Hydrophobic Natural Deep Eutectic Solvents for Extraction of Bioactive Compounds: Multiscale Characterization, Quantum Simulations, and Molecular Interaction Studies with Cry j 1 and Amb a 1 Allergens
Abstract
1. Introduction
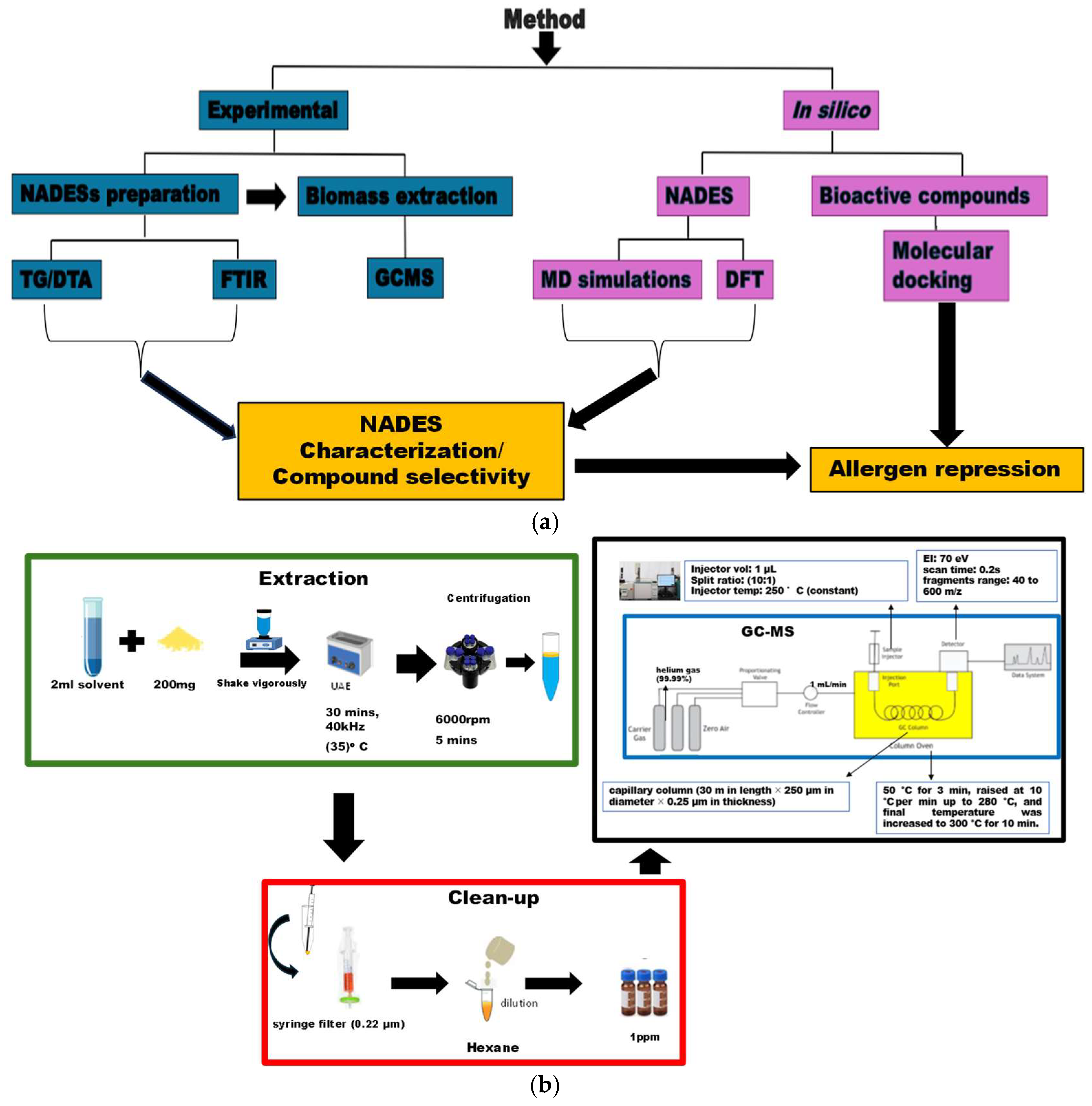
2. Materials and Methods
2.1. Experimental
2.1.1. Chemicals/Reagents
2.1.2. Synthesis of the Natural Deep Eutectic Solvents
2.1.3. NADES Characterization
2.1.4. Ultrasonic Assisted Extraction
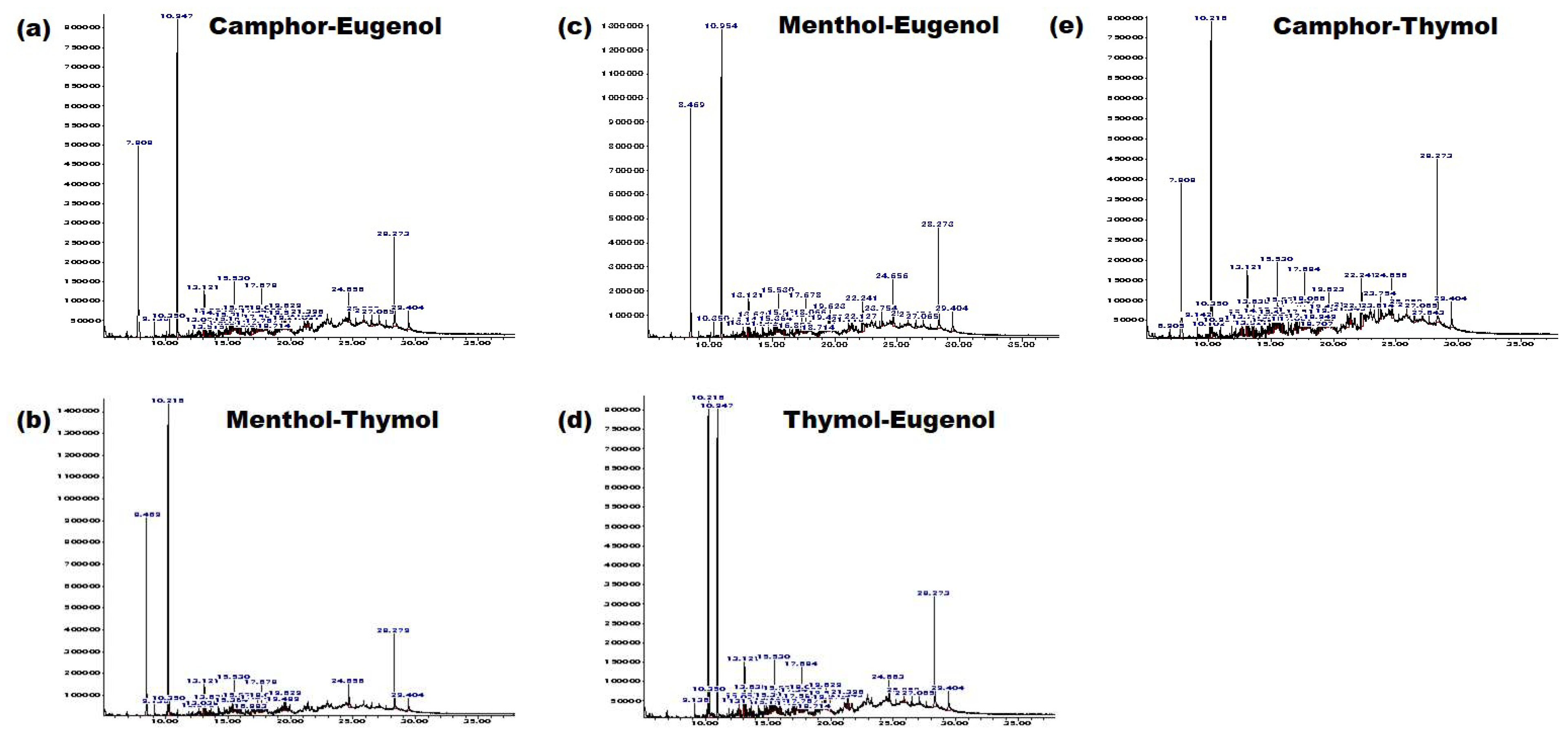
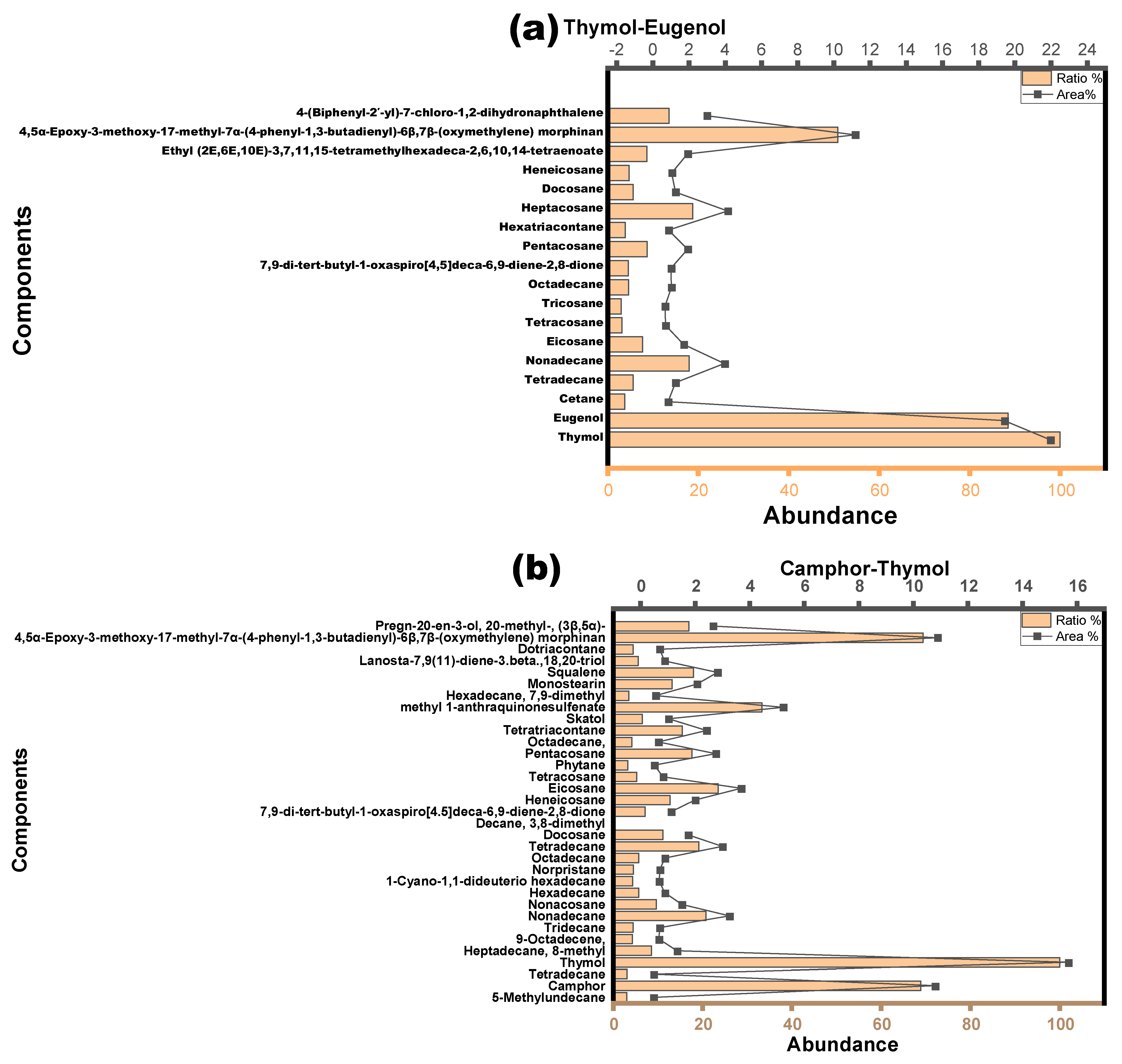
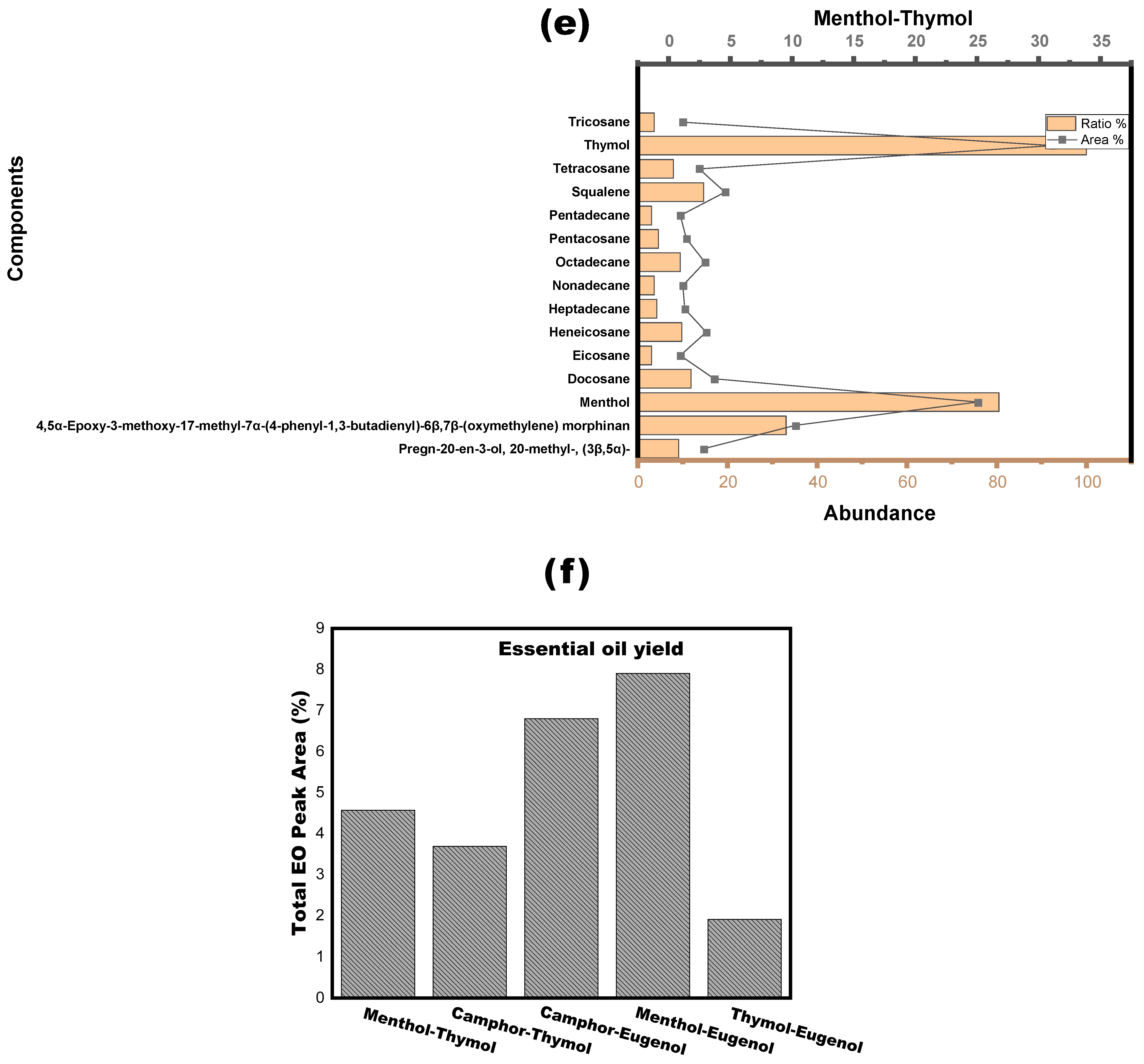
2.1.5. Determination of Components of the Extract
2.2. In Silico Methods
2.2.1. Density Functional Theory Calculations
2.2.2. Molecular Dynamics Simulations
2.2.3. Molecular Docking Simulation
2.3. Statistical Analysis
3. Results and Discussion
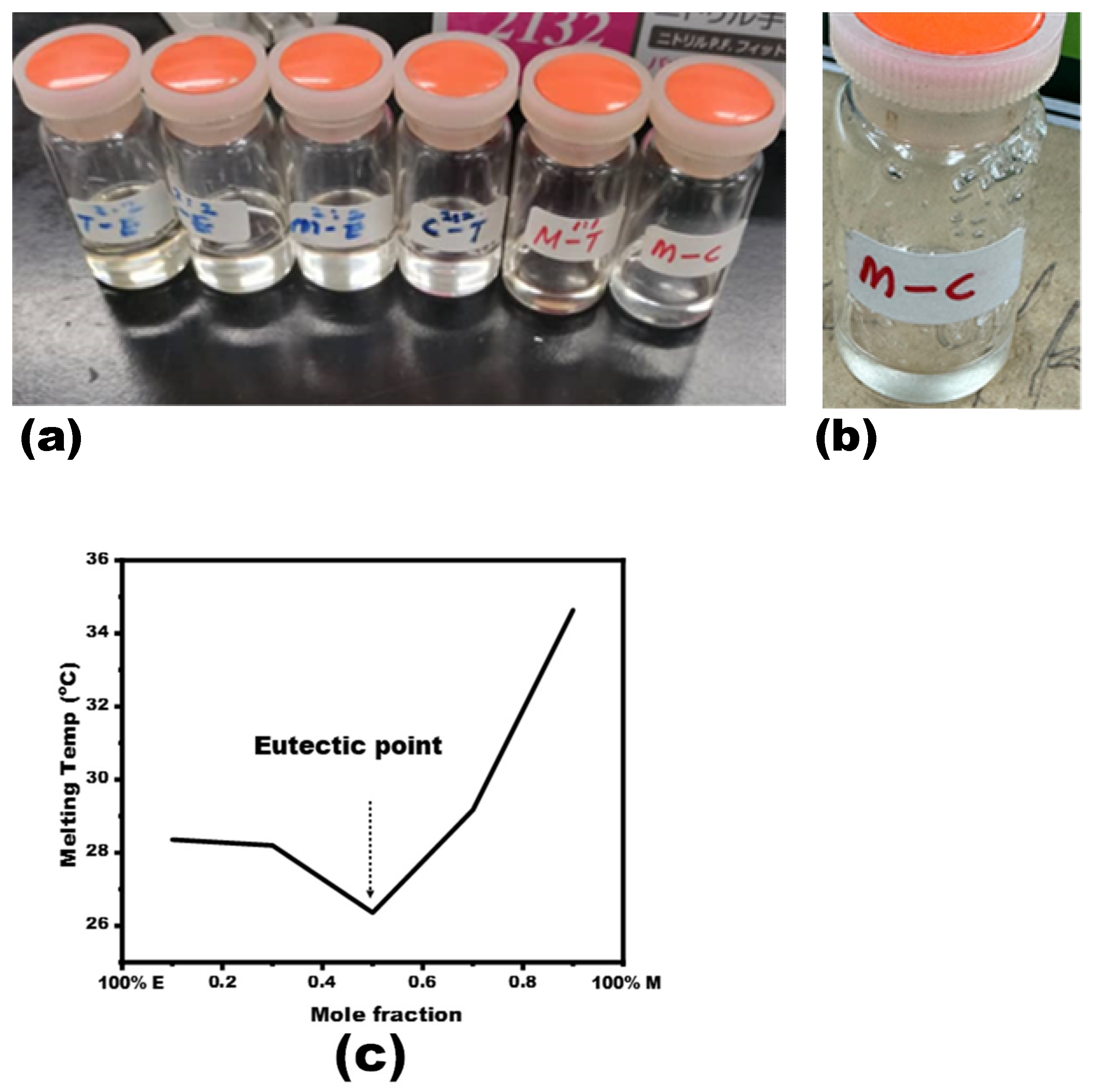
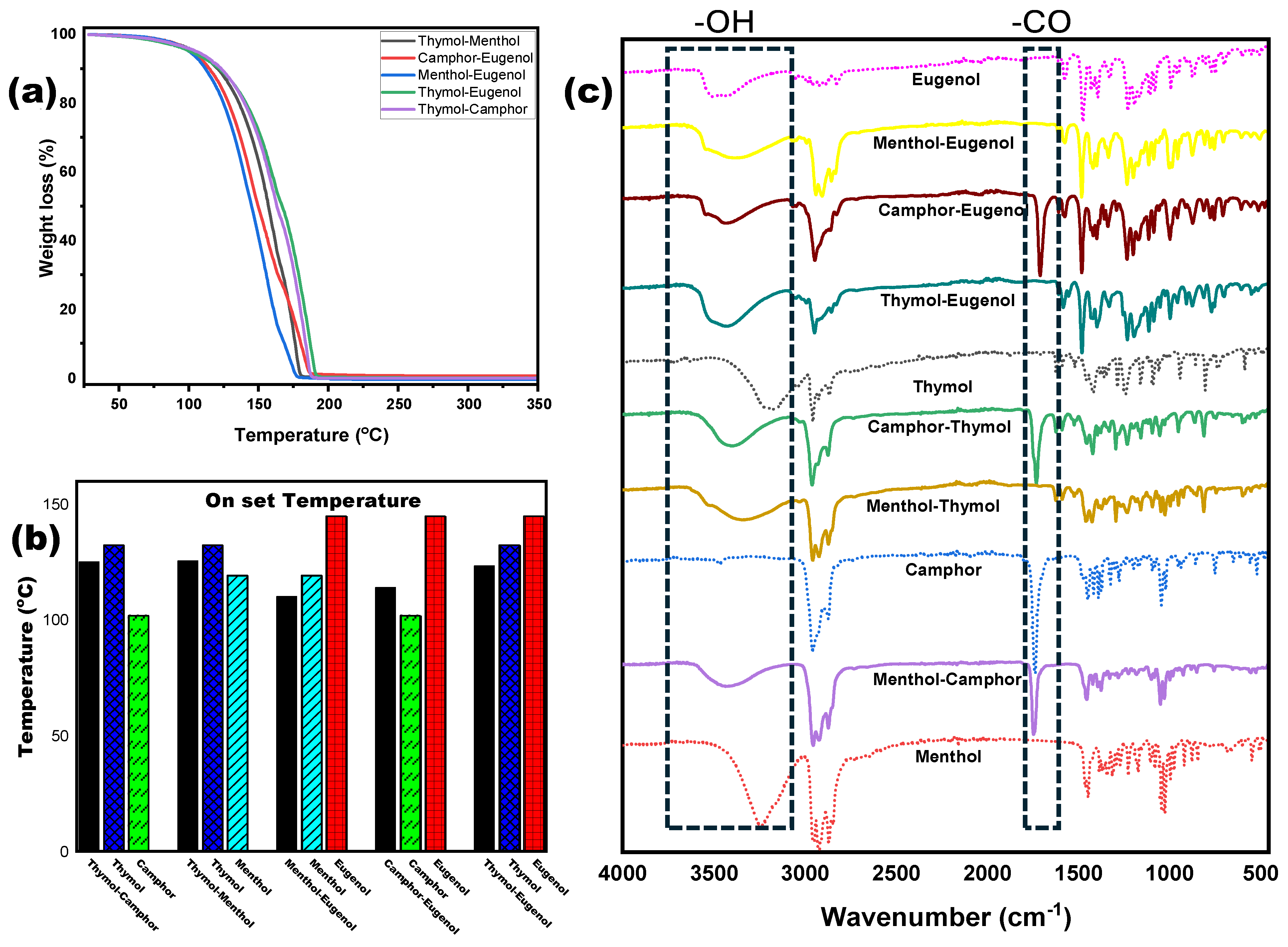
3.1. NADES Synthesis and Characterization
3.2. NADESs as a Green Separation Tool
3.3. In Silico Simulation
3.3.1. Density Functional Theory (DFT)
Frontier Molecular Orbital Analysis
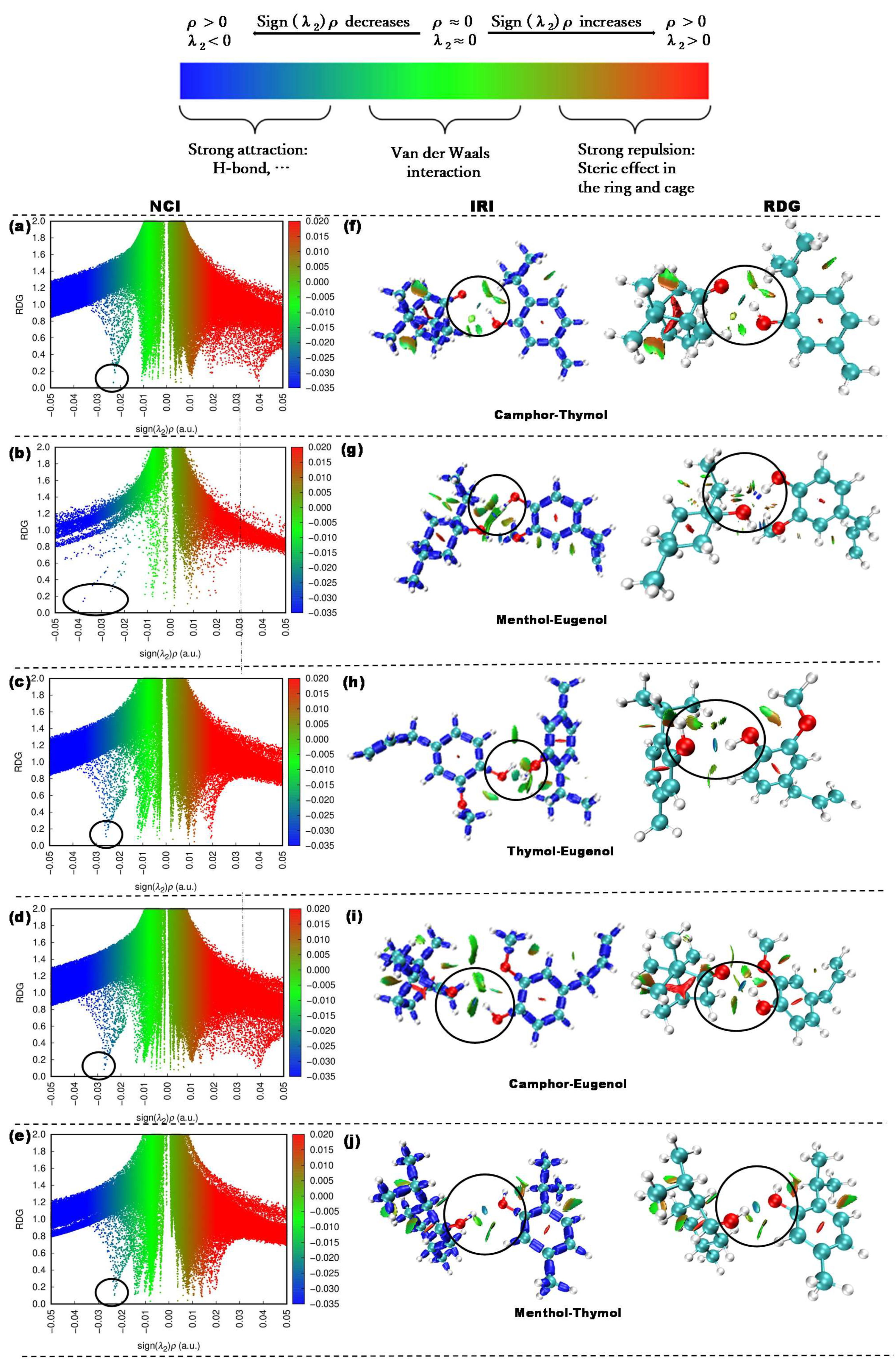
- Investigation of hydrogen bond interactions in the NADES system (QTAIM, NCI, and IRI analysis).
3.3.2. Molecular Dynamics Simulations of the NADESs
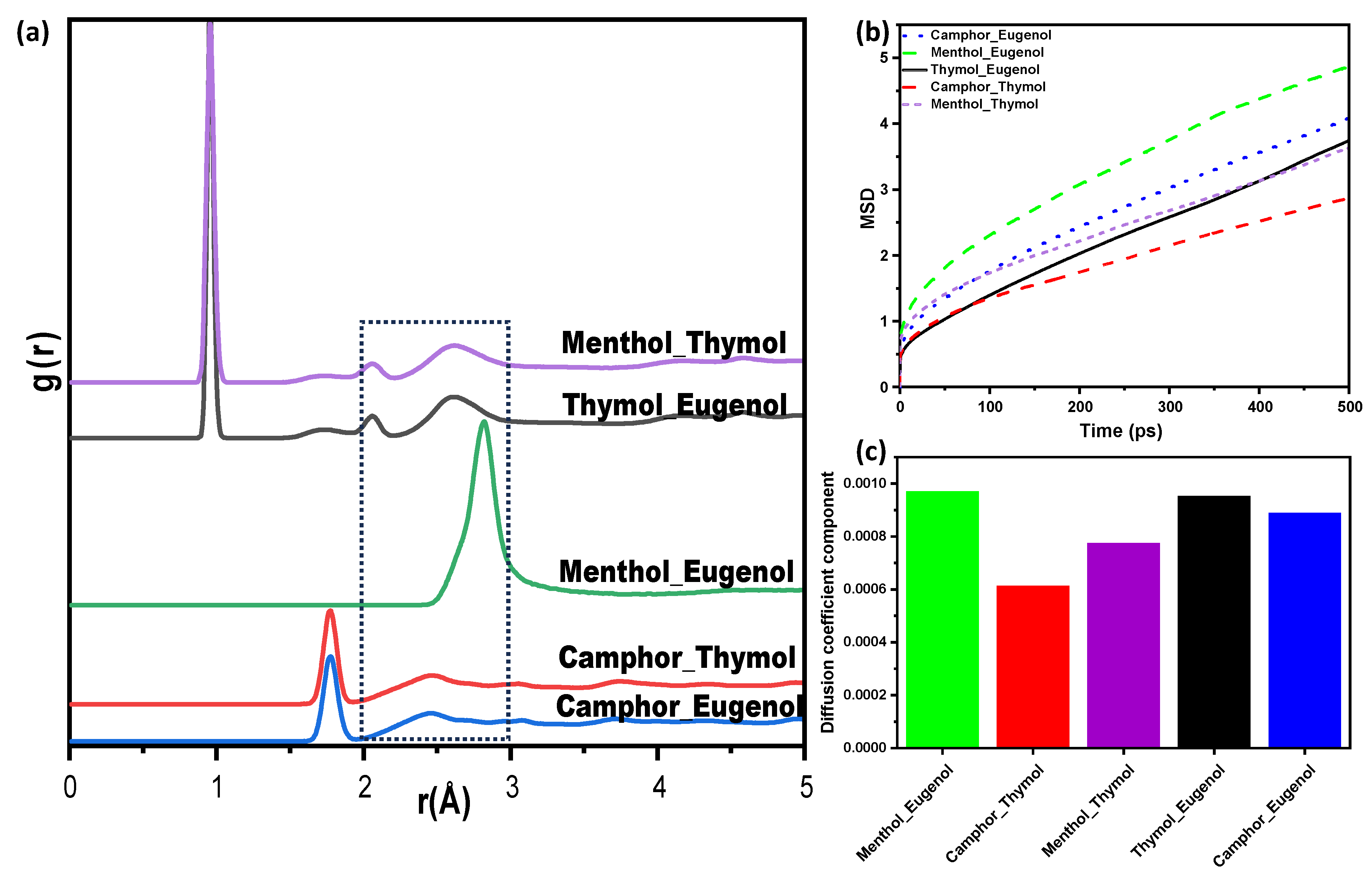
Mean Square Displacement (MSD) and Diffusion Coefficient
Radial Distribution Functions (RDFs)
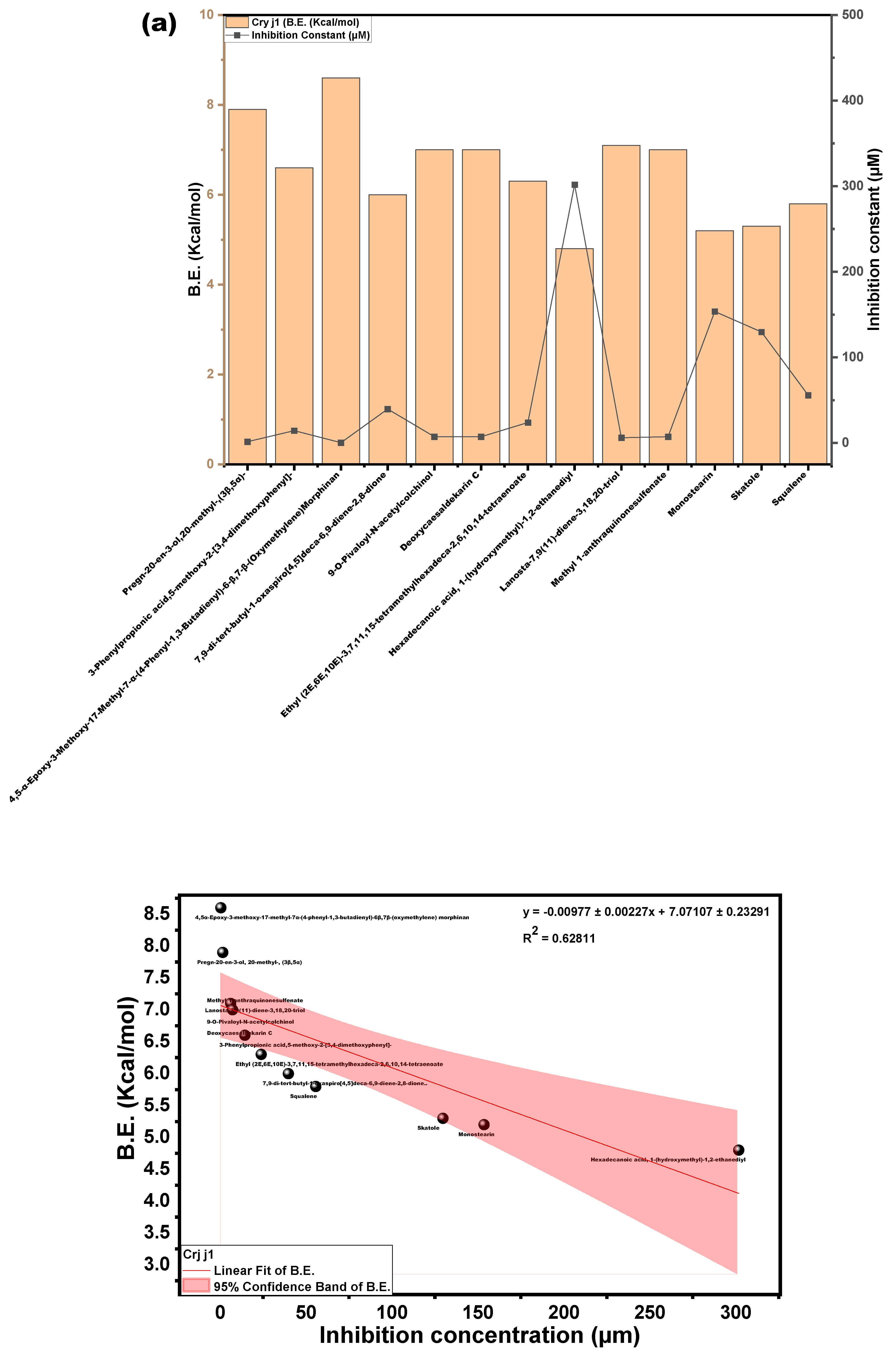
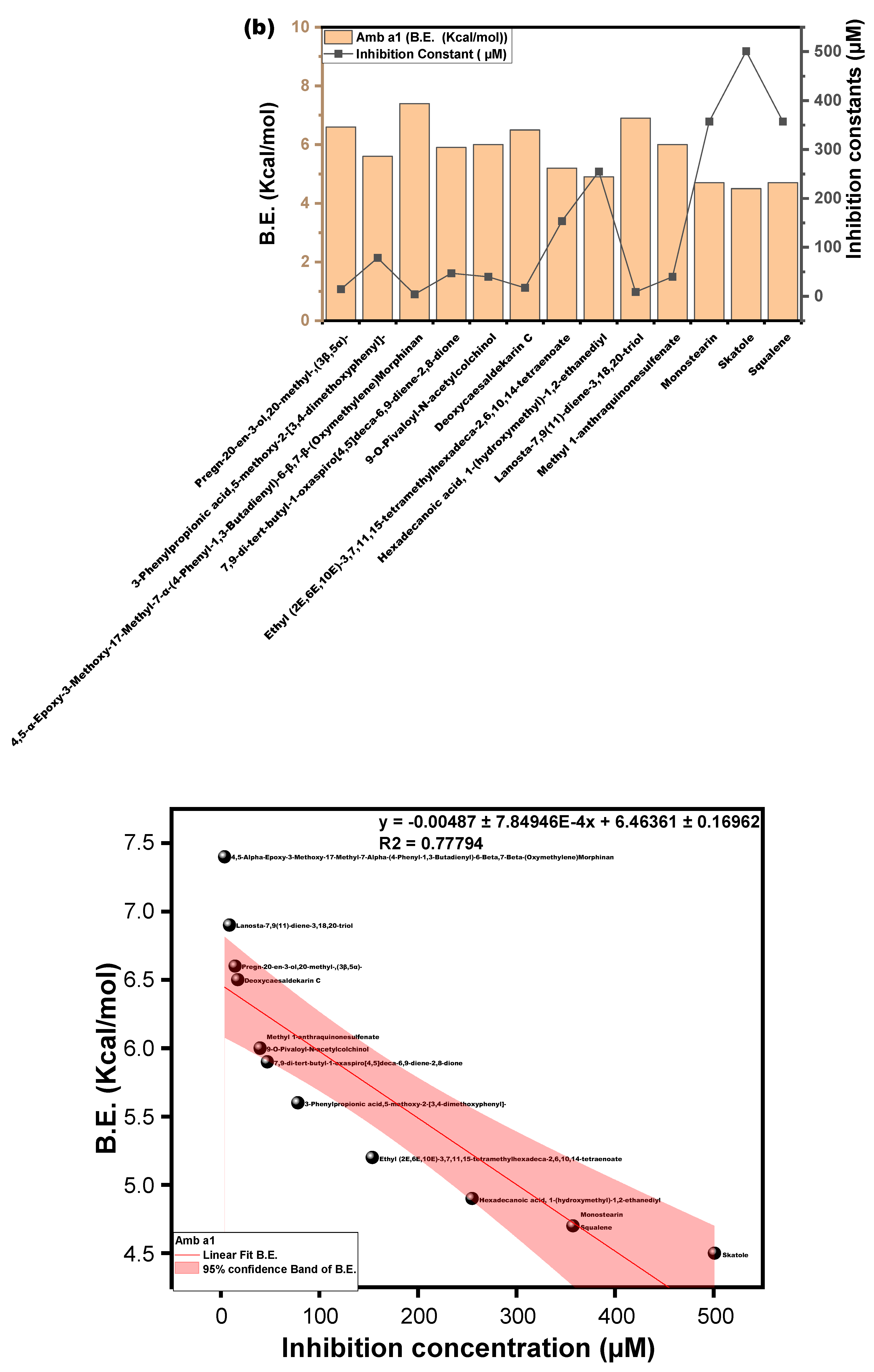
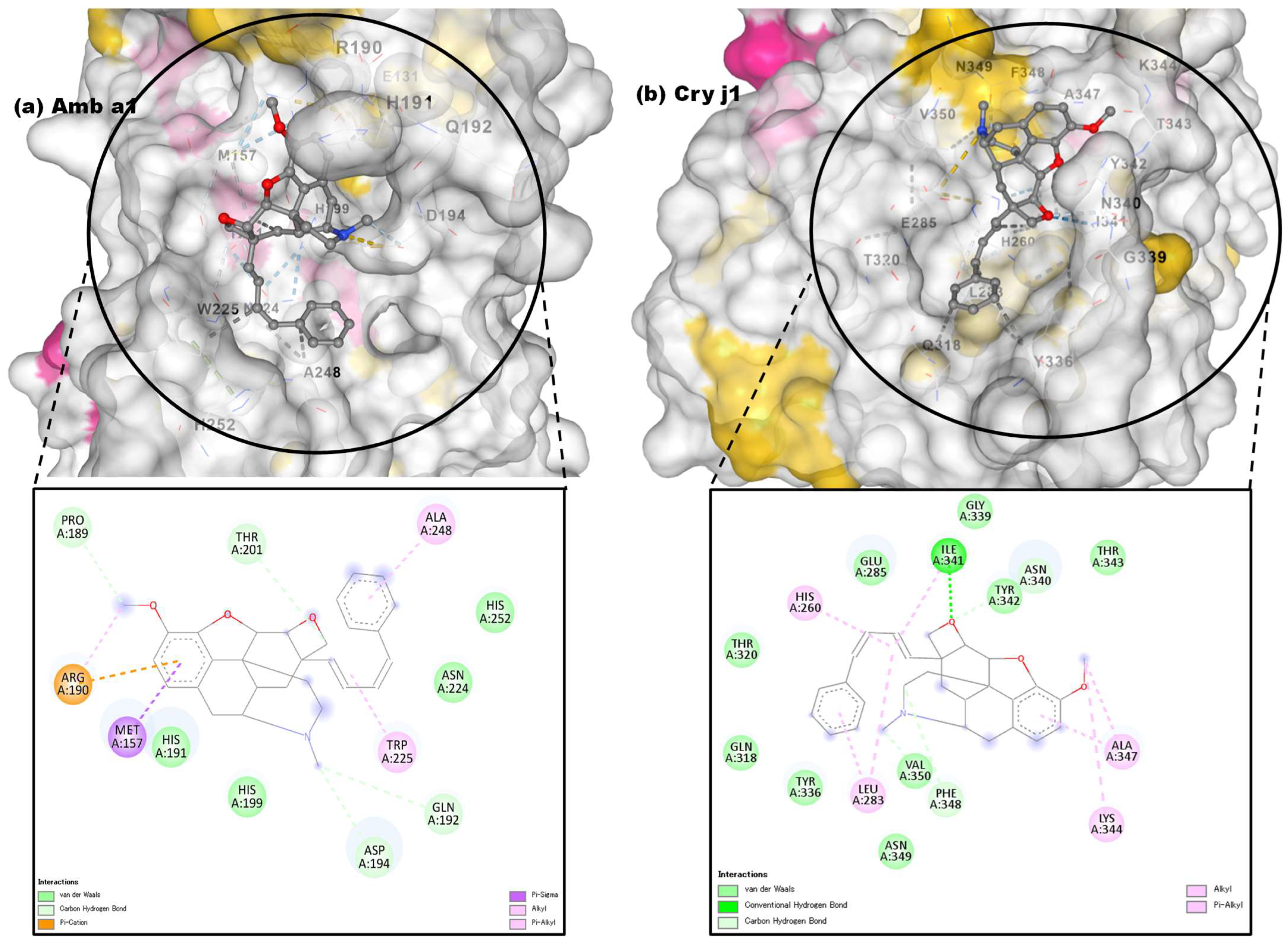
3.3.3. Molecular Docking Approach: Allergen Repressing Potential of the Bioactive Compounds
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Ikpa, C.C.B.; Maduka, T.O.D. Antimicrobial Properties of Methanol Extract of Dacryodes Edulis Seed and Determination of Phytochemical Composition Using FTIR and GCMS. Chem. Afr. 2020, 3, 927–935. [Google Scholar] [CrossRef]
- Huang, M.M.; Yiin, C.L.; Lock, S.S.M.; Chin, B.L.F.; Othman, I.; Chan, Y.H. Natural Deep Eutectic Solvents (NADES) for Sustainable Extraction of Bioactive Compounds from Medicinal Plants: Recent Advances, Challenges, and Future Directions. J. Mol. Liq. 2025, 425, 127202. [Google Scholar] [CrossRef]
- Ullah, A.; Munir, S.; Badshah, S.L.; Khan, N.; Ghani, L.; Poulson, B.G.; Emwas, A.-H.; Jaremko, M. Important Flavonoids and Their Role as a Therapeutic Agent. Molecules 2020, 25, 5243. [Google Scholar] [CrossRef]
- Parham, S.; Kharazi, A.Z.; Bakhsheshi-Rad, H.R.; Nur, H.; Ismail, A.F.; Sharif, S.; RamaKrishna, S.; Berto, F. Antioxidant, Antimicrobial and Antiviral Properties of Herbal Materials. Antioxidants 2020, 9, 1309. [Google Scholar] [CrossRef]
- Kalyniukova, A.; Holuša, J.; Musiolek, D.; Sedlakova-Kadukova, J.; Płotka-Wasylka, J.; Andruch, V. Application of Deep Eutectic Solvents for Separation and Determination of Bioactive Compounds in Medicinal Plants. Ind. Crop. Prod. 2021, 172, 114047. [Google Scholar] [CrossRef]
- Maduka, T.O.; Qingyue, W.; Enyoh, C.E.; Wang, W. Phytochemistry, Traditional Applications, and Pharmacology of Thujopsis Dolabrata Wood: A Comprehensive Review with Emphasis on Extraction Techniques. Ind. Crop. Prod. 2024, 217, 118822. [Google Scholar] [CrossRef]
- Lin, Y.; Xiao, K.; Wang, Q.; Lu, S.; Wang, W.; Seguchi, A. Research on Repressing Allergen Cry j 1 Released from Japanese Cedar Pollen Using Todomatsu Oil. Atmosphere 2023, 14, 991. [Google Scholar] [CrossRef]
- Zbîrcea, L.E.; Buzan, M.R.; Grijincu, M.; Babaev, E.; Stolz, F.; Valenta, R.; Păunescu, V.; Panaitescu, C.; Chen, K.W. Relationship between IgE Levels Specific for Ragweed Pollen Extract, Amb a 1 and Cross-Reactive Allergen Molecules. Int. J. Mol. Sci. 2023, 24, 4040. [Google Scholar] [CrossRef] [PubMed]
