Characterization of Escherichia coli from Edible Insect Species: Detection of Shiga Toxin-Producing Isolate
Abstract
:1. Introduction
2. Materials and Methods
2.1. Insect Samples
2.2. Isolate Collection and Species Identification
2.3. Antimicrobial Susceptibility Testing
2.4. Molecular Analyses
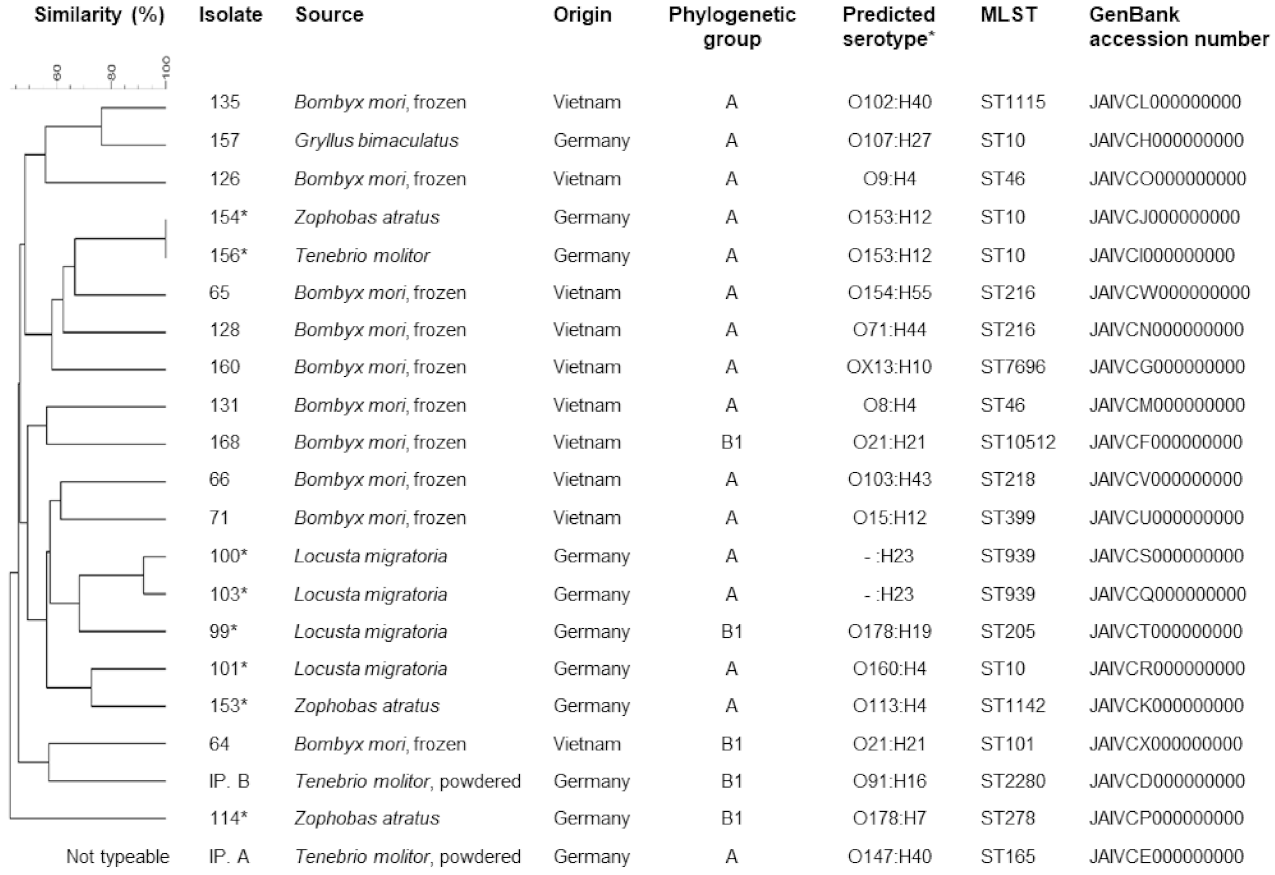
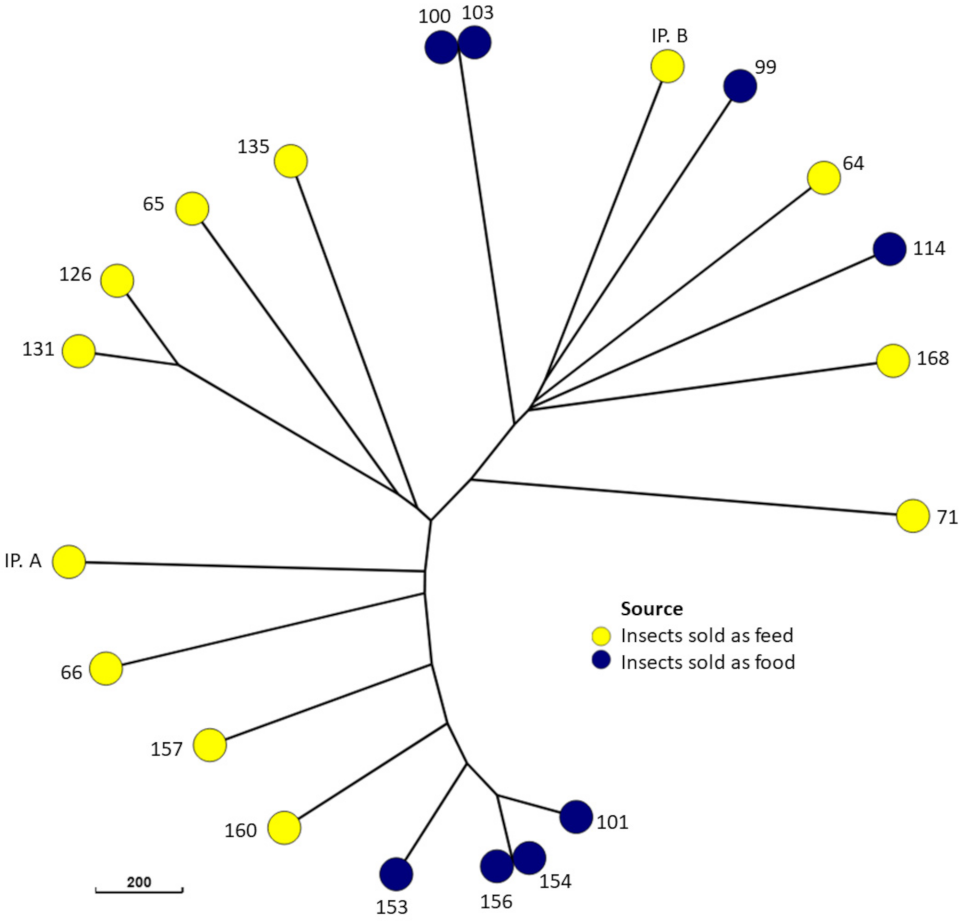
3. Results
4. Discussion
4.1. Detection of STEC in Insects Sold as Feed
4.2. Molecular Typing and Antimicrobial Resistance of E. coli from Edible Insect Species
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- EFSA Scientific Committee. Risk profile related to production and consumption of insects as food and feed. EFSA J. 2015, 13, 4257. [Google Scholar] [CrossRef] [Green Version]
- Zielińska, E.; Karaś, M.; Jakubczyk, A.; Zieliński, D.; Baraniak, B. Edible Insects as Source of Proteins. In Bioactive Molecules in Food. Reference Series in Phytochemistry; Mérillon, J.M., Ramawat, K., Eds.; Springer: Cham, Switzerland, 2018. [Google Scholar]
- EFSA Panel on Nutrition, Novel Foods and Food Allergens (NDA); Turck, D.; Castenmiller, J.; De Henauw, S.; Hirsch-Ernst, K.I.; Kearney, J.; Maciuk, A.; Mangelsdorf, I.; McArdle, H.J.; Naska, A.; et al. Safety of dried yellow mealworm (Tenebrio molitor larva) as a novel food pursuant to Regulation (EU) 2015/2283. EFSA J. 2021, 19, e06343. [Google Scholar] [PubMed]
- Grabowski, N.T.; Ahlfeld, B.; Lis, K.A.; Jansen, W.; Kehrenberg, C. The current legal status of edible insects in Europe. Berl. Und Münchener Tierärztliche Wochenschr. 2019, 132, 312–316. [Google Scholar]
- Arbeitspapier der AFFL Zum Inverkehrbringen von Insekten und Daraus Hergestellten Produkten als Lebensmittel [Working Paper of the AFFL regarding the Placing on the Market of Insects and Products thereof as Food], Internal Administrative Document; Länderarbeitsgemeinschaft Verbraucherschutz (AFFL): Wardenburg, Germany, 2019.
