, 54, 102971. [Google Scholar] [CrossRef]
- Shurson, G.C. Yeast and yeast derivatives in feed additives and ingredients: Sources, characteristics, animal responses, and quantification methods. Anim. Feed Sci. Tech. 2018, 235, 60–76. [Google Scholar] [CrossRef]
- Øverland, M.; Karlsson, A.; Mydland, L.T.; Romarheim, O.H.; Skrede, A. Evaluation of Candida utilis, Kluyveromyces marxianus and Saccharomyces cerevisiae yeasts as protein sources in diets for Atlantic salmon (Salmo salar). Aquaculture 2013, 402–403, 1–7. [Google Scholar] [CrossRef]
- Margareth, Ø.; Anders, S. Yeast derived from lignocellulosic biomass as a sustainable feed resource for use in aquaculture. J. Sci. Food Agric. 2017, 97, 733–742. [Google Scholar] [CrossRef]
- Rosano, G.L.; Rosano, G.L.; Ceccarelli, E.A.; Ceccarelli, E.A. Recombinant protein expression in Escherichia coli: Advances and challenges. Front. Microbiol. 2014, 5, 172. [Google Scholar] [CrossRef] [PubMed]
- Duygu, A.; Özge, Ş.T.; Şerife, A.; Birsen, Y.; Eren, G.B.; Pasquale, R.; Giuseppe, S.; Fatih, Ö. Lactic acid bacteria as pro-technological, bioprotective and health-promoting cultures in the dairy food industry. Food Biosci. 2022, 47, 101617. [Google Scholar] [CrossRef]
- Bothe, H.; Jensen, K.M.; Mergel, A.; Larsen, J.; Jorgensen, C.; Bothe, H.; Jorgensen, L. Heterotrophic bacteria growing in association with Methylococcus capsulatus (Bath) in a single cell protein production process. Appl. Microbiol. Biotechnol. 2002, 59, 33–39. [Google Scholar] [CrossRef]
- Abrini, J.; Naveau, H.; Nyns, E.J. Clostridium autoethanogenum, sp. nov., an anaerobic bacterium that produces ethanol from carbon monoxide. Arch. Microbiol. 1994, 161, 345–351. [Google Scholar] [CrossRef]
- Cui, W.; Han, L.; Suo, F.; Liu, Z.; Zhou, L.; Zhou, Z. Exploitation of Bacillus subtilis as a robust workhorse for production of heterologous proteins and beyond. World J. Microb. Biotechnol. 2018, 34, 145. [Google Scholar] [CrossRef]
- Glencross, B.D.; Huyben, D.; Schrama, J.W. The application of single-cell ingredients in aquaculture feeds—A Review. Fishes 2020, 5, 22. [Google Scholar] [CrossRef]
- Muhammad, Q.W.; Simon, B.; Katerina, K.; Kjersti, U.A.; Dorinde, M.M.K.; Kari, S.; Anne, R. Protein enrichment of wheat bread with microalgae: Microchloropsis gaditana, Tetraselmis chui and Chlorella vulgaris. Foods 2021, 10, 3078. [Google Scholar] [CrossRef]
- David, L.; Gergely, K.; Line, D.H.; Liv, T.M.; Volkmar, P.; Svein, J.H.; Vincent, G.H.E. Production and characterization of yeasts grown on media composed of spruce-derived sugars and protein hydrolysates from chicken by-products. Microb. Cell Fact. 2020, 19, 1. [Google Scholar] [CrossRef]
- Cai, G.L.; Zheng, B.B.; Wang, G.; Cao, Y.; Wang, X.; Lu, J. Improvement of nutritional value of peanut meal by fermentation. China Oils Fats 2010, 35, 31–34. [Google Scholar]
- Zhang, W.W.; Zhang, H.; Zhang, W.; Su, X.R.; Xu, J.J. A marine bacterium with high-efficiency carotenoid production, its cultivation method, and applications. China Patent CN202510000932.4, 2 January 2025. issued 1 May 2025. [Google Scholar]