- Wopfner, N.; Jahn-Schmid, B.; Schmidt, G.; Christ, T.; Hubinger, G.; Briza, P.; Radauer, C.; Bohle, B.; Vogel, V.; Ebner, C. The Alpha and Beta Subchain of Amb a 1, the Major Ragweed-Pollen Allergen Divergent Reactivity at the IgE and T-Cell Level. Mol. Immunol. 2009, 46, 2090–2097. [Google Scholar] [CrossRef] [PubMed]
- Nortjie, E.; Basitere, M.; Moyo, D.; Nyamukamba, P. Extraction Methods, Quantitative and Qualitative Phytochemical Screening of Medicinal Plants for Antimicrobial Textiles: A Review. Plants 2022, 11, 2011. [Google Scholar] [CrossRef] [PubMed]
- Thakur, M.; Pathania, D. Environmental Fate of Organic Pollutants and Effect on Human Health. In Abatement of Environmental Pollutants; Elsevier: Amsterdam, The Netherlands, 2020; pp. 245–262. [Google Scholar]
- Srivastava, R.; Bhardwaj, N.; Jain, S.K.; Metya, A.K.; Parambil, J.V. Unveiling the Potential of Acetic Acid-Based Hydrophobic Natural Deep Eutectic Solvents for Phytochemical Extraction. J. Mol. Liq. 2024, 408, 125314. [Google Scholar] [CrossRef]
- Fourmentin, S.; Gomes, M.C.; Lichtfouse, E. (Eds.) Deep Eutectic Solvents for Medicine, Gas Solubilization and Extraction of Natural Substances; Springer Nature: Cham, Switzerland, 2020; Volume 56. [Google Scholar]
- Martins, M.A.R.; Pinho, S.P.; Coutinho, J.A.P. Insights into the Nature of Eutectic and Deep Eutectic Mixtures. J. Solut. Chem. 2018, 48, 962–982. [Google Scholar] [CrossRef]
- Smith, E.L.; Abbott, A.P.; Ryder, K.S. Deep Eutectic Solvents (DESs) and Their Applications. Chem. Rev. 2014, 114, 11060–11082. [Google Scholar] [CrossRef] [PubMed]
- Rafati, S.; Ebrahimi, N.; Sadeghi, R. New Family of Type V Natural Hydrophobic Deep Eutectic Solvents Based on Thymol-Acetamide/Acetanilide: Characteristics, Intermolecular Interactions and Applications in Liquid–Liquid Extraction. Sep. Purif. Technol. 2025, 359, 130583. [Google Scholar] [CrossRef]
- Omar, K.A.; Sadeghi, R. Database of Deep Eutectic Solvents and Their Physical Properties: A Review. J. Mol. Liq. 2023, 384, 121899. [Google Scholar] [CrossRef]
- Rodríguez-Llorente, D.; Cañada-Barcala, A.; Álvarez-Torrellas, S.; Águeda, V.I.; García, J.; Larriba, M. A Review of the Use of Eutectic Solvents, Terpenes and Terpenoids in Liquid–Liquid Extraction Processes. Processes 2020, 8, 1220. [Google Scholar] [CrossRef]
- Liu, Y.; Friesen, J.B.; McAlpine, J.B.; Lankin, D.C.; Chen, S.N.; Pauli, G.F. Natural Deep Eutectic Solvents: Properties, Applications, and Perspectives. J. Nat. Prod. 2018, 81, 679–690. [Google Scholar] [CrossRef]
- Paiva, A.; Craveiro, R.; Aroso, I.; Martins, M.; Reis, R.L.; Duarte, A.R.C. Natural Deep Eutectic Solvents–Solvents for the 21st Century. ACS Sustain. Chem. Eng. 2014, 2, 1063–1071. [Google Scholar] [CrossRef]
- Martins, M.A.; Silva, L.P.; Schaeffer, N.; Abranches, D.O.; Maximo, G.J.; Pinho, S.P.; Coutinho, J.A. Greener Terpene–Terpene Eutectic Mixtures as Hydrophobic Solvents. ACS Sustain. Chem. Eng. 2019, 7, 17414–17423. [Google Scholar] [CrossRef]
- Zielińska-Błajet, M.; Pietrusiak, P.; Feder-Kubis, J. Selected Monocyclic Monoterpenes and Their Derivatives as Effective Anticancer Therapeutic Agents. Int. J. Mol. Sci. 2021, 22, 4763. [Google Scholar] [CrossRef]
- Padilla, N.; Delso, I.; Bergua, F.; Lafuente, C.; Artal, M. Characterization of Camphor: Thymol or Dl-Menthol Eutectic Mixtures; Structure, Thermophysical Properties, and Lidocaine Solubility. J. Mol. Liq. 2024, 405, 125069. [Google Scholar] [CrossRef]
- Santos, C.D.; Cabot, J.C. Persistent Effects after Camphor Ingestion: A Case Report and Literature Review. J. Emerg. Med. 2015, 48, 298–304. [Google Scholar] [CrossRef]
- Escobar, A.; Perez, M.; Romanelli, G.; Blustein, G. Thymol Bioactivity: A Review Focusing on Practical Applications. Arab. J. Chem. 2020, 13, 9243–9269. [Google Scholar] [CrossRef]
