Genome-Wide Identification and Analysis of SRO Gene Family in Chinese Cabbage (Brassica rapa L.)
Abstract
1. Introduction
2. Result
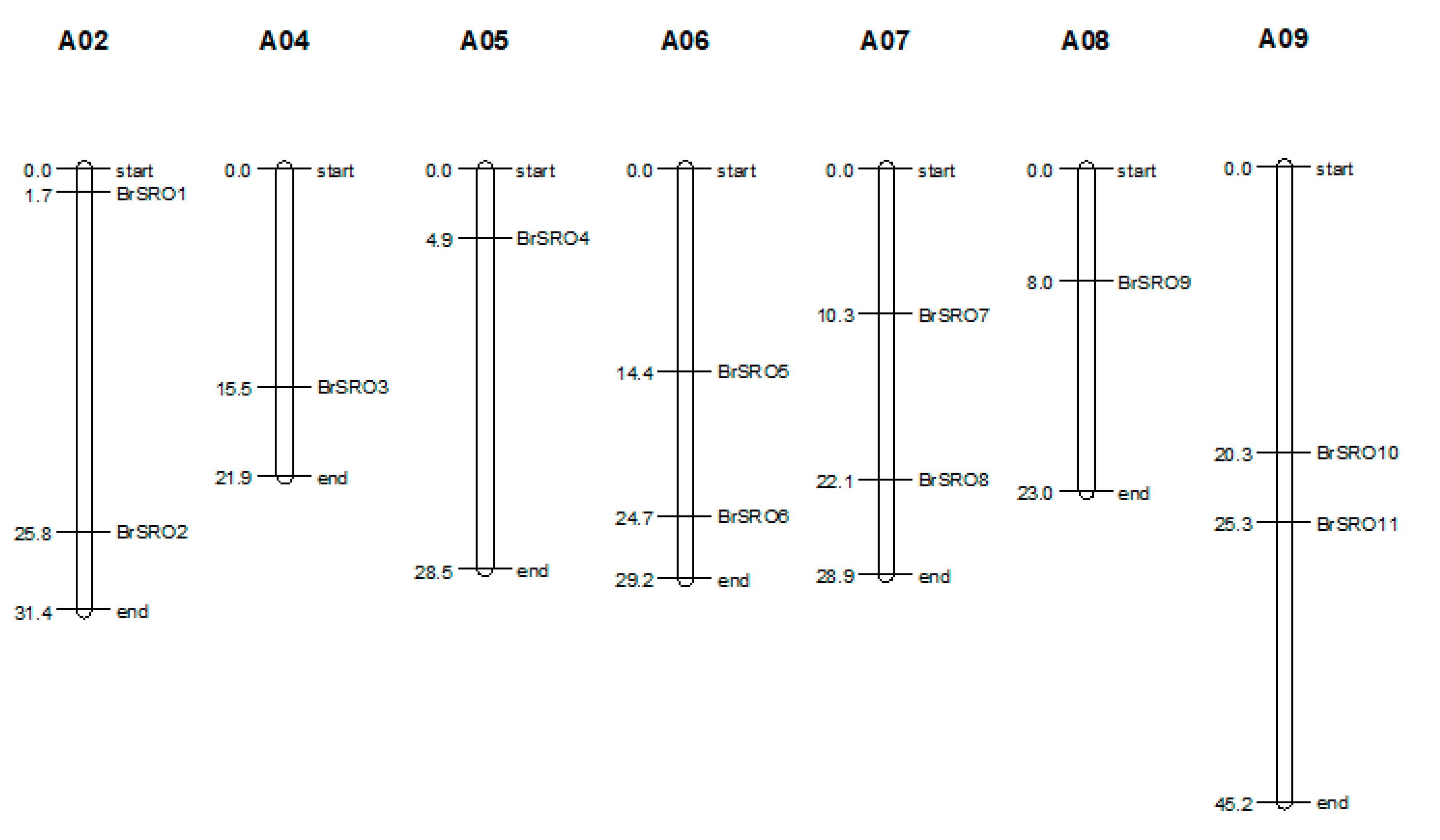
2.1. Identification and Chromosomal Location of the SRO Family Genes in Chinese Cabbage
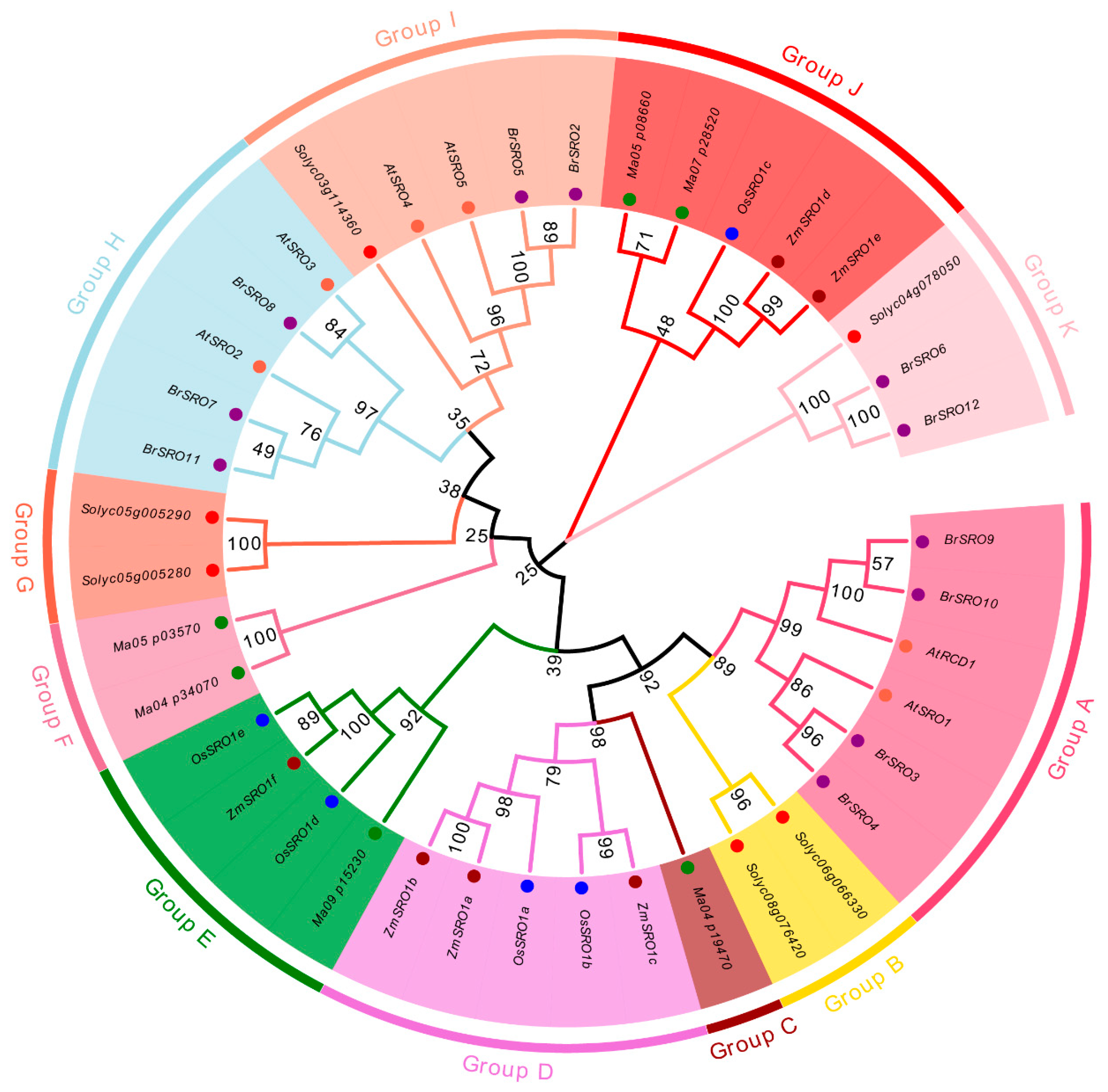
2.2. Phylogenetic Analysis of the SRO Family Genes in Chinese Cabbage
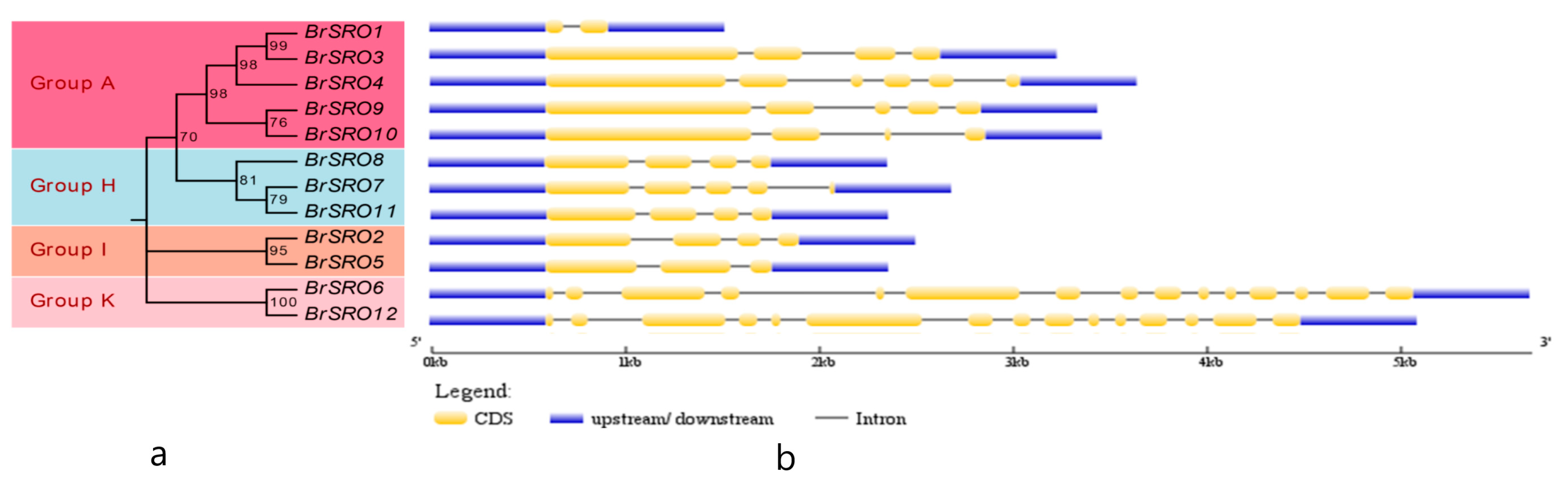
2.3. Gene Structure of the BrSRO Genes
2.4. Conserved Motifs Analysis of BrSRO Proteins
2.5. Cis-Elements in the Promoters of BrSRO Genes
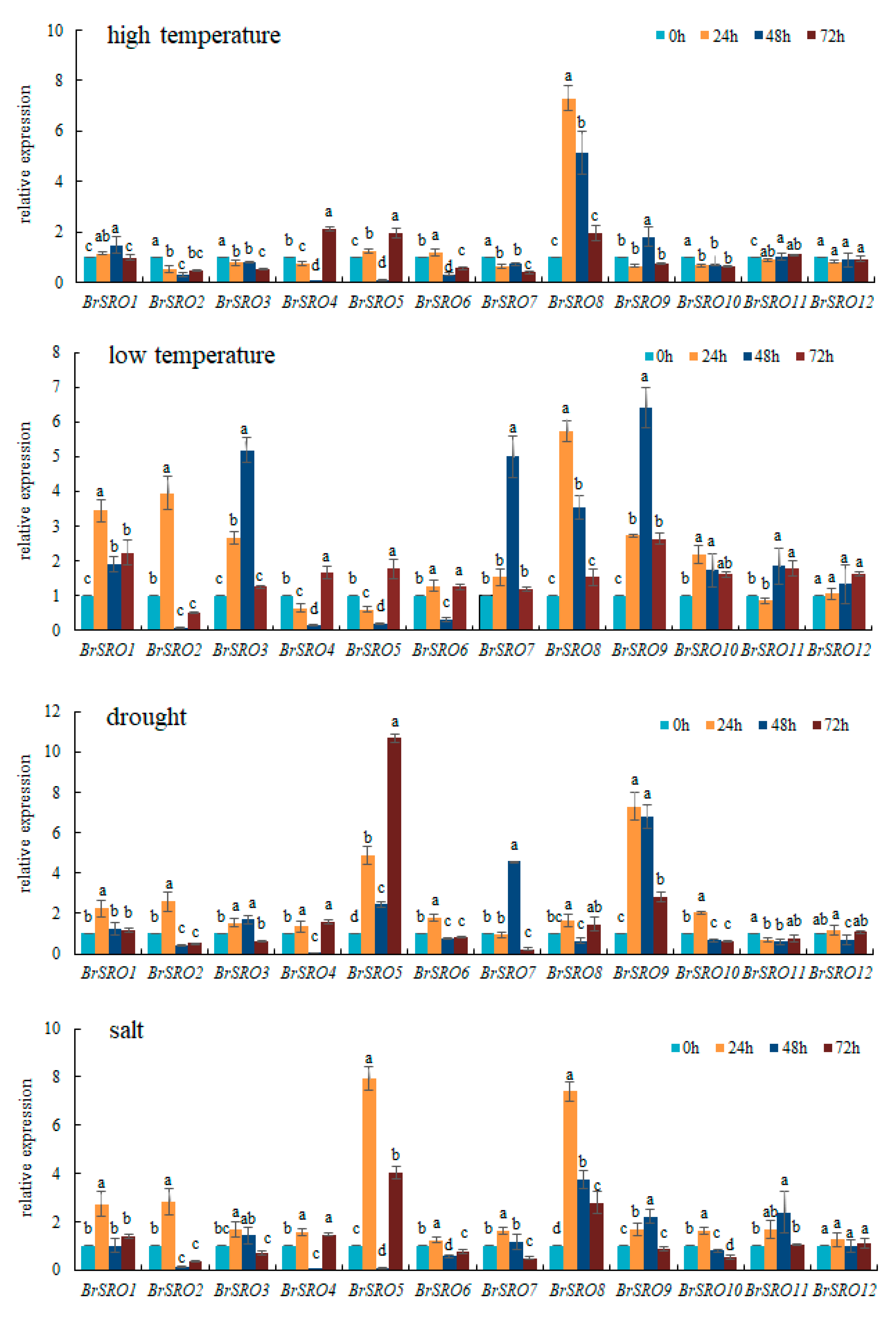
2.6. Relative Expression of 12 BrSRO Genes
3. Discussion
4. Materials and Methods
4.1. Identification and Sequence Analysis of SRO Genes in Chinese Cabbage
4.2. Phylogenetic Analysis of SRO Genes in Chinese Cabbage
4.3. Gene Structure and Conserved Motifs Analysis of BrSROs
4.4. Chromosomal Distribution and Cis-Element Analyses of SRO Genes in Chinese Cabbage
4.5. Plant Materials, Growth Conditions and Treatments
4.6. RNA Isolation and qRT-PCR Analysis
Author Contributions
Funding
Conflicts of Interest
References
- Liu, S.; Liu, S.; Wang, M.; Wei, T.; Meng, C.; Wang, M.; Xia, G. A wheat similar to rcd-one gene enhances seedling growth and abiotic stress resistance by modulating redox homeostasis and maintaining genomic integrity. Plant Cell 2014, 26, 164–180. [Google Scholar] [CrossRef] [PubMed]
