Mesenchymal Stem Cells as a Promising Cell Source for Integration in Novel In Vitro Models
Abstract
1. Introduction
2. Mesenchymal Stem Cells as a Cell Source
2.1. Brief Research History
2.2. Physiological Sources of MSCs
2.3. Characteristics
2.4. Applicability of MSCs in In Vitro Models
2.4.1. MSC Differentiation Capacity
2.4.2. Enhancing MSC Differentiation with Scaffold Techniques
3. Novel In Vitro Technologies and Their Potential for Application with MSCs
3.1. Limitations of Traditional 2D Cultures
3.2. Microfluidics and Organs-on-a-Chip
3.2.1. Organs-on-a-Chip
3.2.2. MSCs in Microfluidics and in Organs-on-a-Chip
3.2.3. Limitations of Microfluidic and OoC Platforms
3.3. Fabricated 3D Structures: Scaffolds and Bioprinting
3.3.1. Scaffolds
Materials Used in Scaffolds
MSCs in Scaffold Models
3.3.2. Bioprinting
Bioprinting Tissues Containing MSCs
Bioprinting, MSCs and Blood Vessel Generation
3.4. Spheroids and Organoids
3.4.1. MSCs in Spheroids
3.4.2. MSCs in Organoids
4. Future Perspectives and Conclusions
Funding
Conflicts of Interest
Abbreviations
| α-MHC | α-myosin heavy chain |
| 5-aza | 5-Azacytidine |
| AAT | alpha-1 anti-trypsin |
| ACAN | Aggrecan |
| ANP | trial natriuretic peptide |
| BDNF | brain-derived neurotrophic factor |
| BMP | Bone morphogenetic proteins |
| CK18 | cytokeratin 18 |
| CK19 | cytokeratin 19 |
| COL2A1 | chondrocyte marker type II collagen |
| cTnT | cardiac troponin T |
| CX43 | connexin 43 |
| ECs | endothelial cells |
| ESCs | embryonic stem cells |
| FABP2 | Fatty acid-binding protein 2 |
| FABP4 | fatty acid binding protein 4 |
| FGF | fibroblasts growth factor |
| GAG | glycosaminoglycan |
| GATA4 | GATA binding protein 4 |
| HA | hyaluronic acid |
| hEGF | human epithelial growth factor |
| HGF | hepatocyte growth factor |
| HSCSs | hematopoietic stem cells |
| HSPCs | hematopoietic stem and progenitor cells |
| HUVEC | human umbilical vein endothelial cells |
| IGF | insulin-like growth factor |
| iPSCs | induced pluripotent stem cells |
| ISCT | International society for cellular therapy |
| KLK | Kallikreins |
| LPL | lipoprotein lipase |
| MAP2 | microtubule-associated protein 2 |
| MSCs | mesenchymal stem cells |
| Nkx2.5 | homeobox protein 2.5 |
| NURR1 | nuclear receptor related 1 protein |
| NVU | neurovascular unit |
| OoC | Organ-on-a-Chip |
| PC | polycarbonate |
| PCL | polycaprolactone |
| PDMS | polydimethylsiloxane |
| PEG | polyethylene glycol |
| PGA | polyglycolic acid |
| PGLA | poly lactic/glycolic acids co-polymers |
| PLA/PDLLA | polylactic acid |
| poly HEMA | poly hydroxyethyl methacrylate |
| PPARγ | peroxisome proliferator-activated receptor γ |
| PVA | polyvinyl alcohol |
| RA | retinoic acid |
| RUNX2 | runt-related transcription factor 2 |
| SF | Silk fibroin |
| TAT | tyrosine amino transferase |
| TGF-β | transforming growth factor beta |
| VE-cadherin | vascular endothelial cadherin |
| VEGF | vascular endothelial growth factor |
| VEGFR-2 | vascular endothelial growth factor receptor 2 |
| vWF | von Willebrand factor |
References
- Ziółkowska, K.; Kwapiszewski, R.; Brzózka, Z. Microfluidic devices as tools for mimicking the in vivo environment. New J. Chem. 2011, 35, 979. [Google Scholar] [CrossRef]
- Bhushan, A.; Senutovitch, N.; Bale, S.S.; McCarty, W.J.; Hegde, M.; Jindal, R.; Golberg, I.; Berk Usta, O.; Yarmush, M.L.; Vernetti, L.; et al. Towards a three-dimensional microfluidic liver platform for predicting drug efficacy and toxicity in humans. Stem Cell Res. 2013, 4, S16. [Google Scholar] [CrossRef]
- Sung, J.H.; Srinivasan, B.; Esch, M.B.; McLamb, W.T.; Bernabini, C.; Shuler, M.L.; Hickman, J.J. Using physiologically-based pharmacokinetic-guided “body-on-a-chip” systems to predict mammalian response to drug and chemical exposure. Exp. Biol. Med. 2014, 239, 1225–1239. [Google Scholar] [CrossRef] [PubMed]
- Polini, A.; Prodanov, L.; Bhise, N.S.; Manoharan, V.; Dokmeci, M.R.; Khademhosseini, A. Organs-on-a-chip: A new tool for drug discovery. Expert Opin. Drug Discov. 2014, 9, 335–352. [Google Scholar] [CrossRef] [PubMed]
- Guillouzo, A.; Guguen-Guillouzo, C. Evolving concepts in liver tissue modeling and implications for in vitro toxicology. Expert Opin. Drug Metab. Toxicol. 2008, 4, 1279–1294. [Google Scholar] [CrossRef] [PubMed]
- Rauti, R.; Renous, N.; Maoz, B.M. Mimicking the Brain Extracellular Matrix in Vitro: A Review of Current Methodologies and Challenges. Isr. J. Chem. 2019, 59, 1–12. [Google Scholar] [CrossRef]
- Friedenstein, A.J.; Petrakova, K.V.; Kurolesova, A.I.; Frolova, G.P. Heterotopic Transplants of Bone Marrow. Transplantation 1968, 6, 230–247. [Google Scholar] [CrossRef]
- Caplan, A.I. Mesenchymal stem cells. J. Orthop. Res. 1991, 9, 641–650. [Google Scholar] [CrossRef]
- Pittenger, M.F.; Mackay, A.M.; Beck, S.C.; Jaiswal, R.K.; Douglas, R.; Mosca, J.D.; Moorman, M.A.; Simonetti, D.W.; Craig, S.; Marshak, D.R. Multilineage potential of adult human mesenchymal stem cells. Science 1999, 284, 143–147. [Google Scholar] [CrossRef]
- Andrzejewska, A.; Lukomska, B.; Janowski, M. Concise Review: Mesenchymal Stem Cells: From Roots to Boost: MSCs: From Roots to Boost. Stem Cells 2019, 37, 855–864. [Google Scholar] [CrossRef]
- Wagner, W.; Wein, F.; Seckinger, A.; Frankhauser, M.; Wirkner, U.; Krause, U.; Blake, J.; Schwager, C.; Eckstein, V.; Ansorge, W.; et al. Comparative characteristics of mesenchymal stem cells from human bone marrow, adipose tissue, and umbilical cord blood. Exp. Hematol. 2005, 33, 1402–1416. [Google Scholar] [CrossRef]
- Huang, G.T.-J.; Gronthos, S.; Shi, S. Mesenchymal Stem Cells Derived from Dental Tissues vs. Those from Other Sources: Their Biology and Role in Regenerative Medicine. J. Dent. Res. 2009, 88, 792–806. [Google Scholar] [CrossRef]
- Allickson, J.G. Recent Studies Assessing the Proliferative Capability of a Novel Adult Stem Cell Identified in Menstrual Blood. TOSC J. 2011, 3, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Riekstina, U.; Muceniece, R.; Cakstina, I.; Muiznieks, I.; Ancans, J. Characterization of human skin-derived mesenchymal stem cell proliferation rate in different growth conditions. Cytotechnology 2008, 58, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Ullah, I.; Subbarao, R.B.; Rho, G.J. Human mesenchymal stem cells - current trends and future prospective. Biosci. Rep. 2015, 35, e00191. [Google Scholar] [CrossRef] [PubMed]
- Abdal Dayem, A.; Lee, S.B.; Kim, K.; Lim, K.M.; Jeon, T.; Seok, J.; Cho, S.-G. Production of Mesenchymal Stem Cells through Stem Cell Reprogramming. IJMS 2019, 20, 1922. [Google Scholar] [CrossRef]
- Horwitz, E.M.; Le Blanc, K.; Dominici, M.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.C.; Deans, R.J.; Krause, D.S.; Keating, A. Clarification of the nomenclature for MSC: The International Society for Cellular Therapy position statement. Cytotherapy 2005, 7, 393–395. [Google Scholar] [CrossRef]
- Dominici, M.; Le Blanc, K.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.C.; Krause, D.S.; Deans, R.J.; Keating, A.; Prockop, D.J.; Horwitz, E.M. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006, 8, 315–317. [Google Scholar] [CrossRef]
- Fitzsimmons, R.E.B.; Mazurek, M.S.; Soos, A.; Simmons, C.A. Mesenchymal Stromal/Stem Cells in Regenerative Medicine and Tissue Engineering. Stem Cells Int. 2018, 2018, 1–16. [Google Scholar] [CrossRef]
- Anisimov, V.N.; Ukraintseva, S.V.; Yashin, A.I. Cancer in rodents: Does it tell us about cancer in humans? Nat. Rev. Cancer 2005, 5, 807–819. [Google Scholar] [CrossRef]
- Moon, S.Y.; Park, Y.B.; Kim, D.-S.; Oh, S.K.; Kim, D.-W. Generation, culture, and differentiation of human embryonic stem cells for therapeutic applications. Mol. Ther. 2006, 13, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Medvedev, S.P.; Shevchenko, A.I.; Zakian, S.M. Induced Pluripotent Stem Cells: Problems and Advantages when Applying them in Regenerative Medicine. Acta Nat. 2010, 2, 18–28. [Google Scholar] [CrossRef]
- Khanabdali, R.; Rosdah, A.A.; Dusting, G.J.; Lim, S.Y. Harnessing the secretome of cardiac stem cells as therapy for ischemic heart disease. Biochem. Pharmacol. 2016, 113, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Davis, D.R. Cardiac stem cells in the post-Anversa era. Eur. Heart J. 2019, 40, 1039–1041. [Google Scholar] [CrossRef]
- Rosen, E.D.; MacDougald, O.A. Adipocyte differentiation from the inside out. Nat. Rev. Mol. Cell Biol. 2006, 7, 885–896. [Google Scholar] [CrossRef]
- Musina, R.A.; Bekchanova, E.S.; Belyavskii, A.V.; Sukhikh, G.T. Differentiation potential of mesenchymal stem cells of different origin. Bull Exp. Biol. Med. 2006, 141, 147–151. [Google Scholar] [CrossRef]
- Danišovič, Ľ.; Varga, I.; Polák, Š. Growth factors and chondrogenic differentiation of mesenchymal stem cells. Tissue Cell 2012, 44, 69–73. [Google Scholar] [CrossRef]
- Banas, A.; Teratani, T.; Yamamoto, Y.; Tokuhara, M.; Takeshita, F.; Quinn, G.; Okochi, H.; Ochiya, T. Adipose tissue-derived mesenchymal stem cells as a source of human hepatocytes. Hepatology 2007, 46, 219–228. [Google Scholar] [CrossRef]
- Ji, R.; Zhang, N.; You, N.; Li, Q.; Liu, W.; Jiang, N.; Liu, J.; Zhang, H.; Wang, D.; Tao, K.; et al. The differentiation of MSCs into functional hepatocyte-like cells in a liver biomatrix scaffold and their transplantation into liver-fibrotic mice. Biomaterials 2012, 33, 8995–9008. [Google Scholar] [CrossRef]