- Kamatou, G.P.; Vermaak, I.; Viljoen, A.M.; Lawrence, B.M. Menthol: A Simple Monoterpene with Remarkable Biological Properties. Phytochemistry 2013, 96, 15–25. [Google Scholar] [CrossRef]
- Nejad, S.M.; Özgüneş, H.; Başaran, N. Pharmacological and Toxicological Properties of Eugenol. Turk. J. Pharm. Sci. 2017, 14, 201. [Google Scholar] [CrossRef] [PubMed]
- Nisar, M.F.; Khadim, M.; Rafiq, M.; Chen, J.; Yang, Y.; Wan, C.C. Pharmacological Properties and Health Benefits of Eugenol: A Comprehensive Review. Oxidative Med. Cell. Longev. 2021, 2021, 2497354. [Google Scholar] [CrossRef] [PubMed]
- Osch, D.J.; Dietz, C.H.; Spronsen, J.; Kroon, M.C.; Gallucci, F.; Sint Annaland, M.; Tuinier, R. A Search for Natural Hydrophobic Deep Eutectic Solvents Based on Natural Components. ACS Sustain. Chem. Eng. 2019, 7, 2933–2942. [Google Scholar] [CrossRef]
- Osiecka, D.; Vakh, C.; Makoś-Chełstowska, P.; Kubica, P. Plant-Based Meat Substitute Analysis Using Microextraction with Deep Eutectic Solvent Followed by LC-MS/MS to Determine Acrylamide, 5-Hydroxymethylfurfural and Furaneol. Anal. Bioanal. Chem. 2024, 416, 1117–1126. [Google Scholar] [CrossRef]
- Goswami, A.; Rahman, S.N.R.; Sree, A.; Shunmugaperumal, T. Solubility of Cinnarizine in Natural Deep Eutectic Solvent (Camphor+ Menthol) and Correlation with Different Solubility Models. Fluid Phase Equilibria 2024, 578, 114008. [Google Scholar] [CrossRef]
- Boudesocque-Delaye, L.; Ardeza, I.M.; Verger, A.; Grard, R.; Théry-Koné, I.; Perse, X.; Munnier, E. Natural Deep Eutectic Solvents as a Novel Bio-Based Matrix for Ready-to-Use Natural Antioxidants-Enriched Ingredients. Extr. Formul. Optim. Cosmet. 2024, 11, 17. [Google Scholar]
- Hanwell, M.D.; Curtis, D.E.; Lonie, D.C.; Vandermeersch, T.; Zurek, E.; Hutchison, G.R. Avogadro: An Advanced Semantic Chemical Editor, Visualization, and Analysis Platform. J. Cheminform. 2012, 4, 17. [Google Scholar] [CrossRef]
- Naseem, Z.; Shehzad, R.A.; Jabeen, S.; Tahir, S.; Mushtaq, F.; Zahid, M.; Iqbal, J. Quantum Chemical Investigation of Choline Chloride-Based Deep Eutectic Solvents. Chem. Phys. 2023, 571, 111936. [Google Scholar] [CrossRef]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A. Gaussian 09, Revision D.01; Gaussian Inc.: Wallingford, CT, USA, 2013. [Google Scholar]
- Bader, R.F.W. Atoms in molecules. Acc. Chem. Res. 1985, 18, 9–15. [Google Scholar] [CrossRef]
- Lu, T.; Chen, Q. Interaction Region Indicator: A Simple Real Space Function Clearly Revealing Both Chemical Bonds and Weak Interactions. Chem. Methods 2021, 1, 231–239. [Google Scholar] [CrossRef]
- Johnson, E.R.; Keinan, S.; Mori-Sánchez, P.; Contreras-García, J.; Cohen, A.J.; Yang, W. Revealing Noncovalent Interactions. J. Am. Chem. Soc. 2010, 132, 6498–6506. [Google Scholar] [CrossRef]
- Fuster, F.; Sevin, A.; Silvi, B. Topological Analysis of the Electron Localization Function (ELF) Applied to the Electrophilic Aromatic Substitution. J. Phys. Chem. A 2000, 104, 852–858. [Google Scholar] [CrossRef]
- Dávila, M.J.; Alcalde, R.; Atilhan, M.; Aparicio, S. PρT measurements and derived properties of liquid 1-alkanols. J. Chem. Thermodyn. 2012, 47, 241–259. [Google Scholar] [CrossRef]
- Rodríguez, Y.F.; Benito, C.; Aparicio, S.; Trenzado, J.L. Unveiling the High-Pressure Behavior of Thymol+ Carvone NADES: A Combined Experimental-Computational Approach. J. Supercrit. Fluids 2025, 215, 106408. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A Smooth Particle Mesh Ewald Method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Hoover, W.G. Canonical Dynamics: Equilibrium Phase-Space Distributions. Phys. Rev. A 1985, 31, 1695–1697. [Google Scholar] [CrossRef] [PubMed]
