Next Generation Sequencing of Single Nucleotide Polymorphic DNA-Markers in Selecting for Intramuscular Fat, Fat Melting Point, Omega-3 Long-Chain Polyunsaturated Fatty Acids and Meat Eating Quality in Tattykeel Australian White MARGRA Lamb
Abstract
1. Introduction
2. Materials and Methods
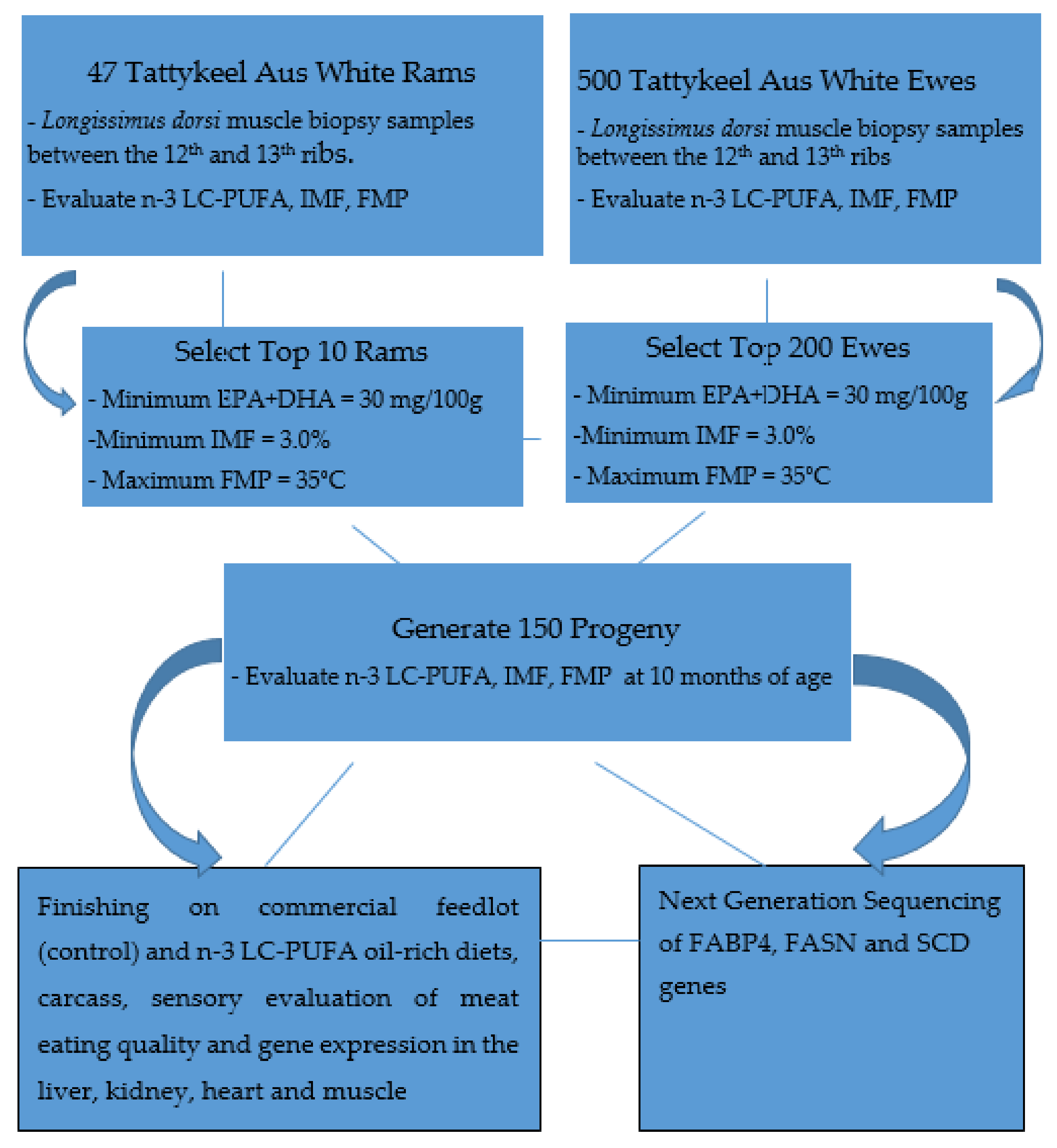
2.1. Animals and Experimental Design
2.2. Muscle Biopsy Sampling Procedure
2.3. Determination of Intramuscular Fat
2.4. Determination of Fat Melting Point
2.5. Determination of Fatty Acid Composition
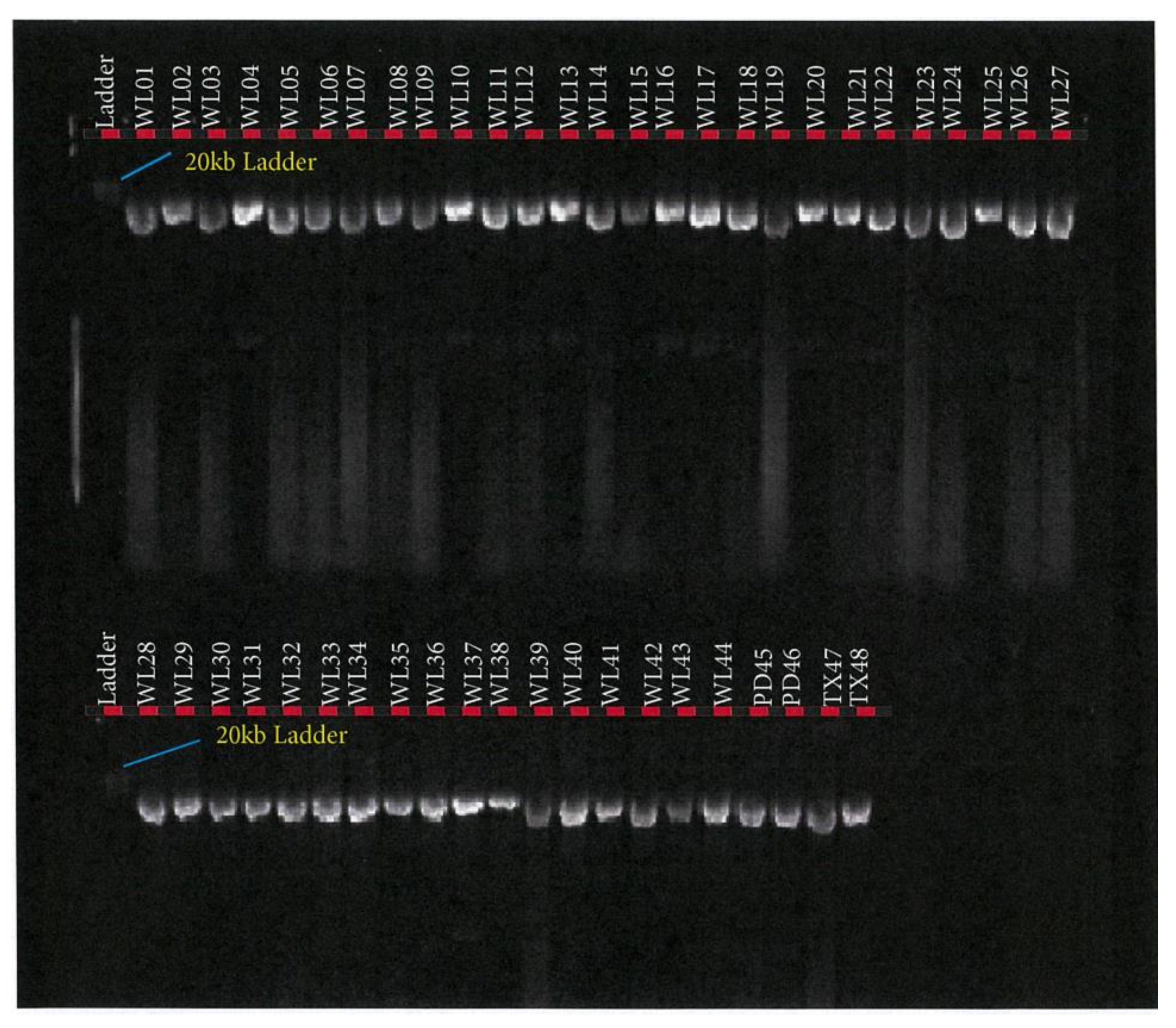
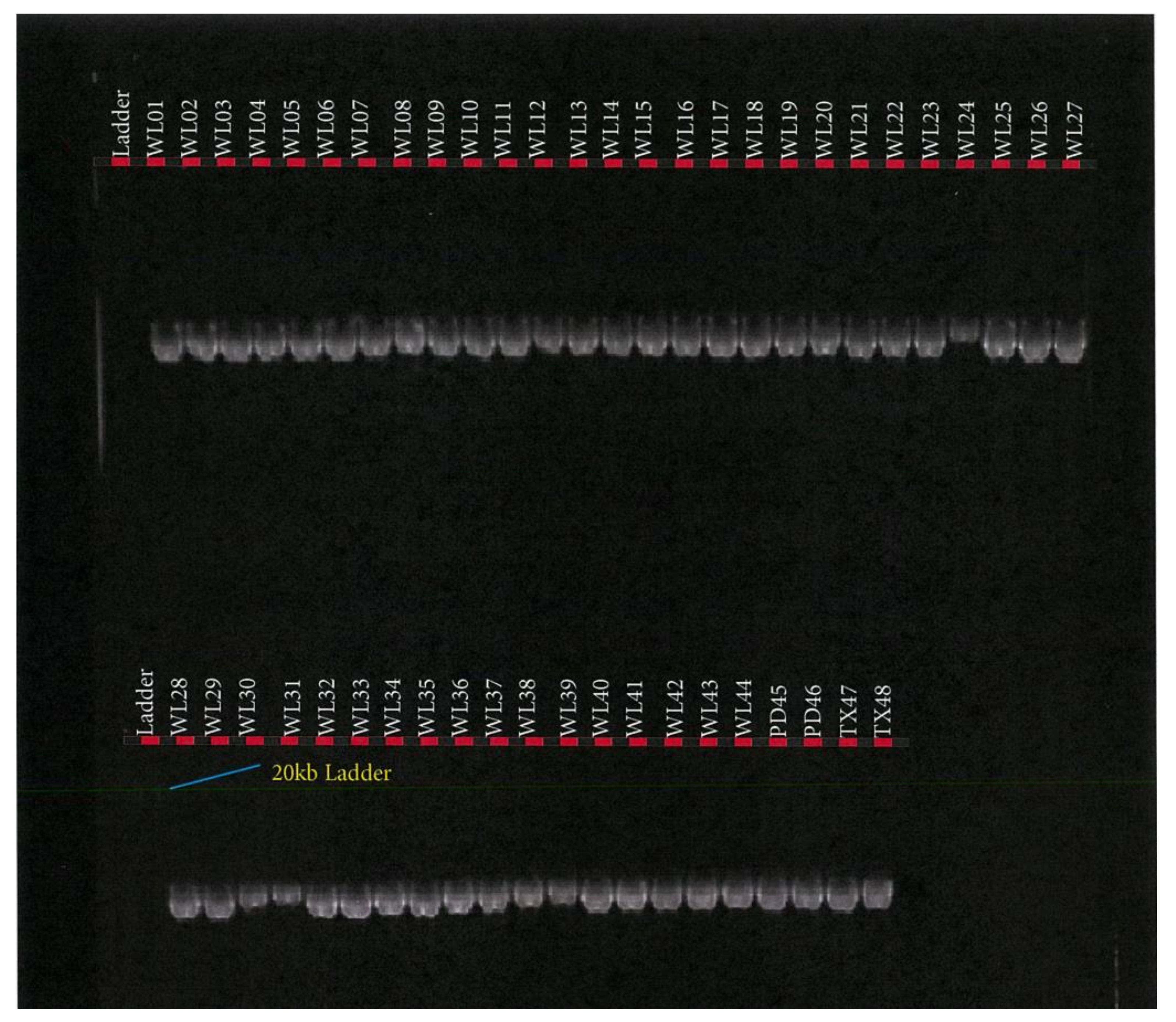
2.6. Blood Collection and Genomic DNA Extraction
2.7. Primer Design
2.7.1. FASN, FABP4, and SCD Primers
2.7.2. Long-Range PCR
2.7.3. FASN Gene
2.7.4. FABP4 and SCD
2.8. PCR Clean-Up
2.9. Library Preparation, Quantification, Normalization, and Sequencing
2.10. Bioinformatics and Next Generation Sequencing Data Analysis
2.11. Statistical Analyses
3. Results
3.1. SCD, FASN, and FABP4 Gene SNP Variants and Genotypes
3.2. Correlations between SCD, FASN, and FABP4 Gene SNP, FMP, IMF, and Fatty Acids
3.3. Associations between SCD, FASN and FABP4 SNP, FMP, IMF, and Fatty Acids
3.4. Tukey-Adjusted Multiple Comparison Tests for Significant SNP, FMP, IMF, and Fatty Acids
4. Discussion
4.1. SCD Gene Polymorphism
4.2. FASN Gene Polymorphism
4.3. FABP Gene Polymorphism
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Thomas, E.M.; Roden, J.A.; Haresign, W.; Richardson, R.; Lambe, N.R.; Clelland, N.; Gardner, G.E.; Scollan, N.D. Meat eating and nutritional quality of lambs sired by high and low muscle density rams. Animal 2021, 15, 100136. [Google Scholar] [CrossRef]
- Pewan, S.B.; Otto, J.R.; Kinobe, R.T.; Adegboye, O.A.; Malau-Aduli, A.E.O. MARGRA lamb eating quality and human health-promoting omega-3 long-chain polyunsaturated fatty acid profiles of Tattykeel Australian White Sheep: Linebreeding and gender effects. Antioxidants 2020, 9, 1118. [Google Scholar] [CrossRef]
- Heck, R.T.; Lorenzo, J.M.; Dos Santos, B.A.; Cichoski, A.J.; de Menezes, C.R.; Campagnol, P.C.B. Microencapsulation of healthier oils: An efficient strategy to improve the lipid profile of meat products. Curr. Opin. Food Sci. 2021, 40, 6–12. [Google Scholar] [CrossRef]
- Pewan, S.B.; Otto, J.R.; Huerlimann, R.; Budd, A.M.; Mwangi, F.W.; Edmunds, R.C.; Holman, B.W.B.; Henry, M.L.E.; Kinobe, R.T.; Adegboye, O.A.; et al. Genetics of omega-3 long-chain polyunsaturated fatty acid metabolism and meat eating quality in Tattykeel Australian White lambs. Genes 2020, 11, 587. [Google Scholar] [CrossRef] [PubMed]
- Gardner, G.E.; Apps, R.; McColl, R.; Craigie, C.R. Objective measurement technologies for transforming the Australian and New Zealand livestock industries. Meat Sci. 2021, 179, 108556. [Google Scholar] [CrossRef] [PubMed]
- Fowler, S.M.; Wheeler, D.; Morris, S.; Mortimer, S.I.; Hopkins, D.L. Partial least squares and machine learning for the prediction of intramuscular fat content of lamb loin. Meat Sci. 2021, 177, 108505. [Google Scholar] [CrossRef] [PubMed]
