What Happened to the Phycobilisome?
Abstract
1. Introduction
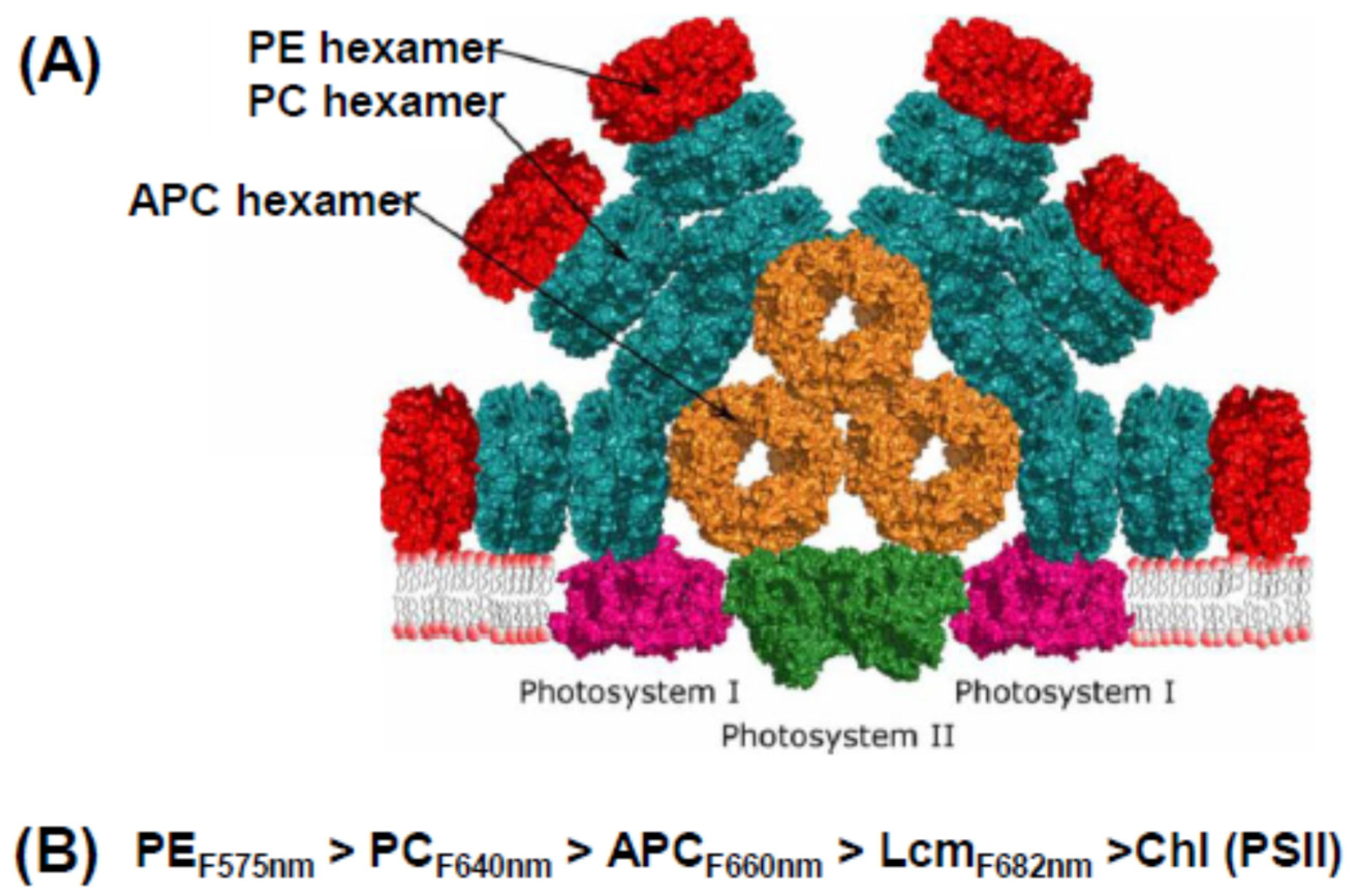
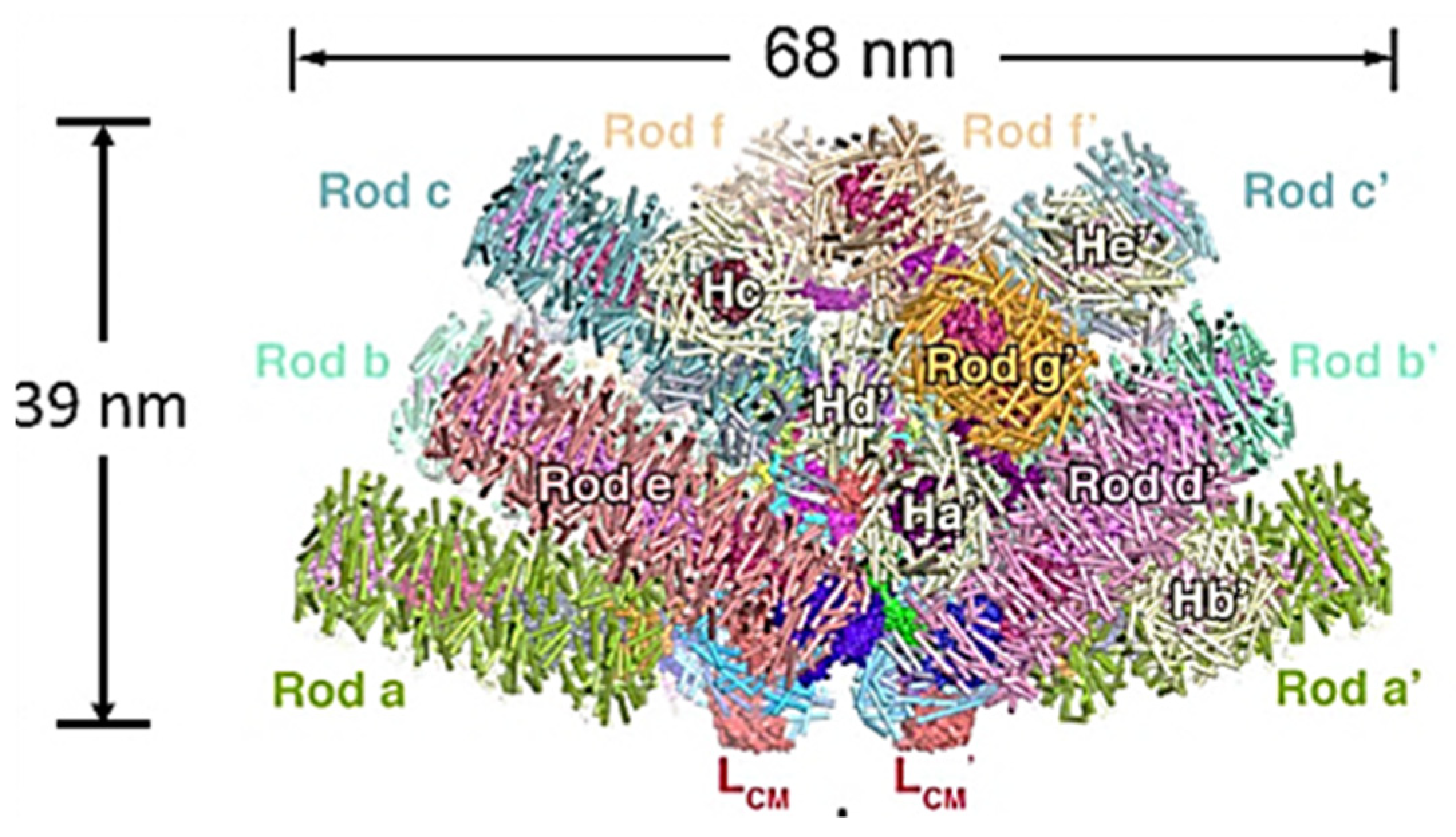
2. Phycobilisome Basics
3. What a Few Deviant Cyanobacteria Might Tell Us about PBS Loss
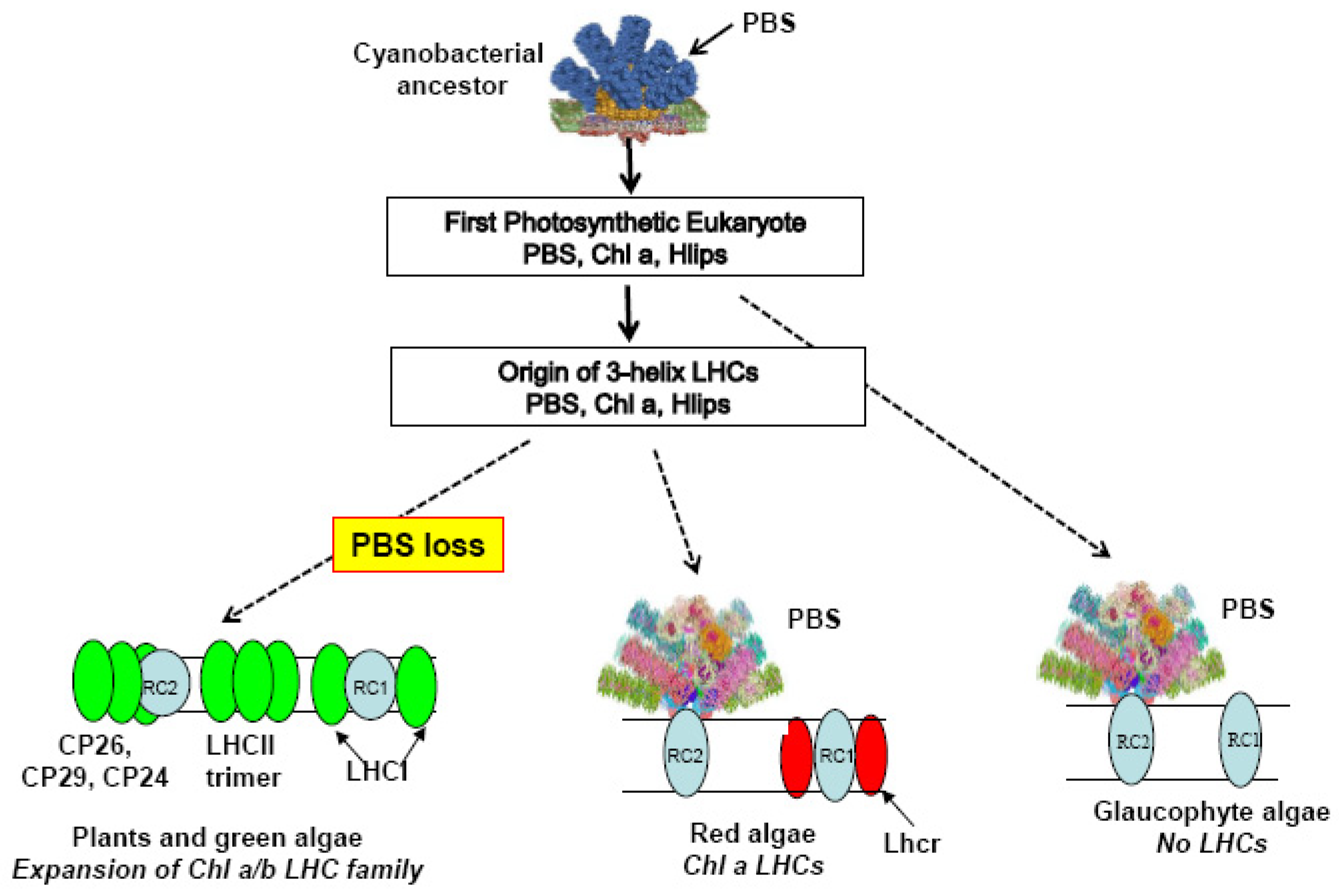
4. Light-Harvesting in the Three Primary Lineages of Photosynthetic Eukaryotes
5. Advantages and Disadvantages of Losing the PBS
6. PBS Loss after Secondary Endosymbiosis
7. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Arbalat, A.; Canestro, C. Evolution by gene loss. Nat. Rev. Genet. 2016, 379–391. [Google Scholar] [CrossRef] [PubMed]
- Olson, M.V. When less is more: Gene loss as an engine of evolutionary change. Am. J. Hum. Genet. 1999, 64, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Wolf, Y.I.; Koonin, E.V. Genome reduction as the dominant mode of evolution. BioEssays 2013, 35, 829–837. [Google Scholar] [CrossRef] [PubMed]
- Monroe, J.G.; Powell, T.; Price, N.; Mullen, J.L.; Howard, A.; Evans, K.; Lovell, J.T.; McKay, J.K. Drought adaptation in Arabidopsis thaliana by extensive genetic loss-of-function. Elife 2018, 7, e41038. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, D.; Qiu, H.; Lee, J.; Yoon, H.S.; Weber, A.P.M.; Price, D.C. When Less is More: Red Algae as Models for Studying Gene Loss and Genome Evolution in Eukaryotes. Crit. Rev. Plant Sci. 2018, 37, 81–99. [Google Scholar] [CrossRef]
- Mimuro, M.; Kikuchi, H. Antenna Systems and Energy Transfer in Cyanophyta and Rhodophyta. In Plant Cell Monographs; Springer: Dordrecht, Netherlands, 2003; pp. 281–306. [Google Scholar]
- Watanabe, M.; Ikeuchi, M. Phycobilisome: Architecture of a light-harvesting supercomplex. Photosynth. Res. 2013, 116, 265–276. [Google Scholar] [CrossRef]
- Gantt, E.; Grabowski, B.; Cunningham, F.X. Antenna systems of red algae: Phycobilisomes with photosystem II and chlorophyll complexes with photosystem I. In Light-Harvesting Antennas in Photosynthesis; Green, B.R., Parson, W.W., Eds.; Kluwer Academic Publishers: Dordrecht, Netherlands, 2003; pp. 307–322. [Google Scholar]
