Transient Unfolding and Long-Range Interactions in Viral BCL2 M11 Enable Binding to the BECN1 BH3 Domain
Abstract
1. Introduction
2. Methods
2.1. System Preparation and Conformational Sampling
2.2. Anharmonic Conformational Analysis (ANCA)
2.3. Temporal Decorrelation of Fourth-Order on Time-Delayed Cumulant Matrices
2.4. Identifying Conformational Sub-States
2.5. Experimental Methods
3. Results
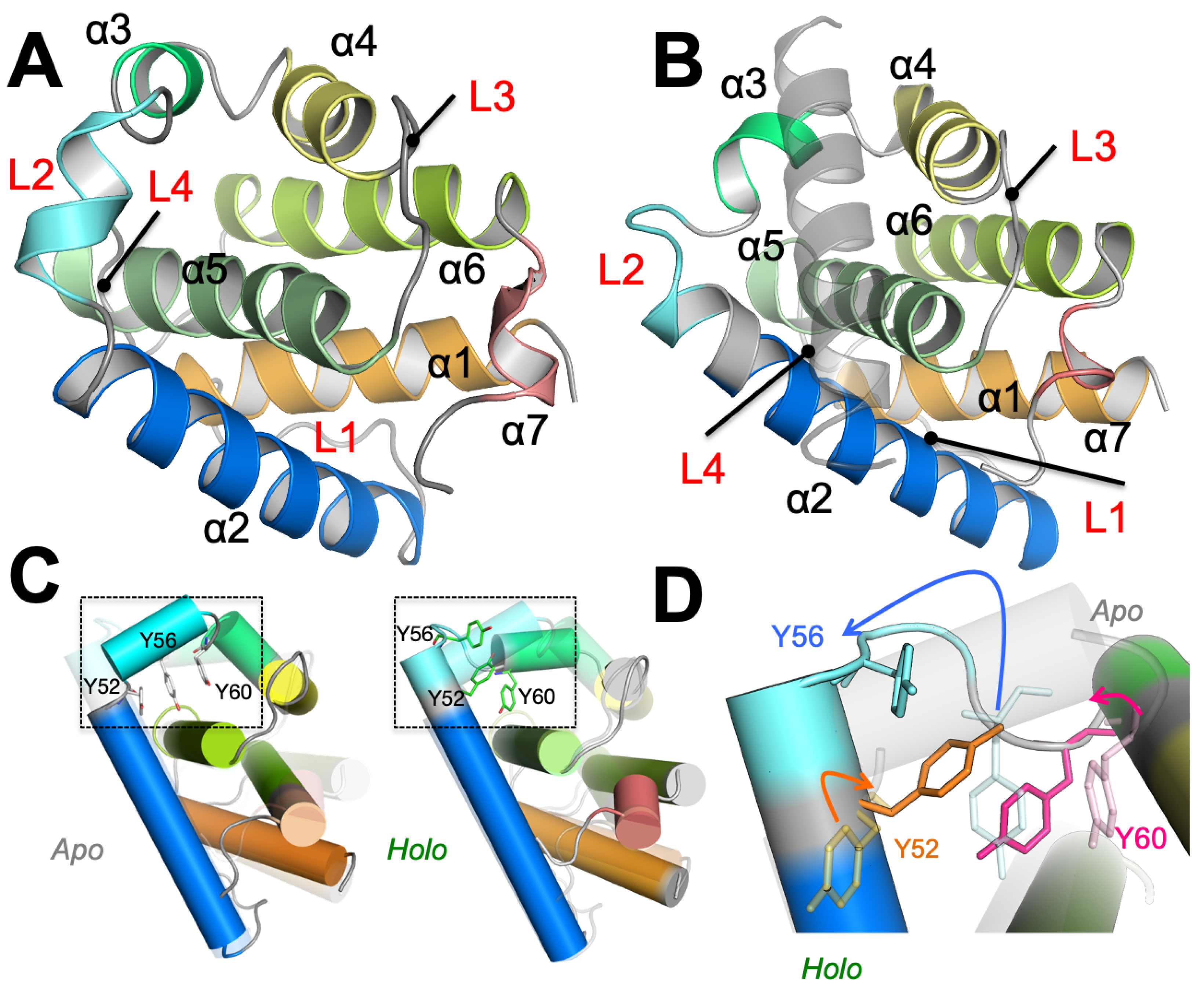
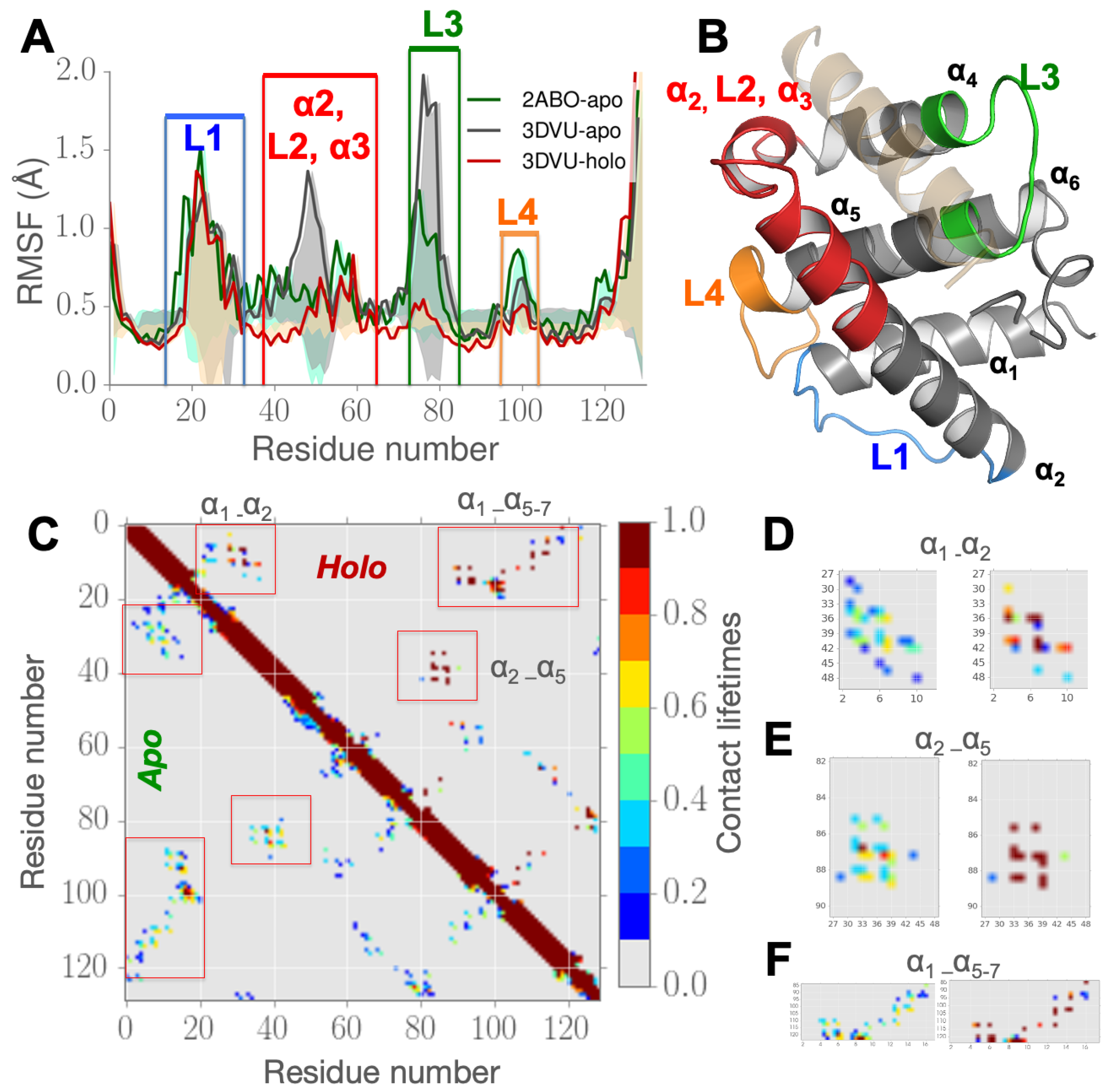
3.1. Partial Unfolding of M11- Facilitates Opening/Closing of BH3D Binding Site
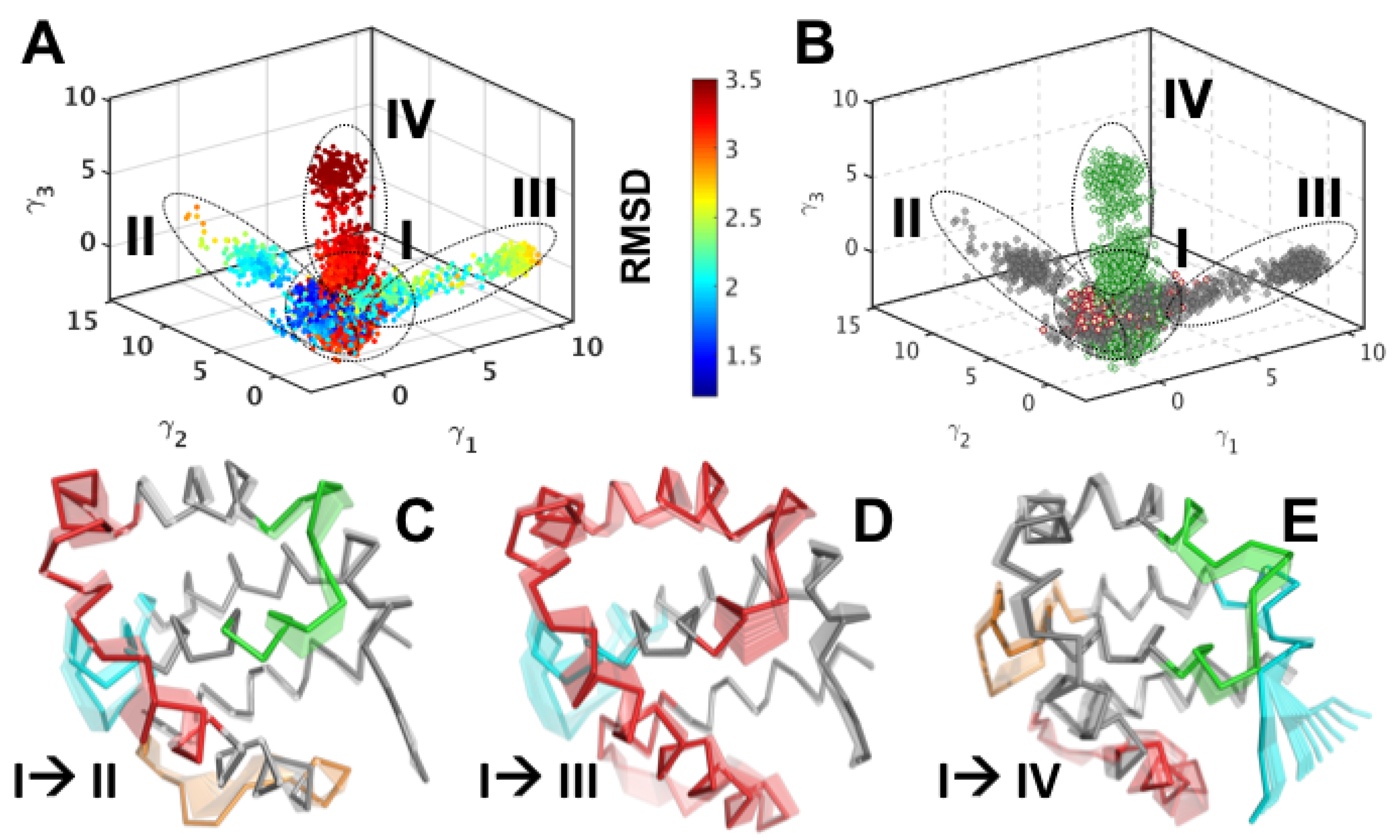
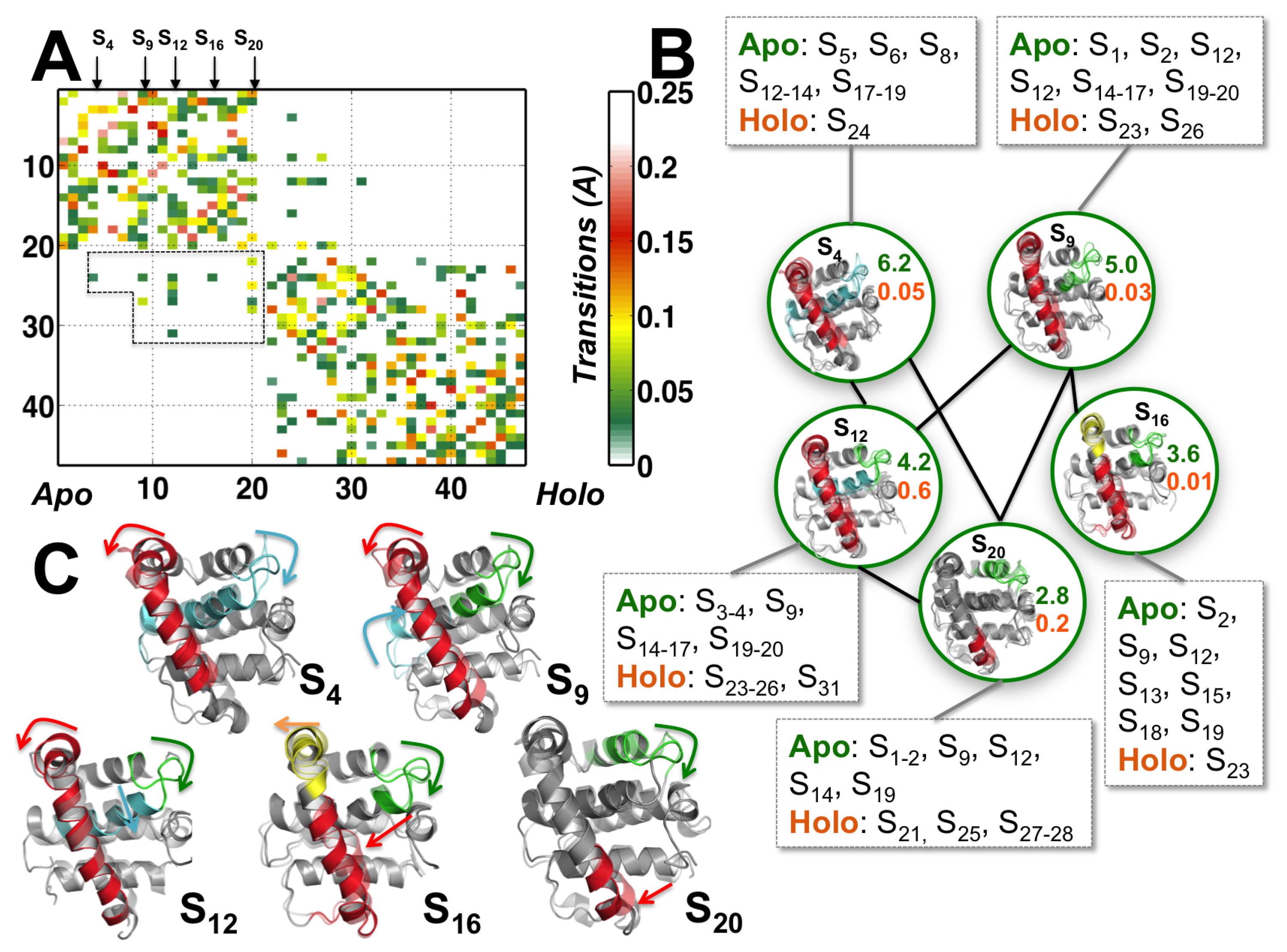
3.2. Structural Intermediates of the Apo- to Holo-M11 Show Transient Unfolding of 2
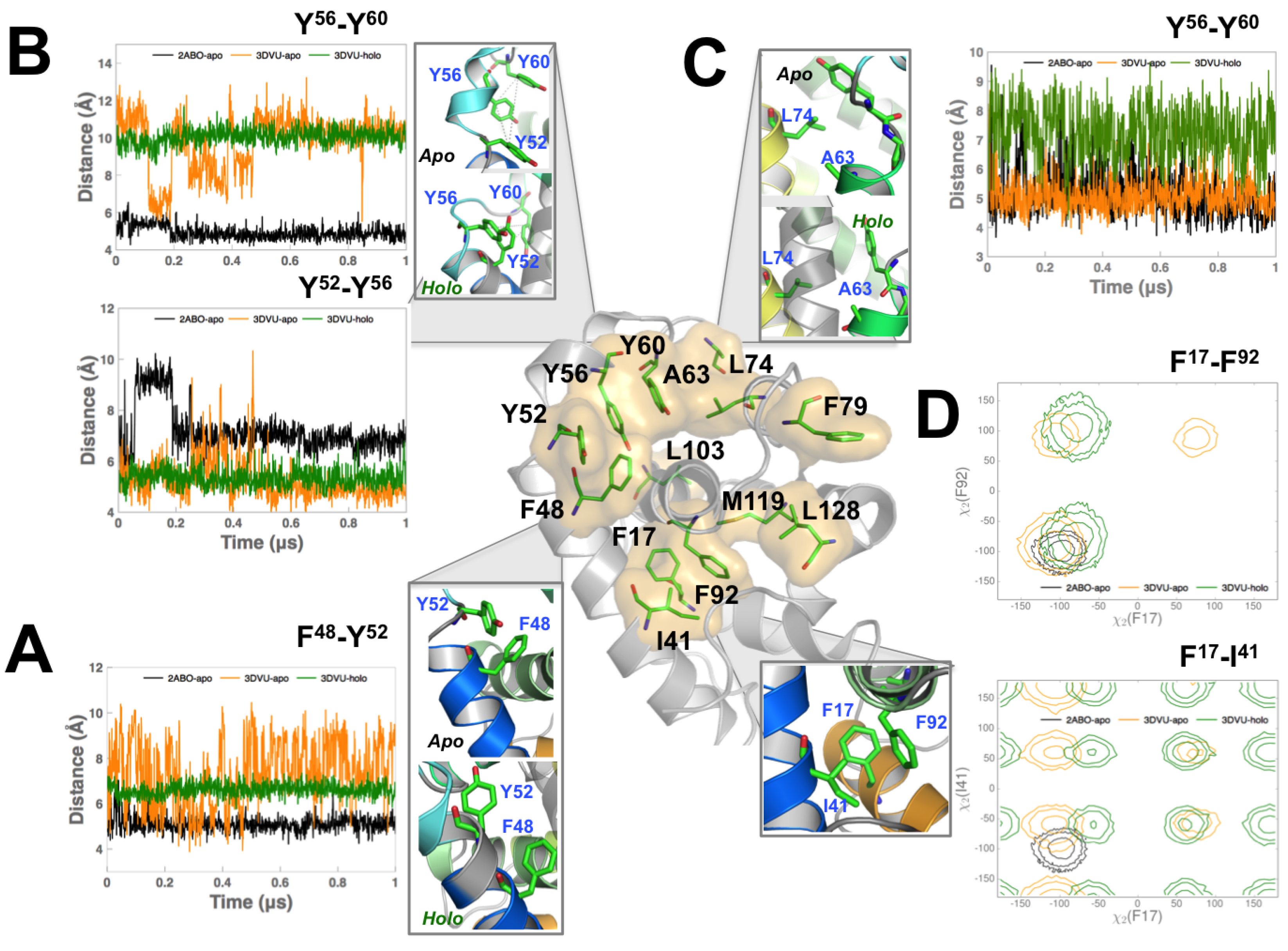
3.3. Perturbing Hydrophobic Interactions Farther from M11 Binding Cleft Impact BECN1 BH3D Binding
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Cory, S.; Adams, J.M. The Bcl2 family: Regulators of the cellular life-or-death switch. Nat. Rev. Cancer 2002, 2, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Cory, S.; Huang, D.C.S.; Adams, J.M. The Bcl-2 family: Roles in cell survival and oncogenesis. Oncogene 2003, 22, 8590–8607. [Google Scholar] [CrossRef] [PubMed]
- Youle, R.J.; Strasser, A. The BCL-2 protein family: Opposing activities that mediate cell death. Nat. Rev. Mol. Cell Biol. 2008, 9, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Hardwick, J.M.; Soane, L. Multiple Functions of BCL-2 Family Proteins. Cold Spring Harb. Perspect Biol. 2013, 5, a008722. [Google Scholar] [CrossRef] [PubMed]
- Hardwick, J.; Bellows, D. Viral versus cellular BCL-2 proteins. Cell Death Differ. 2003, 10, S68–S76. [Google Scholar] [CrossRef]
- Bellows, D.S.; Chau, B.N.; Lee, P.; Lazebnik, Y.; Burns, W.H.; Hardwick, J.M. Antiapoptotic Herpesvirus Bcl-2 Homologs Escape Caspase-Mediated Conversion to Proapoptotic Proteins. J. Virol. 2000, 74, 5024–5031. [Google Scholar] [CrossRef]