- You, J.; Zong, W.; Du, H.; Hu, H.; Xiong, L. A special member of the rice sro family, ossro1c, mediates responses to multiple abiotic stresses through interaction with various transcription factors. Plant Mol. Biol. 2014, 84, 693–705. [Google Scholar] [CrossRef] [PubMed]
- Jaspers, P.; Blomster, T.; Brosché, M.; Salojärvi, J.; Ahlfors, R.; Vainonen, J.P.; Reddy, R.A.; Immink, R.; Angenent, G.; Turck, F.; et al. Unequally redundant rcd1 and sro1 mediate stress and developmental responses and interact with transcription factors. Plant J. 2009, 60, 268–279. [Google Scholar] [CrossRef] [PubMed]
- Citarelli, M.; Teotia, S.; Lamb, R.S. Evolutionary history of the poly(adp-ribose) polymerase gene family in eukaryotes. BMC Evol. Biol. 2010, 10, 308. [Google Scholar] [CrossRef]
- Jaspers, P.; Overmyer, K.; Wrzaczek, M.; Vainonen, J.P.; Blomster, T.; Salojärvi, J.; Reddy, R.A.; Kangasjärvi, J. The rst and parp-like domain containing sro protein family: Analysis of protein structure, function and conservation in land plants. BMC Genom. 2010, 11, 170. [Google Scholar] [CrossRef]
- Katiyar-Agarwal, S.; Zhu, J.; Kim, K.; Agarwal, M.; Fu, X.; Huang, A.; Zhu, J.K. The plasma membrane na+/h+ antiporter sos1 interacts with rcd1 and functions in oxidative stress tolerance in Arabidopsis. Proc. Natl. Acad. Sci. USA 2006, 103, 18816–18821. [Google Scholar] [CrossRef]
- Ahlfors, R.; Lång, S.; Overmyer, K.; Jaspers, P.; Brosché, M.; Tauriainen, A.; Kollist, H.; Tuominen, H.; Belles-Boix, E.; Piippo, M.; et al. Arabidopsis radical-induced cell death1 belongs to the wwe protein-protein interaction domain protein family and modulates abscisic acid, ethylene, and methyl jasmonate responses. Plant Cell 2004, 16, 1925–1937. [Google Scholar] [CrossRef]
