Nicotinamide, Nicotinamide Riboside and Nicotinic Acid—Emerging Roles in Replicative and Chronological Aging in Yeast
Abstract
1. Introduction
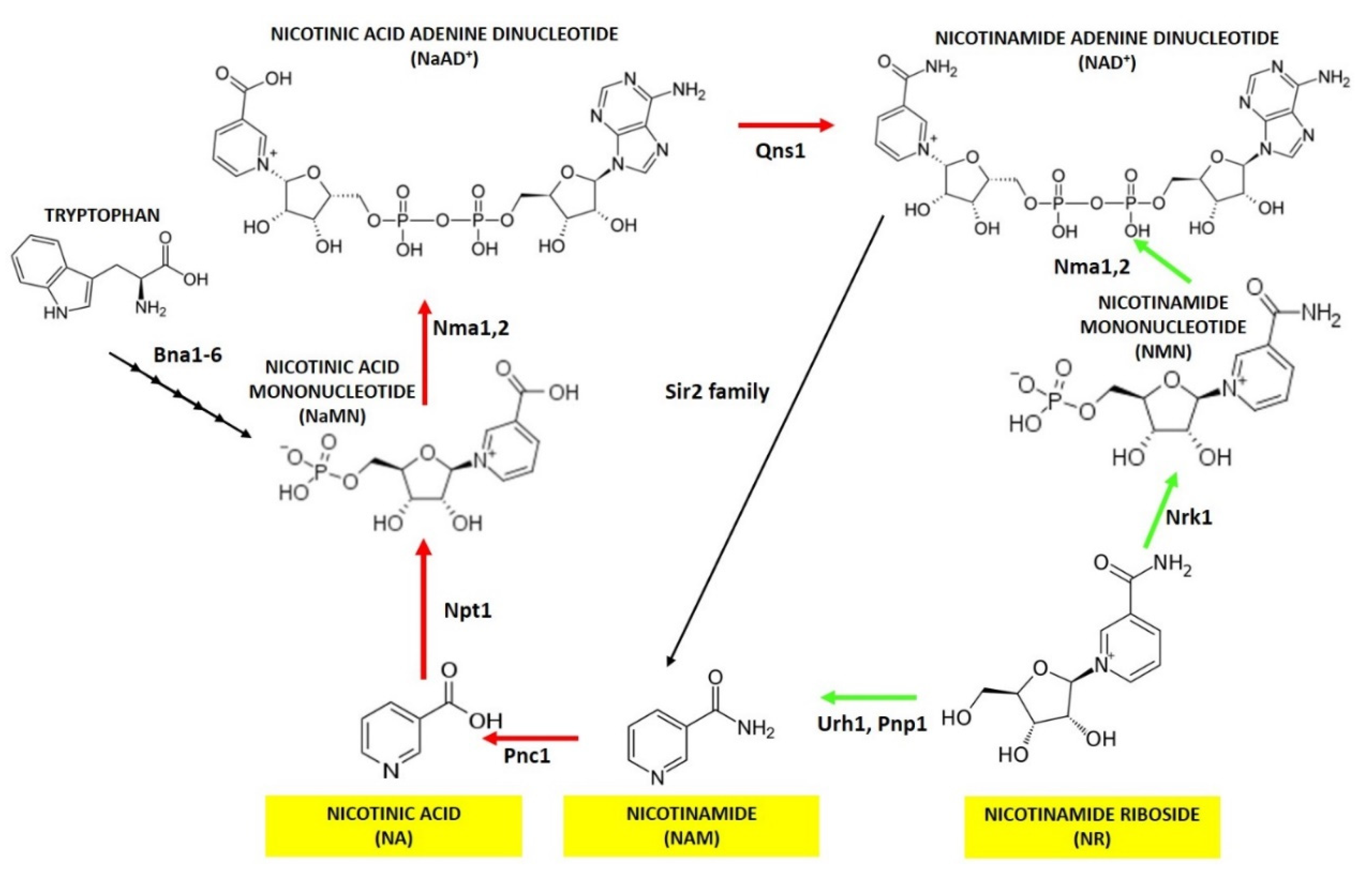
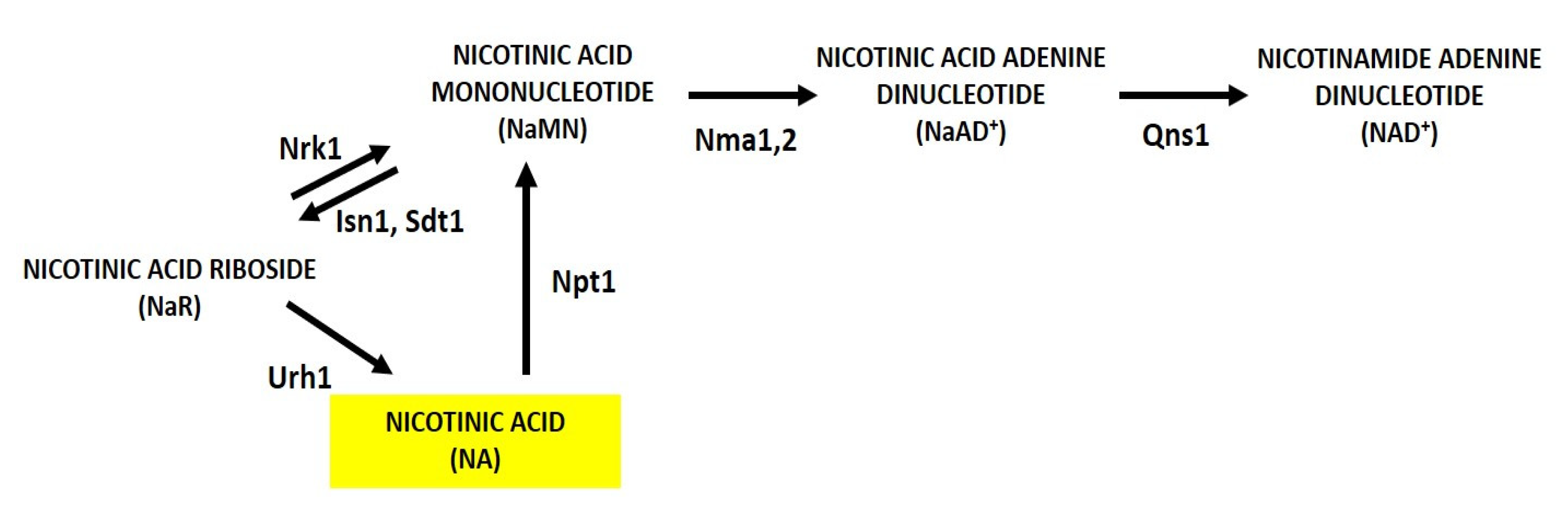
2. Nicotinic Acid, Nicotinamide and Nicotinamide Riboside: Key Metabolites for NAD+ Homeostasis
3. NAD+ Precursors and RLS
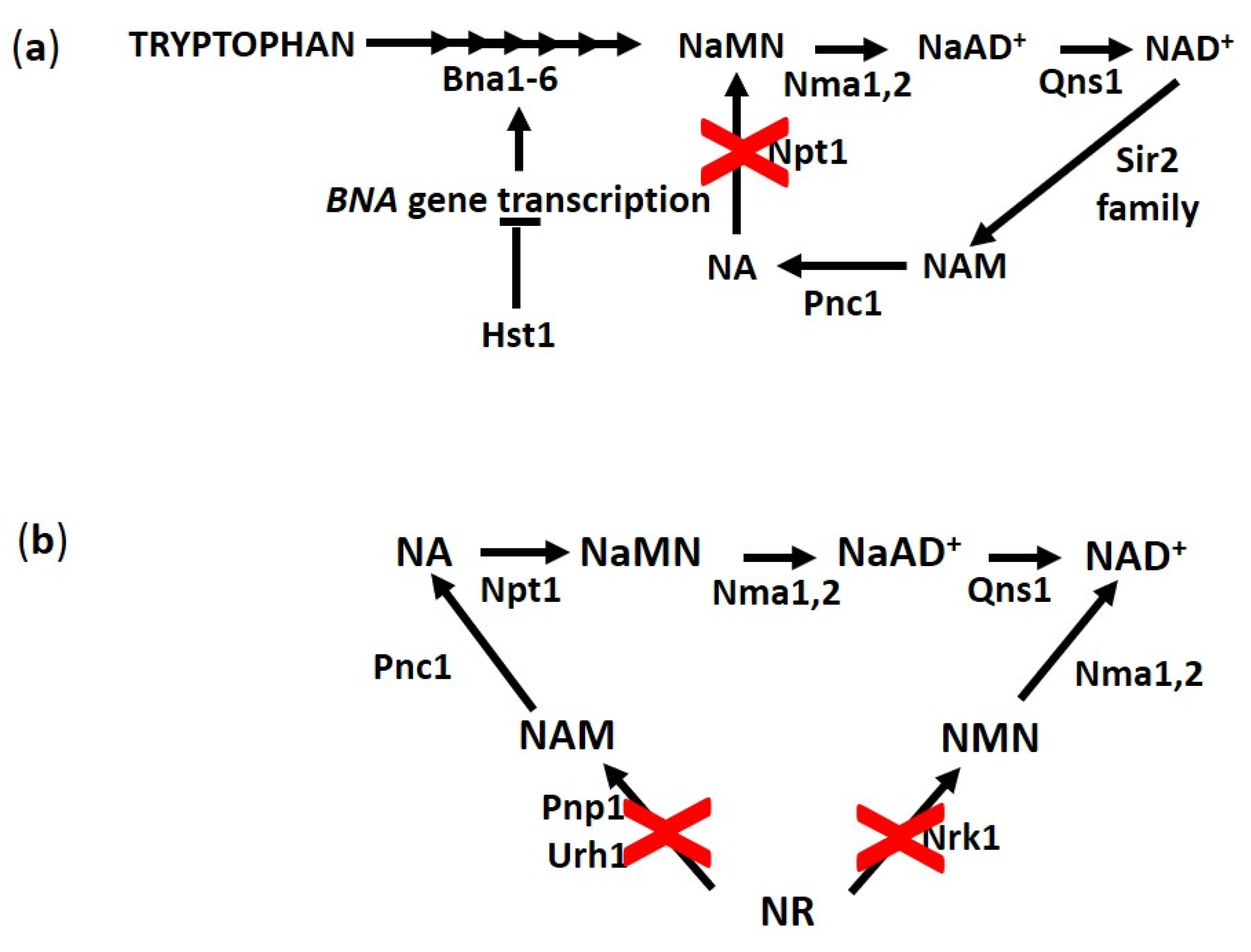
3.1. NA, an NAD+ Precursor Influencing RLS
3.2. NR, a New Vitamin Linking NAD+ Homeostasis and Longevity
3.3. NAM: A Problem or Resource for Replicative Aging?
4. NAD+ Precursors and CLS
Impact of NA, NR and NAM on CLS
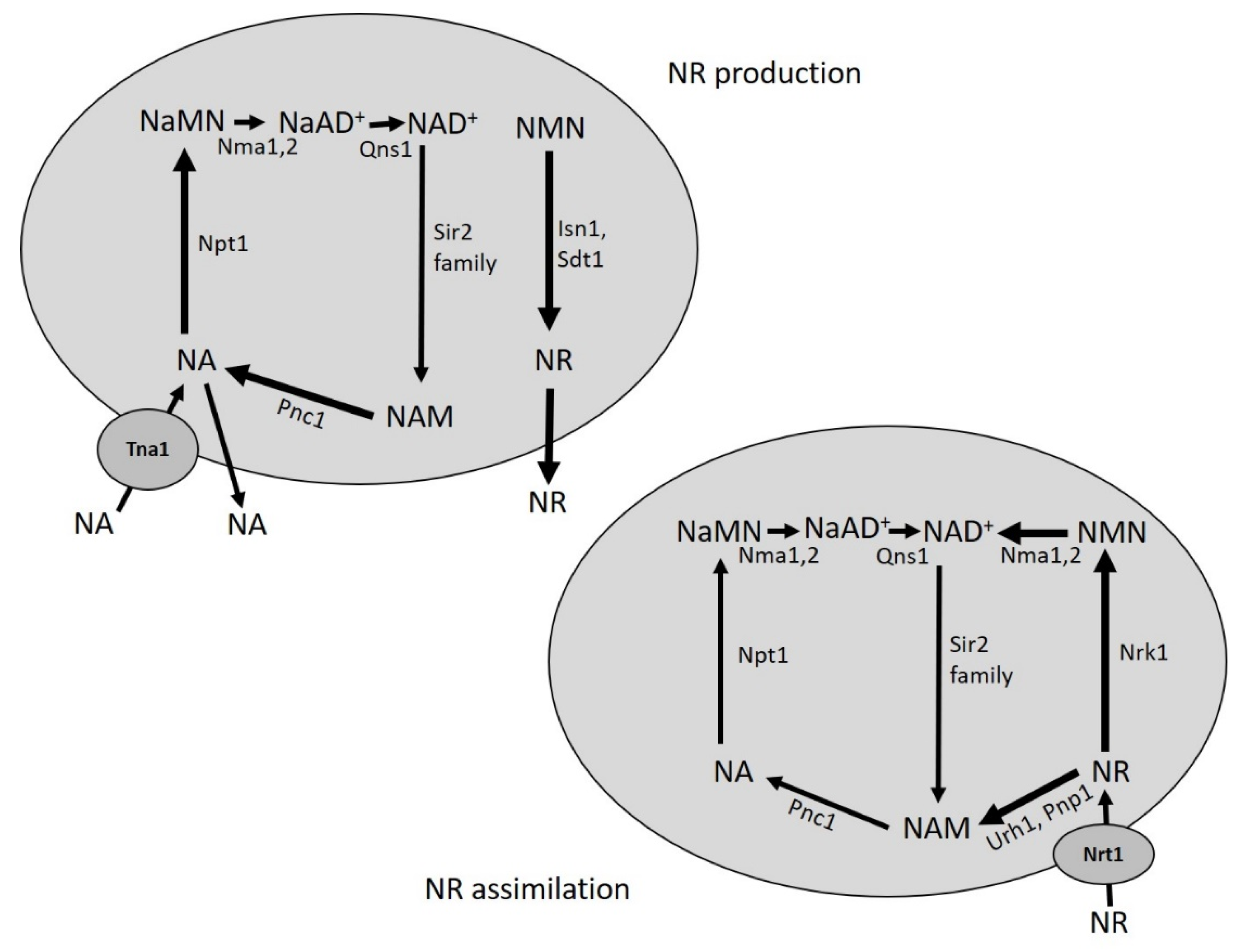
5. Production of NR from Yeast Cultures
6. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Cantó, C.; Menzies, K.J.; Auwerx, J. NAD+ metabolism and the control of energy homeostasis: A balancing act between mitochondria and the nucleus. Cell Metab. 2015, 22, 31–53. [Google Scholar] [CrossRef] [PubMed]