- Gangappa, S.; van Dyk, L.F.; Jewett, T.J.; Speck, S.H.; Virgin, H.W. Identification of the In Vivo Role of a Viral bcl-2. J. Exp. Med. 2002, 195, 931–940. [Google Scholar] [CrossRef]
- Cuconati, A.; White, E. Viral homologs of BCL-2: Role of apoptosis in the regulation of virus infection. Genes Dev. 2002, 16, 2465–2478. [Google Scholar] [CrossRef]
- Altmann, M.; Hammerschmidt, W. Epstein-Barr Virus Provides a New Paradigm: A Requirement for the Immediate Inhibition of Apoptosis. PLoS Biol. 2005, 3, e404. [Google Scholar] [CrossRef]
- Xiaofei, E.; Hwang, S.; Oh, S.; Lee, J.S.; Jeong, J.H.; Gwack, Y.; Kowalik, T.F.; Sun, R.; Jung, J.U.; Liang, C. Viral Bcl-2-Mediated Evasion of Autophagy Aids Chronic Infection of γ Herpesvirus 68. PLoS Pathog. 2009, 5, e1000609. [Google Scholar]
- Tarakanova, V.L.; Suarez, F.; Tibbetts, S.A.; Jacoby, M.A.; Weck, K.E.; Hess, J.L.; Speck, S.H.; Virgin, H.W. Murine Gammaherpesvirus 68 Infection Is Associated with Lymphoproliferative Disease and Lymphoma in BALB β2 Microglobulin-Deficient Mice. J. Virol. 2005, 79, 14668–14679. [Google Scholar] [CrossRef] [PubMed]
- Gershburg, E.; Pagano, J.S. Epstein–Barr virus infections: Prospects for treatment. J. Antimicrob. Chemother. 2005, 56, 277–281. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Mei, Y.; Sanishvili, R.; Levine, B.; Colbert, C.L.; Sinha, S. Targeting γ-herpesvirus 68 Bcl-2 mediated down-regulation of autophagy. J. Biol. Chem. 2014, 289, 8029–8040. [Google Scholar] [CrossRef] [PubMed]
- Procko, E.; Berguig, G.Y.; Shen, B.W.; Song, Y.; Frayo, S.; Convertine, A.J.; Margineantu, D.; Booth, G.; Correia, B.E.; Cheng, Y.; et al. A Computationally Designed Inhibitor of an Epstein-Barr Viral Bcl-2 Protein Induces Apoptosis in Infected Cells. Cell 2014, 157, 1644–1656. [Google Scholar] [CrossRef] [PubMed]