- Watanabe, M.; Sato, M.; Kondo, K.; Narikawa, R.; Ikeuchi, M. Phycobilisome model with novel skeleton-like structures in a glaucocystophyte Cyanophora paradoxa. Biochim. Biophys. Acta Bioenerg. 2012, 1817, 1428–1435. [Google Scholar] [CrossRef]
- Green, B.R.; Anderson, J.M.; Parson, W.W. Photosynthetic Membranes and Their Light-Harvesting Antennas. In Plant Cell Monographs; Springer: Dordrecht, Netherlands, 2003; pp. 1–28. [Google Scholar]
- Lynch, M.; Field, M.C.; Goodson, H.V.; Malik, H.S.; Pereira-Leal, J.B.; Roos, D.S.; Turkewitz, A.P.; Sazer, S. Evolutionary cell biology: Two origins, one objective. Proc. Natl. Acad. Sci. USA 2014, 111, 16990–16994. [Google Scholar] [CrossRef]
- Gould, S.J. Wonderful Life: The Burgess Shale and the Nature of History; WW Norton & Company: New York, NY, USA, 1989. [Google Scholar]
- Blount, Z.D.; Lenski, R.E.; Losos, J.B. Contingency and determinism in evolution: Replaying life’s tape. Science 2018, 362, eaam5979. [Google Scholar] [CrossRef]
- Bridgham, J.T.; Ortlund, E.A.; Thornton, J.W. An epistatic ratchet constrains the direction of glucocorticoid receptor evolution. Nature 2009, 461, 515–519. [Google Scholar] [CrossRef]
- Natarajan, C.; Hoffmann, F.G.; Weber, R.E.; Fago, A.; Witt, C.C.; Storz, J.F. Predictable convergence in hemoglobin function has unpredictable molecular underpinnings. Science 2016, 354, 336–339. [Google Scholar] [CrossRef] [PubMed]
- Bryant, D.A.; Canniffe, D.P. How nature designs light-harvesting antenna systems: Design principles and functional realization in chlorophototrophic prokaryotes. J. Phys. B: At. Mol. Opt. Phys. 2018, 51, 33001. [Google Scholar] [CrossRef]
- Saer, R.G.; Blankenship, R.E. Light harvesting in phototrophic bacteria: Structure and function. Biochem. J. 2017, 474, 2107–2131. [Google Scholar] [CrossRef] [PubMed]