- Puglisi, M.A.; Saulnier, N.; Piscaglia, A.C.; Tondi, P.; Agnes, S.; Gasbarrini, A. Adipose tissue-derived mesenchymal stem cells and hepatic differentiation: Old concepts and future perspectives. Eur. Rev. Med. Pharm. Sci. 2011, 15, 355–364. [Google Scholar]
- Guo, X.; Bai, Y.; Zhang, L.; Zhang, B.; Zagidullin, N.; Carvalho, K.; Du, Z.; Cai, B. Cardiomyocyte differentiation of mesenchymal stem cells from bone marrow: New regulators and its implications. Stem Cell Res. 2018, 9, 44. [Google Scholar] [CrossRef] [PubMed]
- Szaraz, P.; Gratch, Y.S.; Iqbal, F.; Librach, C.L. In Vitro Differentiation of Human Mesenchymal Stem Cells into Functional Cardiomyocyte-like Cells. JoVE 2017, 55757. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Zhang, X.; Qian, H.; Zhu, W.; Sun, X.; Hu, J.; Zhou, H.; Chen, Y. Mesenchymal Stern Cells from Adult Human Bone Marrow Differentiate into a Cardiomyocyte Phenotype In Vitro. Exp. Biol. Med. (Maywood) 2004, 229, 623–631. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, P.H.; Daibert, A.P.F.; Monteiro, B.S.; Okano, B.S.; Lott, J. Differentiation of Adipose Tissue-Derived Mesenchymal Stem Cells into Cardiomyocytes. Arq Bras Cardiol. 2013, 100, 82–89. [Google Scholar] [CrossRef]
- Anghileri, E.; Marconi, S.; Pignatelli, A.; Cifelli, P.; Galié, M.; Sbarbati, A.; Krampera, M.; Belluzzi, O.; Bonetti, B. Neuronal Differentiation Potential of Human Adipose-Derived Mesenchymal Stem Cells. Stem Cells Dev. 2008, 17, 909–916. [Google Scholar] [CrossRef] [PubMed]
- Ghorbani, S.; Tiraihi, T.; Soleimani, M. Differentiation of mesenchymal stem cells into neuron-like cells using composite 3D scaffold combined with valproic acid induction. J. Biomater. Appl. 2018, 32, 702–715. [Google Scholar] [CrossRef]
- Takeda, Y.S.; Xu, Q. Neuronal Differentiation of Human Mesenchymal Stem Cells Using Exosomes Derived from Differentiating Neuronal Cells. PLoS ONE 2015, 10. [Google Scholar] [CrossRef]
- Yim, E.K.; Pang, S.W.; Leong, K.W. Synthetic Nanostructures Inducing Differentiation of Human Mesenchymal Stem Cells into Neuronal Lineage. Exp. Cell Res. 2007, 313, 1820–1829. [Google Scholar] [CrossRef]
- Păunescu, V.; Deak, E.; Herman, D.; Siska, I.R.; T?anasie, G.; Bunu, C.; Anghel, S.; Tatu, C.A.; Oprea, T.I.; Henschler, R.; et al. In vitro differentiation of human mesenchymal stem cells to epithelial lineage. J. Cell. Mol. Med. 2007, 11, 502–508. [Google Scholar] [CrossRef]
- Spees, J.L.; Olson, S.D.; Ylostalo, J.; Lynch, P.J.; Smith, J.; Perry, A.; Peister, A.; Wang, M.Y.; Prockop, D.J. Differentiation, cell fusion, and nuclear fusion during ex vivo repair of epithelium by human adult stem cells from bone marrow stroma. Proc. Natl. Acad. Sci. USA 2003, 100, 2397–2402. [Google Scholar] [CrossRef]
- Bai, K.; Huang, Y.; Jia, X.; Fan, Y.; Wang, W. Endothelium oriented differentiation of bone marrow mesenchymal stem cells under chemical and mechanical stimulations. J. Biomech. 2010, 43, 1176–1181. [Google Scholar] [CrossRef] [PubMed]
- Janeczek Portalska, K.; Leferink, A.; Groen, N.; Fernandes, H.; Moroni, L.; van Blitterswijk, C.; de Boer, J. Endothelial Differentiation of Mesenchymal Stromal Cells. PLoS ONE 2012, 7, e46842. [Google Scholar] [CrossRef] [PubMed]
- Wingate, K.; Bonani, W.; Tan, Y.; Bryant, S.J.; Tan, W. Compressive elasticity of three-dimensional nanofiber matrix directs mesenchymal stem cell differentiation to vascular cells with endothelial or smooth muscle cell markers. Acta Biomater. 2012, 8, 1440–1449. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-B.; Jiang, X.-B.; Yang, L. Differentiation of rat marrow mesenchymal stem cells into pancreatic islet beta-cells. World J. Gastroenterol. 2004, 10, 3016–3020. [Google Scholar] [CrossRef]
- Pavathuparambil Abdul Manaph, N.; Sivanathan, K.N.; Nitschke, J.; Zhou, X.-F.; Coates, P.T.; Drogemuller, C.J. An overview on small molecule-induced differentiation of mesenchymal stem cells into beta cells for diabetic therapy. Stem Cell Res. 2019, 10, 293. [Google Scholar] [CrossRef]
- Shivakumar, S.B.; Lee, H.-J.; Son, Y.-B.; Bharti, D.; Ock, S.A.; Lee, S.-L.; Kang, Y.-H.; Park, B.-W.; Rho, G.-J. In vitro differentiation of single donor derived human dental mesenchymal stem cells into pancreatic β cell-like cells. Biosci. Rep. 2019, 39, BSR20182051. [Google Scholar] [CrossRef]
- Schneider, R.K.M.; Neuss, S.; Stainforth, R.; Laddach, N.; Bovi, M.; Knuechel, R.; Perez-Bouza, A. Three-dimensional epidermis-like growth of human mesenchymal stem cells on dermal equivalents: Contribution to tissue organization by adaptation of myofibroblastic phenotype and function. Differentiation 2008, 76, 156–167. [Google Scholar] [CrossRef]
- Dos Santos, J.F.; Borçari, N.R.; da Silva Araújo, M.; Nunes, V.A. Mesenchymal stem cells differentiate into keratinocytes and express epidermal kallikreins: Towards an in vitro model of human epidermis. J. Cell. Biochem. 2019, 120, 13141–13155. [Google Scholar] [CrossRef]
- Han, C.-m.; Wang, S.-y.; Lai, P.-p.; Cen, H.-h. Human bone marrow-derived mesenchymal stem cells differentiate into epidermal-like cells in vitro. Differentiation 2007, 75, 292–298. [Google Scholar] [CrossRef]
- Yu, Y.; Song, Y.; Chen, Y.; Zhang, F.; Qi, F. Differentiation of umbilical cord mesenchymal stem cells into hepatocytes in comparison with bone marrow mesenchymal stem cells. Mol. Med. Rep. 2018. [Google Scholar] [CrossRef]
- Urrutia, D.N.; Caviedes, P.; Mardones, R.; Minguell, J.J.; Vega-Letter, A.M.; Jofre, C.M. Comparative study of the neural differentiation capacity of mesenchymal stromal cells from different tissue sources: An approach for their use in neural regeneration therapies. PLoS ONE 2019, 14, e0213032. [Google Scholar] [CrossRef]
- Ricciardi, M.; Malpeli, G.; Bifari, F.; Bassi, G.; Pacelli, L.; Kamdje, A.H.N.; Chilosi, M.; Krampera, M. Comparison of Epithelial Differentiation and Immune Regulatory Properties of Mesenchymal Stromal Cells Derived from Human Lung and Bone Marrow. PLoS ONE 2012, 7, e35639. [Google Scholar] [CrossRef] [PubMed]
- Gabr, M.M.; Zakaria, M.M.; Refaie, A.F.; Khater, S.M.; Ashamallah, S.A.; Ismail, A.M.; El-Badri, N.; Ghoneim, M.A. Generation of Insulin-Producing Cells from Human Bone Marrow-Derived Mesenchymal Stem Cells: Comparison of Three Differentiation Protocols. Biomed. Res. Int. 2014. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, Y.; Yang, M.; Zou, Y.; Liu, H.; Liang, Z.; Yin, Y.; Niu, G.; Yan, Z.; Zhang, B. Efficient Differentiation of Bone Marrow Mesenchymal Stem Cells into Endothelial Cells in Vitro. Eur. J. Vasc. Endovasc. Surg. 2018, 55, 257–265. [Google Scholar] [CrossRef]
- Ma, J.; Guo, L.; Fiene, S.J.; Anson, B.D.; Thomson, J.A.; Kamp, T.J.; Kolaja, K.L.; Swanson, B.J.; January, C.T. High purity human-induced pluripotent stem cell-derived cardiomyocytes: Electrophysiological properties of action potentials and ionic currents. Am. J. Physiol. Heart Circ. Physiol. 2011, 301, H2006–H2017. [Google Scholar] [CrossRef]
- Tanthaisong, P.; Imsoonthornruksa, S.; Ngernsoungnern, A.; Ngernsoungnern, P.; Ketudat-Cairns, M.; Parnpai, R. Enhanced Chondrogenic Differentiation of Human Umbilical Cord Wharton’s Jelly Derived Mesenchymal Stem Cells by GSK-3 Inhibitors. PLoS ONE 2017, 12, e0168059. [Google Scholar] [CrossRef] [PubMed]
- Tazhitdinova, R.; Timoshenko, A.V. The Emerging Role of Galectins and O-GlcNAc Homeostasis in Processes of Cellular Differentiation. Cells 2020, 9, 1792. [Google Scholar] [CrossRef] [PubMed]
- Hanna, H.; Mir, L.M.; Andre, F.M. In vitro osteoblastic differentiation of mesenchymal stem cells generates cell layers with distinct properties. Stem Cell Res. 2018, 9, 203. [Google Scholar] [CrossRef]
- Alkhalil, M.; Smajilagic, A.; Redzic, A. Human Dental Pulp Mesenchymal Stem Cells Isolation and Osteoblast Differentiation. Med. Glas (Zenica) 2015, 12, 27–32. [Google Scholar]
- Youssef, A.; Han, V.K.M. Regulation of Osteogenic Differentiation of Placental-Derived Mesenchymal Stem Cells by Insulin-Like Growth Factors and Low Oxygen Tension. Stem Cells Int. 2017, 2017, 1–17. [Google Scholar] [CrossRef]
- Ansari, A.S.; Yazid, M.D.; Sainik, N.Q.A.V.; Razali, R.A.; Saim, A.B.; Idrus, R.B.H. Osteogenic Induction of Wharton’s Jelly-Derived Mesenchymal Stem Cell for Bone Regeneration: A Systematic Review. Stem Cells Int. 2018, 2018, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Fakhry, M. Molecular mechanisms of mesenchymal stem cell differentiation towards osteoblasts. WJSC 2013, 5, 136. [Google Scholar] [CrossRef] [PubMed]
- Hildebrandt, C.; Büth, H.; Thielecke, H. Influence of cell culture media conditions on the osteogenic differentiation of cord blood-derived mesenchymal stem cells. Ann. Anat. Anat. Anz. 2009, 191, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Luke, A.M.; Patnaik, R.; Kuriadom, S.; Abu-Fanas, S.; Mathew, S.; Shetty, K.P. Human dental pulp stem cells differentiation to neural cells, osteocytes and adipocytes-An in vitro study. Heliyon 2020, 6, e03054. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi, Z.; Afshari, J.T.; Keramati, M.R.; Hamidi, D.; Ganjibakhsh, M.; Zarmehri, A.M.; Jangjoo, A.; Hadi, M.; Ameri, M.A.; Moinzadeh, L. Differentiation of adipocytes and osteocytes from human adipose and placental mesenchymal stem cells. Mesenchymal Stem Cells 2015, 18, 8. [Google Scholar]