- Foight, G.W.; Keating, A.E. Locating Herpesvirus Bcl-2 Homologs in the Specificity Landscape of Anti-Apoptotic Bcl-2 Proteins. J. Mol. Biol. 2015, 427, 2468–2490. [Google Scholar] [CrossRef]
- Mei, Y.; Su, M.; Soni, G.; Salem, S.; Colbert, C.L.; Sinha, S.C. Intrinsically disordered regions in autophagy proteins. Proteins Struct. Func. Bioinform. 2014, 82, 565–578. [Google Scholar] [CrossRef]
- Flanagan, A.M.; Letai, A. BH3 domains define selective inhibitory interactions with BHRF-1 and KSHV BCL-2. Cell Death. Differ. 2007, 15, 580–588. [Google Scholar] [CrossRef]
- Kvansakul, M.; Wei, A.H.; Fletcher, J.I.; Willis, S.N.; Chen, L.; Roberts, A.W.; Huang, D.C.S.; Colman, P.M. Structural Basis for Apoptosis Inhibition by Epstein-Barr Virus BHRF1. PLoS Pathog. 2010, 6, e1001236. [Google Scholar] [CrossRef]
- Sinha, S.C.; Colbert, C.L.; Becker, N.; Wei, Y.; Levine, B. Molecular basis of the regulation of Beclin 1-dependent autophagy by the γ-herpesvirus 68 Bcl-2 homolog M11. Autophagy 2008, 4, 989–997. [Google Scholar] [CrossRef]
- Ku, B.; Woo, J.S.; Liang, C.; Lee, K.H.; Hong, H.S.; E, X.; Kim, K.S.; Jung, J.U.; Oh, B.H. Structural and Biochemical Bases for the Inhibition of Autophagy and Apoptosis by Viral BCL-2 of Murine γ-Herpesvirus 68. PLoS Pathog. 2008, 4, e25. [Google Scholar] [CrossRef]
- Kang, R.; Zeh, H.J.; Lotze, M.T.; Tang, D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 2011, 18, 571–580. [Google Scholar] [CrossRef]
- Decuypere, J.P.; Parys, J.; Bultynck, G. Regulation of the autophagic Bcl-2/Beclin interaction. Cells 2012, 1, 284–312. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Willis, S.N.; Wei, A.; Smith, B.J.; Fletcher, J.I.; Hinds, M.G.; Colman, P.M.; Day, C.L.; Adams, J.M.; Huang, D.C.S. Differential Targeting of Prosurvival Bcl-2 Proteins by Their BH3-Only Ligands Allows Complementary Apoptotic Function. Mol. Cell 2005, 17, 393–403. [Google Scholar] [CrossRef]