- Stoops, J.; Crauwels, S.; Waud, M.; Claes, J.; Lievens, B.; Van Campenhout, L. Microbial community assessment of mealworm larvae (Tenebrio molitor) and grasshoppers (Locusta migratoria migratorioides) sold for human consumption. Food Microbiol. 2016, 53, 122–127. [Google Scholar] [CrossRef] [PubMed]
- Van der Fels-Klerx, H.J.; Camenzuli, L.; Belluco, S.; Meijer, N.; Ricci, A. Food safety issues related to uses of insects for feeds and foods. Compr. Rev. Food Sci. Food Saf. 2018, 17, 1172–1183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eilenberg, J.; Vlak, J.M.; Nielsen-LeRoux, C.; Cappellozza, S.; Jensen, A.B. Diseases in insects produced for food and feed. J. Insects Food Feed. 2015, 1, 87–102. [Google Scholar]
- Grau, T.; Vilcinskas, A.; Joop, G. Sustainable farming of the mealworm Tenebrio molitor for the production of food and feed. Z. Für Nat. C 2017, 72, 337–349. [Google Scholar] [CrossRef]
- Garofalo, C.; Milanovic, V.; Cardinali, F.; Aquilanti, L.; Clementi, F.; Osimani, A. Current knowledge on the microbiota of edible insects intended for human consumption: A state-of-the-art review. Food Res. Int. 2019, 125, 108527. [Google Scholar] [CrossRef]
- Osimani, A.; Milanović, V.; Garofalo, C.; Cardinali, F.; Roncolini, A.; Sabbatini, R.; De Filippis, F.; Ercolini, D.; Gabucci, C.; Petruzzelli, A.; et al. Revealing the microbiota of marketed edible insects through PCR-DGGE, metagenomic sequencing and real-time PCR. Int. J. Food Microbiol. 2018, 276, 54–62. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA) and the European Centre for Disease Prevention and Control (ECDC). The European Union One Health 2018 Zoonoses Report. EFSA J. 2019, 17, e05926. [Google Scholar]
- Alonso, M.Z.; Lucchesi, P.M.A.; Rodríguez, E.M.; Parma, A.E.; Padola, N.L. Enteropathogenic (EPEC) and Shigatoxigenic Escherichia coli (STEC) in broiler chickens and derived products at different retail stores. Food Control 2012, 23, 351–355. [Google Scholar]
- European Food Safety Authority (EFSA) and the European Centre for Disease Prevention and Control (ECDC). The European Union summary report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2016. EFSA J. 2018, 16, e05182. [Google Scholar]
- Verraes, C.; Van Boxstael, S.; Van Meervenne, E.; Van Coillie, E.; Butaye, P.; Catry, B.; de Schaetzen, M.A.; Van Huffel, X.; Imberechts, H.; Dierick, K.; et al. Antimicrobial resistance in the food chain: A review. Int. J. Environ. Res. Public Health 2013, 10, 2643–2669. [Google Scholar] [CrossRef] [Green Version]
- Doumith, M.; Day, M.J.; Hope, R.; Wain, J.; Woodford, N. Improved multiplex PCR strategy for rapid assignment of the four major Escherichia coli phylogenetic groups. J. Clin. Microbiol. 2012, 50, 3108–3110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clinical and Laboratory Standards Institute (CLSI). Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically: Approved Standard-Ninth Edition; M07-A9; CLSI: Wayne, PA, USA, 2012. [Google Scholar]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 30th ed.; M100-ED30; CLSI: Wayne, PA, USA, 2020; Available online: http://em100.edaptivedocs.net/dashboard.aspx (accessed on 26 June 2020).
