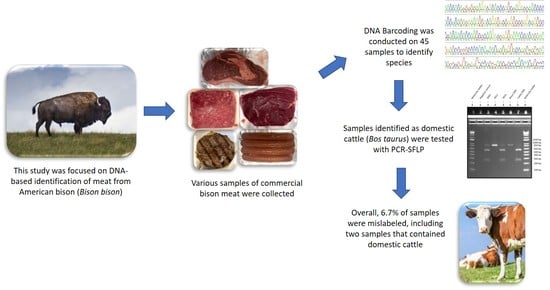
Use of DNA Barcoding Combined with PCR-SFLP to Authenticate Species in Bison Meat Products
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. DNA Extraction
2.3. DNA Barcoding
2.4. Sequence Analysis
2.5. PCR-SFLP
3. Results
3.1. DNA Barcoding
3.2. PCR-SFLP
3.3. Mislabeled Products
3.4. Additional Labeling Concerns
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Derr, J.; Hedrick, P.W.; Halbert, N.D.; Plough, L.; Dobson, L.K.; King, J.; Duncan, C.; Hunter, D.L.; Cohen, N.D.; Hedgecock, D. Phenotypic effects of cattle mitochondrial DNA in American bison. Conserv. Biol. 2012, 26, 1130–1136. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.H. How many bison originally populated western rangelands? Rangelands 1995, 17, 148–150. [Google Scholar]
- Meagher, M.M. The Bison of Yellowstone National Park; National Park Service Scientific Monograph Number One: Washington, DC, USA, 1973. [Google Scholar]
- Boyd, M.M. Crossing bison and cattle: First cross dangerous but results are better in each succeeding generation—Hope of taking fur and hump of bison and placing them upon back of domestic ox. J. Hered. 1914, 5, 189–197. [Google Scholar] [CrossRef]
- Goodnight, C. My experience with bison hybrids. J. Hered. 1914, 5, 197–199. [Google Scholar] [CrossRef]
- Hedrick, P.W. Cattle ancestry in bison: Explanations for higher mtDNA than autosomal ancestry. Mol. Ecol. 2010, 19, 3328–3335. [Google Scholar] [CrossRef] [PubMed]
- Freese, C.; Aune, K.; Boyd, D.; Derr, J.; Forrest, S.; Cormack Gates, C.; Gogan, P.; Grassel, S.; Halbert, N.; Kunkel, K.; et al. Second chance for the plains bison. Biol. Conserv. 2007, 136, 175–184. [Google Scholar] [CrossRef]
- Isenberg, A. The Destruction of the Bison: An Environmental History, 1750–1920; Cambridge University Press: Cambridge, UK, 2000; p. 222. [Google Scholar]
- Aune, K.; Jørgensen, D.; Gates, C. Bison Bison (Errata Version Published in 2018). The IUCN Red List of Threatened Species 2017: E.T2815A123789863. Available online: https://dx.doi.org/10.2305/IUCN.UK.2017-3.RLTS.T2815A45156541.en (accessed on 5 January 2021).
- Gates, C.; Freese, C.; Gogan, P.; Kotzman, M. American Bison: Status Survey and Conservation Guidelines 2010; IUCN: Gland, Switzerland, 2010; p. 134. [Google Scholar]
- National Bison Association. Current Status. Available online: https://bisoncentral.com/current-status/ (accessed on 5 January 2021).
- Tuchscherer, R. As bison sales plummet, U.S. government set to buy millions of pounds of meat. Argus Leader, 4 December 2020. [Google Scholar]
- U.S. Department of Agriculture. Economics, Statistics and Market Information System. USDA Monthly Bison (Carass and Cuts). Available online: https://usda.library.cornell.edu/concern/publications/sf268510c (accessed on 5 January 2021).
- U.S. Department of Agriculture Economic Research Service. Meat Price Spreads. Available online: https://www.ers.usda.gov/data-products/meat-price-spreads/ (accessed on 5 January 2021).