- Gavathiotis, E.; Suzuki, M.; Davis, M.L.; Pitter, K.; Bird, G.H.; Katz, S.G.; Tu, H.C.; Kim, H.; Cheng, E.H.Y.; Tjandra, N.; et al. BAX activation is initiated at a novel interaction site. Nature 2008, 455, 1076–1081. [Google Scholar] [CrossRef]
- Gavathiotis, E.; Reyna, D.; Davis, M.L.; Bird, G.H.; Walensky, L. BH3-Triggered Structural Reorganization Drives the Activation of Proapoptotic {BAX}. Mol. Cell 2010, 40, 481–492. [Google Scholar] [CrossRef] [PubMed]
- George, N.M.; Evans, J.J.; Luo, X. A three-helix homo-oligomerization domain containing BH3 and BH1 is responsible for the apoptotic activity of Bax. Genes Dev. 2007, 21, 1937–1948. [Google Scholar] [CrossRef] [PubMed]
- Ionescu, C.M.; Svobodová Vařeková, R.; Prehn, J.H.M.; Huber, H.J.; Koča, J. Charge Profile Analysis Reveals That Activation of Pro-apoptotic Regulators Bax and Bak Relies on Charge Transfer Mediated Allosteric Regulation. PLoS Comput. Biol. 2012, 8, e1002565. [Google Scholar] [CrossRef]
- Follis, A.V.; Chipuk, J.E.; Fisher, J.C.; Yun, M.K.; Grace, C.R.; Nourse, A.; Baran, K.; Ou, L.; Min, L.; White, S.W.; et al. PUMA binding induces partial unfolding within BCL-xL to disrupt p53 binding and promote apoptosis. Nat. Chem. Biol. 2013, 9, 163–168. [Google Scholar] [CrossRef]
- Kvansakul, M.; Hinds, M.G. The Bcl-2 family: Structures, interactions and targets for drug discovery. Apoptosis 2015, 20, 136–150. [Google Scholar] [CrossRef]
- Ramanathan, A.; Savol, A.J.; Langmead, C.J.; Agarwal, P.K.; Chennubhotla, C.S. Discovering conformational sub-states relevant to protein function. PLoS ONE 2011, 6, e15827. [Google Scholar] [CrossRef]