- Chen, M.-Y.; Lie, P.-C.; Li, Z.-L.; Wei, X. Endothelial differentiation of Wharton’s jelly–derived mesenchymal stem cells in comparison with bone marrow–derived mesenchymal stem cells. Exp. Hematol. 2009, 37, 629–640. [Google Scholar] [CrossRef]
- Saben, J.; Thakali, K.M.; Lindsey, F.E.; Zhong, Y.; Badger, T.M.; Andres, A.; Shankar, K. Distinct adipogenic differentiation phenotypes of human umbilical cord mesenchymal cells dependent on adipogenic conditions. Exp. Biol. Med. (Maywood) 2014, 239, 1340–1351. [Google Scholar] [CrossRef] [PubMed]
- Pournasr, B.; Mohamadnejad, M.; Bagheri, M.; Aghdami, N.; Malekzadeh, R.; Baharvand, H. In Vitro Differentiation of Human Bone Marrow Mesenchy- mal Stem Cells into Hepatocyte-like Cells. Arch. Iran. Med. 2011, 14, 244–249. [Google Scholar]
- Snykers, S.; Vanhaecke, T.; De Becker, A.; Papeleu, P.; Vinken, M.; Van Riet, I.; Rogiers, V. Chromatin remodeling agent trichostatin A: A key-factor in the hepatic differentiation of human mesenchymal stem cells derived of adult bone marrow. BMC Dev. Biol. 2007, 7, 24. [Google Scholar] [CrossRef]
- Ohkoshi, S.; Hara, H.; Hirono, H.; Watanabe, K.; Hasegawa, K. Regenerative medicine using dental pulp stem cells for liver diseases. WJGPT 2017, 8, 1. [Google Scholar] [CrossRef]
- Bornstein, R.; Macias, M.I.; de la Torre, P.; Grande, J.; Flores, A.I. Human decidua-derived mesenchymal stromal cells differentiate into hepatic-like cells and form functional three-dimensional structures. Cytotherapy 2012, 14, 1182–1192. [Google Scholar] [CrossRef] [PubMed]
- Talaei-Khozani, T.; Borhani-Haghighi, M.; Ayatollahi, M.; Vojdani, Z. An in vitro model for hepatocyte-like cell differentiation from Wharton’s jelly derived-mesenchymal stem cells by cell-base aggregates. Gastroenterol. Hepatol. Bed Bench 2015, 8, 188–199. [Google Scholar] [PubMed]
- Roberts, E.G.; Piekarski, B.L.; Huang, K.; Emani, S.; Wong, J.Y.; Emani, S.M. Evaluation of Placental Mesenchymal Stem Cell Sheets for Myocardial Repair and Regeneration. Tissue Eng. Part A 2019, 25, 867–877. [Google Scholar] [CrossRef] [PubMed]
- Nartprayut, K.; U-Pratya, Y.; Kheolamai, P.; Manochantr, S.; Chayosumrit, M.; Issaragrisil, S.; Supokawej, A. Cardiomyocyte differentiation of perinatally-derived mesenchymal stem cells. Mol. Med. Rep. 2013, 7, 1465–1469. [Google Scholar] [CrossRef] [PubMed]
- Franco Lambert, A.P.; Fraga Zandonai, A.; Bonatto, D.; Cantarelli Machado, D.; Pêgas Henriques, J.A. Differentiation of human adipose-derived adult stem cells into neuronal tissue: Does it work? Differentiation 2009, 77, 221–228. [Google Scholar] [CrossRef]
- Portmann-Lanz, C.B.; Schoeberlein, A.; Portmann, R.; Mohr, S.; Rollini, P.; Sager, R.; Surbek, D.V. Turning placenta into brain: Placental mesenchymal stem cells differentiate into neurons and oligodendrocytes. Am. J. Obstet. Gynecol. 2010, 202, 294.e1–294.e11. [Google Scholar] [CrossRef]
- Bae, K.S.; Park, J.B.; Kim, H.S.; Kim, D.S.; Park, D.J.; Kang, S.J. Neuron-Like Differentiation of Bone Marrow-Derived Mesenchymal Stem Cells. Yonsei Med. J. 2011, 52, 401. [Google Scholar] [CrossRef]
- Shi, Q.; Gao, J.; Jiang, Y.; Sun, B.; Lu, W.; Su, M.; Xu, Y.; Yang, X.; Zhang, Y. Differentiation of human umbilical cord Wharton’s jelly-derived mesenchymal stem cells into endometrial cells. Stem Cell Res. 2017, 8, 246. [Google Scholar] [CrossRef]
- Engler, A.J.; Sen, S.; Sweeney, H.L.; Discher, D.E. Matrix elasticity directs stem cell lineage specification. Cell 2006, 126, 677–689. [Google Scholar] [CrossRef]
- Agrawal, P.; Pramanik, K. Enhanced chondrogenic differentiation of human mesenchymal stem cells in silk fibroin/chitosan/glycosaminoglycan scaffolds under dynamic culture condition. Differentiation 2019, 110, 36–48. [Google Scholar] [CrossRef]
- Agrawal, P.; Pramanik, K.; Vishwanath, V.; Biswas, A.; Bissoyi, A.; Patra, P.K. Enhanced chondrogenesis of mesenchymal stem cells over silk fibroin/chitosan-chondroitin sulfate three dimensional scaffold in dynamic culture condition. J. Biomed. Mater. Res. Part B Appl. Biomater. 2018, 106, 2576–2587. [Google Scholar] [CrossRef]
- Asadpour, S.; Kargozar, S.; Moradi, L.; Ai, A.; Nosrati, H.; Ai, J. Natural biomacromolecule based composite scaffolds from silk fibroin, gelatin and chitosan toward tissue engineering applications. Int. J. Biol. Macromol. 2020, 154, 1285–1294. [Google Scholar] [CrossRef] [PubMed]
- Luetchford, K.A.; Chaudhuri, J.B.; De Bank, P.A. Silk fibroin/gelatin microcarriers as scaffolds for bone tissue engineering. Mater. Sci. Eng. C 2020, 106, 110116. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Xu, P.; Shen, T.; Zhang, Y.; Ye, J.; Gao, C. Influence of pore architectures of silk fibroin/collagen composite scaffolds on the regeneration of osteochondral defects in vivo. J. Mater. Chem. B 2020, 8, 391–405. [Google Scholar] [CrossRef]
- Li, W.-J.; Tuli, R.; Huang, X.; Laquerriere, P.; Tuan, R.S. Multilineage differentiation of human mesenchymal stem cells in a three-dimensional nanofibrous scaffold. Biomaterials 2005, 26, 5158–5166. [Google Scholar] [CrossRef] [PubMed]
- Xin, X.; Hussain, M.; Mao, J.J. Continuing differentiation of human mesenchymal stem cells and induced chondrogenic and osteogenic lineages in electrospun PLGA nanofiber scaffold. Biomaterials 2007, 28, 316–325. [Google Scholar] [CrossRef] [PubMed]
- Donzelli, E.; Salvadè, A.; Mimo, P.; Viganò, M.; Morrone, M.; Papagna, R.; Carini, F.; Zaopo, A.; Miloso, M.; Baldoni, M.; et al. Mesenchymal stem cells cultured on a collagen scaffold: In vitro osteogenic differentiation. Arch. Oral Biol. 2007, 52, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Richardson, S.M.; Curran, J.M.; Chen, R.; Vaughan-Thomas, A.; Hunt, J.A.; Freemont, A.J.; Hoyland, J.A. The differentiation of bone marrow mesenchymal stem cells into chondrocyte-like cells on poly-l-lactic acid (PLLA) scaffolds. Biomaterials 2006, 27, 4069–4078. [Google Scholar] [CrossRef]
- Uebersax, L.; Merkle, H.P.; Meinel, L. Insulin-like growth factor I releasing silk fibroin scaffolds induce chondrogenic differentiation of human mesenchymal stem cells. J. Control. Release 2008, 127, 12–21. [Google Scholar] [CrossRef]
- Thibault, R.A.; Scott Baggett, L.; Mikos, A.G.; Kasper, F.K. Osteogenic Differentiation of Mesenchymal Stem Cells on Pregenerated Extracellular Matrix Scaffolds in the Absence of Osteogenic Cell Culture Supplements. Tissue Eng. Part A 2009, 16, 431–440. [Google Scholar] [CrossRef]
- Meinel, L.; Hofmann, S.; Karageorgiou, V.; Zichner, L.; Langer, R.; Kaplan, D.; Vunjak-Novakovic, G. Engineering cartilage-like tissue using human mesenchymal stem cells and silk protein scaffolds. Biotechnol. Bioeng. 2004, 88, 379–391. [Google Scholar] [CrossRef]
- Kazemnejad, S.; Allameh, A.; Seoleimani, M.; Gharehbaghian, A.; Mohammadi, Y.; Amirizadeh, N.; Esmaeili, S. Functional hepatocyte-like cells derived from human bone marrow mesenchymal stem cells on a novel 3-dimensional biocompatible nanofibrous scaffold. Int. J. Artif. Organs 2008, 31, 500–507. [Google Scholar] [CrossRef]
- Aleahmad, F.; Ebrahimi, S.; Salmannezhad, M.; Azarnia, M.; Jaberipour, M.; Hoseini, M.; Talaei-Khozani, T. Heparin/Collagen 3D Scaffold Accelerates Hepatocyte Differentiation of Wharton’s Jelly-Derived Mesenchymal Stem Cells. Tissue Eng. Regen. Med. 2017, 14, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Gu, H.; Yue, Z.; Leong, W.S.; Nugraha, B.; Tan, L.P. Control of in vitro neural differentiation of mesenchymal stem cells in 3D macroporous, cellulosic hydrogels. Regen. Med. 2010, 5, 245–253. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.; Laco, F.; Ramakrishna, S.; Liao, S.; Chan, C.K. Differentiation of bone marrow-derived mesenchymal stem cells into multi-layered epidermis-like cells in 3D organotypic coculture. Biomaterials 2009, 30, 3251–3258. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Ma, J.; Gao, Y.; Dong, M.; Zheng, Z.; Li, Y.; Tan, R.; She, Z.; Yang, L. Epithelial differentiation of human adipose-derived stem cells (hASCs) undergoing three-dimensional (3D) cultivation with collagen sponge scaffold (CSS) via an indirect co-culture strategy. Stem Cell Res. Ther. 2020, 11, 141. [Google Scholar] [CrossRef]
- Li, Z.; Guo, X.; Palmer, A.F.; Das, H.; Guan, J. High-efficiency matrix modulus-induced cardiac differentiation of human mesenchymal stem cells inside a thermosensitive hydrogel. Acta Biomater. 2012, 8, 3586–3595. [Google Scholar] [CrossRef]
- Crowder, S.W.; Liang, Y.; Rath, R.; Park, A.M.; Maltais, S.; Pintauro, P.N.; Hofmeister, W.; Lim, C.C.; Wang, X.; Sung, H.-J. Poly(ε-caprolactone)-carbon nanotube composite scaffolds for enhanced cardiac differentiation of human mesenchymal stem cells. Nanomedicine (Lond.) 2013, 8. [Google Scholar] [CrossRef]
- Safaeijavan, R.; Soleimani, M.; Divsalar, A.; Eidi, A.; Ardeshirylajimi, A. Comparison of random and aligned PCL nanofibrous electrospun scaffolds on cardiomyocyte differentiation of human adipose-derived stem cells. Iran. J. Basic Med. Sci. 2014, 17, 903–911. [Google Scholar]