- Ribot, E.M.; Fair, M.A.; Gautom, R.; Cameron, D.N.; Hunter, S.B.; Swaminathan, B.; Barrett, T.J. Standardization of pulsed-field gel electrophoresis protocols for the subtyping of Escherichia coli O157:H7, Salmonella, and Shigella for PulseNet. Foodbourne Pathog. Dis. 2006, 3, 59–67. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.F.; et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef]
- Joensen, K.G.; Scheutz, F.; Lund, O.; Hasman, H.; Kaas, R.S.; Nielsen, E.M.; Aarestrup, F.M. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Microbiol. 2014, 52, 1501–1510. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Z.; Alikhan, N.F.; Sergeant, M.J.; Luhmann, N.; Vaz, C.; Francisco, A.P.; Carriço, J.A.; Achtman, M. GrapeTree: Visualization of core genomic relationships among 100,000 bacterial pathogens. Genome Res. 2018, 28, 1395–1404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scheutz, F.; Teel, L.D.; Beutin, L.; Piérard, D.; Buvens, G.; Karch, H.; Mellmann, A.; Caprioli, A.; Tozzoli, R.; Morabito, S.; et al. Multicenter evaluation of a sequence-based protocol for subtyping Shiga toxins and standardizing Stx nomenclature. J. Clin. Microbiol. 2012, 50, 2951–2963. [Google Scholar] [CrossRef] [Green Version]
- Grabowski, N.T.; Klein, G. Microbiology of processed edible insect products-Results of a preliminary survey. Int. J. Food Microbiol. 2017, 243, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Franzetti, E.; Huang, Z.J.; Shi, Y.X.; Xie, K.; Deng, X.J.; Li, J.P.; Li, Q.R.; Yang, W.Y.; Zeng, W.N.; Casartelli, M.; et al. Autophagy precedes apoptosis during the remodeling of silkworm larval midgut. Apoptosis 2012, 17, 305–324. [Google Scholar] [CrossRef] [PubMed]
- European Rapid Alert System for Food and Feed (RASFF). 2013. Available online: https://webgate.ec.europa.eu/rasff-window/portal/?event=notificationDetail&NOTIF_REFERENCE=2013.1211 (accessed on 25 January 2021).
- EFSA BIOHAZ Panel: Koutsoumanis, K.; Allende, A.; Alvarez-Ordóñez, A.; Bover-Cid, S.; Chemaly, M.; Davies, R.; De Cesare, A.; Herman, L.; Hilbert, F.; Lindqvist, R.; et al. Scientific Opinion on the pathogenicity assessment of Shiga toxin-producing Escherichia coli (STEC ) and the public health risk posed by contamination of food with STEC. EFSA J. 2020, 18, e05967. [Google Scholar]
- Persad, A.K.; LeJeune, J.T. Animal reservoirs of Shiga toxin-producing Escherichia coli. Microbiol. Spectr. 2014, 2, EHEC-0027-2014. [Google Scholar] [CrossRef]
- Tschinkel, W.R. Larval dispersal and cannibalism in a natural population of Zophobas atratus (Coleoptera: Tenebrionidae). Anim. Behav. 1981, 29, 990–996. [Google Scholar] [CrossRef]
- Miko, A.; Rivas, M.; Bentancor, A.; Delannoy, S.; Fach, P.; Beutin, L. Emerging types of Shiga toxin-producing E. coli (STEC) O178 present in cattle, deer, and humans from Argentina and Germany. Front. Cell. Infect. Microbiol. 2014, 4, 78. [Google Scholar] [CrossRef] [Green Version]
- Scheutz, F.; Cheasty, T.; Woodward, D.; Smith, H.R. Designation of O174 and O175 to temporary O groups OX3 and OX7, and six new E. coli O groups that include Verocytotoxin-producing E. coli (VTEC): O176, O177, O178, O179, O180 and O181. Apmis 2004, 112, 569–584. [Google Scholar] [CrossRef] [PubMed]