- Burger, V.M.; Ramanathan, A.; Savol, A.J.; Stanley, C.B.; Agarwal, P.K.; Chennubhotla, C.S. Quasi-anharmonic analysis reveals intermediate States in the nuclear co-activator receptor binding domain ensemble. Pac. Symp. Biocomput. 2012, 12, 70–81. [Google Scholar]
- Ramanathan, A.; Savol, A.; Burger, V.; Chennubhotla, C.S.; Agarwal, P.K. Protein Conformational Populations and Functionally Relevant Substates. Acc. Chem. Res. 2013, 47, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, A.; Savol, A.J.; Agarwal, P.K.; Chennubhotla, C.S. Event detection and sub-state discovery from biomolecular simulations using higher-order statistics: Application to enzyme adenylate kinase. Proteins Struct. Funct. Bioinform. 2012, 80, 2536–2551. [Google Scholar] [CrossRef] [PubMed]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, A.; Agarwal, P.K. Computational Identification of Slow Conformational Fluctuations in Proteins. J. Phys. Chem. B 2009, 113, 16669–16680. [Google Scholar] [CrossRef][Green Version]
- Ryckaert, J.P.; Ciccotti, G.; Berendsen, H.J.C. Numerical integration of the cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. J. Comput. Phys. 1977, 23, 327–341. [Google Scholar] [CrossRef]
- Mao, B.; Pear, M.R.; McCammon, J.A.; Northrup, S.H. Molecular dynamics of ferrocytochrome c: Anharmonicity of atomic displacements. Biopolymers 1982, 21, 1979–1989. [Google Scholar] [CrossRef]
- Ichiye, T.; Karplus, M. Anisotropy and anharmonicity of atomic fluctuations in proteins: Analysis of a molecular dynamics simulation. Proteins Struct. Func. Bioinform. 1987, 2, 236–259. [Google Scholar] [CrossRef]
- Savol, A.J.; Burger, V.M.; Agarwal, P.K.; Ramanathan, A.; Chennubhotla, C.S. QAARM: Quasi-anharmonic autoregressive model reveals molecular recognition pathways in ubiquitin. Bioinformatics 2011, 27, 52–60. [Google Scholar] [CrossRef][Green Version]