- Sridhar, S.; Venugopal, J.R.; Sridhar, R.; Ramakrishna, S. Cardiogenic differentiation of mesenchymal stem cells with gold nanoparticle loaded functionalized nanofibers. Colloids Surf. B Biointerfaces 2015, 134, 346–354. [Google Scholar] [CrossRef]
- Ravichandran, R.; Venugopal, J.R.; Sundarrajan, S.; Mukherjee, S.; Ramakrishna, S. Cardiogenic differentiation of mesenchymal stem cells on elastomeric poly (glycerol sebacate)/collagen core/shell fibers. World J. Cardiol. 2013, 5, 28–41. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.-L.; Chen, C.-P.; Lo, C.-M.; Wang, H.-S. Stiffness-controlled three-dimensional collagen scaffolds for differentiation of human Wharton’s jelly mesenchymal stem cells into cardiac progenitor cells. J. Biomed. Mater. Res. Part A 2016, 104, 2234–2242. [Google Scholar] [CrossRef]
- Nadri, S.; Barati, G.; Mostafavi, H.; Esmaeilzadeh, A.; Enderami, S.E. Differentiation of conjunctiva mesenchymal stem cells into secreting islet beta cells on plasma treated electrospun nanofibrous scaffold. Artif. Cells Nanomed. Biotechnol. 2018, 46, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Enderami, S.E.; Soleimani, M.; Mortazavi, Y.; Nadri, S.; Salimi, A. Generation of insulin-producing cells from human adipose-derived mesenchymal stem cells on PVA scaffold by optimized differentiation protocol. J. Cell. Physiol. 2018, 233, 4327–4337. [Google Scholar] [CrossRef]
- Khorsandi, L.; Nejad-Dehbashi, F.; Ahangarpour, A.; Hashemitabar, M. Three-dimensional differentiation of bone marrow-derived mesenchymal stem cells into insulin-producing cells. Tissue Cell 2015, 47, 66–72. [Google Scholar] [CrossRef]
- Khorsandi, L.; Khodadadi, A.; Nejad-Dehbashi, F.; Saremy, S. Three-dimensional differentiation of adipose-derived mesenchymal stem cells into insulin-producing cells. Cell Tissue Res. 2015, 361, 745–753. [Google Scholar] [CrossRef]
- Ahn, E.H.; Kim, Y.; Kshitiz; An, S.S.; Afzal, J.; Lee, S.; Kwak, M.; Suh, K.-Y.; Kim, D.-H.; Levchenko, A. Spatial control of adult stem cell fate using nanotopographic cues. Biomaterials 2014, 35, 2401–2410. [Google Scholar] [CrossRef]
- Silva, G.A.; Czeisler, C.; Niece, K.L.; Beniash, E.; Harrington, D.A.; Kessler, J.A.; Stupp, S.I. Selective differentiation of neural progenitor cells by high-epitope density nanofibers. Science 2004, 303, 1352–1355. [Google Scholar] [CrossRef] [PubMed]
- Gurkan, U.A.; El Assal, R.; Yildiz, S.E.; Sung, Y.; Trachtenberg, A.J.; Kuo, W.P.; Demirci, U. Engineering Anisotropic Biomimetic Fibrocartilage Microenvironment by Bioprinting Mesenchymal Stem Cells in Nanoliter Gel Droplets. Mol. Pharm. 2014, 11, 2151–2159. [Google Scholar] [CrossRef] [PubMed]
- Sieber, S.; Wirth, L.; Cavak, N.; Koenigsmark, M.; Marx, U.; Lauster, R.; Rosowski, M. Bone marrow-on-a-chip: Long-term culture of human haematopoietic stem cells in a three-dimensional microfluidic environment. J. Tissue Eng. Regen. Med. 2018, 12, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Occhetta, P.; Centola, M.; Tonnarelli, B.; Redaelli, A.; Martin, I.; Rasponi, M. High-Throughput Microfluidic Platform for 3D Cultures of Mesenchymal Stem Cells, Towards Engineering Developmental Processes. Sci. Rep. 2015, 5, 10288. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Park, H.-J.; Han, S.; Lee, J.; Ko, E.; Kim, J.; Lee, J.S.; Yu, J.H.; Song, K.Y.; Cheong, E.; et al. Recapitulation of in vivo-like paracrine signals of human mesenchymal stem cells for functional neuronal differentiation of human neural stem cells in a 3D microfluidic system. Biomaterials 2015, 63, 177–188. [Google Scholar] [CrossRef] [PubMed]
- Vorwald, C.E.; Joshee, S.; Leach, J.K. Spatial localization of endothelial cells in heterotypic spheroids influences Notch signaling. J. Mol. Med. 2020, 98, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Badiavas, E.V.; Abedi, M.; Butmarc, J.; Falanga, V.; Quesenberry, P. Participation of bone marrow derived cells in cutaneous wound healing. J. Cell. Physiol. 2003, 196, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Selimović, Š.; Dokmeci, M.R.; Khademhosseini, A. Organs-on-a-chip for drug discovery. Curr. Opin. Pharmacol. 2013, 13, 829–833. [Google Scholar] [CrossRef] [PubMed]
- Nikolakopoulou, P.; Rauti, R.; Voulgaris, D.; Shlomy, I.; Maoz, M.B.; Herland, A. Recent progress in translational engineered in vitro models of the central nervous system. Brain 2020. [Google Scholar] [CrossRef]
- Sim, W.Y.; Park, S.W.; Park, S.H.; Min, B.H.; Park, S.R.; Yang, S.S. A pneumatic micro cell chip for the differentiation of human mesenchymal stem cells under mechanical stimulation. Lab Chip 2007, 7, 1775. [Google Scholar] [CrossRef]
- Yang, X.; Xu, X.; Zhang, Y.; Wen, W.; Gao, X. 3D Microstructure Inhibits Mesenchymal Stem Cells Homing to the Site of Liver Cancer Cells on a Microchip. Genes 2017, 8, 218. [Google Scholar] [CrossRef]
- Zheng, W.; Xie, Y.; Zhang, W.; Wang, D.; Ma, W.; Wang, Z.; Jiang, X. Fluid flow stress induced contraction and re-spread of mesenchymal stem cells: A microfluidic study. Integr. Biol. 2012, 4, 1102. [Google Scholar] [CrossRef]
- Jeon, J.S.; Bersini, S.; Whisler, J.A.; Chen, M.B.; Dubini, G.; Charest, J.L.; Moretti, M.; Kamm, R.D. Generation of 3D functional microvascular networks with human mesenchymal stem cells in microfluidic systems. Integr. Biol. 2014, 6, 555–563. [Google Scholar] [CrossRef]
- Nelson, M.R.; Ghoshal, D.; Mejías, J.C.; Rubio, D.F.; Keith, E.; Roy, K. A Multi-Niche Microvascularized Human Bone-Marrow-on-a-Chip. bioRxiv 2019. [Google Scholar] [CrossRef]
- Marturano-Kruik, A.; Nava, M.M.; Yeager, K.; Chramiec, A.; Hao, L.; Robinson, S.; Guo, E.; Raimondi, M.T.; Vunjak-Novakovic, G. Human bone perivascular niche-on-a-chip for studying metastatic colonization. PNAS 2018, 115, 1256–1261. [Google Scholar] [CrossRef] [PubMed]
- Leisten, I.; Kramann, R.; Ventura Ferreira, M.S.; Bovi, M.; Neuss, S.; Ziegler, P.; Wagner, W.; Knüchel, R.; Schneider, R.K. 3D co-culture of hematopoietic stem and progenitor cells and mesenchymal stem cells in collagen scaffolds as a model of the hematopoietic niche. Biomaterials 2012, 33, 1736–1747. [Google Scholar] [CrossRef]
- Ventura Ferreira, M.S.; Jahnen-Dechent, W.; Labude, N.; Bovi, M.; Hieronymus, T.; Zenke, M.; Schneider, R.K.; Neurs, S. Cord blood-hematopoietic stem cell expansion in 3D fibrin scaffolds with stromal support. Biomaterials 2012, 33, 6987–6997. [Google Scholar] [CrossRef] [PubMed]
- Raic, A.; Rödling, L.; Kalbacher, H.; Lee-Thedieck, C. Biomimetic macroporous PEG hydrogels as 3D scaffolds for the multiplication of human hematopoietic stem and progenitor cells. Biomaterials 2014, 35, 929–940. [Google Scholar] [CrossRef]
- Dhami, S.P.S.; Kappala, S.S.; Thompson, A.; Szegezdi, E. Three-dimensional ex vivo co-culture models of the leukaemic bone marrow niche for functional drug testing. Drug Discov. Today 2016, 21, 1464–1471. [Google Scholar] [CrossRef]
- Ham, J.; Lever, L.; Fox, M.; Reagan, M.R. In Vitro 3D Cultures to Reproduce the Bone Marrow Niche. JBMR Plus 2019, 3. [Google Scholar] [CrossRef]
- Amadori, S.; Torricelli, P.; Panzavolta, S.; Parrilli, A.; Fini, M.; Bigi, A. Multi-Layered Scaffolds for Osteochondral Tissue Engineering: In Vitro Response of Co-Cultured Human Mesenchymal Stem Cells. Macromol. Biosci. 2015, 15, 1535–1545. [Google Scholar] [CrossRef]
- Çakmak, S.; Çakmak, A.S.; Kaplan, D.L.; Gümüşderelioğlu, M. A Silk Fibroin and Peptide Amphiphile-Based Co-Culture Model for Osteochondral Tissue Engineering. Macromol. Biosci. 2016, 16, 1212–1226. [Google Scholar] [CrossRef]
- Zhou, J.; Xu, C.; Wu, G.; Cao, X.; Zhang, L.; Zhai, Z.; Zheng, Z.; Chen, X.; Wang, Y. In vitro generation of osteochondral differentiation of human marrow mesenchymal stem cells in novel collagen–hydroxyapatite layered scaffolds. Acta Biomater. 2011, 7, 3999–4006. [Google Scholar] [CrossRef]
- Lozito, T.P.; Alexander, P.G.; Lin, H.; Gottardi, R.; Cheng, A.W.-M.; Tuan, R.S. Three-dimensional osteochondral microtissue to model pathogenesis of osteoarthritis. Stem Cell Res. 2013, 4, S6. [Google Scholar] [CrossRef] [PubMed]
- Moore, C.A.; Shah, N.N.; Smith, C.P.; Rameshwar, P. 3D Bioprinting and Stem Cells. In Somatic Stem Cells: Methods and Protocols; Singh, S.R., Rameshwar, P., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2018; pp. 93–103. ISBN 978-1-4939-8697-2. [Google Scholar]
- Jang, E.H.; Kim, J.-H.; Lee, J.H.; Kim, D.-H.; Youn, Y.-N. Enhanced Biocompatibility of Multi-Layered, 3D Bio-Printed Artificial Vessels Composed of Autologous Mesenchymal Stem Cells. Polymers 2020, 12, 538. [Google Scholar] [CrossRef]
- Hung, S.-C.; Pochampally, R.R.; Chen, S.-C.; Hsu, S.-C.; Prockop, D.J. Angiogenic Effects of Human Multipotent Stromal Cell Conditioned Medium Activate the PI3K-Akt Pathway in Hypoxic Endothelial Cells to Inhibit Apoptosis, Increase Survival, and Stimulate Angiogenesis. Stem Cells 2007, 25, 2363–2370. [Google Scholar] [CrossRef] [PubMed]
- Bourget, J.-M.; Kérourédan, O.; Medina, M.; Rémy, M.; Thébaud, N.B.; Bareille, R.; Chassande, O.; Amédée, J.; Catros, S.; Devillard, R. Patterning of Endothelial Cells and Mesenchymal Stem Cells by Laser-Assisted Bioprinting to Study Cell Migration. Available online: https://www.hindawi.com/journals/bmri/2016/3569843/ (accessed on 14 May 2020).