- Lianou, A.; Papakonstantinou, M.; Nychas, G.E.; Stoitsis, J. Fraud in meat and poultry products. In Food Fraud; Hellberg, R.S., Everstine, K., Sklare, S.A., Eds.; Academic Press: Cambridge, MA, USA, 2021; pp. 85–108. [Google Scholar]
- Nagarajan, M.; Nimisha, K.; Thomas, S. Molecular characterisation of ruminant mammals using DNA barcodes. In DNA Barcoding and Molecular Phylogeny; Trivedi, S., Rehman, H., Saggu, S., Panneerselvam, C., Ghosh, S., Eds.; Springer: Cham, Switzerland, 2020. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; deWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. B Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Kane, D.E.; Hellberg, R.S. Identification of species in ground meat products sold on the U.S. commercial market using DNA-based methods. Food Control 2016, 59, 158–163. [Google Scholar] [CrossRef]
- Quinto, C.A.; Tinoco, R.; Hellberg, R.S. DNA barcoding reveals mislabeling of game meat species on the U.S. commercial market. Food Control 2016, 59, 386–392. [Google Scholar] [CrossRef]
- Hellberg, R.S.; Hernandez, B.C.; Hernandez, E.L. Identification of meat and poultry species in food products using DNA barcoding. Food Control 2017, 80, 23–28. [Google Scholar] [CrossRef]
- Verkaar, E.L.C.; Nijman, I.J.; Boutaga, K.; Lenstra, J.A. Differentiation of cattle species in beef by PCR-RFLP of mitochondrial and satellite DNA. Meat Sci. 2002, 60, 365–369. [Google Scholar] [CrossRef]
- Okuma, T.; Hellberg, R. Identification of meat species in pet foods using a real-time polymerase chain reaction (PCR) assay. Food Control 2015, 50, 9–17. [Google Scholar] [CrossRef]
- Ivanova, N.; Clare, E.; Borisenko, A. DNA barcoding in mammals. Methods Mol. Biol. 2012, 858, 153–182. [Google Scholar] [CrossRef] [PubMed]
- Hellberg, R.S.; Kawalek, M.D.; Van, K.T.; Shen, Y.; Williams-Hill, D.M. Comparison of DNA extraction and PCR setup methods for use in high-throughput DNA barcoding of fish species. Food Anal. Methods 2014, 7, 1950–1959. [Google Scholar] [CrossRef]
- Marsolier-Kergoat, M.; Palacio, P.; Berthonaud, V.; Maksud, F.; Stafford, T.; Begouen, R.; Elalouf, J. Hunting the extinct steppe bison (Bison priscus) mitochondrial genome in the Trois-Freres paleolithic painted cave. PLoS ONE 2015, 10, e0128267. [Google Scholar] [CrossRef] [PubMed]
- U.S. Food and Drug Administration. Food and Drug Administration Warning Letter (CHI-03-11); FDA: Chicago, IL, USA, 2011; Czimer’s Foods, Inc 2/4/11.
- Naaum, A.; Shehata, H.; Chen, S.; Li, J.; Tabujara, N.; Awmack, D.; Lutze-Wallace, C.; Hanner, R. Complementary molecular methods detect undeclared species in sausage products at retail markets in Canada. Food Control 2018, 84, 339–344. [Google Scholar] [CrossRef]
- Shehata, H.; Naaum, A.; Chen, S.; Murphy, T.; Li, J.; Shannon, K.; Awmack, D.; Locas, A.; Hanner, R. Re-visiting the occurrence of undeclared species in sausage products sold in Canada. Food Res. Int. 2019, 122, 593–598. [Google Scholar] [CrossRef] [PubMed]
- U.S. Department of Agriculture Food Safety and Inspection Service. Bison from Farm to Table. Available online: https://www.fsis.usda.gov/wps/portal/fsis/topics/food-safety-education/get-answers/food-safety-fact-sheets/meat-preparation/focus-on-bison/ct_index (accessed on 6 January 2021).