- Lambe, N.R.; Clelland, N.; Draper, J.; Smith, E.M.; Yates, J.; Bunger, L. Prediction of intramuscular fat in lamb by visible and near-infrared spectroscopy in an abattoir environment. Meat Sci. 2021, 171, 108286. [Google Scholar] [CrossRef] [PubMed]
- Alvarenga, T.I.R.C.; Hopkins, D.L.; Morris, S.; McGilchrist, P.; Fowler, S.M. Intramuscular fat prediction of the semimembranosus muscle in hot lamb carcases using NIR. Meat Sci. 2021, 181, 108404. [Google Scholar] [CrossRef] [PubMed]
- Andueza, D.; Picard, F.; Hocquette, J.F.; Listrat, A. Prediction of the intramuscular connective tissue components of fresh and freeze-dried samples by near infrared spectroscopy. Meat Sci. 2021, 179, 108537. [Google Scholar] [CrossRef]
- Patel, N.; Toledo-Alvarado, H.; Bittante, G. Performance of different portable and hand-held near-infrared spectrometers for predicting beef composition and quality characteristics in the abattoir without meat sampling. Meat Sci. 2021, 178, 108518. [Google Scholar] [CrossRef]
- Dixit, Y.; Pham, H.Q.; Realini, C.E.; Agnew, M.P.; Craigie, C.R.; Reis, M.M. Evaluating the performance of a miniaturized NIR spectrophotometer for predicting intramuscular fat in lamb: A comparison with benchtop and hand-held Vis-NIR spectrophotometers. Meat Sci. 2020, 162, 108026020. [Google Scholar] [CrossRef]
- Dixit, Y.; Hitchman, S.; Hicks, T.M.; Lim, P.; Wong, C.K.; Holibar, L.; Gordon, K.C.; Loeffen, M.; Farouk, M.M.; Craigie, C.R.; et al. Non-invasive spectroscopic and imaging systems for prediction of beef quality in a meat processing pilot plant. Meat Sci. 2021, 181, 108410. [Google Scholar] [CrossRef]
- Knight, M.I.; Linden, N.; Ponnampalam, E.N.; Kerr, M.G.; Brown, W.G.; Hopkins, D.L.; Baud, S.; Ball, A.J.; Borggaard, C.; Wesley, I. Development of VISNIR predictive regression models for ultimate pH, meat tenderness (shear force) and intramuscular fat content of Australian lamb. Meat Sci. 2019, 155, 102–108. [Google Scholar] [CrossRef]
- Knight, M.I.; Butler, K.L.; Linden, N.P.; Burnett, V.F.; Ball, A.J.; McDonagh, M.B.; Behrendt, R. Understanding the impact of sire lean meat yield breeding value on carcass composition, meat quality, nutrient and mineral content of Australian lamb. Meat Sci. 2020, 170, 108236. [Google Scholar] [CrossRef]
- Anderson, F.; Williams, A.; Pannier, L.; Pethick, D.W.; Gardner, G.E. Sire carcass breeding values affect body composition in lambs—2. Effects on fat and bone weight and their distribution within the carcass as measured by computed tomography. Meat Sci. 2016, 116, 243–252. [Google Scholar] [CrossRef]
- Stewart, S.M.; Gardner, G.E.; Williams, A.; Pethick, D.W.; McGilchrist, P.; Kuchida, K. Association between visual marbling score and chemical intramuscular fat with camera marbling percentage in Australian beef carcasses. Meat Sci. 2021, 181, 108369. [Google Scholar] [CrossRef] [PubMed]
- Connaughton, S.L.; Williams, A.; Anderson, F.; Kelman, K.R.; Peterse, J.; Gardner, G.E. Dual energy X-ray absorptiometry predicts lamb carcass composition at abattoir chain speed with high repeatability across varying processing factors. Meat Sci. 2021, 181, 108413. [Google Scholar] [CrossRef] [PubMed]