- Olsen, M.F.L.; Pedersen, J.S.; Thomsen, S.T.; Martens, H.J.; Petersen, A.; Jensen, P.E. Outdoor cultivation of a novel isolate of the microalgae Scenedesmus sp. and the evaluation of its potential as a novel protein crop. Physiol. Plant. 2021, 173, 483–494. [Google Scholar] [CrossRef]
- Hewavitharana, G.G.; Perera, D.N.; Navaratne, S.B.; Wickramasinghe, I. Extraction methods of fat from food samples and preparation of fatty acid methyl esters for gas chromatography: A review. Arab. J. Chem. 2020, 13, 6865–6875. [Google Scholar] [CrossRef]
- Yemm, E.W.; Willis, A.J. The estimation of carbohydrates in plant extracts by anthrone. Biochem. J. 1954, 57, 508–514. [Google Scholar] [CrossRef]
- Ye, Y.; Cai, Y.; Wang, F.; He, Y.; Yang, Y.; Guo, Z.; Liu, M.; Ren, H.; Wang, S.; Liu, D. Industrial microbial technologies for feed protein production from non-protein nitrogen. Microorganisms 2025, 13, 742. [Google Scholar] [CrossRef]
- Li, Y.P.; Ahmadi, F.; Kariman, K.; Lackner, M. Recent advances and challenges in single cell protein (SCP) technologies for food and feed production. npj Sci. Food 2024, 8, 66. [Google Scholar] [CrossRef]
- Patelski, P.; Berlowska, J.; Dziugan, P.; Pielech-Przybylska, K.; Balcerek, M.; Dziekonska, U.; Kalinowska, H. Utilisation of sugar beet bagasse for the biosynthesis of yeast SCP. J. Food Eng. 2015, 167, 32–37. [Google Scholar] [CrossRef]
- Shifa, A.B.; Meshal, H.K.A.; Othman, A.A.; Sayed, S.S.; Yasir, M.; Esam, I.A.; Hichem, C. Bacillus species as direct-fed microbial antibiotic alternatives for monogastric production. Probiotics Antimicrob. Proteins 2022, 15, 1–16. [Google Scholar] [CrossRef]
- Lokapirnasari, W.P.; Sahidu, A.M.; Nurhajati, T.; Soepranianondo, K.; Yulianto, A.B. Potency of Bacillus cereus WPL 415 to increase crude protein and decrease crude fiber of animal feed stuff. KnE Life Sci. 2017, 2017, 579–587. [Google Scholar] [CrossRef]
- Bratosin, B.C.; Darjan, S.; Vodnar, D.C. Single cell protein: A potential substitute in human and animal nutrition. Sustainability 2021, 13, 9284. [Google Scholar] [CrossRef]
- Helen, O.; Christian, K.A.; Calistus, O.; Arthur, O.; Francis, I.; Taghi, M.; Olumide, A.O.; Amarachukwu, A. Single cell protein for foods and feeds: A review of trends. Open Microbiol. J. 2022, 16, 1–17. [Google Scholar] [CrossRef]
- Raziq, A. Single cell protein (SCP) production and potential substrates: A comprehensive review. Pure Appl. Biol. 2020, 9, 1743–1754. [Google Scholar] [CrossRef]
- Montenegro-Herrera, C.A.; Portillo, F.V.L.; Hernández-Chávez, G.T.; Martinez, A. Single-cell protein production potential with the extremophilic red microalgae Galdieria sulphuraria: Growth and biochemical characterization. J. Appl. Phycol. 2022, 34, 1341–1352. [Google Scholar] [CrossRef]
- Kornochalert, N.; Kantachote, D.; Chaiprapat, S.; Techkarnjanaruk, S. Use of Rhodopseudomonas palustris P1 stimulated growth by fermented pineapple extract to treat latex rubber sheet wastewater to obtain single cell protein. Ann. Microbiol. 2014, 64, 1021–1032. [Google Scholar] [CrossRef]