- Yaku, K.; Okabe, K.; Nakagawa, T. NAD metabolism: Implications in aging and longevity. Ageing Res. Rev. 2018, 47, 1–17. [Google Scholar] [CrossRef]
- James Theoga Raj, C.; Lin, S.J. Cross-talk in NAD+ metabolism: Insights from Saccharomyces cerevisiae. Curr. Genet. 2019, 65, 1113–1119. [Google Scholar] [CrossRef]
- McClure, J.M.; Gallo, C.M.; Smith, D.L.; Matecic, M.; Hontz, R.D.; Buck, S.W.; Racette, F.G.; Smith, J.S. Pnc1p-mediated nicotinamide clearance modifies the epigenetic properties of rDNA silencing in Saccharomyces cerevisiae. Genetics 2008, 180, 797–810. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Marmorstein, R. Structural basis for Sirtuin activity and inhibition. J. Biol. Chem. 2012, 287, 42428–42435. [Google Scholar] [CrossRef] [PubMed]
- Kosciuk, T.; Wang, M.; Hong, J.Y.; Lin, H. Updates on the epigenetic roles of Sirtuins. Curr. Opin. Chem. Biol. 2019, 51, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Imai, S.I.; Guarente, L. It takes two to tango: NAD+ and Sirtuins in aging/longevity control. NPS Aging Mech. Dis. 2016, 2, e16017. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Lee, J.H.; Lee, H.Y.; Min, K.J. Sirtuin signaling in cellular senescence and aging. BMB Rep. 2019, 52, 24–34. [Google Scholar] [CrossRef]
- Imai, S.I.; Guarente, L. NAD+ and Sirtuins in aging and disease. Trends Cell Biol. 2014, 24, 464–471. [Google Scholar] [CrossRef]
- Verdin, E. NAD+ in aging, metabolism, and neurodegeneration. Science 2015, 350, 1208–1213. [Google Scholar] [CrossRef]
- Fang, E.F.; Lautrup, S.; Hou, Y.; Demarest, T.G.; Croteau, D.L.; Mattson, M.P.; Bohr, V.A. NAD+ in aging: Molecular mechanisms and translational implications. Trends Mol. Med. 2017, 23, 899–916. [Google Scholar] [CrossRef]