- Pannier, L.; Gardner, G.E.; Pearce, K.L.; McDonagh, M.; Ball, A.J.; Jacob, R.H.; Pethick, D.W. Associations of sire estimated breeding values and objective meat quality measurements with sensory scores in Australian lamb. Meat Sci. 2014, 96, 1076–1087. [Google Scholar] [CrossRef] [PubMed]
- Holman, B.W.B.; Hopkins, D.L. The use of conventional laboratory-based methods to predict consumer acceptance of beef and sheep meat: A review. Meat Sci. 2021, 181, 108586. [Google Scholar] [CrossRef] [PubMed]
- Malau-Aduli, A.E.O.; Siebert, B.D.; Bottema, C.D.K.; Pitchford, W.S. Breed comparison of the fatty acid composition of muscle phospholipids in Jersey and Limousin cattle. J. Anim. Sci. 1998, 76, 766–773. [Google Scholar] [CrossRef][Green Version]
- Holman, B.; Flakemore, A.; Kashan, I.A.; Malau-Aduli, A.E.O. Spirulina supplementation, sire breed, sex and basal diet effects on lamb intramuscular fat percentage and fat melting points. Int. J. Vet. Med. Res. Rep. 2014, 2014, 1–9. [Google Scholar] [CrossRef]
- Flakemore, A.R.; Balogun, R.O.; McEvoy, P.D.; Malau-Aduli, B.S.; Nichols, P.; Malau-Aduli, A.E.O. Genetic variation in intramuscular fat of prime lambs supplemented with varying concentrations of degummed crude canola oil. Int. J. Nutr. Food Sci. 2014, 3, 203–209. [Google Scholar] [CrossRef]
- Malau-Aduli, A.E.O.; Holman, B.W.B.; Kashani, A.; Nichols, P.D. Sire breed and sex effects on the fatty acid composition and content of heart, kidney, liver, adipose and muscle tissues of purebred and first-cross prime lambs. Anim. Prod. Sci. 2016, 56, 2122–2132. [Google Scholar] [CrossRef]
- Bligh, E.G.; Dyer, W.J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Phys. 1959, 37, 911–917. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.R.; Nichols, P.D.; Barnes, J.; Davies, N.W.; Peacock, E.J.; Carter, C.G. Regiospecificity profiles of storage and membrane lipids from the gill and muscle tissue of atlantic salmon (Salmo salar L.) grown at elevated temperature. Lipids 2006, 41, 865–876. [Google Scholar] [CrossRef] [PubMed]
- Clayton, E. Graham Centre Monograph No. 4: Long-Chain Omega-3 Polyunsaturated Fatty Acids in Ruminant Nutrition: Benefits to Animals and Humans; Nugent, T., Nicholls, C., Eds.; Charles Sturt University: Wagga-Wagga, NSW, Australia, 2014. [Google Scholar]
- Faircloth, B.C.; Glenn, T.C.; White, N.D. Illumina Library Prep Protocol. Release 2.1 23 February 2014. Available online: https://buildmedia.readthedocs.org/media/pdf/protocols-libprep/latest/protocols-libprep.pdf (accessed on 8 June 2021).
- Graffelman, J.; Jain, D.; Weir, B. A genome-wide study of Hardy–Weinberg equilibrium with next generation sequence data. Hum. Genet. 2017, 136, 727–741. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; Rstudio Version 1.3.1056; R Foundation for Statistical Computing: Vienna, Austria, 2021; ISBN 3-900051-07-0. Available online: http://www.R-project.org/ (accessed on 10 August 2021).