- Vainonen, J.P.; Jaspers, P.; Wrzaczek, M.; Lamminmäki, A.; Reddy, R.A.; Vaahtera, L.; Brosché, M.; Kangasjärvi, J. Rcd1-dreb2a interaction in leaf senescence and stress responses in Arabidopsis thaliana. Biochem. J. 2012, 442, 573–581. [Google Scholar] [CrossRef]
- Teotia, S.; Lamb, R.S. The paralogous genes radical-induced cell death1 and similar to rcd one1 have partially redundant functions during Arabidopsis development. Plant Physiol. 2009, 151, 180–198. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Gao, L.; Jin, P.; Cui, L. The similar to rcd-one 1 protein sro1 interacts with gpx3 and functions in plant tolerance of mercury stress. Biosci. Biotechnol. Biochem. 2018, 82, 74–80. [Google Scholar] [CrossRef]
- Borsani, O.; Zhu, J.; Verslues, P.E.; Sunkar, R.; Zhu, J.K. Endogenous sirnas derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 2005, 123, 1279–1291. [Google Scholar] [CrossRef] [PubMed]
- Li, B.Z.; Zhao, X.; Zhao, X.L.; Peng, L. Structure and function analysis of Arabidopsis thaliana sro protein family. Yi Chuan Hered. 2013, 35, 1189–1197. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Li, R.; Qu, F.; Yao, J.; Hao, Y.; Wang, X.; You, C. Identification of the sro gene family in apples (malus × domestica) with a functional characterization of mdrcd1. Tree Genet. Genomes 2017, 13, 94. [Google Scholar] [CrossRef]