- Parrinello, M.; Rahman, A. Polymorphic Transitions in Single Crystals: A New Molecular Dynamics Method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Liang, X.; Zhang, J.; Huang, Z.; Guo, Y. Sustainable Recovery and Recycling of Natural Deep Eutectic Solvent for Biomass Fractionation via Industrial Membrane-Based Technique. Ind. Crop. Prod. 2023, 194, 116351. [Google Scholar] [CrossRef]
- Harifi-Mood, A.R.; Sarafrazi, M. CO2 Solubility in Terpenoid-Based Hydrophobic Deep Eutectic Solvents: An Experimental and Theoretical Study. J. Environ. Chem. Eng. 2023, 11, 109177. [Google Scholar] [CrossRef]
- Phaechamud, T.; Tuntarawongsa, S.; Charoensuksai, P. Evaporation Behavior and Characterization of Eutectic Solvent and Ibuprofen Eutectic Solution. AAPS PharmSciTech 2016, 17, 1213–1220. [Google Scholar] [CrossRef]
- Fattahi, N.; Shamsipur, M.; Nematifar, Z.; Babajani, N.; Moradi, M.; Soltani, S.; Akbari, S. Novel Deep Eutectic Solvent-Based Liquid Phase Microextraction for the Extraction of Estrogenic Compounds from Environmental Samples. RSC Adv. 2022, 12, 14467–14476. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Zhang, H.; Lou, X.; Wu, X.; Wang, Y.; Zhao, K.; Du, X.; Xia, X. Analysis of NADES and Its Water Tailoring Effects Constructed from Inulin and L-Proline Based on Structure, Physicochemical and Antifreeze Properties. Int. J. Biol. Macromol. 2024, 277, 134049. [Google Scholar] [CrossRef]
- Kianpoor, A.; Sadeghi, R. Novel Hydrophobic Acetanilide-Based Deep Eutectic Solvents. Preparation, Properties, Their Applications in Liquid–Liquid Extraction. Chem. Eng. Sci. 2024, 288, 119866. [Google Scholar] [CrossRef]
- Rozas, S.; Zamora, L.; Benito, C.; Atilhan, M.; Aparicio, S. A Study on Monoterpenoid-Based Natural Deep Eutectic Solvents. Green Chem. Eng. 2023, 4, 99–114. [Google Scholar] [CrossRef]
- Zahid, S.; Rasool, A.; Ans, M.; Yaseen, M.; Iqbal, J. Quantum Chemical Approach of Donor− π–Acceptor Based Arylborane–Arylamine Macrocycles with Outstanding Photovoltaic Properties toward High-Performance Organic Solar Cells. Energy Fuels 2021, 35, 15018–15032. [Google Scholar] [CrossRef]
- Ishaq, M.; Gilani, M.A.; Ahmad, F.; Afzal, Z.M.; Arshad, I.; Bilad, M.R.; Ayub, K.; Khan, A.L. Theoretical and Experimental Investigation of CO2 Capture through Choline Chloride Based Supported Deep Eutectic Liquid Membranes. J. Mol. Liq. 2021, 335, 116234. [Google Scholar] [CrossRef]
- Parsaee, Z.; Mohammadi, K.; Ghahramaninezhad, M.; Hosseinzadeh, B. A Novel Nano-Sized Binuclear Nickel (II) Schiff Base Complex as a Precursor for NiO Nanoparticles: Synthesis, Characterization, DFT Study and Antibacterial Activity. New J. Chem. 2016, 40, 10569–10583. [Google Scholar] [CrossRef]
- Lashgari, A.; Ghamami, S.; Govindarajan, M.; Salgado-Morán, G.; Montes Romero, P.; Gerli Candia, L. A Theoretical Quantum Study of the Electronic Properties of Mentoxy Dichloro Phosphorous (C10H19OPCl2). J. Chil. Chem. Soc. 2018, 63, 3887–3897. [Google Scholar] [CrossRef]
- Khan, P.; Jamshaid, M.; Tabassum, S.; Perveen, S.; Mahmood, T.; Ayub, K.; Yang, J.; Gilani, M.A. Exploring the Interaction of Ionic Liquids with Al12N12 and Al12P12 Nanocages for Better Electrode-Electrolyte Materials in Super Capacitors. J. Mol. Liq. 2021, 344, 117828. [Google Scholar] [CrossRef]
- Parr, R.G.; Szentpály, L.V.; Liu, S. Electrophilicity index. J. Am. Chem. Soc. 1999, 121, 1922–1924. [Google Scholar] [CrossRef]
- Popelier, P.L.A.; Aicken, F.M.; O’Brien, S.E. Atoms in Molecules; Prentice Hall: Hoboken, NJ, USA, 2000; Volume 188. [Google Scholar]
- Kumar, P.S.V.; Raghavendra, V.; Subramanian, V. Bader’s Theory of Atoms in Molecules (AIM) and Its Applications to Chemical Bonding. J. Chem. Sci. 2016, 128, 1527–1536. [Google Scholar] [CrossRef]
- Bader, R.F.W.; Johnson, S.; Tang, H.T.; Popelier, P.L.A. The Electron Pair. J. Phys. Chem. 1996, 100, 15398. [Google Scholar] [CrossRef]