- Fuller, C.A.; Pellino, C.A.; Flagler, M.J.; Strasser, J.E.; Weiss, A.A. Shiga toxin subtypes display dramatic differences in potency. Infect. Immun. 2011, 79, 1329–1337. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newton, H.J.; Sloan, J.; Bulach, D.M.; Seemann, T.; Allison, C.C.; Tauschek, M.; Robins-Browne, R.M.; Paton, J.C.; Whittam, T.S.; Paton, A.W.; et al. Shiga toxin-producing Escherichia coli strains negative for locus of enterocyte effacement. Emerg. Infect. Dis. 2009, 15, 372–380. [Google Scholar] [CrossRef] [PubMed]
- García-Meniño, I.; García, V.; Mora, A.; Díaz-Jiménez, D.; Flament-Simon, S.C.; Alonso, M.P.; Blanco, J.E.; Blanco, M.; Blanco, J. Swine enteric colibacillosis in Spain: Pathogenic potential of mcr-1 ST10 and ST131 E. coli isolates. Front. Microbiol. 2018, 9, 2659. [Google Scholar] [CrossRef] [PubMed]
- Yamaji, R.; Rubin, J.; Thys, E.; Friedman, C.R.; Riley, L.W. Persistent Pandemic Lineages of Uropathogenic Escherichia coli in a College Community from 1999 to 2017. J. Clin. Microbiol. 2018, 56, e01834-17. [Google Scholar] [PubMed] [Green Version]
- Qiu, J.; Jiang, Z.; Ju, Z.; Zhao, X.; Yang, J.; Guo, H.; Sun, S. Molecular and Phenotypic characteristics of Escherichia coli isolates from farmed minks in Zhucheng, China. BioMed Res. Int. 2019, 2019, 3917841. [Google Scholar] [CrossRef] [Green Version]
- Poirel, L.; Madec, J.Y.; Lupo, A.; Schink, A.K.; Kieffer, N.; Nordmann, P.; Schwarz, S. Antimicrobial resistance in Escherichia coli. Microbiol. Spectr. 2018, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jacoby, G.A.; Strahilevitz, J.; Hooper, D.C. Plasmid-mediated quinolone resistance. Microbiol. Spectr. 2014, 2, 475–503. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fukuda, A.; Usui, M.; Okamura, M.; Dong-Liang, H.; Tamura, Y. Role of Flies in the maintenance of antimicrobial resistance in farm environments. Microb. Drug Resist. 2019, 25, 127–132. [Google Scholar] [CrossRef]
- Zurek, L.; Ghosh, A. Insects represent a link between food animal farms and the urban environment for antibiotic resistance traits. Appl. Environ. Microbiol. 2014, 80, 3562–3567. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Milanović, V.; Osimani, A.; Pasquini, M.; Aquilanti, L.; Garofalo, C.; Taccari, M.; Cardinali, F.; Riolo, P.; Clementi, F. Getting insight into the prevalence of antibiotic resistance genes in specimens of marketed edible insects. Int. J. Food Microbiol. 2016, 227, 22–28. [Google Scholar] [CrossRef]
- Osimani, A.; Cardinali, F.; Aquilanti, L.; Garofalo, C.; Roncolini, A.; Milanović, V.; Pasquini, M.; Tavoletti, S.; Clementi, F. Occurrence of transferable antibiotic resistances in commercialized ready-to-eat mealworms (Tenebrio molitor L.). Int. J. Food Microbiol. 2017, 263, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Osimani, A.; Garofalo, C.; Aquilanti, L.; Milanović, V.; Cardinali, F.; Taccari, M.; Pasquini, M.; Tavoletti, S.; Clementi, F. Transferable antibiotic resistances in marketed edible grasshoppers (Locusta migratoria migratorioides). J. Food Sci. 2017, 82, 1184–1192. [Google Scholar] [CrossRef]