- You, J.; Zong, W.; Li, X.; Ning, J.; Hu, H.; Li, X.; Xiao, J.; Xiong, L. The snac1-targeted gene ossro1c modulates stomatal closure and oxidative stress tolerance by regulating hydrogen peroxide in rice. J. Exp. Bot. 2013, 64, 569–583. [Google Scholar] [CrossRef]
- Cheng, F.; Liu, S.; Wu, J.; Fang, L.; Sun, S.; Liu, B.; Li, P.; Hua, W.; Wang, X. Brad, the genetics and genomics database for brassica plants. BMC Plant Biol. 2011, 11, 136. [Google Scholar] [CrossRef]
- Huang, X.Y.; Tao, P.; Li, B.Y.; Wang, W.H.; Yue, Z.C.; Lei, J.L.; Zhong, X.M. Genome-wide identification, classification, and analysis of heat shock transcription factor family in chinese cabbage (Brassica rapa pekinensis). Genet. Mol. Res. GMR 2015, 14, 2189–2204. [Google Scholar] [CrossRef]
- Tao, P.; Zhong, X.; Li, B.; Wang, W.; Yue, Z.; Lei, J.; Guo, W.; Huang, X. Genome-wide identification and characterization of aquaporin genes (aqps) in chinese cabbage (Brassica rapa ssp. Pekinensis). Mol. Genet. Genom. MGG 2014, 289, 1131–1145. [Google Scholar] [CrossRef]
- Liu, Y.; Guan, X.; Liu, S.; Yang, M.; Ren, J.; Guo, M.; Huang, Z.; Zhang, Y. Genome-wide identification and analysis of tcp transcription factors involved in the formation of leafy head in chinese cabbage. Int. J. Mol. Sci. 2018, 19, 847. [Google Scholar] [CrossRef]
- Wang, Z.; Tang, J.; Hu, R.; Wu, P.; Hou, X.L.; Song, X.M.; Xiong, A.S. Genome-wide analysis of the r2r3-myb transcription factor genes in chinese cabbage (Brassica rapa ssp. Pekinensis) reveals their stress and hormone responsive patterns. BMC Genom. 2015, 16, 17. [Google Scholar] [CrossRef]