- Amari, S.; Cichocki, A.; Yang, H.H. A New Learning Algorithm for Blind Signal Separation. In Advances in Neural Information Processing Systems 9, NIPS; Denver, C.O., Mozer, M., Jordan, M.I., Petsche, T., Eds.; MIT Press: Cambridge, MA, USA, 1996; pp. 757–763. [Google Scholar]
- Burger, V.; Chennubhotla, C. Nhs: Network-based hierarchical segmentation for cryo-electron microscopy density maps. Biopolymers 2012, 97, 732–741. [Google Scholar] [CrossRef][Green Version]
- Loh, J.; Huang, Q.; Petros, A.M.; Nettesheim, D.; van Dyk, L.F.; Labrada, L.; Speck, S.H.; Levine, B.; Olejniczak, E.T.; Virgin, H.W., IV. A Surface Groove Essential for Viral Bcl-2 Function During Chronic Infection In Vivo. PLoS Pathog. 2005, 1, e10. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Darden, T.A.; Cheatham, T.E.I.; Simmerling, C.L.; Wang, J.; Duke, R.E.; Luo, R.; Merz, K.M.; Wang, B.; Pearlman, D.A.; et al. AMBER 12; University of California: San Francisco, CA, USA, 2012. [Google Scholar]
- Touw, W.; Baakman, C.; Black, J.; te Beek, T.A.H.; Krieger, E.; Joosten, R.; Vriend, G. A series of PDB related databases for everyday needs. Nucl. Acids Res. 2015, 43, D364–D368. [Google Scholar] [CrossRef]
- Kabsch, W.; Sander, C. Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometric features. Biopolymers 1983, 22, 2577–2637. [Google Scholar] [CrossRef] [PubMed]
- Chennubhotla, C.; Bahar, I. Signal Propagation in Proteins and Relation to Equilibrium Fluctuations. PLoS Comput. Biol. 2007, 3, e172. [Google Scholar]
- Chennubhotla, C.; Bahar, I. Markov Methods for Hierarchical Coarse-Graining of Large Protein Dynamics. J. Comput. Biol. 2007, 14, 765–776. [Google Scholar] [CrossRef]
- Chavez, L.L.; Onuchic, J.; Clementi, C. Quantifying the Roughness on the Free Energy Landscape: Entropic Bottlenecks and Protein Folding Rates. J. Am. Chem. Soc. 2004, 126, 8426–8432. [Google Scholar] [CrossRef]