| Collection Site | Number of Sites Visited | Number of Each Product Type Collected | Total Number of Products Collected | |||
|---|---|---|---|---|---|---|
| Cooked Burgers | Uncooked Burger Patties/Ground Meat | Uncooked Whole Cuts/Steaks | Hot Dogs | |||
| Grocery stores | 9 | 0 | 11 | 6 | 0 | 17 |
| Restaurants | 21 | 21 | 0 | 0 | 0 | 21 |
| Butchers | 4 | 0 | 1 | 3 | 0 | 4 |
| Online vendors | 1 | 0 | 1 | 1 | 1 | 3 |
| Overall | 35 | 21 | 13 | 10 | 1 | 45 |
| Primer Set | Primer Name | Primer Direction | Primer Sequence (3′-5′) 1 | Ratio in Cocktail | Product Length | Reference |
|---|---|---|---|---|---|---|
| Mammalian primer cocktail for DNA barcoding | LepF1_t1 | Forward | TGTAAAACGACGGCCAGTATTCAACCAATCATAAAGATATTGG | 1 | 658 bp 2 | Ivanova et al. [23] |
| VF1_t1 | Forward | TGTAAAACGACGGCCAGTTCTCAACCAACCACAAAGACATTGG | 1 | |||
| VF1d_t1 | Forward | TGTAAAACGACGGCCAGTTCTCAACCAACCACAA RGAYATYGG | 1 | |||
| VF1i_t1 | Forward | TGTAAAACGACGGCCAGTTCTCAACCAACCAIAAI GAIATIGG | 3 | |||
| LepR1_t1 | Reverse | CAGGAAACAGCTATGACTAAACTTCTGGATGTCC AAAAAATCA | ||||
| VR1d_t1 | Reverse | CAGGAAACAGCTATGACTAGACTTCTGGGTGGCC RAARAAYCA | 1 | |||
| VR1_t1 | Reverse | CAGGAAACAGCTATGACTAGACTTCTGGGTGGCCAAAGAATCA | 1 | |||
| VR1i_t1 | Reverse | CAGGAAACAGCTATGACTAGACTTCTGGGTGICCI AAIAAICA | 3 | |||
| Satellite 1.711b | 1.711b_F | Forward | CTGGGTGTGACAGTGTTTAAC | 1 | 822 bp | Verkaar et al. [21] |
| 1.711b_R | Reverse | TGATCCAGGGTATTCGAAGGA | 1 |
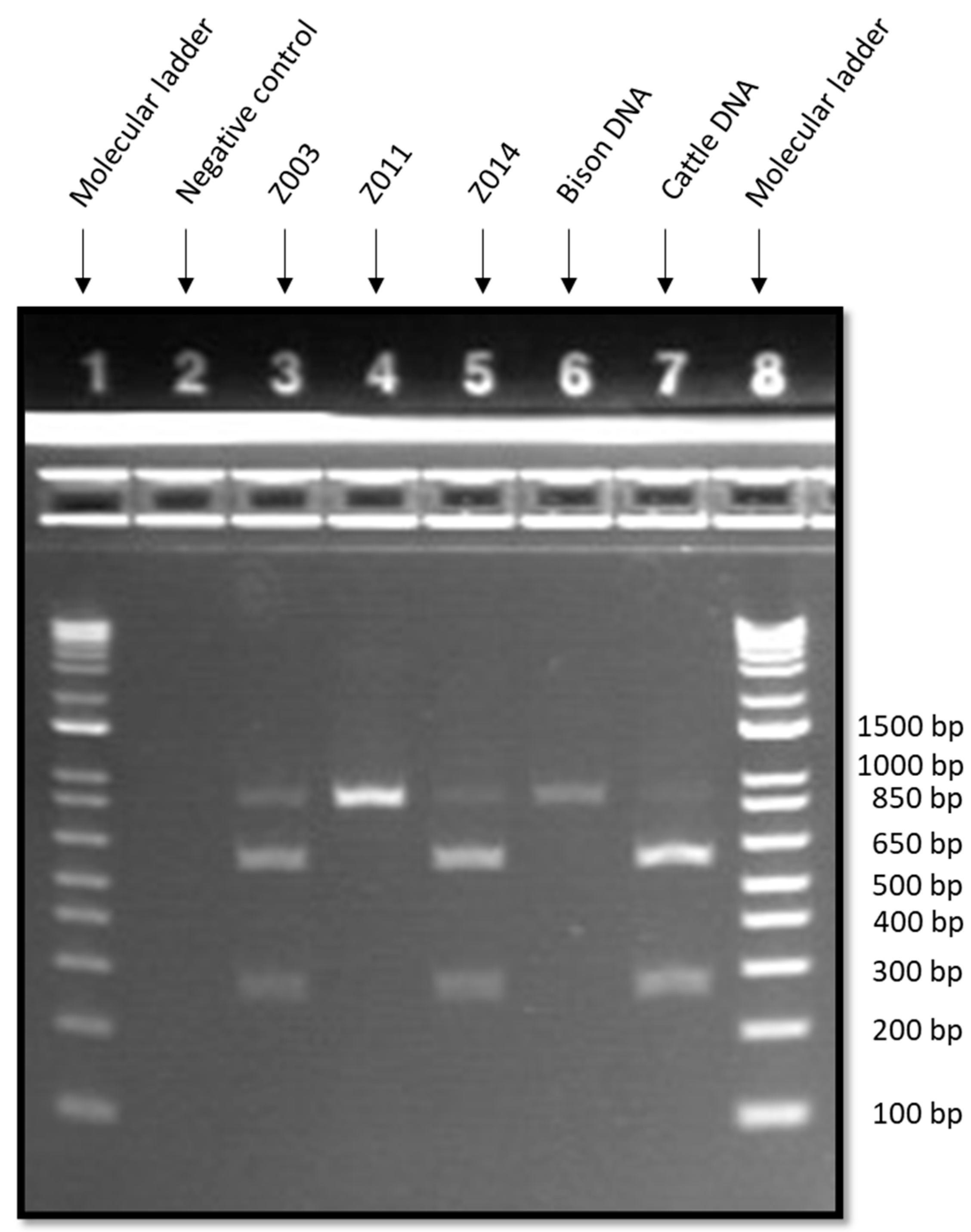
| Sample | Estimated Length of PCR-SFLP Fragments (bp) | Species Determination |
|---|---|---|
| Z003 | 257, 552, 809 | Domestic cattle (Bos taurus) |
| Z011 | 822 | American bison (Bison bison) |