- Apt, K.E.; Collier, J.L.; Grossman, A.R. Evolution of the phycobiliproteins. J. Mol. Biol. 1995, 248, 79–96. [Google Scholar] [CrossRef]
- Liu, H.; Zhang, H.; Niedzwiedzki, D.M.; Prado, M.; He, G.; Gross, M.L.; Blankenship, R.E. Phycobilisomes supply excitations to both photosystems in a megacomplex in cyanobacteria. Science 2013, 342, 1104–1107. [Google Scholar] [CrossRef]
- Chang, L.; Liu, X.; Li, Y.; Liu, C.-C.; Yang, F.; Zhao, J.; Sui, S.-F. Structural organization of an intact phycobilisome and its association with photosystem II. Cell Res. 2015, 25, 726–737. [Google Scholar] [CrossRef]
- Zhang, J.; Ma, J.; Liu, D.; Qin, S.; Sun, S.; Zhao, J.; Sui, S.-F. Structure of phycobilisome from the red alga Griffithsia pacifica. Nature 2017, 551, 57–63. [Google Scholar] [CrossRef]
- Chen, M.; Bibby, T.S. Photosynthetic apparatus of antenna-reaction centre supercomplexes in oxyphotobacteria: Insight through significance of Pcb/IsiA proteins. Photosyn. Res. 2017, 86, 165–173. [Google Scholar] [CrossRef]
- Green, B.R.; Gantt, E. Distal and extrinsic photosytem II antennas. In Photosystem II: The Light-Driven Water: Plastoquinone Oxidoreductase; Wydrzynski, T., Satoh, K., Eds.; Springer: Dordrecht, Netherlands, 2005; pp. 23–44. [Google Scholar]
- Hu, Q.; Marquardt, J.; Iwasaki, I.; Miyashita, H.; Kurano, N.; Mörschel, E.; Miyachi, S. Molecular structure, localization and function of biliproteins in the chlorophyll a/d containing oxygenic photosynthetic prokaryote Acaryochloris marina. Biochim. Biophys. Acta Bioenerg. 1999, 1412, 250–261. [Google Scholar] [CrossRef]
- La Roche, J.; van der Staay, G.W.M.; Partensky, F.; Ducret, A.; Aebersold, R.; Li, R.; Golden, S.S.; Hiller, R.G.; Wrench, P.M.; Larkum, A.W.D.; et al. Independent evolution of the prochlorophyte and green plant chlorophyll a/b light-harvesting proteins. Proc. Natl. Acad. Sci. USA 1996, 93, 15244–15248. [Google Scholar] [CrossRef]