- Emamian, S.; Lu, T.; Kruse, H.; Emamian, H. Exploring Nature and Predicting Strength of Hydrogen Bonds: A Correlation Analysis between Atoms-in-Molecules Descriptors, Binding Energies, and Energy Components of Symmetry-Adapted Perturbation Theory. J. Comput. Chem. 2019, 40, 2868–2881. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.; Liu, Y.; Sebbah, T.; Cao, X. A Theoretical Study on Terpene-Based Natural Deep Eutectic Solvent: Relationship between Viscosity and Hydrogen-Bonding Interactions. Glob. Chall. 2021, 5, 2000103. [Google Scholar] [CrossRef]
- Yao, J.; Xiao, L.; Li, C.; Wang, B.; Chen, Y.; Yan, X.; Cui, Z. Exploration of the Multiscale Interaction Mechanism between Natural Deep Eutectic Solvents and Silybin by QC Calculation and MD Simulation. J. Mol. Liq. 2022, 363, 119768. [Google Scholar] [CrossRef]
- Klix, C.L.; Maret, G.; Keim, P. Discontinuous Shear Modulus Determines the Glass Transition Temperature. Phys. Rev. X 2015, 5, 041033. [Google Scholar] [CrossRef][Green Version]
- Enyoh, C.E.; Wang, Q.; Wang, W.; Chowdhury, T.; Rabin, M.H.; Islam, R.; Lin, Y.; Xiao, K. Sorption of Per-and Polyfluoroalkyl Substances (PFAS) Using Polyethylene (PE) Microplastics as Adsorbent: Grand Canonical Monte Carlo and Molecular Dynamics (GCMC-MD) Studies. Int. J. Environ. Anal. Chem. 2024, 104, 2719–2735. [Google Scholar] [CrossRef]
- Farooq, M.Q.; Odugbesi, G.A.; Abbasi, N.M.; Anderson, J.L. Elucidating the Role of Hydrogen Bond Donor and Acceptor on Solvation in Deep Eutectic Solvents Formed by Ammonium/Phosphonium Salts and Carboxylic Acids. ACS Sustain. Chem. Eng. 2020, 8, 18286–18296. [Google Scholar] [CrossRef]
- Pádua, A.A.; Costa Gomes, M.F.; Canongia Lopes, J.N. Molecular Solutes in Ionic Liquids: A Structural Perspective. Acc. Chem. Res. 2007, 40, 1087–1096. [Google Scholar] [CrossRef]
- Quesada-Moreno, M.M.; Fatima, M.; Medel, R.; Pérez, C.; Schnell, M. Sniffing out Camphor: The Fine Balance between Hydrogen Bonding and London Dispersion in the Chirality Recognition with α-Fenchol. Phys. Chem. Chem. Phys. 2022, 24, 12849–12859. [Google Scholar] [CrossRef] [PubMed]
- Perkins, S.L.; Painter, P.; Colina, C.M. Experimental and Computational Studies of Choline Chloride-Based Deep Eutectic Solvents. J. Chem. Eng. Data 2014, 59, 3652–3662. [Google Scholar] [CrossRef]
- Sun, X.; Luo, H.; Dai, S. Ionic Liquids-Based Extraction: A Promising Strategy for the Advanced Nuclear Fuel Cycle. Chem. Rev. 2012, 112, 2100–2128. [Google Scholar] [CrossRef]
- Mahdy, A.H.S.; Zayed, S.E.; Abo-Bakr, A.M.; Hassan, E.A. Camphor: Synthesis, Reactions and Uses as a Potential Moiety in the Development of Complexes and Organocatalysts. Tetrahedron 2022, 121, 132913. [Google Scholar] [CrossRef]
- Kusano, S.; Kukimoto-Niino, M.; Satta, Y.; Ohsawa, N.; Uchikubo-Kamo, T.; Wakiyama, M.; Ikeda, M.; Terada, T.; Yamamoto, K.; Nishimura, Y.; et al. Structural Basis for the Specific Recognition of the Major Antigenic Peptide from the Japanese Cedar Pollen Allergen Cry j 1 by HLA-DP5. J. Mol. Biol. 2014, 426, 3016–3027. [Google Scholar] [CrossRef] [PubMed]
- Fujimura, T.; Fujinami, K.; Ishikawa, R.; Tateno, M.; Tahara, Y.; Okumura, Y.; Ohta, H.; Miyazaki, H.; Taniguchi, M.R.F.A. Cry j 1 and Cry j 2 from Japanese Cedar Pollen, Conjugated with Polyethylene Glycol Potentiate the Attenuation of Cry j 1-Specific IgE Production in Cry j 1-Sensitized Mice and Japanese Cedar Pollen Allergen-Sensitized Monkeys. Int. Arch. Allergy Immunol. 2015, 168, 32–43. [Google Scholar] [CrossRef] [PubMed]