- Otto, H.; Reche, P.A.; Bazan, F.; Dittmar, K.; Haag, F.; Koch-Nolte, F. In silico characterization of the family of parp-like poly(adp-ribosyl)transferases (parts). BMC Genom. 2005, 6, 139. [Google Scholar] [CrossRef]
- Hassa, P.O.; Hottiger, M.O. The diverse biological roles of mammalian parps, a small but powerful family of poly-adp-ribose polymerases. Front. Biosci. J. Virtual Libr. 2008, 13, 3046–3082. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.Y.; Zhang, T.; Kraus, W.L. Poly(adp-ribosyl)ation by parp-1: ‘Par-laying’ nad+ into a nuclear signal. Genes Dev. 2005, 19, 1951–1967. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Guo, C.; Shan, H.; Kong, H. Divergence of duplicate genes in exon-intron structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1187–1192. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.W.; Penny, D. Patterns of intron loss and gain in plants: Intron loss-dominated evolution and genome-wide comparison of o. Sativa and a. Thaliana. Mol. Biol. Evol. 2007, 24, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Cheong, J.J.; Choi, Y.D. Methyl jasmonate as a vital substance in plants. Trends Genet. TIG 2003, 19, 409–413. [Google Scholar] [CrossRef]
- Chini, A.; Fonseca, S.; Fernández, G.; Adie, B.; Chico, J.M.; Lorenzo, O.; García-Casado, G.; López-Vidriero, I.; Lozano, F.M.; Ponce, M.R.; et al. The jaz family of repressors is the missing link in jasmonate signalling. Nature 2007, 448, 666–671. [Google Scholar] [CrossRef]