- Van der Staay, G.W.M.; Yurkova, N.; Green, B.R. The 38 kDa chlorophyll a/b protein of the prokaryote Prochlorothrix hollandica is encoded by a divergent pcb gene. Plant Mol. Biol. 1998, 36, 709–716. [Google Scholar] [CrossRef]
- Chen, M.; Hiller, R.G.; Howe, C.J.; Larkum, A.W.D. Unique origin and lateral transfer of prokaryotic chlorophyll-b and chlorophyll-d light-harvesting systems. Mol. Biol. Evol. 2005, 22, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.-Y.; Bandyopadhyay, A.; Pakrasi, H.B. Function, regulation and distribution of IsiA, a membrane-bound chlorophyll a-antenna protein in cyanobacteria. Photosynthetica 2018, 56, 322–333. [Google Scholar] [CrossRef]
- Shih, P.M.; Wu, D.; Latifi, A.; Axen, S.D.; Fewer, D.P.; Talla, E.; Calteau, A.; Cai, F.; Tandeau de Marsac, N.; Rippka, R.; et al. Improving the coverage of the cyanobcterial phylum using diversity-driven genome sequencing. Proc. Natl. Acad. Sci. USA 2013, 110, 1053–1058. [Google Scholar] [CrossRef] [PubMed]
- Pennisi, E. Making waves. Science 2017, 355, 1006–1009. [Google Scholar] [CrossRef]
- Hess, W.R. Genome analysis of marine photosynthetic microbes and their global role. Curr. Opin. Biotechnol. 2004, 15, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Oborník, M. Endosymbiotic Evolution of Algae, Secondary Heterotrophy and Parasitism. Biomolecules 2019, 9, 266. [Google Scholar] [CrossRef] [PubMed]
- de Vries, J.; Archibald, J.M. Endosymbiosis: Did plastids evolve from a freshwater cyanobacterium? Curr. Biol. 2017, 27, R103–R122. [Google Scholar] [CrossRef]
- Tomitani, A.; Okada, K.; Miyashita, H.; Matthijs, H.C.P.; Ohno, T.; Tanaka, A. Chlorophyll b and phycobilins in the common ancestor of cyanobacteria and chloroplasts. Nature 1999, 400, 159–162. [Google Scholar] [CrossRef]
- Kirilovsky, D.; Kerfeld, C.A. Cyanobacterial photoprotection by the orange carotenoid protein. Nat. Plants 2016, 2, 16180. [Google Scholar] [CrossRef]