- Slatkin, M. Linkage disequilibrium—Understanding the evolutionary past and mapping the medical future. Nat. Rev. Genet. 2008, 9, 477–485. [Google Scholar] [CrossRef]
- Khatib, H. Transgenerational epigenetic inheritance in farm animals: How substantial is the evidence? Livest. Sci. 2021, 250, 104557. [Google Scholar] [CrossRef]
- Sharifi, R.S.; Noshahr, F.A.; Seifdavati, J.; Evrigh, N.H.; Cipriano-Salazar, M.; Mariezcurrena-Berasain, M.A. Comparison of haplotype method using for genomic prediction versus single SNP genotypes in sheep breeding programs. Small Rum. Res. 2021, 199, 106380. [Google Scholar] [CrossRef]
- Deus, A.R.S.D.; Silva, G.R.; Sena, L.S.; Britto, F.B.; Carvalho, D.A.D.; Freitas, J.V.G.D.; Sarmento, J.L.R. Comparison of kinship estimates in Santa Inês sheep using microsatellite and genome-wide SNP markers. Small Rum. Res. 2021, 201, 106399. [Google Scholar] [CrossRef]
- Xu, Z.; Diao, S.; Teng, J.; Chen, Z.; Feng, X.; Cai, X.; Yuan, X.; Zhang, H.; Li, J.; Zhang, Z. Breed identification of meat using machine learning and breed tag SNPs. Food Control 2021, 125, 107971. [Google Scholar] [CrossRef]
- Krivoruchko, A.; Sermyagin, A.; Saprikina, T.; Golovanova, N.; Kvochko, A.; Yatsyk, O. Genome wide associations study of single nucleotide polymorphisms with productivity parameters in Jalgin merino for identification of new candidate genes. Gene Rep. 2021, 23, 101065. [Google Scholar] [CrossRef]
- Kumar, H.; Panigrahi, M.; Saravanan, K.A.; Parida, S.; Bhushan, B.; Gaur, G.K.; Dutt, T.; Mishra, B.P.; Singh, R.K. SNPs with intermediate minor allele frequencies facilitate accurate breed assignment of Indian Tharparkar cattle. Gene 2021, 777, 145473. [Google Scholar] [CrossRef] [PubMed]
- Long, J. Parentage analysis using genome-wide high-density SNP microarray. Gene 2021, 785, 145605. [Google Scholar] [CrossRef] [PubMed]
- Tao, R.; Wang, S.; Chen, A.; Xia, R.; Zhang, X.; Yang, Q.; Qu, Y.; Zhang, S.; Li, C. Parallel sequencing of 87 STR and 294 SNP markers using the prototype of the SifaMPS panel on the MiSeq FGx™ system. Forensic Sci. Int. Genet. 2021, 52, 102490. [Google Scholar] [CrossRef] [PubMed]
- Grochowska, E.; Lisiak, D.; Akram, M.Z.; Adeniyi, O.O.; Luhken, G.; Borys, B. Association of a polymorphism in exon 3 of the IGF1R gene with growth, body size, slaughter and meat quality traits in Colored Polish Merino sheep. Meat Sci. 2021, 172, 108314. [Google Scholar] [CrossRef]
- Lopes, F.B.; Baldi, F.; Passafaro, T.L.; Brunes, L.C.; Costa, M.F.O.; Eifert, E.C.; Narciso, M.G.; Rosa, G.J.M.; Lobo, R.B.; Magnabosco, C.U. Genome-enabled prediction of meat and carcass traits using Bayesian regression, single-step genomic best linear unbiased prediction and blending methods in Nelore cattle. Animal 2021, 15, 100006. [Google Scholar] [CrossRef]
- Marín-Garzón, N.A.; Magalhães, A.F.B.; Mota, L.F.M.; Fonseca, L.F.S.; Chardulo, L.A.L.; Albuquerque, L.G. Genome-wide association study identified genomic regions and putative candidate genes affecting meat color traits in Nellore cattle. Meat Sci. 2021, 171, 108288. [Google Scholar] [CrossRef]
- Perkins, L.; Cantwell, M.; Hows, S.; Scollan, N. The impact of nutrition on intramuscular omega-3 fatty acid composition of lamb meat: A systematic review and meta-analysis. Anim. Sci. Proc. 2021, 12, 28. [Google Scholar] [CrossRef]
- Maggiolino, A.; Bragaglio, A.; Salzano, A.; Rufrano, D.; Claps, S.; Sepe, L.; Damiano, S.; Ciarcia, R.; Dinardo, F.R.; Hopkins, D.L.; et al. Dietary supplementation of suckling lambs with anthocyanins: Effects on growth, carcass, oxidative and meat quality traits. Anim. Feed Sci. Technol. 2021, 276, 114925. [Google Scholar] [CrossRef]
- Moloney, A.P.; O’Riordan, E.G.; McGee, M.; Carberry, C.M.; Moran, L.; Menamin, K.M.; Monahan, F.J. Growth, efficiency and the fatty acid composition of blood and muscle from previously grazed late-maturing bulls fed rumen protected fish oil in a high concentrate finishing ration. Livest. Sci. 2021, 244, 104344. [Google Scholar] [CrossRef]
- Realini, C.E.; Pavan, E.; Johnson, P.L.; Font-i-Furnols, M.; Jacob, N.; Agnew, M.; Craigie, C.R.; Moon, C.D. Consumer liking of M. longissimus lumborum from New Zealand pasture-finished lamb is influenced by intramuscular fat. Meat Sci. 2021, 173, 108380. [Google Scholar] [CrossRef]
- Stampa, E.; Schipmann-Schwarze, C.; Hamm, U. Consumer perceptions, preferences, and behavior regarding pasture-raised livestock products: A review. Food Qual. Prefer. 2020, 82, 103872. [Google Scholar] [CrossRef]
- López-Pedrouso, M.; Lorenzo, J.M.; Gullón, B.; Campagnol, P.C.B.; Franco, D. Novel strategy for developing healthy meat products replacing saturated fat with oleogels. Curr. Opin. Food Sci. 2021, 40, 40–45. [Google Scholar] [CrossRef]
- Liu, G.; Ding, Y.; Chen, Y.; Yang, Y. Effect of energy intake and L-carnitine on fattening performance, carcass traits, meat quality, blood metabolites, and gene expression of lamb. Small Rum. Res. 2020, 183, 106025. [Google Scholar] [CrossRef]
- Calvo, J.H.; González-Calvo, L.; Dervishi, E.; Blanco, M.; Iguácel, L.P.; Sarto, P.; Pérez-Campo, F.M.; Serrano, M.; Bolado-Carrancio, A.; Rodríguez-Rey, J.C.; et al. A functional variant in the stearoyl-CoA desaturase (SCD) gene promoter affects gene expression in ovine muscle. Livest. Sci. 2019, 219, 62–70. [Google Scholar] [CrossRef]
- Wang, X.; Fang, C.; He, H.; Cao, H.; Liu, L.; Jiang, L.; Ma, Y.; Liu, W. Identification of key genes in sheep fat tail evolution based on RNA-seq. Gene 2021, 781, 145492. [Google Scholar] [CrossRef] [PubMed]