- Eriksson, S.; Böhlenius, H.; Moritz, T.; Nilsson, O. Ga4 is the active gibberellin in the regulation of leafy transcription and Arabidopsis floral initiation. Plant Cell 2006, 18, 2172–2181. [Google Scholar] [CrossRef]
- Liang, W.; Ma, X.; Wan, P.; Liu, L. Plant salt-tolerance mechanism: A review. Biochem. Biophys. Res. Commun. 2018, 495, 286–291. [Google Scholar] [CrossRef]
- Wang, Q.; Guan, C.; Wang, P.; Ma, Q.; Bao, A.K.; Zhang, J.L.; Wang, S.M. The effect of Athkt1;1 or AtSOS1 mutation on the expressions of Na⁺ or K⁺ transporter genes and ion homeostasis in Arabidopsis thaliana under salt stress. Int. J. Mol. Sci. 2019, 20, 1085. [Google Scholar] [CrossRef]
- Ahuja, I.; de Vos, R.C.; Bones, A.M.; Hall, R.D. Plant molecular stress responses face climate change. Trends Plant Sci. 2010, 15, 664–674. [Google Scholar] [CrossRef]
- TAIR. Available online: http://www.Arabidopsis.org (accessed on 11 November 2015).
- Wang, X.; Wang, H.; Wang, J.; Sun, R.; Wu, J.; Liu, S.; Bai, Y.; Mun, J.H.; Bancroft, I.; Cheng, F.; et al. The genome of the mesopolyploid crop species brassica rapa. Nat. Genet. 2011, 43, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Banana Genome Hub. Available online: http://banana-genome-hub.southgreen.fr/ (accessed on 17 September 2020).
- RGAP. Available online: http://rice.plantbiology.msu.edu (accessed on 11 November 2015).
- Phytozome. Available online: https://phytozome.jgi.doe.gov/pz/portal.html (accessed on 17 September 2020).