- Komenda, J.; Sobotka, R. Cyanobacterial high-light-inducible proteins—Protectors of chlorophyll–protein synthesis and assembly. Biochim. Biophys. Acta Bioenerg. 2016, 1857, 288–295. [Google Scholar] [CrossRef] [PubMed]
- Neilson, J.A.D.; Durnford, D.G. Evolutionary distribution of light-harvesting complex-like proteins in photosynthetic eukaryotes. Genome 2010, 53, 68–78. [Google Scholar] [CrossRef] [PubMed]
- Green, B.R. Sequence conservation of light-harvesting and stress-response proteins in relation to the three-dimensional molecular structure of LHCII. Photosynth. Res. 1995, 44, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Green, B.R. The Evolution of Light-harvesting Antennas. In Plant Cell Monographs; Springer: Dordrecht, Netherlands, 2003; pp. 129–168. [Google Scholar]
- Durnford, D.; Deane, J.; Tan, S.; McFadden, G.; Gantt, E.; Green, B.; Deane, J. A Phylogenetic Assessment of the Eukaryotic Light-Harvesting Antenna Proteins, with Implications for Plastid Evolution. J. Mol. Evol. 1999, 48, 59–68. [Google Scholar] [CrossRef]
- Neilson, J.A.D.; Durnford, D.G. Structural and functional diversification of the light-harvesting complexes in photosynthetic eukaryotes. Photosyn. Res. 2010, 106, 57–71. [Google Scholar] [CrossRef]
- Brawley, S.H.; Blouin, N.A.; Ficko-Blean, E.; Wheeler, G.L.; Lohr, M.; Goodson, H.V.; Jenkins, J.W.; Blaby-Haas, C.E.; Helliwell, K.E.; Chan, C.X.; et al. Insights into the red algae and eukaryotic evolution from the genome ofPorphyra umbilicalis (Bangiophyceae, Rhodophyta). Proc. Natl. Acad. Sci. USA 2017, 114, E6361–E6370. [Google Scholar] [CrossRef]
- Pi, X.; Tian, L.; Dai, H.-E.; Qin, X.; Cheng, L.; Kuang, T.; Sui, S.-F.; Shen, J.-R. Unique organization of photosystem I–light-harvesting supercomplex revealed by cryo-EM from a red alga. Proc. Natl. Acad. Sci. USA 2018, 115, 4423–4428. [Google Scholar] [CrossRef]