- Bordas-Le Floch, V.; Groeme, R.; Chabre, H.; Baron-Bodo, V.; Nony, E.; Mascarell, L.; Moingeon, P. New Insights into Ragweed Pollen Allergens. Curr. Allergy Asthma Rep. 2015, 15, 63. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, A.R.; Scheurer, S.; Vieths, S. Food Allergens: Molecular and Immunological Aspects, Allergen Databases and Cross-Reactivity. Chem. Immunol. Allergy 2015, 101, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.H.; Lee, T.; Oh, I.; Nam, K.W.; Kim, K.H.; Oh, K.B.; Shin, J.; Mar, W. Mast Cell Stabilizing Effect of (−)-Elema-1, 3, 11 (13)-Trien-12-Ol and Thujopsene from Thujopsis Dolabrata Is Mediated by Down-Regulation of Interleukin-4 Secretion in Antigen-Induced RBL-2H3 Cells. Biol. Pharm. Bull. 2013, 36, 339–345. [Google Scholar] [CrossRef] [PubMed]
- Javeed, A.; Sarfraz, M.; Bhutta, N.K.; Han, B. Alkaloids as Natural Anti-Allergy Agents: A Mini Review. Allergy Med. 2024, 2, 100014. [Google Scholar] [CrossRef]
- Oppong, E.; Flink, N.; Cato, A.C. Molecular Mechanisms of Glucocorticoid Action in Mast Cells. Mol. Cell. Endocrinol. 2013, 380, 119–126. [Google Scholar] [CrossRef]
- Kusano, S.; Ueda, S.; Oryoji, D.; Toyoumi, A.; Hashimoto-Tane, A.; Kishi, H.; Hamana, H.; Muraguchi, A.; Jin, H.; Arase, H. Contributions of the N-Terminal Flanking Residues of an Antigenic Peptide from the Japanese Cedar Pollen Allergen Cry j 1 to the T-Cell Activation by HLA-DP5. Int. Immunol. 2023, 35, 447–458. [Google Scholar] [CrossRef]
- Su, H.G.; Peng, X.R.; Shi, Q.Q.; Huang, Y.J.; Zhou, L.; Qiu, M.H. Lanostane triterpenoids with anti-inflammatory activities from Ganoderma lucidum. Phytochemistry 2020, 173, 112256. [Google Scholar] [CrossRef] [PubMed]


| s/n | Chemical | MF | M/gmol−1 | Purity | Phase at RT | Structure |
|---|---|---|---|---|---|---|
| 1 | Camphor | C10H16O | 152.23 | >0.98 | Solid | 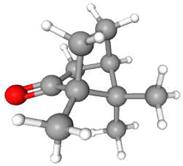 |
| 2 | Thymol | C10H14O | 150.22 | >0.98 | Solid | 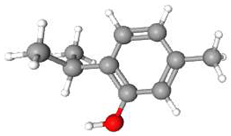 |
| 3 | Eugenol | C10H12O2 | 164.20 | >0.98 | Liquid | 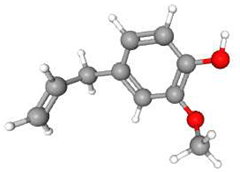 |
| 4 | DL-Menthol | C10H20O | 156.27 | >0.98 | Solid |  |
| (a) | ||
| Compound/NADES | Onset Temperature (°C) | Peak Decomposition Temperature (°C) |
| Thymol–Camphor | 125.2 | 174.3 |
| Thymol–Eugenol | 123.4 | 185.1 |
| Menthol–Eugenol | 110.2 | 155.4 |
| Camphor–Eugenol | 114 | 178 |
| Thymol–Menthol | 125.5 | 176.9 |
| Thymol (pure) | 132.5 | 181.8 |
| Camphor (pure) | 102 | 165.4 |
| Eugenol (pure) | 145 | 206.8 |
| Menthol (pure) | 119.3 | 167 |
| (b) | ||
| Mole Fraction of Menthol | 1st Peak Temp (°C) | DTA Min (µV) |
| 0.1 | 28.36 | –1.28 |
| 0.3 | 28.2 | –1.38 |
| 0.5 | 26.36 | –1.19 |
| 0.7 | 29.17 | –2.07 |
| 0.9 | 34.64 | –14.48 |
| Phytochemical Family | Compounds |
|---|---|
| Alkane-related compounds | (I) 1-Cyano-1,1-dideuterio hexadecane; (II) 2-Methylhexadecane; (III) 2-Methylpentadecane; (IV) 3,8-Dimethyldecane; (V) 3-Methylheneicosane; (VI) 3-Methyloctadecane; (VII) 5-Methylundecane; (VIII) Cetane; (IX) Decane, 3,8-dimethyl-; (X) Docosane; (XI) Eicosane; (XII) Heneicosane; (XIII) Heptadecane; (XIV) Heptadecane, 8-methyl-; (XV) 8-methyl-Heptacosane; (XVI) Hentriacontane; (XVII) Hexadecane; (XVIII) Hexadecane, 7,9-dimethyl-; (XIX) Hexatriacontane; (XX) Nonadecane; (XXI) Nonacosane; (XXII) Norpristane; (XXIII) Octadecane; (XXIV) Octadecane, 2-methyl-; (XXV) Pentacosane; (XXVI) Pentadecane; (XXVII) Phytane; (XXVIII) Tetracosane; (XXIX) Tetradecane; (XXX) Tricosane; (XXXI) Triacontane |
| Polycyclic aromatic compound derivatives | 4-(Biphenyl-2′-yl)-7-chloro-1,2-dihydronaphthalene |
| Alkaloids | (I) 4,5α-Epoxy-3-methoxy-17-methyl-7α-(4-phenyl-1,3-butadienyl)-6β,7β-(oxymethylene) morphinan; (II) Skatole |
| Terpenes and terpenoids | (I) Ethyl (2E,6E,10E)-3,7,11,15-tetramethylhexadeca-2,6,10,14-tetraenoate; (II) Deoxycaesaldekarin C; (III) Lanosta-7,9(11)-diene-3,18,20-triol; (IV) Squalene |
| Fatty acids and esters | (I) Hexadecanoic acid, 1-(hydroxymethyl)-1,2-ethanediyl ester; (II) Monostearin; (III) 7,9-di-tert-butyl-1-oxaspiro[4,5]deca-6,9-diene-2,8-dione |