- Aali, M.; Moradi-Shahrbabak, H.; Moradi-Shahrbabak, M.; Sadeghi, M.; Kohram, H. Polymorphism in the SCD gene is associated with meat quality and fatty acid composition in Iranian fat- and thin-tailed sheep breeds. Livest. Sci. 2016, 188, 81–90. [Google Scholar] [CrossRef]
- Alvarenga, T.I.R.C.; Chen, Y.; Lewandowski, P.; Ponnampalam, E.N.; Sadiq, S.; Clayton, E.H.; van de Ven, R.J.; Perez, J.R.O.; Hopkins, D.L. The expression of genes encoding enzymes regulating fat metabolism is affected by maternal nutrition when lambs are fed algae high in omega-3. Livest. Sci. 2016, 187, 53–60. [Google Scholar] [CrossRef]
- García-Fernández, M.; Gutiérrez-Gil, B.; García-Gámez, E.; Arranz, J.J. Genetic variability of the Stearoyl-CoA desaturase gene in sheep. Mol. Cell. Probes. 2009, 23, 107–111. [Google Scholar] [CrossRef]
- Avilés, C.; Horcada, A.; Polvillo, O.; Membrillo, A.; Anaya, G.; Molina, A.; Alcalde, M.J.; Panea, B. Association study between variability in the SCD gene and the fatty acid profile in perirenal and intramuscular fat deposits from Spanish goat populations. Small Rum. Res. 2016, 136, 127–131. [Google Scholar] [CrossRef]
- Mortimer, S.; Van der Werf, J.; Jacob, R.H.; Hopkins, D.; Pannier, L.; Pearce, K.; Gardner, G.E.; Warner, R.D.; Geesink, G.H.; Hocking Edwards, J.E.; et al. Genetic parameters for meat quality traits of Australian lamb meat. Meat Sci. 2014, 96, 1016–1024. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Chriki, S.; Ellies-Oury, M.-P.; Legrand, I.; Pogorzelski, G.; Wierzbicki, J.; Farmer, L.; Troy, D.; Polkinghorne, R.; Hocquette, J.-F.; et al. European conformation and fat scores of bovine carcasses are not good indicators of marbling. Meat Sci. 2020, 170, 108233. [Google Scholar] [CrossRef] [PubMed]
- Pannier, L.; Pethick, D.; Geesink, G.; Ball, A.; Jacob, R.; Gardner, G. Intramuscular fat in the longissimus muscle is reduced in lambs from sires selected for leanness. Meat Sci. 2014, 96, 1068–1075. [Google Scholar] [CrossRef]
- Raza, S.H.A.; Gui, L.; Khan, R.; Schreurs, N.M.; Xiaoyu, W.; Wu, S.; Mei, C.; Wang, L.; Ma, X.; Wei, D.; et al. Association between FASN gene polymorphisms, ultrasound carcass traits and intramuscular fat in Qinchuan cattle. Gene 2018, 645, 55–59. [Google Scholar] [CrossRef]
- Armstrong, E.; Ciappesoni, G.; Iriarte, W.; Da Silva, C.; Macedo, F.; Navajas, E.A.; Brito, G.; San Julián, R.; Gimeno, D.; Postiglioni, A. Novel genetic polymorphisms associated with carcass traits in grazing Texel sheep. Meat Sci. 2018, 145, 202–208. [Google Scholar] [CrossRef]
- Dervishi, E.; Serrano, C.; Joy, M.; Serrano, M.; Rodellar, C.; Calvo, J.H. The effect of feeding system in the expression of genes related with fat metabolism in semitendinous muscle in sheep. Meat Sci. 2011, 89, 91–97. [Google Scholar] [CrossRef]
- Sanz, A.; Serrano, C.; Ranera, B.; Dervishi, E.; Zaragoza, P.; Calvo, J.H.; Rodellar, C. Novel polymorphisms in the 5′UTR of FASN, GPAM, MC4R and PLIN1 ovine candidate genes: Relationship with gene expression and diet. Small Rum. Res. 2015, 123, 70–74. [Google Scholar] [CrossRef]
- Fang, X.; Zhao, Z.; Jiang, P.; Haibin, X.; Hang, Y.; Yang, R. Identification of the bovine HSL gene expression profiles and its association with fatty acid composition and fat deposition traits. Meat Sci. 2017, 131, 107–118. [Google Scholar] [CrossRef]
- Papaleo-Mazzucco, J.; Goszczynski, D.E.; Ripoli, M.V.; Meluccia, L.M.; Pardo, A.M.; Colatto, E.; Rogberg-Muñoz, A.; Mezzadra, C.A.; Depetris, G.J.; Giovambattista, G.; et al. Growth, carcass and meat quality traits in beef from Angus, Hereford and cross-breed grazing steers, and their association with SNPs in genes related to fat deposition metabolism. Meat Sci. 2016, 114, 121–129. [Google Scholar] [CrossRef]
- Barton, L.; Bures, D.; Kott, T.; Rehak, D. Associations of polymorphisms in bovine DGAT1, FABP4, FASN, and PPARGC1A genes with intramuscular fat content and the fatty acid composition of muscle and subcutaneous fat in Fleckvieh bulls. Meat Sci. 2016, 114, 18–23. [Google Scholar] [CrossRef]
- Yeon, S.H.; Lee, S.H.; Choi, B.H.; Lee, H.J.; Jang, G.W.; Lee, K.T.; Kim, K.H.; Lee, J.H.; Chung, H.Y. Genetic variation of FASN is associated with fatty acid composition of Hanwoo. Meat Sci. 2013, 94, 133–138. [Google Scholar] [CrossRef]
- Zappaterra, M.; Luis, D.; Zambonelli, P.; Mele, M.; Serra, A.; Costa, L.N.; Davoli, R. Association study between backfat fatty acid composition and SNPs in candidate genes highlights the effect of FASN polymorphism in large white pigs. Meat Sci. 2019, 156, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, J.; Gong, H.; Cui, L.; Zhang, W.; Ma, J.; Chen, C.; Ai, H.; Xiao, S.; Huang, L.; et al. Genetic correlation of fatty acid composition with growth, carcass, fat deposition and meat quality traits based on GWAS data in six pig populations. Meat Sci. 2019, 150, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Renaville, B.; Bacciu, N.; Lanzoni, M.; Mossa, F.; Piasentier, E. Association of single nucleotide polymorphisms in fat metabolism candidate genes with fatty acid profiles of muscle and subcutaneous fat in heavy pigs. Meat Sci. 2018, 139, 220–227. [Google Scholar] [CrossRef] [PubMed]
- Zappaterra, M.; Deserti, M.; Mazza, R.; Braglia, S.; Zambonelli, P.; Davoli, R. A gene and protein expression study on four porcine genes related to intramuscular fat deposition. Meat Sci. 2016, 121, 27–32. [Google Scholar] [CrossRef]