- Antoshvili, M.; Caspy, I.; Hippler, M.; Nelson, N. Structure and function of photosystem I in Cyanidioschyzon merolae. Photosyn. Res. 2019, 139, 499–508. [Google Scholar] [CrossRef]
- Green, B.R. Evolution of light-harvestng antennas in an oxygen world. In Evolution of Aquatic Photoautotrophs; Falkowski, P.G., Knoll, A.H., Eds.; Academic Press: New York, NY, USA, 2008; pp. 37–53. [Google Scholar]
- Ballottari, M.; Girardon, J.; Dall’Osto, L.; Bassi, R. Evolution and functional properties of Photosystem II light harvesting complexes in eukaryotes. Biochim. Biophys. Acta Bioenerg. 2012, 1817, 143–157. [Google Scholar] [CrossRef]
- Stadnichuk, I.N.; Tropin, I.V. Antenna replacement in the evolutionary origin of chloroplasts. Microbiology 2014, 83, 299–314. [Google Scholar] [CrossRef]
- Levy, I.; Gantt, E. Development of photosynthetic activity in porphyridium purpureum (Rhodophyta) following nitrogen starvation1,2. J. Phycol. 1990, 26, 62–68. [Google Scholar] [CrossRef]
- Grossman, A.R.; Schaefer, M.R.; Chiang, G.G.; Collier, J.L. The phycobilisome, a light-harvesting complex responsive to environmental conditions. Microbiol. Rev. 1993, 57, 725–749. [Google Scholar] [PubMed]
- Derks, A.; Schaven, K.; Bruce, D. Diverse mechanisms for photoprotection in photosynthesis. Dynamic regulation of photosystem II excitation in response to rapid environmental change. Biochim. Biophys. Acta Bioenerg. 2015, 1847, 468–485. [Google Scholar] [CrossRef] [PubMed]
- Ueno, Y.; Aikawa, S.; Kondo, A.; Akimoto, S. Energy Transfer in Cyanobacteria and Red Algae: Confirmation of Spillover in Intact Megacomplexes of Phycobilisome and Both Photosystems. J. Phys. Chem. Lett. 2016, 7, 3567–3571. [Google Scholar] [CrossRef] [PubMed]
- Kaňa, R.; Kotabová, E.; Lukeš, M.; Papáček, Š.; Matonoha, C.; Liu, L.-N.; Prášil, O.; Mullineaux, C.W. Phycobiisome mobility and its role in the regulation of light harvesting in red algae. Plant Physiol. 2014, 165, 1618–1631. [Google Scholar] [CrossRef] [PubMed]