- Ritala, A.; Häkkinen, T.S.; Toivari, M.; Wiebe, M.G. Single Cell Protein—State-of-the-Art, industrial landscape and patents 2001-2016. Front. Microbiol. 2017, 8, 2009. [Google Scholar] [CrossRef]
- Thu, N.T.H.; Thuý H, Đ. Isolation of a methane-oxydizing bacterium for the study on single cell protein production from methane. Vietnam. J. Biotechnol. 2016, 14, 581–588. [Google Scholar] [CrossRef]
- Patthawaro, S.; Saejung, C. Production of single cell protein from manure as animal feed by using photosynthetic bacteria. Microbiologyopen 2019, 8, 12. [Google Scholar] [CrossRef]
- Shi, S.B.; Valle-Rodríguez, J.O.; Siewers, V.; Nielsen, J. Prospects for microbial biodiesel production. Biotechnol. J. 2011, 6, 277–285. [Google Scholar] [CrossRef]
- Lee, J.A.; Kim, H.U.; Na, J.G.; Ko, Y.S.; Cho, J.S.; Lee, S.Y. Factors affecting the competitiveness of bacterial fermentation. Trends Biotechnol. 2022, 41, 798–816. [Google Scholar] [CrossRef]
- Hannah, A.; Thomas, R.; Abhijeet, B.; Arthur, R.; Dan, C. Toxicological challenges to microbial bioethanol production and strategies for improved tolerance. Ecotoxicology 2015, 24, 2156–2174. [Google Scholar] [CrossRef]
- Xu, G.; Zhao, X.; Zhao, S.; Rogers, M.J.; He, J. Salinity determines performance, functional populations, and microbial ecology in consortia attenuating organohalide pollutants. ISME J. 2023, 17, 660–670. [Google Scholar] [CrossRef]
- Fang, J.; Zhou, J.; Zeng, W. Breeding and process optimization of Yarrowia lipolytica for high-yield α-ketoglutaric acid production. Food Ferment. Ind. 2021, 47, 137–144. [Google Scholar] [CrossRef]
- Yuan, H.; Sun, Q.; Wang, L.; Fu, Z.; Zhou, T.; Ma, J.; Liu, X.; Fan, G.; Teng, C. Optimization of high-density fermentation conditions for Saccharomycopsis fibuligera Y1402 through response surface analysis. Foods 2024, 13, 1546. [Google Scholar] [CrossRef]
- Cao, G.; Guan, Z.; Liu, F.; Liao, X.; Cai, Y. Arachidonic acid production by Mortierella alpina using raw crop materials. Acta Sci. Pol. Technol. Aliment. 2015, 14, 133–143. [Google Scholar] [CrossRef]
- Su, X.; Han, Y.; Dodd, D.; Hwan, M.Y.; Shosuke, Y.; Roderick, I.M.; Isaac, K.O.C. Reconstitution of a thermostable xylan-degrading enzyme mixture from the bacterium Caldicellulosiruptor bescii. Appl. Environ. Microb. 2013, 79, 1481–1490. [Google Scholar] [CrossRef]
- Flint, H.J.; Scott, K.P.; Duncan, S.H.; Louis, P.; Forano, E. Microbial degradation of complex carbohydrates in the gut. Gut Microbes 2012, 3, 289–306. [Google Scholar] [CrossRef]
- Yang, Q.; Tian, M.; Dong, P.; Zhao, Y.; Deng, Y. Engineering Yarrowia lipolytica to enhance the production of malonic acid via malonyl-CoA pathway at high titer. Adv. Sci. 2025, 12, e2411665. [Google Scholar] [CrossRef]
- Huang, J.; An, Y.; Hossain, M.Z.; Yuvaraj, R.; Zhao, M.; Yun, J.; Zhang, G.; Zhang, Y.; Li, X.; Qi, X. Enhanced biosynthesis of D-Arabitol by Metschnikowia reukaufii through optimizing medium composition and fermentation conditions. Appl. Biochem. Biotechnol. 2022, 194, 3119–3135. [Google Scholar] [CrossRef]