- Ates, S.; Keles, G.; Demirci, U.; Dogan, S.; Kirbas, M.; Filley, S.J.; Parker, N.B. The effects of feeding system and breed on the performance and meat quality of weaned lambs. Small Rum. Res. 2020, 192, 106225. [Google Scholar] [CrossRef]
- Yan, W.; Zhou, H.; Hu, J.; Luo, Y.; Hickford, J.G.H. Variation in the FABP4 gene affects carcass and growth traits in sheep. Meat Sci. 2018, 145, 334–339. [Google Scholar] [CrossRef] [PubMed]
- Bai, J.L.; Xu, H.W.; Zang, R.X.; He, H.J.; Cai, Y.; Cao, X.; Peng, F.J.; Han, J.; Wu, J.P.; Yang, J.T. Cloning of the heart fatty acid-binding protein (H-FABP) gene and its tissue-specific expression profile in the Lanzhou fat-tailed sheep, Ovis aries. Small Rum. Res. 2013, 112, 114–122. [Google Scholar] [CrossRef]
- Malau-Aduli, A.E.O.; Nguyen, D.V.; Le, V.H.; Nguyen, Q.V.; Otto, J.R.; Malau-Aduli, B.S.; Nichols, P.D. Correlations between growth and wool quality traits of genetically divergent Australian lambs in response to canola or flaxseed oil supplementation. PLoS ONE 2019, 14, e0208229. [Google Scholar] [CrossRef]
- Le, H.V.; Nguyen, D.V.; Nguyen, Q.V.; Malau-Aduli, B.S.; Nichols, P.D.; Malau-Aduli, A.E.O. Fatty acid profiles of muscle, liver, heart and kidney of Australian prime lambs fed different polyunsaturated fatty acids enriched pellets in a feedlot system. Sci. Rep. 2019, 9, 1238. [Google Scholar] [CrossRef]
- Nguyen, Q.V.; Malau-Aduli, B.S.; Cavalieri, J.; Nichols, P.D.; Malau-Aduli, A.E.O. Enhancing omega-3 long-chain polyunsaturated fatty acid content of dairy-derived foods for human consumption. Nutrients 2019, 11, 743. [Google Scholar] [CrossRef]
- Nguyen, Q.V.; Le, H.V.; Nguyen, D.V.; Malau-Aduli, B.S.; Nichols, P.D.; Malau-Aduli, A.E.O. Enhancement of dairy sheep cheese eating quality with increased omega-3 long-chain polyunsaturated fatty acids. J. Dairy Sci. 2019, 102, 211–222. [Google Scholar] [CrossRef]
- Nguyen, Q.V.; Le, V.H.; Nguyen, D.V.; Malau-Aduli, B.S.; Nichols, P.D.; Malau-Aduli, A.E.O. Supplementing grazing dairy ewes with oil and rumen-protected EPA + DHA pellets enhances health-beneficial n-3 long-chain polyunsaturated fatty acids in sheep milk. Eur. J. Lipid Sci. Technol. 2018, 120, 1700256. [Google Scholar] [CrossRef]
- Nguyen, Q.V.; Le, H.V.; Nguyen, D.V.; Nish, P.; Otto, J.R.; Malau-Aduli, B.S.; Nichols, P.D.; Malau-Aduli, A.E.O. Supplementing dairy ewes grazing low quality pastures with plant-derived and rumen-protected oils containing eicosapentaenoic acid and docosahexaenoic acid pellets increases body condition score and milk, fat, and protein yields. Animals 2018, 8, 241. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.V.; Malau-Aduli, B.S.; Cavalieri, J.; Nichols, P.D.; Malau-Aduli, A.E.O. Supplementation with plant-derived oils rich in omega-3 polyunsaturated fatty acids for lamb production. Vet. Anim. Sci. 2018, 6, 29–40. [Google Scholar] [CrossRef]
- Nguyen, D.V.; Le, V.H.; Nguyen, Q.V.; Malau-Aduli, B.S.; Nichols, P.D.; Malau-Aduli, A.E.O. Omega–3 long-chain fatty acids in the heart, kidney, liver and plasma metabolite profiles of Australian prime lambs supplemented with pelleted canola and flaxseed oils. Nutrients 2017, 9, 893. [Google Scholar] [CrossRef]
- Nguyen, D.V.; Flakemore, A.R.; Otto, J.R.; Ives, S.W.; Smith, R.W.; Nichols, P.D.; Malau-Aduli, A.E.O. Nutritional value and sensory characteristics of meat eating quality of Australian prime lambs supplemented with pelleted canola and flaxseed oils: Fatty acid profiles of muscle and adipose tissues. Intern. Med. Rev. 2017, 3, 1–21. [Google Scholar] [CrossRef]
- Flakemore, A.R.; Malau-Aduli, B.S.; Nichols, P.D.; Malau-Aduli, A.E.O. Omega-3 fatty acids, nutrient retention values, and sensory meat eating quality in cooked and raw Australian lamb. Meat Sci. 2017, 123, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Fowler, S.M.; Morris, S.; Hopkins, D.L. Nutritional composition of lamb retail cuts from the carcases of extensively finished lambs. Meat Sci. 2019, 154, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, D.B.; Medeiros, G.R.; Guim, A.; Azevedo, P.S.; Suassuna, D.M.L., Jr.; Maciel, M.V.; Costa, C.A.; Lopes, L.A.; Silva, J.L.; Véras, A.S.C.; et al. Growth performance, carcass traits and meat quality of lambs fed with increasing levels of spineless cactus. Anim. Feed Sci. Technol. 2021, 272, 114788. [Google Scholar] [CrossRef]
- Díaz, M.T.; Pérez, C.; Sánchez, C.I.; Lauzurica, S.; Cañeque, V.; González, C.; De La Fuente, J. Feeding microalgae increases omega 3 fatty acids of fat deposits and muscles in light lambs. J. Food Compos. Anal. 2017, 56, 115–123. [Google Scholar] [CrossRef]
- Ponnampalam, E.N.; Butler, K.L.; Jacob, R.H.; Pethick, D.W.; Ball, A.J.; Hocking Edwards, J.E.; Geesink, G.; Hopkins, D.L. Health beneficial long chain omega-3 fatty acid levels in Australian lamb managed under extensive finishing systems. Meat Sci. 2014, 96, 1104–1110. [Google Scholar] [CrossRef]
- Ponnampalam, E.N.; Butler, K.L.; Pearce, K.M.; Mortimer, S.I.; Pethick, D.W.; Ball, A.J.; Hopkins, D.L. Sources of variation of health claimable long chain omega-3 fatty acids in meat from Australian lamb slaughtered at similar weights. Meat Sci. 2014, 96, 1095–1103. [Google Scholar] [CrossRef]
- Ponnampalam, E.N.; Sinclair, A.J.; Egan, A.R.; Ferrier, G.R.; Leury, B.J. Dietary manipulation of muscle long-chain omega-3 and omega-6 fatty acids and sensory properties of lamb meat. Meat Sci. 2002, 60, 125–132. [Google Scholar] [CrossRef]