| Z014 | 257, 552, 809 | Domestic cattle (Bos taurus) |
| Bison DNA positive control | 822 | American bison (Bison bison) |
| Cattle DNA positive control | 257, 552, 809 (faint) | Domestic cattle (Bos taurus) |
| Non-template control | No fragments | Negative |
| Sample ID | Product Description on Label/Menu | Purchase Location | Price Paid (USD) | Amount Purchased | Product Type | Identified Species |
|---|---|---|---|---|---|---|
| Z021 | Buffalo burger | Restaurant | $11.84 | 1 burger (no weight given) | Cooked burger | Red deer (Cervus elaphus) |
| Z003 | Fresh 90/10 ground bison | Grocery store | $9.99 | 0.45 kg prepackaged | Uncooked ground | Domestic cattle (Bos taurus) |
| Z014 | Bison burger | Restaurant | $17.64 | 1 burger (no weight given) | Cooked burger | Domestic cattle (Bos taurus) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Scales, Z.M.; Narbay, E.; Hellberg, R.S. Use of DNA Barcoding Combined with PCR-SFLP to Authenticate Species in Bison Meat Products. Foods 2021, 10, 347. https://doi.org/10.3390/foods10020347
Scales ZM, Narbay E, Hellberg RS. Use of DNA Barcoding Combined with PCR-SFLP to Authenticate Species in Bison Meat Products. Foods. 2021; 10(2):347. https://doi.org/10.3390/foods10020347
Chicago/Turabian StyleScales, Zerika M., Elif Narbay, and Rosalee S. Hellberg. 2021. "Use of DNA Barcoding Combined with PCR-SFLP to Authenticate Species in Bison Meat Products" Foods 10, no. 2: 347. https://doi.org/10.3390/foods10020347
APA StyleScales, Z. M., Narbay, E., & Hellberg, R. S. (2021). Use of DNA Barcoding Combined with PCR-SFLP to Authenticate Species in Bison Meat Products. Foods, 10(2), 347. https://doi.org/10.3390/foods10020347