- SMART. Available online: http://smart.embl.de/ (accessed on 11 November 2015).
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. Expasy: Sib bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.S.; Lin, C.J.; Hwang, J.K. Predicting subcellular localization of proteins for gram-negative bacteria by support vector machines based on n-peptide compositions. Protein Sci. A Publ. Protein Soc. 2004, 13, 1402–1406. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. Mega 7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evol. Int. J. Org. Evol. 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. Meme suite: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. Gsds 2.0: An upgraded gene feature visualization server. Bioinformatics (Oxford, England) 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Voorrips, R.E. Mapchart: Software for the graphical presentation of linkage maps and qtls. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. Plantcare, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative pcr and the 2(-delta delta c(t)) method. Methods (San Diego, CA) 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]

| Gene Name | Gene ID | Chromosome Location | Protein Length (aa) | Molecular Weight (kd) | PI | Total Number of Atoms | INSTABILITY Index | Fat Index | Predicted Subcellular Localization |
|---|---|---|---|---|---|---|---|---|---|
| BrSRO1 | Bra033139 | Chromosome A02: 16,846,412–16,846,733 | 77 | 8830.55 | 9.6 | 1272 | 33.9 | 100.13 | Chloroplast. Nucleus. |
| BrSRO2 | Bra029254 | Chromosome A02: 26,296,663–26,297,969 | 303 | 34,150.08 | 8.68 | 4793 | 59.1 | 82.71 | Chloroplast. |
| BrSRO3 | Bra017317 | Chromosome A04: 15,393,395–15,395,430 | 530 | 58,637.64 | 6.99 | 8192 | 34.19 | 80.55 | Nucleus. |
| BrSRO4 | Bra005336 | Chromosome A05: 4,905,877–4,908,322 | 524 | 58,577.61 | 6.1 | 8178 | 37.71 | 80.29 | Chloroplast. |
| BrSRO5 | Bra010096 | Chromosome A06: 19,383,958–19,385,124 | 313 | 34,876.77 | 8.59 | 4889 | 57.49 | 80 | Nucleus. |
| BrSRO6 | Bra033662 | Chromosome A06: 25,753,358–25,757,833 | 771 | 85,056.17 | 9.07 | 11,856 | 57.48 | 63.97 | Nucleus. |
| BrSRO7 | Bra012380 | Chromosome A07: 8,098,261–8,099,752 | 310 | 34,575.22 | 8.86 | 4865 | 45.5 | 85.29 | Chloroplast. |
| BrSRO8 | Bra016219 | Chromosome A07: 18,821,147–18,822,313 | 304 | 33,789.47 | 8.15 | 4744 | 39 | 88.16 | Chloroplast. |
| BrSRO9 | Bra035511 | Chromosome A08: 7,983,100–7,985,345 | 558 | 62,697.49 | 6.59 | 8754 | 37.36 | 79 | Chloroplast. Nucleus. |
| BrSRO10 | Bra023252 | Chromosome A09: 20,223,502–20,225,770 | 482 | 54,230.68 | 6.02 | 7557 | 42.34 | 78.28 | Nucleus. |
| BrSRO11 | Bra024609 | Chromosome A09: 24,077,869–24,079,029 | 308 | 33,418.73 | 6.19 | 4682 | 40.86 | 84.84 | Chloroplast. |
| BrSRO12 | Bra035961 | Scaffold000111: 11,933–15,826 | 779 | 85,523.47 | 8.98 | 11,902 | 55.21 | 61.69 | Nucleus. |