- Santoro, M.; Awosika, T.O.; Snodderly, K.L.; Hurley-Novatny, A.C.; Lerman, M.J.; Fisher, J.P. Endothelial/Mesenchymal Stem Cell Crosstalk Within Bioprinted Cocultures. Tissue Eng. Part A 2019, 26, 339–349. [Google Scholar] [CrossRef] [PubMed]
- Figueroa, R.J.; Koch, T.G.; Betts, D.H. Osteogenic differentiation of equine cord blood multipotent mesenchymal stromal cells within coralline hydroxyapatite scaffolds in vitro. Vet. Comp. Orthop. Traumatol. 2011, 24, 354–362. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Langenbach, F.; Berr, K.; Naujoks, C.; Hassel, A.; Hentschel, M.; Depprich, R.; Kubler, N.R.; Meyer, U.; Wiesmann, H.-P.; Kögler, G.; et al. Generation and differentiation of microtissues from multipotent precursor cells for use in tissue engineering. Nat. Protoc. 2011, 6, 1726–1735. [Google Scholar] [CrossRef]
- Wang, W.; Itaka, K.; Ohba, S.; Nishiyama, N.; Chung, U.; Yamasaki, Y.; Kataoka, K. 3D spheroid culture system on micropatterned substrates for improved differentiation efficiency of multipotent mesenchymal stem cells. Biomaterials 2009, 30, 2705–2715. [Google Scholar] [CrossRef]
- Cheng, N.-C.; Wang, S.; Young, T.-H. The influence of spheroid formation of human adipose-derived stem cells on chitosan films on stemness and differentiation capabilities. Biomaterials 2012, 33, 1748–1758. [Google Scholar] [CrossRef]
- Guo, L.; Zhou, Y.; Wang, S.; Wu, Y. Epigenetic changes of mesenchymal stem cells in three-dimensional (3D) spheroids. J. Cell. Mol. Med. 2014, 18, 2009–2019. [Google Scholar] [CrossRef]
- Bartosh, T.J.; Ylöstalo, J.H.; Mohammadipoor, A.; Bazhanov, N.; Coble, K.; Claypool, K.; Lee, R.H.; Choi, H.; Prockop, D.J. Aggregation of human mesenchymal stromal cells (MSCs) into 3D spheroids enhances their antiinflammatory properties. Proc. Natl. Acad. Sci. USA 2010, 107, 13724–13729. [Google Scholar] [CrossRef]
- Saleh, F.A.; Whyte, M.; Genever, P. Effects of endothelial cells on human mesenchymal stem cell activity in a three-dimensional in vitro model. Eur. Cells Mater. 2011. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.; Lee, H.; Park, Y.H.; Jeon, E.; Chung, H.K.; Lee, E.S.; Shim, J.H.; Kang, K.-T. Three-dimensional Angiogenesis Assay System using Co-culture Spheroids Formed by Endothelial Colony Forming Cells and Mesenchymal Stem Cells. JoVE (J. Vis. Exp.) 2019, e60032. [Google Scholar] [CrossRef]
- Shah, S.; Kang, K.-T. Two-Cell Spheroid Angiogenesis Assay System Using Both Endothelial Colony Forming Cells and Mesenchymal Stem Cells. Biomol. Ther. (Seoul) 2018, 26, 474–480. [Google Scholar] [CrossRef]
- Bhang, S.H.; Cho, S.-W.; La, W.-G.; Lee, T.-J.; Yang, H.S.; Sun, A.-Y.; Baek, S.-H.; Rhie, J.-W.; Kim, B.-S. Angiogenesis in ischemic tissue produced by spheroid grafting of human adipose-derived stromal cells. Biomaterials 2011, 32, 2734–2747. [Google Scholar] [CrossRef] [PubMed]
- Bhang, S.H.; Lee, S.; Shin, J.-Y.; Lee, T.-J.; Kim, B.-S. Transplantation of Cord Blood Mesenchymal Stem Cells as Spheroids Enhances Vascularization. Tissue Eng. Part A 2012, 18, 2138–2147. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Han, Y.-S.; Lee, S.H. Long-Duration Three-Dimensional Spheroid Culture Promotes Angiogenic Activities of Adipose-Derived Mesenchymal Stem Cells. B Biomol. Ther. (Seoul) 2016, 24, 260–267. [Google Scholar] [CrossRef]
- Song, L.; Yuan, X.; Jones, Z.; Griffin, K.; Zhou, Y.; Ma, T.; Li, Y. Assembly of Human Stem Cell-Derived Cortical Spheroids and Vascular Spheroids to Model 3-D Brain-like Tissues. Sci. Rep. 2019, 9, 5977. [Google Scholar] [CrossRef]
- Varzideh, F.; Pahlavan, S.; Ansari, H.; Halvaei, M.; Kostin, S.; Feiz, M.-S.; Latifi, H.; Aghdami, N.; Braun, T.; Baharvand, H. Human cardiomyocytes undergo enhanced maturation in embryonic stem cell-derived organoid transplants. Biomaterials 2019, 192, 537–550. [Google Scholar] [CrossRef]
- Leeman, K.T.; Pessina, P.; Lee, J.-H.; Kim, C.F. Mesenchymal Stem Cells Increase Alveolar Differentiation in Lung Progenitor Organoid Cultures. Sci. Rep. 2019, 9, 6479. [Google Scholar] [CrossRef]
- Devarasetty, M.; Wang, E.; Soker, S.; Skardal, A. Mesenchymal stem cells support growth and organization of host-liver colorectal-tumor organoids and possibly resistance to chemotherapy. Biofabrication 2017, 9, 021002. [Google Scholar] [CrossRef]
- Mark, D.; Haeberle, S.; Roth, G.; von Stetten, F.; Zengerle, R. Microfluidic lab-on-a-chip platforms: Requirements, characteristics and applications. Chem. Soc. Rev. 2010, 39, 1153. [Google Scholar] [CrossRef]
- Chován, T.; Guttman, A. Microfabricated devices in biotechnology and biochemical processing. Trends Biotechnol. 2002, 20, 116–122. [Google Scholar] [CrossRef]
- Bhatia, S.N.; Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 2014, 32, 760–772. [Google Scholar] [CrossRef]
- Ahadian, S.; Civitarese, R.; Bannerman, D.; Mohammadi, M.H.; Lu, R.; Wang, E.; Davenport-Huyer, L.; Lai, B.; Zhang, B.; Zhao, Y.; et al. Organ-On-A-Chip Platforms: A Convergence of Advanced Materials, Cells, and Microscale Technologies. Adv. Healthc. Mater. 2018, 7, 1700506. [Google Scholar] [CrossRef]
- Benam, K.H.; Novak, R.; Nawroth, J.; Hirano-Kobayashi, M.; Ferrante, T.C.; Choe, Y.; Prantil-Baun, R.; Weaver, J.C.; Bahinski, A.; Parker, K.K.; et al. Matched-Comparative Modeling of Normal and Diseased Human Airway Responses Using a Microengineered Breathing Lung Chip. Cell Syst. 2016, 3, 456–466.e4. [Google Scholar] [CrossRef]
- Jang, K.-J.; Otieno, M.A.; Ronxhi, J.; Lim, H.-K.; Ewart, L.; Kodella, K.R.; Petropolis, D.B.; Kulkarni, G.; Rubins, J.E.; Conegliano, D.; et al. Reproducing human and cross-species drug toxicities using a Liver-Chip. Sci. Transl. Med. 2019, 11, 517. [Google Scholar] [CrossRef]
- Lee, H.; Cho, D.-W. One-step fabrication of an organ-on-a-chip with spatial heterogeneity using a 3D bioprinting technology. Lab Chip 2016, 16, 2618–2625. [Google Scholar] [CrossRef]
- Huh, D.; Matthews, B.D.; Mammoto, A.; Montoya-Zavala, M.; Hsin, H.Y.; Ingber, D.E. Reconstituting Organ-Level Lung Functions on a Chip. Science 2010, 328, 1662–1668. [Google Scholar] [CrossRef]