- Ponnampalam, E.N.; Trout, G.R.; Sinclair, A.J.; Egan, A.R.; Leury, B.J. Comparison of the color stability and lipid oxidative stability of fresh and vacuum packaged lamb muscle containing elevated omega-3 and omega-6 fatty acid levels from dietary manipulation. Meat Sci. 2001, 58, 151–161. [Google Scholar] [CrossRef]
- Álvarez-Rodríguez, J.; Ripoll, G.; Lobón, S.; Sanz, A.; Blanco, M.; Joy, M. Alfalfa but not milk in lamb’s diet improves meat fatty acid profile and α-tocopherol content. Food Res. Int. 2018, 107, 708–716. [Google Scholar] [CrossRef]
- Knight, M.I.; Daetwyler, H.D.; Hayes, B.J.; Hayden, M.J.; Ball, A.J.; Pethick, D.W.; McDonagh, M.B. An independent validation association study of carcass quality, shear force, intramuscular fat percentage and omega-3 polyunsaturated fatty acid content with gene markers in Australian lamb. Meat Sci. 2014, 96, 1025–1033. [Google Scholar] [CrossRef]
- Malau-Aduli, A.E.O.; Kashani, A. Molecular genetics-nutrition interactions in the expression of AANAT, ADRB3, BTG2 and FASN genes in the heart, kidney and liver of Australian lambs supplemented with Spirulina (Arthrospira platensis). Genes Genom. 2015, 37, 633–644. [Google Scholar] [CrossRef]
- Kashani, A.; Holman, B.W.B.; Malau-Aduli, A.E.O. Single nucleotide polymorphisms of the ovine ADRB3 gene in crossbred Australian sheep supplemented with Spirulina (Arthrospira platensis) cyanobacterial microalgae. Biomed. J. Sci. Tech. Res. 2017, 1, 462–467. [Google Scholar] [CrossRef]
- Mallick, R.; Basak, S.; Duttaroy, A.K. Docosahexaenoic acid, 22:6n-3: Its roles in the structure and function of the brain. Int. J. Dev. Neurosci. 2019, 79, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Shang, P.; Zhang, B.; Zhang, J.; Duan, M.; Wu, L.; Gong, X.; Tang, K.; Zhang, H.; Chamba, Y. Expression and single-nucleotide polymorphisms of the H-FABP gene in pigs. Gene 2019, 710, 156–160. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, Y.H.; Zhang, S.; Li, F.; Wang, S.; Dai, L.; Jiang, H.; Xia, S.; Liu, D.; Sun, B.; et al. Association of A-FABP gene polymorphism in intron 1 with meat quality traits in Junmu No. 1 white swine. Gene 2011, 487, 170–173. [Google Scholar] [CrossRef] [PubMed]






| Gene | Sequence | Length (bp) | Ta (°C) | Fragment Length (bp) | |
|---|---|---|---|---|---|
| FASN 1 | Forward | CCTACTTTCCCATGCTCAGAGAA | 23 | 68 | 7890 |
| Reverse | CTACGTTGCTGAGGAAGAACTCTA | 24 | 68 | ||
| FASN 2 | Forward | ACCGTCTCTCCTTCTTCTTTGAC | 23 | 68 | 8798 |
| Reverse | GAAGTTGAGGGAGGCGTAATAGAT | 24 | 68 | ||
| FASN 3 | Forward | CTAGAGTTCTTCCTCAGCAACGTA | 24 | 68 | 9288 |
| Reverse | GCCAGGGAGCTGTGAATAATACTA | 24 | 68 | ||
| FABP4 | Forward | TTGTTGAATGGCTGGGCTTATAAC | 24 | 60 | 4107 |
| Reverse | TAAGAAAATACTTCCTGGGGCACA | 24 | 60 | ||
| SCD | Forward | CAAACTTAGGTCTGCAACTTTCGT | 24 | 65 | 11,545 |
| Reverse | TTTCCCACTTCAACTCACCCTATT | 24 | 65 |
| Lamb breed, generation, type of control and genotypes (major allele frequencies in brackets) Parental composites 1st (F1) and 2nd (F2) composites Positive (+) and negative (−) controls | ||||||
| SNP Locus | TAW Parents (n = 147) | TAW F1 (n = 75) | TAW F2 (n = 75) | Poll Dorset (+n = 2) | Texel (+n = 2) | Rambouillet (−n = 2) |
| g.23880613A>G | GG (0.82) | GG (0.93) | GG (0.73) | GG | GG | AA |
| g.23881050T>C | CT (0.58) | CT (0.54) | CT (0.90) | CC | CC | TT |
| g.23883280G>A | AG (0.53) | AG (0.71) | AG (0.60) | AA | AA | GG |
| g.23885910C>A | AC (0.57) | AC (0.71) | AC (0.53) | CC | CC | CC |
| g.23887165A>G | GA (0.69) | GG (0.82) | GG (0.70) | GG | GG | AA |
| g.23888763C>T | TC (0.58) | TC (0.54) | CC (0.93) | CC | CC | CC |
| g.23889346T>G | GT (0.68) | GG (0.82) | GG (0.70) | GG | GG | TT |
| g.23890209T>C | CT (0.67) | CC (0.82) | CC (0.70) | CC | CC | TT |
| Lamb breed, generation, type of control, and genotypes (major allele frequencies in brackets) Parental composite 1st (F1) and 2nd (F2) composites, Positive (+) and negative (−) controls | ||||||
| SNP Locus | TAW Parents (n = 147) | TAW F1 (n = 75) | TAW F2 (n = 75) | Poll Dorset (+n = 2) | Texel (+n = 2) | Rambouillet (−n = 2) |
| g.12316077T>G | GG (0.89) | GG (0.86) | GG (0.95) | GG | GG | TT |
| g.12318491A>G | GG (0.89) | GG (0.86) | GG (0.95) | GG | GG | AA |
| g.12320583T>C | CC (0.89) | CC (0.86) | CC (0.97) | CC | CC | TT |
| g.12321671T>C | CC (0.89) | CC (0.86) | CC (0.97) | CC | CC | TT |
| g.12323864A>G | GA (0.70) | GA (0.69) | GA (0.70) | GG | GG | AA |
| g.12324288G>A | AG (0.69) | AG (0.68) | AG (0.69) | AA | AA | GG |
| g.12326992T>C | CC (0.88) | CC (0.79) | CC (0.90) | CC | CC | TT |
| g.12327084->CT | CT (0.50) | CT (0.50) | CT (0.50) | CT | CT | TT |
| g.12328120T>C | CC (0.89) | CC (0.86) | CC (0.97) | CC | CC | TT |
| Lamb breed, generation, type of control, and genotypes (major allele frequencies in brackets) Parental composites 1st (F1) and 2nd (F2) composites Positive (+) and negative (−) controls | ||||||
| SNP Locus | TAW Parents (n = 147) | TAW F1 (n = 75) | TAW F2 (n = 75) | Poll Dorset (+n = 2) | Texel (+n = 2) | Rambouillet (−n = 2) |
| g.62826961T>C | CT (0.61) | TT (0.64) | CT (0.60) | TT | TT | TT |
| g.62826965C>G | GC (0.61) | GC (0.57) | GC (0.60) | GG | GG | CC |
| g.62829478A>T | AT (0.55) | AT (0.61) | AT (0.53) | AA | AA | AA |
| SNP effect (p-values) SCD FABP4 FASN | ||||||
| Variable | Mean | SD | CV (%) | g.23881050T>C | g.62829478A>T | g.12323864A>G |
| FMP (°C) | 33.65 | 2.74 | 8.14 | 0.2700 | 0.6115 | 0.0544 * |
| IMF (%) | 4.43 | 1.31 | 29.58 | 0.0089 ** | 0.0539 * | 0.1915 |
| Fatty acids (mg/100 g) | ||||||