| Gene | Hormonal Response Cis-Elements | Stress Response Cis-Elements | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Abscisic Acid Response Element | Methyl Jasmonate Response Element | Salicylic Acid Response Element | Auxin Response Element | Gibberellin Response Element | Anaerobic Induction Response Element | Drought Response Element | Low-Temperature Response Element | Defense and Stress Response Element | ||||
| ABRE | CGTCA-Motif | TGACG-Motif | TCA-Element | TGA-Element | GARE-Motif | TATC-Box | P-Box | ARE | MBS | LTR | TC-Rich Repeats | |
| BrSRO1 | 0 | 3 | 3 | 0 | 0 | 1 | 0 | 0 | 2 | 3 | 0 | 0 |
| BrSRO2 | 4 | 2 | 2 | 0 | 3 | 0 | 2 | 1 | 3 | 0 | 0 | 1 |
| BrSRO3 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 4 | 4 | 1 | 0 |
| BrSRO4 | 5 | 0 | 0 | 1 | 2 | 0 | 0 | 0 | 4 | 1 | 1 | 0 |
| BrSRO5 | 5 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 1 | 1 | 0 |
| BrSRO6 | 1 | 2 | 2 | 0 | 2 | 0 | 0 | 0 | 5 | 0 | 0 | 0 |
| BrSRO7 | 2 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 0 |
| BrSRO8 | 3 | 5 | 2 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 0 | 1 |
| BrSRO9 | 0 | 3 | 3 | 0 | 1 | 1 | 0 | 0 | 4 | 2 | 0 | 0 |
| BrSRO10 | 1 | 4 | 5 | 1 | 1 | 1 | 0 | 3 | 0 | 0 | 1 | 0 |
| BrSRO11 | 3 | 3 | 3 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
| BrSRO12 | 5 | 4 | 4 | 1 | 1 | 0 | 0 | 2 | 0 | 2 | 0 | 0 |
| Gene Name | Forward Primer Sequence (5′-3′) | Reverse Primer Sequence (5′-3′) |
|---|---|---|
| BrSRO01 | AAGCTGAGGATGATTGTTGGAGA | CAAAGCAGTGTGTGGTAAGCG |
| BrSRO02 | GGGTTTGCCGCCGTTGGATC | TTTGCCGCCGCCTTCTTCAC |
| BrSRO03 | AAGCCTGCTGAGGAGGAAGACC | CGACGCCACCTGAAAACCTATACG |
| BrSRO04 | GAACTCACGGCTCACCTTGGAAG | GAGCAGAGGGTAAGGCATCAAAGC |
| BrSRO05 | AGCTGCGGAGTCGGAAGATGG | CCTCGTGGAACAACCTCAGACTTC |
| BrSRO06 | AATGAATGCTCGTGGTCCGTTGG | GCTTGGTGGTGGCGGTGAAG |
| BrSRO07 | GCGATCACCACGAGAGCCAAG | AGCCAGCGTACCAACCGTATTTG |
| BrSRO08 | GCGGAGGCTATGAAGAGGAAGAAC | CGACCTCGCTGCTGCTAAACC |
| BrSRO09 | CACCAAACCCGCAGACCCAAG | TGACCAGCGACTTCCCAGAGC |
| BrSRO10 | TCTGGTGTCAAGCCTGCTGGAG | CGAGCTTCCGCAATCTCACTGG |
| BrSRO11 | GCGGTTGTGTCAGTGCTGTCC | GCCACTTGTCTCATCTTCCGAACC |
| BrSRO12 | GTGTGGAAGAAAGGATGCGAGGAC | CGTTGATTTGCTGCCGAACATCTG |
| actin | CCAGGAATCGCTGACCGTAT | CTGTTGGAAAGTGCTGAGGGA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qiao, Y.; Gao, X.; Liu, Z.; Wu, Y.; Hu, L.; Yu, J. Genome-Wide Identification and Analysis of SRO Gene Family in Chinese Cabbage (Brassica rapa L.). Plants 2020, 9, 1235. https://doi.org/10.3390/plants9091235
Qiao Y, Gao X, Liu Z, Wu Y, Hu L, Yu J. Genome-Wide Identification and Analysis of SRO Gene Family in Chinese Cabbage (Brassica rapa L.). Plants. 2020; 9(9):1235. https://doi.org/10.3390/plants9091235
Chicago/Turabian StyleQiao, Yali, Xueqin Gao, Zeci Liu, Yue Wu, Linli Hu, and Jihua Yu. 2020. "Genome-Wide Identification and Analysis of SRO Gene Family in Chinese Cabbage (Brassica rapa L.)" Plants 9, no. 9: 1235. https://doi.org/10.3390/plants9091235
APA StyleQiao, Y., Gao, X., Liu, Z., Wu, Y., Hu, L., & Yu, J. (2020). Genome-Wide Identification and Analysis of SRO Gene Family in Chinese Cabbage (Brassica rapa L.). Plants, 9(9), 1235. https://doi.org/10.3390/plants9091235