- Kimura, H.; Sakai, Y.; Fujii, T. Organ/body-on-a-chip based on microfluidic technology for drug discovery. Drug Metab. Pharmacokinet. 2018, 33, 43–48. [Google Scholar] [CrossRef]
- Powers, M.J.; Domansky, K.; Kaazempur-Mofrad, M.R.; Kalezi, A.; Capitano, A.; Upadhyaya, A.; Kurzawski, P.; Wack, K.E.; Stolz, D.B.; Kamm, R.; et al. A microfabricated array bioreactor for perfused 3D liver culture. Biotechnol. Bioeng. 2002, 78, 257–269. [Google Scholar] [CrossRef]
- Kimura, H.; Yamamoto, T.; Sakai, H.; Sakai, Y.; Fujii, T. An integrated microfluidic system for long-term perfusion culture and on-line monitoring of intestinal tissue models. Lab Chip 2008, 8, 741. [Google Scholar] [CrossRef]
- Jang, K.-J.; Suh, K.-Y. A multi-layer microfluidic device for efficient culture and analysis of renal tubular cells. Lab Chip 2010, 10, 36–42. [Google Scholar] [CrossRef]
- Wu, Q.; Liu, J.; Wang, X.; Feng, L.; Wu, J.; Zhu, X.; Wen, W.; Gong, X. Organ-on-a-chip: Recent breakthroughs and future prospects. Biomed. Eng. Online 2020, 19, 9. [Google Scholar] [CrossRef]
- Moya, A. Online oxygen monitoring using integrated inkjet-printed sensors in a liver-on-a-chip system. Lab Chip 2018, 18, 2023–2035. [Google Scholar] [CrossRef]
- Herland, A.; Maoz, B.M.; Das, D.; Somayaji, M.R.; Prantil-Baun, R.; Novak, R.; Cronce, M.; Huffstater, T.; Jeanty, S.S.F.; Ingram, M.; et al. Quantitative prediction of human pharmacokinetic responses to drugs via fluidically coupled vascularized organ chips. Nat. Biomed. Eng. 2020, 4, 421–436. [Google Scholar] [CrossRef]
- Lee, S.H.; Sung, J.H. Organ-on-a-Chip Technology for Reproducing Multiorgan Physiology. Adv. Healthc. Mater. 2018, 7, 1700419. [Google Scholar] [CrossRef]
- Maoz, B.M.; Herland, A.; FitzGerald, E.A.; Grevesse, T.; Vidoudez, C.; Pacheco, A.R.; Sheehy, S.P.; Park, T.-E.; Dauth, S.; Mannix, R.; et al. A linked organ-on-chip model of the human neurovascular unit reveals the metabolic coupling of endothelial and neuronal cells. Nat. Biotechnol. 2018, 36, 865–874. [Google Scholar] [CrossRef]
- Si, L.; Bai, H.; Rodas, M.; Cao, W.; Oh, C.Y.; Jiang, A.; Nurani, A.; Zhu, D.Y.; Goyal, G.; Gilpin, S.E.; et al. Human organs-on-chips as tools for repurposing approved drugs as potential influenza and COVID19 therapeutics in viral pandemics. bioRxiv 2020. [Google Scholar] [CrossRef]
- Raimondi, M.T.; Albani, D.; Giordano, C. An Organ-On-A-Chip Engineered Platform to Study the Microbiota–Gut–Brain Axis in Neurodegeneration. Trends Mol. Med. 2019, 25, 737–740. [Google Scholar] [CrossRef]
- Kang, Y.B.A.; Sodunke, T.R.; Lamontagne, J.; Cirillo, J.; Rajiv, C.; Bouchard, M.J.; Noh, M. Liver sinusoid on a chip: Long-term layered co-culture of primary rat hepatocytes and endothelial cells in microfluidic platforms: Liver Sinusoid on a Chip. Biotechnol. Bioeng. 2015, 112, 2571–2582. [Google Scholar] [CrossRef]
- Zhou, Q.; Patel, D.; Kwa, T.; Haque, A.; Matharu, Z.; Stybayeva, G.; Gao, Y.; Diehl, A.M.; Revzin, A. Liver injury-on-a-chip: Microfluidic co-cultures with integrated biosensors for monitoring liver cell signaling during injury. Lab Chip 2015, 15, 4467–4478. [Google Scholar] [CrossRef]
- Benam, K.H.; Villenave, R.; Lucchesi, C.; Varone, A.; Hubeau, C.; Lee, H.-H.; Alves, S.E.; Salmon, M.; Ferrante, T.C.; Weaver, J.C.; et al. Small airway-on-a-chip enables analysis of human lung inflammation and drug responses in vitro. Nat. Methods 2016, 13, 151–157. [Google Scholar] [CrossRef]
- Marsano, A.; Conficconi, C.; Lemme, M.; Occhetta, P.; Gaudiello, E.; Votta, E.; Cerino, G.; Redaelli, A.; Rasponi, M. Beating heart on a chip: A novel microfluidic platform to generate functional 3D cardiac microtissues. Lab Chip 2016, 16, 599–610. [Google Scholar] [CrossRef]
- Haddrick, M.; Simpson, P.B. Organ-on-a-chip technology: Turning its potential for clinical benefit into reality. Drug Discov. Today 2019, 24, 1217–1223. [Google Scholar] [CrossRef] [PubMed]
- Karimi, M.; Bahrami, S.; Mirshekari, H.; Basri, S.M.M.; Nik, A.B.; Aref, A.R.; Akbari, M.; Hamblin, M.R. Microfluidic systems for stem cell-based neural tissue engineering. Lab Chip 2016, 16, 2551–2571. [Google Scholar] [CrossRef]
- Wiesmann, A.; Bühring, H.-J.; Mentrup, C.; Wiesmann, H.-P. Decreased CD90 expression in human mesenchymal stem cells by applying mechanical stimulation. Head Face Med. 2006, 2, 8. [Google Scholar] [CrossRef]
- Guilak, F.; Cohen, D.M.; Estes, B.T.; Gimble, J.M.; Liedtke, W.; Chen, C.S. Control of Stem Cell Fate by Physical Interactions with the Extracellular Matrix. Cell Stem Cell 2009, 5, 17–26. [Google Scholar] [CrossRef]
- Gaebel, R.; Ma, N.; Liu, J.; Guan, J.; Koch, L.; Klopsch, C.; Gruene, M.; Toelk, A.; Wang, W.; Mark, P.; et al. Patterning human stem cells and endothelial cells with laser printing for cardiac regeneration. Biomaterials 2011, 32, 9218–9230. [Google Scholar] [CrossRef]
- Jain, R.K. Molecular regulation of vessel maturation. Nat. Med. 2003, 9, 685–693. [Google Scholar] [CrossRef]
- Velve-Casquillas, G.; Le Berre, M.; Piel, M.; Tran, P.T. Microfluidic tools for cell biological research. Nano Today 2010, 5, 28–47. [Google Scholar] [CrossRef]
- Sung, J.H.; Kam, C.; Shuler, M.L. A microfluidic device for a pharmacokinetic–pharmacodynamic (PK–PD) model on a chip. Lab Chip 2010, 10, 446. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.-Y.; Dai, H.; Leong, K.W. Inducing hepatic differentiation of human mesenchymal stem cells in pellet culture. Biomaterials 2006, 27, 4087–4097. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Su, P.; Xu, C.; Yang, J.; Yu, W.; Huang, D. Chondrogenic differentiation of human mesenchymal stem cells: A comparison between micromass and pellet culture systems. Biotechnol. Lett. 2010, 32, 1339–1346. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Hudalla, G.A. Using Self-Assembling Peptides to Integrate Biomolecules into Functional Supramolecular Biomaterials. Molecules 2019, 24, 1450. [Google Scholar] [CrossRef]