- Kamat, A.P.; Lesk, A.M. Contact patterns between helices and strands of sheet define protein folding patterns. Proteins Struct. Func. Bioinform. 2007, 66, 869–876. [Google Scholar] [CrossRef]
- Mizushima, N.; Levine, B.; Cuervo, A.M.; Klionsky, D.J. Autophagy fights disease through cellular self-digestion. Nature 2008, 451, 1069–1075. [Google Scholar] [CrossRef]
- Levine, B.; Sinha, S.C.; Kroemer, G. Bcl-2 family members: Dual regulators of apoptosis and autophagy. Autophagy 2008, 4, 600–606. [Google Scholar] [CrossRef]
- Kvansakul, M.; Hinds, M.G. Structural biology of the Bcl-2 family and its mimicry by viral proteins. Cell Death Dis. 2013, 4, e909. [Google Scholar] [CrossRef]
- Fire, E.; Gulla, S.V.; Grant, R.A.; Keating, A.E. Mcl-1-Bim complexes accommodate surprising point mutations via minor structural changes. Protein Sci. 2010, 19, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Pan, A.C.; Jacobson, D.; Yatsenko, K.; Sritharan, D.; Weinreich, T.M.; Shaw, D.E. Atomic-level characterization of protein–protein association. Proc. Natl. Acad. Sci. USA 2019, 116, 4244–4249. [Google Scholar] [CrossRef] [PubMed]
- Kharche, S.A.; Sengupta, D. Dynamic protein interfaces and conformational landscapes of membrane protein complexes. Curr. Opin. Struct. Biol. 2020, 61, 191–197. [Google Scholar] [CrossRef]
- Siebenmorgen, T.; Engelhard, M.; Zacharias, M. Prediction of protein–protein complexes using replica exchange with repulsive scaling. J. Comput. Chem. 2020, 41, 1436–1447. [Google Scholar] [CrossRef] [PubMed]
- Dhusia, K.; Su, Z.; Wu, Y. Using Coarse-Grained Simulations to Characterize the Mechanisms of Protein– Protein Association. Biomolecules 2020, 10, 1056. [Google Scholar] [CrossRef] [PubMed]
- Gapsys, V.; de Groot, B.L. On the importance of statistics in molecular simulations for thermodynamics, kinetics and simulation box size. eLife 2020, 9, e57589. [Google Scholar] [CrossRef]