- Mingwang, H.; Haiqin, C.; Xin, T.; Hengqian, L.; Jianxin, Z.; Hao, Z.; Yongquan, C.; Wei, C. Two-stage pH control combined with oxygen-enriched air strategies for the highly efficient production of EPA by Mortierella alpina CCFM698 with fed-batch fermentation. Bioprocess. Biosyst. Eng. 2020, 43, 1725–1733. [Google Scholar] [CrossRef]
- Plackett, R.L.; Burman, J.P. The design of optimum multifactorial experiments. Biometrika 1946, 33, 305–325. [Google Scholar] [CrossRef]
- Ji, X.J.; Huang, H.; Du, J.; Zhu, J.G.; Ren, L.J.; Li, S.; Nie, Z.K. Development of an industrial medium for economical 2,3-butanediol production through co-fermentation of glucose and xylose by Klebsiella oxytoca. Bioresour. Technol. 2009, 100, 5214–5218. [Google Scholar] [CrossRef]
- Liu, Q.L.; Wang, S.L.; Zhi, J.F.; Ming, H.L.; Teng, D.W. Efficient production of lactic acid from sweet sorghum juice by a newly isolated Lactobacillus salivarius CGMCC 7.75. Indian J. Microbiol. 2013, 53, 332–336. [Google Scholar] [CrossRef]
- Bezerra, M.A.; Santelli, R.E.; Oliveira, E.P.; Villar, L.S.; Escaleira, L.A. Response surface methodology (RSM) as a tool for optimization in analytical chemistry. Talanta 2008, 76, 965–977. [Google Scholar] [CrossRef]
- Box, G.E.P.; Wilson, K.B. On the experimental attainment of optimum conditions. J. R. Stat. Soc. B. 1951, 13, 1–38. [Google Scholar] [CrossRef]
| Proximate Composition | Content (Wet Weight, WW) | Content (Dry Weight, DW) | Unit |
|---|---|---|---|
| Moisture | 79.1900 ± 0.0216 | 0 | % |
| Crude ash | 2.3767 ± 0.0205 | 11.4263 ± 0.0988 | % |
| Crude protein | 15.6013 ± 0.0082 | 75.0064 ± 0.0395 | % |
| Crude lipid | 0.1023 ± 0.0026 | 0.4920 ± 0.0126 | % |
| Carbohydrates | 2.6997 ± 0.0021 | 12.9792 ± 0.0099 | % |
| Amino Acid Types | Content (Wet Weight, WW) | Content (Dry Weight, DW) | Unit |
|---|---|---|---|
| Lysine | 1.3617 ± 0.0039 | 6.5472 ± 0.0193 | % |
| Leucine | 0.8917 ± 0.0012 | 4.2869 ± 0.0066 | % |
| Valine | 0.6713 ± 0.0009 | 3.2283 ± 0.0051 | % |
| Isoleucine | 0.6293 ± 0.0041 | 3.0259 ± 0.0208 | % |
| Threonine | 0.5530 ± 0.0022 | 2.6582 ± 0.0102 | % |
| Phenylalanine | 0.5123 ± 0.0040 | 2.4639 ± 0.0189 | % |
| Tryptophan | 0.5127 ± 0.0084 | 2.4656 ± 0.0396 | % |
| Histidine | 0.4813 ± 0.0082 | 2.3110 ± 0.0413 | % |
| Methionine | 0.2820 ± 0.0036 | 1.3556 ± 0.0183 | % |
| Glutamic acid | 2.5110 ± 0.0086 | 12.0731 ± 0.0419 | % |
| Alanine | 1.1717 ± 0.0024 | 5.6330 ± 0.0110 | % |
| Aspartic acid | 1.1390 ± 0.0008 | 5.4755 ± 0.0038 | % |
| Proline | 1.0910 ± 0.0119 | 5.2446 ± 0.0560 | % |
| Glycine | 0.6677 ± 0.0026 | 3.2086 ± 0.0134 | % |
| Arginine | 0.5813 ± 0.0012 | 2.7943 ± 0.0060 | % |
| Tyrosine | 0.4680 ± 0.0051 | 2.2506 ± 0.0248 | % |
| Serine | 0.4497 ± 0.0021 | 2.1625 ± 0.0094 | % |
| Cystine | 0.0913 ± 0.0026 | 0.4390 ± 0.0119 | % |
| Total amino acids | 14.0660 ± 0.0218 | 67.6238 ± 0.1050 | % |
| Fatty Acid Types | Content (Wet Weight, WW) | Content (Dry Weight, DW) | Unit |
|---|---|---|---|
| C14:0 | 3.7375 ± 0.0065 | 17.9247 ± 0.0261 | mg/100 g |
| C14:1n5 | 141.9750 ± 0.2420 | 682.6122 ± 0.9517 | mg/100 g |
| C15:0 | 2.3615 ± 0.0055 | 11.3510 ± 0.0219 | mg/100 g |
| C15:1n5 | 49.0535 ± 0.1175 | 235.9087 ± 0.4731 | mg/100 g |