- Chancey, S.T.; Zahner, D.; Stephens, D.S. Acquired inducible antimicrobial resistance in Gram-positive bacteria. Future Microbiol. 2012, 7, 959–978. [Google Scholar] [CrossRef] [Green Version]
- Milanović, V.; Osimani, A.; Roncolini, A.; Garofalo, C.; Aquilanti, L.; Pasquini, M.; Tavoletti, S.; Vignaroli, C.; Canonico, L.; Ciani, M.; et al. Investigation of the dominant microbiota in ready-to-eat grasshoppers and mealworms and quantification of carbapenem resistance genes by qPCR. Front. Microbiol. 2018, 9, 3036. [Google Scholar] [CrossRef]
- Ferri, M.; Di Federico, F.; Damato, S.; Proscia, F.; Grabowski, N.T. Insects as feed and human food and the public health risk-a review. Berl. Münchener Tierärztliche Wochenschr. 2019, 132, 191–218. [Google Scholar]
| Source | Isolate | Resistance Phenotype | Resistance Genotype | Virulence Genes |
|---|---|---|---|---|
| Insects sold as food | 64 | - | mdf(A) | gad, lpfA, terC |
| 65 | - | mdf(A) | gad, terC, traT | |
| 66 | - | mdf(A) | gad, kpsE, kpsMII, terC | |
| 71 | - | mdf(A) | gad, lpfA, terC | |
| 126 | - | qnrS2, mdf(A) | gad, celb, terC | |
| 128 | - | mdf(A) | gad, astA, terC | |
| 131 | - | mdf(A), sitABC | gad, sitA, terC, traT | |
| 135 | - | mdf(A) | gad, terC | |
| 157 | - | mdf(A) | gad, iss, terC | |
| 160 | - | mdf(A) | gad, astA, celb, fyuA, irp2, terC, traT | |
| 168 | - | mdf(A) | gad, lpfA, terC | |
| IP. A | Tetracyclines | tet(B), mdf(A) | gad, hra, terC | |
| IP. B | - | mdf(A) | gad, lpfA, terC | |
| Insects sold as feed | 99 | - | mdf(A) | gad, hra, iss, lpfA, papC, terC |
| 100 | - | mdf(A) | gad, hra, lpfA, terC | |
| 101 | - | mdf(A) | gad, iss, terC | |
| 103 | - | mdf(A) | gad, hra, lpfA, terC | |
| 114 | - | mdf(A) | gad, cba, celb, cia, cma, cvaC, ehxA, iha, ireA, iss, kpsE, lpfA, mchF, ompT, senB, stx1A, stx1B, stx2A, stx2B, subAB, terC, traT | |
| 153 | - | mdf(A) | gad, iss, ompT, terC | |
| 154 | - | mdf(A) | gad, ccl, iss, terC | |
| 156 | - | mdf(A) | gad, ccl, iss, terC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Müller, A.; Seinige, D.; Grabowski, N.T.; Ahlfeld, B.; Yue, M.; Kehrenberg, C. Characterization of Escherichia coli from Edible Insect Species: Detection of Shiga Toxin-Producing Isolate. Foods 2021, 10, 2552. https://doi.org/10.3390/foods10112552
Müller A, Seinige D, Grabowski NT, Ahlfeld B, Yue M, Kehrenberg C. Characterization of Escherichia coli from Edible Insect Species: Detection of Shiga Toxin-Producing Isolate. Foods. 2021; 10(11):2552. https://doi.org/10.3390/foods10112552
Chicago/Turabian StyleMüller, Anja, Diana Seinige, Nils T. Grabowski, Birte Ahlfeld, Min Yue, and Corinna Kehrenberg. 2021. "Characterization of Escherichia coli from Edible Insect Species: Detection of Shiga Toxin-Producing Isolate" Foods 10, no. 11: 2552. https://doi.org/10.3390/foods10112552
APA StyleMüller, A., Seinige, D., Grabowski, N. T., Ahlfeld, B., Yue, M., & Kehrenberg, C. (2021). Characterization of Escherichia coli from Edible Insect Species: Detection of Shiga Toxin-Producing Isolate. Foods, 10(11), 2552. https://doi.org/10.3390/foods10112552