| Phenolic compounds | (I) Methyl 1-anthraquinonesulfenate; (II) 3-Phenylpropionic acid, 5-methoxy-2-[3,4-dimethoxyphenyl]- |
| Steroids | (I) Pregn-20-en-3-ol, 20-methyl-, (3β,5α)- |
| Components | E Gap | I (eV) | A (eV) | µ (eV) | η (eV) | S (eV) | Χ (eV) | ω (eV) |
|---|---|---|---|---|---|---|---|---|
| Cam-Eug | 3.83 | 8.64 | 4.81 | −6.72 | 1.92 | 0.96 | 6.72 | 11.80 |
| Cam-Thy | 4.26 | 9.20 | 4.94 | −7.07 | 2.13 | 1.07 | 7.07 | 11.72 |
| Men-Eug | 4.14 | 8.99 | 4.84 | −6.91 | 2.07 | 1.04 | 6.91 | 11.53 |
| Thy-Eug | 3.98 | 9.00 | 5.02 | −7.01 | 1.99 | 0.99 | 7.01 | 12.35 |
| Men-Thy | 4.29 | 9.24 | 4.95 | −7.10 | 2.14 | 1.07 | 7.10 | 11.74 |
| Camphor | 5.02 | 9.76 | 4.74 | −7.25 | 2.51 | 1.26 | 7.25 | 10.46 |
| eugenol | 4.13 | 8.93 | 1.68 | −5.30 | 3.62 | 1.81 | 5.30 | 3.89 |
| menthol | 8.85 | 10.54 | 4.80 | −7.67 | 2.87 | 1.43 | 7.67 | 10.25 |
| thymol | 4.27 | 9.21 | 4.94 | −7.07 | 2.13 | 1.07 | 7.07 | 11.72 |
| NADES | (ρe) (a.u.) | (∇2ρe) (a.u.) | CVB |
|---|---|---|---|
| Camphor_Thymol | 0.023 | 0.078 | 0.031 |
| Eugenol_Camphor | 0.027 | 0.086 | 0.009 |
| Eugenol_Thymol | 0.026 | 0.082 | 0.015 |
| Thymol_Menthol | 0.023 | 0.074 | 0.010 |
| Menthol_Eugenol | 0.039 | 0.117 | −0.051 |
| s/n | Compound | Structure | Pub Chem ID | Amb a 1 (B.E) kcal/mol | Cry j 1 (B.E) kcal/mol | Amb a 1 IC (µm) | Cry j 1 (I.C) µm | Cry j 1 Bonding: Interaction Site | Amb a 1 Bonding: Interaction Site |
|---|---|---|---|---|---|---|---|---|---|
| 1 | Pregn-20-en-3-ol,20-methyl-,(3β,5α)- | 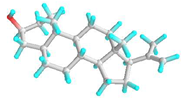 | 22296144 | −6.6 | −7.9 | 14.44 | 1.61 | ALK: LEU 283 VDW: GLN 318, TYR 336, ILE 341, HIS 260, ASN 340, TYR 342, ALA 347, PHE 348, GLU 285, THR 320 | CHB: ALA 368, LYS 77, ALA 78 VDW: GLY 74, GLY 369, PHE 75, GLY 229, THR 228, SER 204, ASP 205 |
| 2 | 3-Phenylpropionic acid,5-methoxy-2-[3,4-dimethoxyphenyl]- | 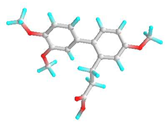 | 628993 | −5.6 | −6.6 | 78.18 | 14.44 | CVB: THR 320, SER 284, TYR 336 CHB: GLU 285 π-T: TYR 336 ALK/π-ALKLYS 335, LEU 283, ILE 341, TYR 342 VDW: ASN 340, GLN 318, GLU 285, SER 319 | CHB: ASP 194, HIS 191, GLU 131 π-S: TRP 225, MET 157 π-π-T: HIS 199 ALK/π-ALK: ALA 248, ARG 190 VDW: ASN 224, THR 201, ASP 196, GLY 195, THR 155, SER 193 |
| 3 | 4,5-Alpha-Epoxy-3-Methoxy-17-Methyl-7-Alpha-(4-Phenyl-1,3-Butadienyl)-6-Beta,7-Beta-(Oxymethylene)Morphinan | 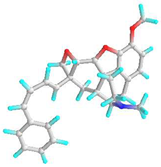 | - | −7.4 | −8.6 | 3.74 | 0.49 | CHB: GLY 339, GLY 338 CVB: THR 320 π-CAT: LYS 335 ALK/π-ALK: ILE 341, VAL 350, HIS 260 VDW: THR 342, PHE 348, GLY 285, LEU 283, GLN 318, TYR 336, ASN 340 | CHB: ASP 194, GLN 192, THR 201 π-ALK: TRP 225, ALA 248 π-CAT: MET 157 VDW: HIS 191, HIS 199, ASN 224, HIS 253 |
| 4 | 7,9-di-tert-butyl-1-oxaspiro[4,5]deca-6,9-diene-2,8-dione |  | 545303 | −5.9 | −6.0 | 47.10 | 39.79 | CVB: ILE 341, TYR 342 ALK: LEU 283 VDW: HIS 260, GLU 285, THR 320, ASN 340, GLY 339, GLY 338, TYR 336 | CVB: THR 201 π-ALK: ALA 248, TRP 225 VDW: ASN 158, MET 157, HIS 199, ASN 224, ASP 218, ASP 194, ASP 196, LEU 245 |
| 5 | 9-O-Pivaloyl-N-acetylcolchinol | 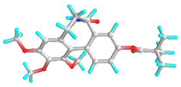 | 634928 | −6.0 | −7.0 | 39.79 | 7.35 | CVB: GLU 285 CHB: PHE 348 π-π-T: HIS 260, VAL 350, ILE 341, LEU 283 VDW: TYR 342, ASN 340, TYR 336, GLN 318, LYS 335, GLN 321, THR 320 | CVB: THR 201 π-π: TRP 225 π-ALK: ALA 248 |
| 6 | Deoxycaesaldekarin C | 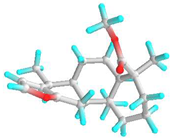 | 572829 | −6.5 | −7.0 | 17.10 | 7.35 | CVB: THR 320, TYR 342 ALK/π-ALK: ALA 347, LEU 283, ILE 341 VDW: GLU 285, GLN 318, TYR 336, ASN 340 | ALK: ALA 78, LYS 77, ALA 368 π-AN: ASP 205 VDW: GLY 74, GLY 369, THR 228, HIS 259, GLY 229, PHE 75, GLU 365 |