| C16:0 | 14.2405 ± 0.0255 | 68.3990 ± 0.1358 | mg/100 g |
| C18:0 | 1.6220 ± 0.0030 | 7.7949 ± 0.0126 | mg/100 g |
| C18:1n9c | 5.6290 ± 0.0020 | 27.0641 ± 0.0785 | mg/100 g |
| C18:2n6c | 8.5200 ± 0.0010 | 40.9615 ± 0.0039 | mg/100 g |
| C18:3n3 | 1.0350 ± 0.0110 | 4.9679 ± 0.0446 | mg/100 g |
| SFAs | 21.9615 ± 0.0270 | 105.4696 ± 0.1406 | mg/100 g |
| MUFAs | 196.6575 ± 0.2690 | 945.5850 ± 1.0657 | mg/100 g |
| PUFAs | 9.5550 ± 0.0110 | 45.9294 ± 0.0448 | mg/100 g |
| TFAs | 228.1740 ± 0.2706 | 1096.9840 ± 1.0758 | mg/100 g |
| Microorganism | Content (DW) | Unit | References |
|---|---|---|---|
| Fungi | |||
| Pleurotus florida | 63 | % | [33] |
| Aspergillus niger | 17–50 | % | [33] |
| Saccharomyces cerevisiae | 24–50 | % | [34] |
| Yarrowia lipolytica | 48–54 | % | [34] |
| Kluyveromyces marxianus | 59 | % | [35] |
| Geotrichum candidum | 40 | % | [36] |
| Bacteria | |||
| Rhodopseudomonas palustris | 65 | % | [37] |
| Methylococcus capsulatus | 70 | % | [38] |
| Methylomonas sp. DM580 | 76 | % | [39] |
| Rhodopseudomonas faecalis PA2 | 62.7 | % | [40] |
| Aifella marina STW181 | 46.4 | % | [40] |
| Microalgae | |||
| Galdieria sulphuraria | 44 | % | [36] |
| Nannochloris sp. | 31–68 | % | [41] |
| Phaeodactylum tricornutum | 18–57 | % | [41] |
| Archaea | |||
| Haloarcula sp. IRU1 | 76 | % | [35] |
| Parameter | Optimized Condition |
|---|---|
| Salinity | 20‰ NaCl |
| Temperature | 32 °C |
| Initial pH | 7.3 |
| Medium composition | 1% (w/v) corn flour, 1% peptone 0.3% beef extract, 0.2% KCl |
| Fermentation duration | 22 h (Flask)/32 h (10 L bioreactor, total time) |
| pH control | Uncontrolled (allowed to fluctuate freely) |
| Agitator speed | 150 rpm for initial 20 h, then 180 rpm for the remaining 12 h (10 L bioreactor) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Zhang, W.; Zhang, W.; Yin, M.; Jiao, L.; Ming, T.; Jia, X.; Gouife, M.; Xu, J.; Kong, F. Optimization of Fermentation Conditions for Enhanced Single Cell Protein Production by Rossellomorea marisflavi NDS and Nutritional Composition Analysis. Foods 2025, 14, 3066. https://doi.org/10.3390/foods14173066
Zhang H, Zhang W, Zhang W, Yin M, Jiao L, Ming T, Jia X, Gouife M, Xu J, Kong F. Optimization of Fermentation Conditions for Enhanced Single Cell Protein Production by Rossellomorea marisflavi NDS and Nutritional Composition Analysis. Foods. 2025; 14(17):3066. https://doi.org/10.3390/foods14173066
Chicago/Turabian StyleZhang, Hui, Wenwen Zhang, Wen Zhang, Minghan Yin, Lefei Jiao, Tinghong Ming, Xiwen Jia, Moussa Gouife, Jiajie Xu, and Fei Kong. 2025. "Optimization of Fermentation Conditions for Enhanced Single Cell Protein Production by Rossellomorea marisflavi NDS and Nutritional Composition Analysis" Foods 14, no. 17: 3066. https://doi.org/10.3390/foods14173066
APA StyleZhang, H., Zhang, W., Zhang, W., Yin, M., Jiao, L., Ming, T., Jia, X., Gouife, M., Xu, J., & Kong, F. (2025). Optimization of Fermentation Conditions for Enhanced Single Cell Protein Production by Rossellomorea marisflavi NDS and Nutritional Composition Analysis. Foods, 14(17), 3066. https://doi.org/10.3390/foods14173066