- O’Brien, F.J. Biomaterials & scaffolds for tissue engineering. Mater. Today 2011, 14, 88–95. [Google Scholar] [CrossRef]
- Pina, S.; Ribeiro, V.P.; Marques, C.F.; Maia, F.R.; Silva, T.H.; Reis, R.L.; Oliveira, J.M. Scaffolding Strategies for Tissue Engineering and Regenerative Medicine Applications. Materials (Basel) 2019, 12, 1824. [Google Scholar] [CrossRef]
- Zhao, P.; Gu, H.; Mi, H.; Rao, C.; Fu, J.; Turng, L. Fabrication of scaffolds in tissue engineering: A review. Front. Mech. Eng. 2018, 13, 107–119. [Google Scholar] [CrossRef]
- Nichols, J.E.; Cortiella, J.; Lee, J.; Niles, J.A.; Cuddihy, M.; Wang, S.; Bielitzki, J.; Cantu, A.; Mlcak, R.; Valdivia, E.; et al. In vitro analog of human bone marrow from 3D scaffolds with biomimetic inverted colloidal crystal geometry. Biomaterials 2009, 30, 1071–1079. [Google Scholar] [CrossRef]
- Di Maggio, N.; Piccinini, E.; Jaworski, M.; Trumpp, A.; Wendt, D.J.; Martin, I. Toward modeling the bone marrow niche using scaffold-based 3D culture systems. Biomaterials 2011, 32, 321–329. [Google Scholar] [CrossRef]
- Derakhshanfar, S.; Mbeleck, R.; Xu, K.; Zhang, X.; Zhong, W.; Xing, M. 3D bioprinting for biomedical devices and tissue engineering: A review of recent trends and advances. Bioact. Mater. 2018, 3, 144–156. [Google Scholar] [CrossRef]
- Ong, C.S.; Yesantharao, P.; Huang, C.Y.; Mattson, G.; Boktor, J.; Fukunishi, T.; Zhang, H.; Hibino, N. 3D bioprinting using stem cells. Pediatric Res. 2018, 83, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Rider, P.; Kačarević, Ž.P.; Alkildani, S.; Retnasingh, S.; Barbeck, M. Bioprinting of tissue engineering scaffolds. J. Tissue Eng. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Fennema, E.; Rivron, N.; Rouwkema, J.; van Blitterswijk, C.; de Boer, J. Spheroid culture as a tool for creating 3D complex tissues. Trends Biotechnol. 2013, 31, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.; Mead, B.E.; Safaee, H.; Langer, R.; Karp, J.M.; Levy, O. Engineering Stem Cell Organoids. Cell Stem Cell 2016, 18, 25–38. [Google Scholar] [CrossRef]
- Ranga, A.; Gjorevski, N.; Lutolf, M.P. Drug discovery through stem cell-based organoid models. Adv. Drug Deliv. Rev. 2014. [Google Scholar] [CrossRef]
- Cui, X.; Hartanto, Y.; Zhang, H. Advances in multicellular spheroids formation. J. R. Soc. Interface 2017, 14. [Google Scholar] [CrossRef]
- Grebenyuk, S.; Ranga, A. Engineering Organoid Vascularization. Front. Bioeng. Biotechnol. 2019, 7. [Google Scholar] [CrossRef]
- Hirschhaeuser, F.; Menne, H.; Dittfeld, C.; West, J.; Mueller-Klieser, W.; Kunz-Schughart, L.A. Multicellular tumor spheroids: An underestimated tool is catching up again. J. Biotechnol. 2010, 148, 3–15. [Google Scholar] [CrossRef]
- Fatehullah, A.; Tan, S.H.; Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 2016, 18, 246–254. [Google Scholar] [CrossRef]
- Pașca, S.P. The rise of three-dimensional human brain cultures. Nature 2018, 553, 437–445. [Google Scholar] [CrossRef]
- Lee, Y.B.; Kim, E.M.; Byun, H.; Chang, H.; Jeong, K.; Aman, Z.M.; Choi, Y.S.; Park, J.; Shin, H. Engineering spheroids potentiating cell-cell and cell-ECM interactions by self-assembly of stem cell microlayer. Biomaterials 2018, 165, 105–120. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.-C.; Chen, C.-H.; Hwang, S.-M.; Lin, W.-W.; Huang, C.-H.; Lee, W.-Y.; Chang, Y.; Sung, H.-W. Spherically Symmetric Mesenchymal Stromal Cell Bodies Inherent with Endogenous Extracellular Matrices for Cellular Cardiomyoplasty. Stem Cells 2009, 27, 724–732. [Google Scholar] [CrossRef] [PubMed]
- Takebe, T.; Sekine, K.; Enomura, M.; Koike, H.; Kimura, M.; Ogaeri, T.; Zhang, R.-R.; Ueno, Y.; Zheng, Y.-W.; Koike, N.; et al. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 2013, 499, 481–484. [Google Scholar] [CrossRef] [PubMed]

| Cell Type | Source of MSCs | Key Differentiation Factors | Markers Expressed | Comments | Ref. |
|---|---|---|---|---|---|
| Chondrocytes | Bone marrow, Adipose tissue, Natal dental pulp, Placenta, Umbilical Cord, Wharton’s jelly | Transforming growth factor beta (TGF-β), insulin-like growth factor (IGF), Bone morphogenetic proteins (BMP), fibroblasts growth factor (FGF) families and galectines. | Type II collagen, Sox9, ACAN, Col2a1, β-catenin, GAG accumulation. | •Natural differentiation pathway. | [9,27,56,57] |
| Osteoblasts | Bone marrow, Adipose tissue, Natal dental pulp, Placenta, Umbilical Cord, Wharton jelly | Ascorbic acid, β-glycerol phosphate, dexamethasone/vitamin D3 and galectines. | Increase in alkaline phosphatase, calcium accumulation, RUNX2 | •Natural differentiation pathway. | [9,57,58,59,60,61,62,63,64,65] |
| Adipocytes | Bone marrow, Adipose tissue, Natal dental pulp, Placenta, Umbilical Cord, Wharton jelly | Dexamethasone, indomethacin, insulin and isobutylmethylxanthin. | PPARγ2, LPL, FABP2, FABP4 | •Natural differentiation pathway. | [9,59,64,65,66,67] |
| Hepatocytes | Bone marrow, Adipose tissue, Dental pulp, Placenta, Umbilical Cord, Wharton jelly | dexamethasone, FGFs, bone morphogenetic proteins, hepatocyte growth factor (HGF), epidermal growth factor, oncostatin M and insulin | CYP3A4, CYP1A1, CYP2C9, albumin, CK-18, CK-19 AAT, TAT, heppar antigen | •Characterization analysis of hepatocytes markers is not exclusive for them. Therefore, functional assays are needed to ensure successful differentiation. •Lack of standardization. | [28,50,68,69,70,71,72] |
| Cardiomyocytes | Bone marrow, Adipose tissue, Placenta, Umbilical Cord Blood, Wharton jelly | 5-Azacytidine (5-aza), miRNAs, IGF-1, insulin gene enhancer binding protein ISL-1, basic FGF (bFGF), TGF-β family, BMP-2, Caveolin-1, Vanilloid receptor 1 and Histone deacetylase 1. | ANP, cTnT, α-MHC, GATA4, Nkx2.5, CX43 | •5-aza considered as a carcinogen. •Some of the inducers have low differentiation efficiency. | [31,34,73,74] |
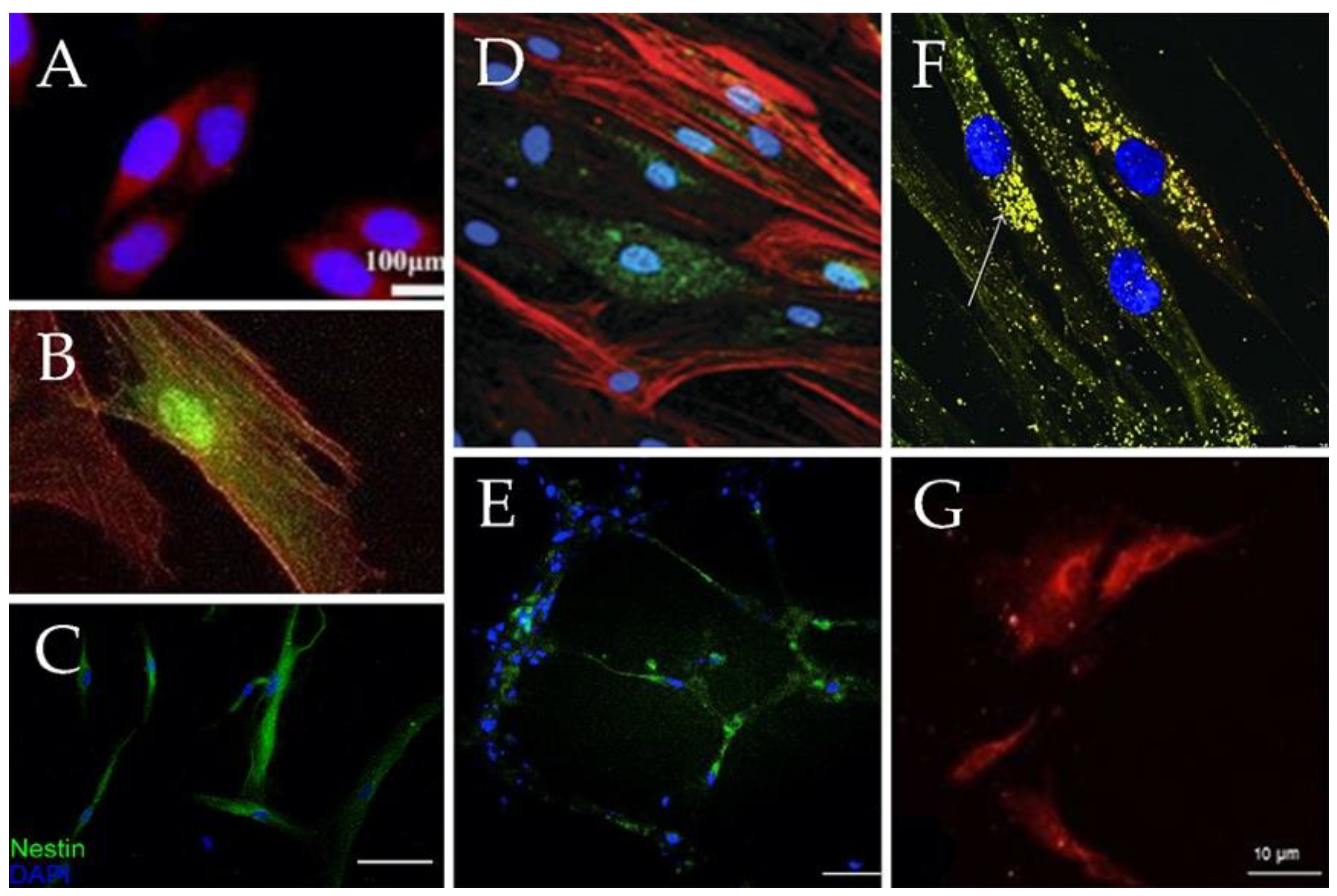