| 7 | Ethyl (2E,6E,10E)-3,7,11,15-tetramethylhexadeca-2,6,10,14-tetraenoate | 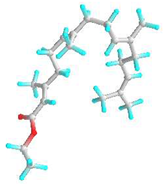 | 5366011 | −5.2 | −6.3 | 153.61 | 23.97 | CVB: LYS 132, GLN 169 ALK/π-ALK: HIS 167, PHE 130, VAL 134, PHE 161, ALA 108, ILE 157, ARG 133 VDW: ASN 158, GLU 164, PRO 165, TYR 106, VAL 166, ARG 177 | CHB: LYS 77, GLN 309 π-ALK: ALA 78, PHE 75, LEU 362, ARG 341, PRO 360 VDW: GLY 74, ALA 368, GLU 365, GLY 228, ASP 204, HIS 80 |
| 8 | Hexadecanoic acid, 1-(hydroxymethyl)-1,2-ethanediyl |  | 99931 | −4.9 | −4.8 | 254.94 | 301.84 | CVB: ILE 341, TYR 342 ALK/π-ALK: LEU 283, TYR 336 VDW: LEU 283, TYR 336 GLN 318, ALA 347, VAL 350, THR 320, GLY 339, ASN 340, GLN 285, PHE 348 | CVB: GLN 309 (ester and OH) ALK: LEU 362, LEU 307, PRO 360, ARG 341 VDW: GLN 284, ALA 373, MET 371, GLN 372, MET 370, ASP 359 |
| 9 | Lanosta-7,9(11)-diene-3,18,20-triol |  | 91695604 | −6.9 | −7.1 | 8.70 | 6.21 | CVB: PHE 348 UDD: PHE 348 ALK/π-ALK: LEU 283, TYR 336 VDW: GLN 318, SER 284, GLU 285, THR 320, GLY 338, ILE 341, GLY 339, ASN 340, TYR 342, ASN 349, ALA 347 | CHB: SER 204 ALK: LYS 257 AC-AC: ASP 205 VDW: HIS 259, GLU 365, THR 228, GLY 229, PHE 75, ALA 78, GLY 369, ALA 368 |
| 10 | Methyl 1-anthraquinonesulfenate |  | 349031192 | −6.0 | −7.0 | 39.79 | 7.35 | CVB: ILE 341 ALK/π-ALK: LEU 283, ILE 341 π-HB: THR 320 VDW: GLN 318, SER 284, HIS 260, GLU 285, TYR 342, ASN 340, GLY 339, TYR 336 | CHB: GLN 309, LEU 362 π-S: LEU 307 π-ALK: ARG 341, PRO 360 VDW: ASP 359, GLN 343, MET 370, ASN 367, VAL 361 |
| 11 | Monostearin |  | 24699 | −4.7 | −5.2 | 357.37 | 153.61 | CVB: GLU 285, VAL 350 CHB: HIS 260 VDW: ASN 340, GLY 339, TYR 342, GLN 321, GLN 318, THR 320, ASN 349 ALK/π-ALK: ILE 341, LEU 283, TYR 336 | CVB: THR 155, ASP 194, HIS 199 ALK/π-ALK: MET 157, TRP 225 VDW: GLY 195, GLY 369, GLU 365, THR 228 |
| 12 | Skatole |  | 6736 | −4.5 | −5.3 | 500.95 | 129.74 | π-CAT: LYS 132 VDW: ARG 133, GLU 164 π-S: ILE 157 π-π-T: PHE 161 ALK/π-ALK: VAL 166, TYR 106 | CHB: ASP 205 π-ALK: ALA 78, ALA 368, PHE 75, HIS 259 VDW: GLY 369, GLY 229, THR 228, GLU 365 |
| 13 | Squalene |  | 638072 | −4.7 | −5.8 | 357.37 | 55.77 | ALK/π-ALK: TYR 336, LEU 283, ILE 341, TYR 342, ALA 347 VDW: GLN 318, GLY 339, THR 320, PHE 348, VAL 350, ASN 340, GLU 285, HIS 260 | ALK: LEU 307, PRO 360, MET 370, LEU 362, ARG 341, ALA 373 VDW: MET 371, GLN 372, GLN 309, ASP 359, GLN 343 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maduka, T.O.; Wang, Q.; Suzuki, M.; Enyoh, C.E.; Wang, W.; Rana, M.S. Hydrophobic Natural Deep Eutectic Solvents for Extraction of Bioactive Compounds: Multiscale Characterization, Quantum Simulations, and Molecular Interaction Studies with Cry j 1 and Amb a 1 Allergens. Separations 2025, 12, 214. https://doi.org/10.3390/separations12080214
Maduka TO, Wang Q, Suzuki M, Enyoh CE, Wang W, Rana MS. Hydrophobic Natural Deep Eutectic Solvents for Extraction of Bioactive Compounds: Multiscale Characterization, Quantum Simulations, and Molecular Interaction Studies with Cry j 1 and Amb a 1 Allergens. Separations. 2025; 12(8):214. https://doi.org/10.3390/separations12080214
Chicago/Turabian StyleMaduka, Tochukwu Oluwatosin, Qingyue Wang, Miho Suzuki, Christian Ebere Enyoh, Weiqian Wang, and Md. Sohel Rana. 2025. "Hydrophobic Natural Deep Eutectic Solvents for Extraction of Bioactive Compounds: Multiscale Characterization, Quantum Simulations, and Molecular Interaction Studies with Cry j 1 and Amb a 1 Allergens" Separations 12, no. 8: 214. https://doi.org/10.3390/separations12080214
APA StyleMaduka, T. O., Wang, Q., Suzuki, M., Enyoh, C. E., Wang, W., & Rana, M. S. (2025). Hydrophobic Natural Deep Eutectic Solvents for Extraction of Bioactive Compounds: Multiscale Characterization, Quantum Simulations, and Molecular Interaction Studies with Cry j 1 and Amb a 1 Allergens. Separations, 12(8), 214. https://doi.org/10.3390/separations12080214