- Lee, H.; Turilli, M.; Jha, S.; Bhowmik, D.; Ma, H.; Ramanathan, A. DeepDriveMD: Deep-Learning Driven Adaptive Molecular Simulations for Protein Folding. In Proceedings of the 2019 IEEE/ACM Third Workshop on Deep Learning on Supercomputers (DLS), Denver, CO, USA, 17 November 2019; pp. 12–19. [Google Scholar]
- Mei, Y.; Glover, K.; Su, M.; Sinha, S.C. Conformational flexibility of BECN1: Essential to its key role in autophagy and beyond. Protein Sci. 2016, 25, 1767–1785. [Google Scholar] [CrossRef]
| (M) | () | H (kJ/mol) | G (kJ/mol) | S (J/K·mol) | Distance (Å) | |
|---|---|---|---|---|---|---|
| WT | - | |||||
| F48A | 9.4 | |||||
| A63K | 11.9 | |||||
| Y56A | 9.7 | |||||
| F17A | N/A | N/A | N/A | N/A | N/A | 19.7 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramanathan, A.; Parvatikar, A.; Chennubhotla, S.C.; Mei, Y.; Sinha, S.C. Transient Unfolding and Long-Range Interactions in Viral BCL2 M11 Enable Binding to the BECN1 BH3 Domain. Biomolecules 2020, 10, 1308. https://doi.org/10.3390/biom10091308
Ramanathan A, Parvatikar A, Chennubhotla SC, Mei Y, Sinha SC. Transient Unfolding and Long-Range Interactions in Viral BCL2 M11 Enable Binding to the BECN1 BH3 Domain. Biomolecules. 2020; 10(9):1308. https://doi.org/10.3390/biom10091308
Chicago/Turabian StyleRamanathan, Arvind, Akash Parvatikar, Srinivas C. Chennubhotla, Yang Mei, and Sangita C. Sinha. 2020. "Transient Unfolding and Long-Range Interactions in Viral BCL2 M11 Enable Binding to the BECN1 BH3 Domain" Biomolecules 10, no. 9: 1308. https://doi.org/10.3390/biom10091308
APA StyleRamanathan, A., Parvatikar, A., Chennubhotla, S. C., Mei, Y., & Sinha, S. C. (2020). Transient Unfolding and Long-Range Interactions in Viral BCL2 M11 Enable Binding to the BECN1 BH3 Domain. Biomolecules, 10(9), 1308. https://doi.org/10.3390/biom10091308