| Neurons | Bone marrow, Adipose tissue, Dental pulp, Placenta, Skin, Umbilical cord | bFGF, human epithelial growth factor (hEGF), Brain-derived neurotrophic factor (BDNF), all trans retinoic acid (RA). Can be also derived with neuronal progenitors’ exosomes. | Nestin, βIII tubulin, tyrosine hidroxilase, synapthophysin, NURR1, MAP2 | •Show robust neuronal electrical activity. •The neuronal functionality of those differentiated cells is controversial. | [35,37,51,64,75,76,77] |
| Epithelium | Bone marrow, Wharton jelly, Lung | hEGF, RA, keratinocyte growth factor, HGF and IGF-II. | CK-18, CK-19, occluding, CD9 | •Protocols based on co-cultures cause undesired cell-to-cell interactions. | [39,52,78] |
| Endothelial cells | Bone marrow | Vascular endothelial growth factor (VEGF), bFGF, IGF, EGF, ascorbic acid and heparin. | vWF, VE-cadherin, VEGFR-2 | •After differentiation, induced-MSCs successfully created vessel like structures. •More research is needed until applicate it in clinic. | [54] |
| Pancreatic β cells | Bone marrow, Adipose tissue, Urine, Dental, Pancreas, Wharton jelly, Placenta, Umbilical cord, Amniotic fluid | Nicotineamide, L-taurine, sodium butyrate, exedin and glucagon-like peptide-1. | Insulin, glucagon, Glut-2, PDX1, NKX6.1, NEUROD1, NGN3, MAFA | •Differentiated cells can even integrate to pancreatic tissue and become mature. •Less tumorigenic effect than other differentiated cells. | [45] |
| Epidermis | Bone marrow, Umbilical cord | EGF, FGF, Insulin, RA and CaCl2, Keratinocyte serum-free medium | P63, CK19, pan-cytokeratin, beta1-integrin, involucrin, KRT5, KRT10, KRT14 | •The cells might show epidermal-like morphology but they do not differentiate into keratinocytes. | [47,48,49] |
| Cell Type | Scaffold Type | Ref. |
|---|---|---|
| Chondrocytes | Silk fibroin (SF), SF/collagen, SF/chitosan/GAGs, PLGA, PLA | [80,81,84,86,88,89] |
| Osteoblasts | Silk fibroin/gelatin, SF/collagen, collagen, PLGA, ECM-based structures | [83,84,86,87,90] |
| Hepatocytes | PCL/collagen, collagen/with heparin | [92,93] |
| Neurons | Nano-grafts from PDMS (with RA), cellulosic hydrogels (with EGF, GFG, BDNF, RA), electrically conductive PLA with alginate, gelatin and carbon nanotubes | [37,38,94] |
| Epidermis | Collagen—as a gel (with EGF and vitamin D3) or as a spongy scaffold (with HaCaT cells) | [95,96] |
| Cardiomyocytes | HEMA hydrogels, PCL, collagen | [97,98,99,100,101,102] |
| Pancreatic β cells | PCL, PVA, fibrin glue, collagen/HA | [103,104,105,106] |
| Microfluidics and Organs-on-a-Chip | |
| Key features | Dynamic flow, mechanical and shear stresses, miniaturization, biosensors, organ-organ interaction, precise control of differentiation influences. |
| Limitations | More expensive than the regular 2D culture, unwanted absorption of materials, require expertise and special equipment, not all organs exist. |
| General use | Biotransformation, analytics, cellular assays, large-scale experiments in which many microfluidic units are combined (for high-throughput screening), OoC platforms, organ-organ interaction, observation of pharmacokinetic processes. |
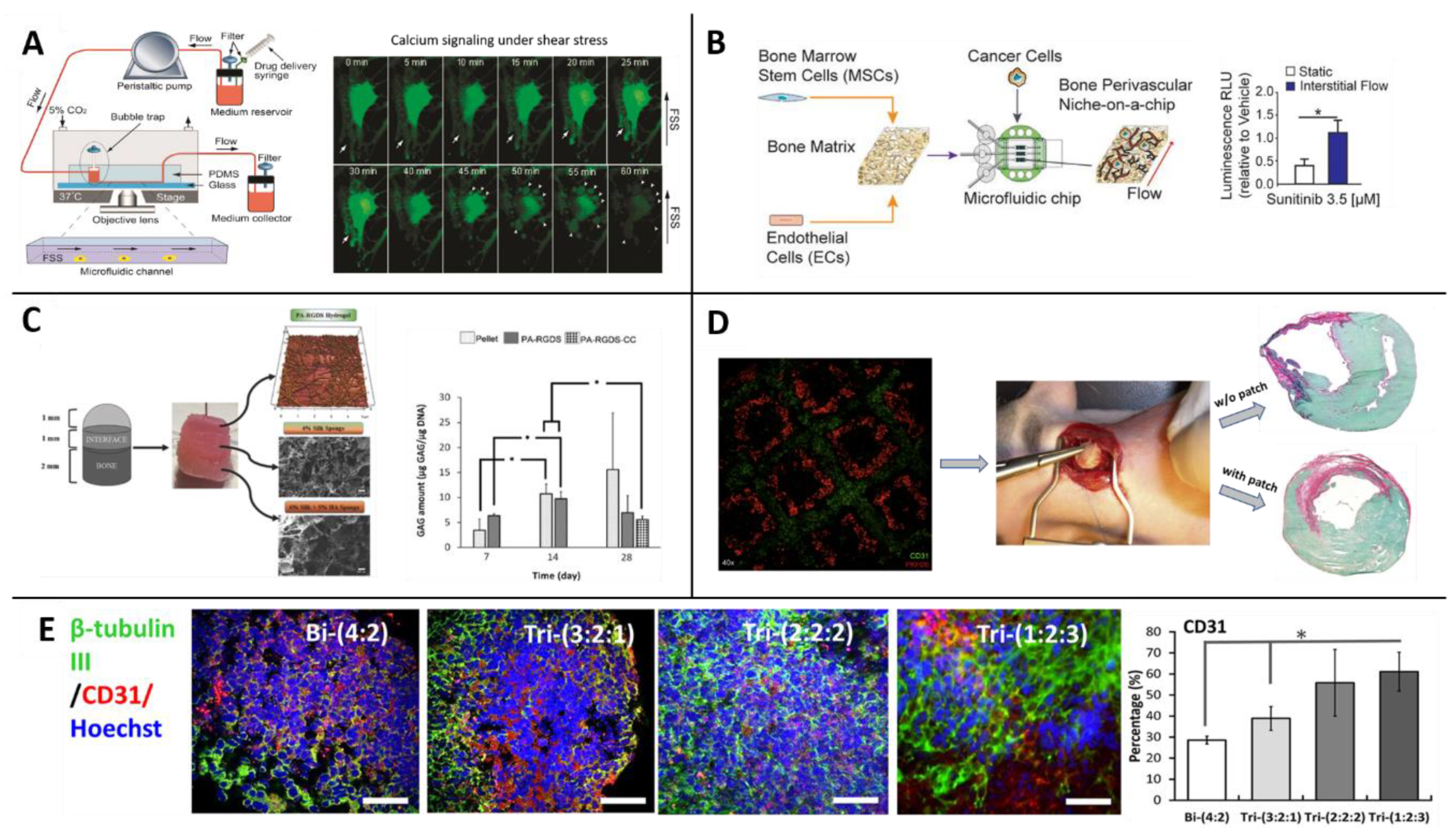
| Use with MSCs | Osteogenesis ([117]), interaction with liver cancer ([118]), vascularization ([119,120]), neuronal differentiation ([112]), bone regeneration ([111]), Bone Marrow-on-a-Chip ([110,121]), perivascular cancer metastasis ([122]). |
| Fabricated 3D structures—Scaffolds and Bioprinting | |
| Key features | 3D porous structures, biomechanical forces, ECM mimicking, printing with high spatial accuracy, biodegradability. |
| Limitations | Lack of vascularization and flow, conventional fabrication methods of scaffolds do not provide high accuracy in the microenvironment, large scale biomanufacturing issues, a challenge to incorporate multiple cell types. |
| General use | Enhancing differentiation, diverse complex geometric structures, regenerative medicine, tissue engineering. |
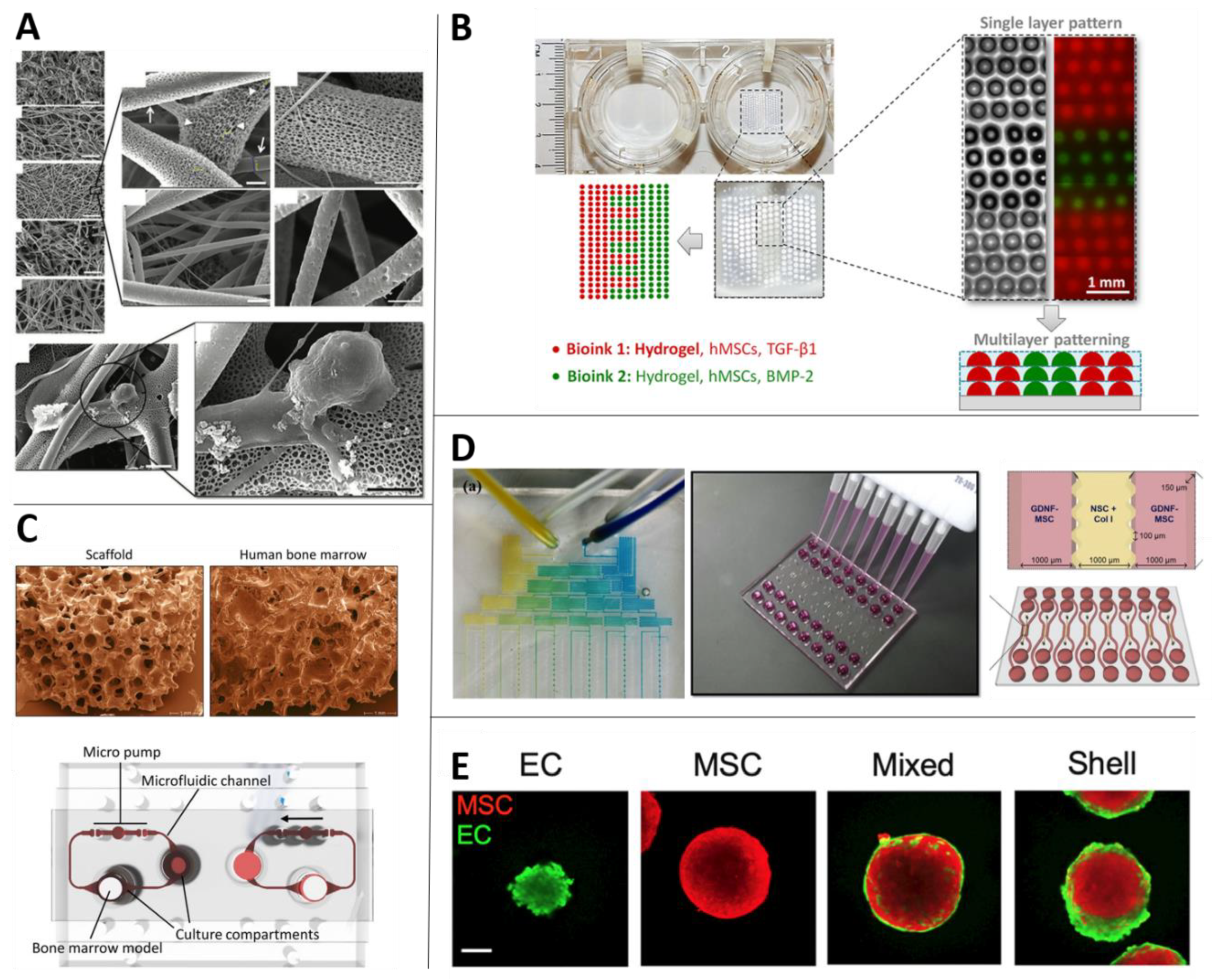
| Use with MSCs | Enhancing differentiation to various lineages (Section 2.4.2), hematopoietic niche in health ([123,124,125]) and disease ([126,127]), osteochondral niche in health ([128,129,130]) and disease ([131]), cancer metastasis in the bone marrow ([132]), multiphase cartilage tissue ([109]), bioprinting blood vessels for implantations ([133,134]) and research ([135,136]). |
| Spheroids and Organoids | |
| Key features | Self-assembly, self ECM fabrication, organoids are defined by functionality of the organ. |
| Limitations | Poor cell viability, long incubation times for some organoid models, not all organs are mimicked, limited functional readouts, usually lack of vascularization and stroma. |
| General use | Organogenesis, drug discovery, regenerative medicine. |
| Use with MSCs | Enhancing properties of differentiation ([137,138,139]), stemness ([140,141]) and immunomodulation ([142]), studying interactions with endothelial cells ([143,144,145,146,147,148]), stromal role in organoids of neurons ([149]), cardiomyocytes ([150]), lung epithelium ([151]) and liver colorectal tumor ([152]). |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Afflerbach, A.-K.; Kiri, M.D.; Detinis, T.; Maoz, B.M. Mesenchymal Stem Cells as a Promising Cell Source for Integration in Novel In Vitro Models. Biomolecules 2020, 10, 1306. https://doi.org/10.3390/biom10091306
Afflerbach A-K, Kiri MD, Detinis T, Maoz BM. Mesenchymal Stem Cells as a Promising Cell Source for Integration in Novel In Vitro Models. Biomolecules. 2020; 10(9):1306. https://doi.org/10.3390/biom10091306
Chicago/Turabian StyleAfflerbach, Ann-Kristin, Mark D. Kiri, Tahir Detinis, and Ben M. Maoz. 2020. "Mesenchymal Stem Cells as a Promising Cell Source for Integration in Novel In Vitro Models" Biomolecules 10, no. 9: 1306. https://doi.org/10.3390/biom10091306
APA StyleAfflerbach, A.-K., Kiri, M. D., Detinis, T., & Maoz, B. M. (2020). Mesenchymal Stem Cells as a Promising Cell Source for Integration in Novel In Vitro Models. Biomolecules, 10(9), 1306. https://doi.org/10.3390/biom10091306