| ALA (C18:3n-3) | 163.03 | 192.27 | 117.94 | 0.7755 | 0.1419 | 0.0033 ** |
| EPA (C20:5n-3) | 25.20 | 11.62 | 46.10 | 0.7683 | 0.1023 | 0.9810 |
| DHA (C22:6n-3) | 8.43 | 4.16 | 49.27 | 0.0111 * | 0.2145 | 0.9480 |
| DPA (C22:5n-3) | 23.85 | 13.70 | 57.44 | 0.0532 * | 0.3894 | 0.0927 |
| EPA + DHA | 33.64 | 14.75 | 43.84 | 0.2036 | 0.4794 | 0.9915 |
| EPA + DHA + DPA | 57.49 | 26.97 | 46.92 | 0.0728 | 0.8958 | 0.2004 |
| MUFA | 3694.70 | 4099.08 | 110.94 | 0.6824 | 0.3949 | 0.0025 ** |
| SFA | 4392.18 | 5238.81 | 119.28 | 0.4000 | 0.5472 | 0.0029 ** |
| C18:2n-6 | 253.68 | 247.70 | 97.64 | 0.6781 | 0.0647 | 0.0138 * |
| C14:0 | 287.92 | 437.58 | 151.98 | 0.0632 | 0.7354 | 0.1190 |
| C16:0 | 2076.17 | 2419.46 | 116.53 | 0.5414 | 0.3751 | 0.0039 ** |
| C18:0 | 1683.83 | 2065.71 | 122.68 | 0.3891 | 0.9125 | 0.0012 ** |
| C18:1n-9 | 2901.10 | 3212.65 | 110.74 | 0.8555 | 0.3696 | 0.0023 ** |
| Multiple Genotype Comparisons | ||||||
| SNP Locus | Variable | Mean ± SE | Genotypes | Difference ± SE | p-Value | |
| SCD g.23881050T>C | DHA (C22:6n-3) (mg/100 g) | |||||
| CC | 7.00 ± 2.11 | CC vs. | CT | −0.639 ± 0.834 | 0.7247 | |
| CT | 7.64 ± 2.09 | CC vs. | TT | −3.998 ± 1.334 | 0.0105 * | |
| TT | 11.00 ± 2.34 | CT vs. | TT | −3.359 ± 1.235 | 0.0223 * | |
| IMF (%) | ||||||
| CC | 3.98 ± 0.312 | CC vs. | CT | −0.407 ± 0.323 | 0.4224 | |
| CT | 4.39 ± 0.287 | CC vs. | TT | −1.446 ± 0.532 | 0.0222 * | |
| TT | 5.43 ± 0.516 | CT vs. | TT | −1.038 ± 0.502 | 0.1041 | |
| DPA (C22:5n-3) (mg/100 g) | ||||||
| CC | 17.9 ± 6.81 | CC vs. | CT | −1.56 ± 2.65 | 0.8270 | |
| CT | 19.4 ± 6.74 | CC vs. | TT | −9.19 ± 4.25 | 0.0850 | |
| TT | 27.1 ± 3.26 | CT vs. | TT | −7.63 ± 3.93 | 0.0356 * | |
| FASN g.12323864A>G | FMP (°C) | |||||
| GG | 34.2 ± 0.4 | GG vs. | GA | 0.81 ± 0.64 | 0.4201 | |
| GA | 33.4 ± 0.3 | GG vs. | AA | 2.98 ± 1.61 | 0.0536 * | |
| AA | 31.5 ± 1.5 | GA vs. | AA | 2.16 ± 1.60 | 0.3685 | |
| ALA (C18:3n-3) (mg/100 g) | ||||||
| GG | 188.7 ± 67.6 | GG vs. | GA | 114.7 ± 39.9 | 0.0149 * | |
| GA | 74.0 ± 66.7 | GG vs. | AA | 147.2 ± 100.1 | 0.3115 | |
| AA | 41.5 ± 113.7 | GA vs. | AA | 32.6 ± 99.8 | 0.9430 | |
| MUFA (mg/100 g) | ||||||
| GG | 4524 ± 1384 | GG vs. | GA | 2617 ± 867 | 0.0099 ** | |
| GA | 1907 ± 1361 | GG vs. | AA | 3089 ± 2175 | 0.3363 | |
| AA | 1436 ± 2415 | GA vs. | AA | 472 ± 2168 | 0.9742 | |
| SFA (mg/100 g) | ||||||
| GG | 5479 ± 1715 | GG vs. | GA | 3270 ± 1121 | 0.0132 * | |
| GA | 2208 ± 1684 | GG vs. | AA | 4162 ± 2812 | 0.3068 | |
| AA | 1317 ± 3086 | GA vs. | AA | 892 ± 2803 | 0.9458 | |
| C18:2n-6 (mg/100 g) | ||||||
| GG | 281 ± 84.8 | GG vs. | GA | 142.5 ± 52.2 | 0.0216 * | |
| GA | 139 ± 83.4 | GG vs. | AA | 105.8 ± 130.8 | 0.6988 | |
| AA | 175 ± 146.4 | GA vs. | AA | −36.7 ± 130.4 | 0.9573 | |
| C16:0 (mg/100 g) | ||||||
| GG | 2539 ± 800 | GG vs. | GA | 1475 ± 518 | 0.0158 * | |
| GA | 1063 ± 786 | GG vs. | AA | 1826 ± 1298 | 0.3433 | |
| AA | 713 ± 1429 | GA vs. | AA | 350 ± 1294 | 0.9604 | |
| C18:0 (mg/100 g) | ||||||
| GG | 2227 ± 658 | GG vs. | GA | 1419 ± 441 | 0.0056 ** | |
| GA | 809 ± 646 | GG vs. | AA | 1711 ± 1106 | 0.2756 | |
| AA | 516 ± 1205 | GA vs. | AA | 292 ± 1102 | 0.9620 | |
| C18:1n-9 (mg/100 g) | ||||||
| GG | 3589 ± 1078 | GG vs. | GA | 2103 ± 679 | 0.0080 ** | |
| GA | 1486 ± 1060 | GG vs. | AA | 2353 ± 1704 | 0.3566 | |
| AA | 1236 ± 1892 | GA vs. | AA | 250 ± 1698 | 0.9882 | |
| FABP4 g.62829478A>T | IMF (%) | |||||
| A | 4.57 ± 0.39 | A vs. | AA | 0.07 ± 0.344 | 0.0556 | |
| AA | 3.92 ± 0.39 | |||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pewan, S.B.; Otto, J.R.; Huerlimann, R.; Budd, A.M.; Mwangi, F.W.; Edmunds, R.C.; Holman, B.W.B.; Henry, M.L.E.; Kinobe, R.T.; Adegboye, O.A.; et al. Next Generation Sequencing of Single Nucleotide Polymorphic DNA-Markers in Selecting for Intramuscular Fat, Fat Melting Point, Omega-3 Long-Chain Polyunsaturated Fatty Acids and Meat Eating Quality in Tattykeel Australian White MARGRA Lamb. Foods 2021, 10, 2288. https://doi.org/10.3390/foods10102288
Pewan SB, Otto JR, Huerlimann R, Budd AM, Mwangi FW, Edmunds RC, Holman BWB, Henry MLE, Kinobe RT, Adegboye OA, et al. Next Generation Sequencing of Single Nucleotide Polymorphic DNA-Markers in Selecting for Intramuscular Fat, Fat Melting Point, Omega-3 Long-Chain Polyunsaturated Fatty Acids and Meat Eating Quality in Tattykeel Australian White MARGRA Lamb. Foods. 2021; 10(10):2288. https://doi.org/10.3390/foods10102288
Chicago/Turabian StylePewan, Shedrach Benjamin, John Roger Otto, Roger Huerlimann, Alyssa Maree Budd, Felista Waithira Mwangi, Richard Crawford Edmunds, Benjamin William Behrens Holman, Michelle Lauren Elizabeth Henry, Robert Tumwesigye Kinobe, Oyelola Abdulwasiu Adegboye, and et al. 2021. "Next Generation Sequencing of Single Nucleotide Polymorphic DNA-Markers in Selecting for Intramuscular Fat, Fat Melting Point, Omega-3 Long-Chain Polyunsaturated Fatty Acids and Meat Eating Quality in Tattykeel Australian White MARGRA Lamb" Foods 10, no. 10: 2288. https://doi.org/10.3390/foods10102288
APA StylePewan, S. B., Otto, J. R., Huerlimann, R., Budd, A. M., Mwangi, F. W., Edmunds, R. C., Holman, B. W. B., Henry, M. L. E., Kinobe, R. T., Adegboye, O. A., & Malau-Aduli, A. E. O. (2021). Next Generation Sequencing of Single Nucleotide Polymorphic DNA-Markers in Selecting for Intramuscular Fat, Fat Melting Point, Omega-3 Long-Chain Polyunsaturated Fatty Acids and Meat Eating Quality in Tattykeel Australian White MARGRA Lamb. Foods, 10(10), 2288. https://doi.org/10.3390/foods10102288