- Niyogi, K.K.; Truong, T.B. Evolution of flexible non-photochemical quenching mechanisms that regulate light harvesting in oxygenic photsynthesis. Curr. Opin. Plant Biol. 2013, 16, 307–314. [Google Scholar] [CrossRef]
- Magdaong, N.C.M.; Blankenship, R.E. Photoprotective, excited-state quenching mechanisms in diverse photosynthetic organisms. J. Boil. Chem. 2018, 293, 5018–5025. [Google Scholar] [CrossRef]
- Delphen, E.; Duval, J.-C.; Etienne, A.-L.; Kirilovsky, D. State transitions of ΔpH-dependent quenching of photosystem II fluorescence in red algae. Biochemistry 1996, 35, 9435–9445. [Google Scholar] [CrossRef]
- Haniewicz, P.; Abram, M.; Nosek, L.; Kirkpatrick, J.; El-Mohsnawy, E.; Olmos, J.D.J.; Kouril, R.; Kargul, J.M. Molecular mechanisms of photoadaptation of photosystem I supercomplex from an evolutionary cyanobacterial/algal intermediate. Plant Physiol. 2018, 176, 1433–1451. [Google Scholar] [CrossRef]
- Butterfield, N.J. Proterozoic photosynthesis—A critical review. Palaeontology 2015, 58, 953–972. [Google Scholar] [CrossRef]
- Gibson, T.M.; Shih, P.M.; Cumming, V.M.; Fischer, W.W.; Crockford, P.W.; Hodgskiss, M.S.W.; Wörndie, S.; Creaser, R.A.; Rainbird, R.H.; Skulski, T.M.; et al. Precise age of Bangiomorpha pubescens dates the origin of eukaryotic photosynthesis. Geology 2018, 46, 135–138. [Google Scholar] [CrossRef]
- MacPherson, A.N.; Hiller, R.G. Light-Harvesting Systems in Chlorophyll c-Containing Algae. In Advances in Photosynthesis and Respiration; Springer: Dordrecht, Netherlands, 2003; pp. 323–352. [Google Scholar]
- Secq, M.-P.O.-L.; Grimwood, J.; Shapiro, H.; Armbrust, E.V.; Bowler, C.; Green, B.R. Chloroplast genomes of the diatoms Phaeodactylum tricornutum and Thalassiosira pseudonana: Comparison with other plastid genomes of the red lineage. Mol. Genet. Genom. 2007, 277, 427–439. [Google Scholar] [CrossRef] [PubMed]
- Jiroutová, K.; Kořený, L.; Bowler, C.; Oborník, M. A Gene in the Process of Endosymbiotic Transfer. PLoS ONE 2010, 5, 13234. [Google Scholar] [CrossRef] [PubMed]
- Funk, C.; Alami, M.; Tibiletti, T.; Green, B.R. High light stress and the one-helix LHC-like proteins of the cryptophyte Guillardia theta. Biochim. Biophys. Acta Gen. Subj. 2011, 1807, 841–846. [Google Scholar] [CrossRef] [PubMed]
- Wilk, K.E.; Harrop, S.J.; Edler, D.; Keenan, G.; Sharples, F.; Hiller, R.G.; Curmi, P.M.G. Evolution of a light-harvesting protein by addition of new subunits and rearrangement of conserved elements: Crystal structure of a cryptophyte p;hycoerythrin at 1.63-Å resolution. Proc. Natl. Acad. Sci. USA 1999, 96, 8901–8906. [Google Scholar] [CrossRef] [PubMed]
- Wit, C.D.V.D.W.-D.; Doust, A.B.; Van Stokkum, I.H.M.; Dekker, J.P.; Wilk, K.E.; Curmi, P.M.G.; Scholes, G.D.; Van Grondelle, R. How Energy Funnels from the Phycoerythrin Antenna Complex to Photosystem I and Photosystem II in CryptophyteRhodomonasCS24 Cells. J. Phys. Chem. 2006, 110, 25066–25073. [Google Scholar]
- Kaňa, R.; Kotabová, E.; Sobotka, R.; Prášil, O. Non-photochemical quenching in cryptophyte algal Rhodomonas salina is located in chlorophyll a/c antennae. PLoS ONE 2012, 7, e29700. [Google Scholar] [CrossRef]
- Cheregi, O.; Kotabová, E.; Prášil, O.; Schröder, W.P.; Kaňa, R.; Funk, C. Presence of state transitions in the cryptophyte alga Guillardia theta. J. Esp. Bot. 2015, 66, 6461–6470. [Google Scholar] [CrossRef]
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Green, B.R. What Happened to the Phycobilisome? Biomolecules 2019, 9, 748. https://doi.org/10.3390/biom9110748
Green BR. What Happened to the Phycobilisome? Biomolecules. 2019; 9(11):748. https://doi.org/10.3390/biom9110748
Chicago/Turabian StyleGreen, Beverley R. 2019. "What Happened to the Phycobilisome?" Biomolecules 9, no. 11: 748. https://doi.org/10.3390/biom9110748
APA StyleGreen, B. R. (2019). What Happened to the Phycobilisome? Biomolecules, 9(11), 748. https://doi.org/10.3390/biom9110748