- Johnson, S.; Imai, S.I. NAD (+) biosynthesis, aging, and disease. F1000Research 2018, 7, e132. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, S.J.; Bernier, M.; Aon, M.A.; Cortassa, S.; Kim, E.Y.; Fang, E.F.; Palacios, H.H.; Ali, A.; Navas-Enamorado, I.; Di Francesco, A.; et al. Nicotinamide improves aspects of healthspan, but not lifespan, in mice. Cell Metab. 2018, 27, 667–676. [Google Scholar] [CrossRef]
- Rajman, L.; Chwalek, K.; Sinclair, D.A. Therapeutic potential of NAD-boosting molecules: The in vivo evidence. Cell Metab. 2018, 27, 529–547. [Google Scholar] [CrossRef]
- Lautrup, S.; Sinclair, D.A.; Mattson, M.P.; Fang, E.F. NAD+ in brain aging and neurodegenerative disorders. Cell Metab. 2019, 30, 630–655. [Google Scholar] [CrossRef] [PubMed]
- Gilmour, B.C.; Gudmundsrud, R.; Frank, J.; Hov, A.; Lautrup, S.; Aman, Y.; Røsjø, H.; Brenner, C.; Ziegler, M.; Tysnes, O.B.; et al. Targeting NAD+ in translational research to relieve diseases and conditions of metabolic stress and ageing. Mech. Ageing Dev. 2020, 186, e111208. [Google Scholar] [CrossRef] [PubMed]
- Bieganowski, P.; Brenner, C. Discoveries of nicotinamide riboside as a nutrient and conserved NRK genes establish a Preiss-Handler independent route to NAD+ in fungi and humans. Cell 2004, 117, 495–502. [Google Scholar] [CrossRef]
- Bogan, K.L.; Brenner, C. Nicotinic acid, nicotinamide, and nicotinamide riboside: A molecular evaluation of NAD+ precursor vitamins in human nutrition. Annu. Rev. Nutr. 2008, 28, 115–130. [Google Scholar] [CrossRef] [PubMed]
- Chi, Y.; Sauve, A.A. Nicotinamide riboside, a trace nutrient in foods, is a Vitamin B3 with effects on energy metabolism and neuroprotection. Curr. Opin. Clin. Nutr. Metab. Care 2013, 16, 657–661. [Google Scholar] [CrossRef] [PubMed]
- MacLean, M.; Harris, N.; Piper, P.W. Chronological lifespan of stationary phase yeast cells; a model for investigating the factors that might influence the ageing of postmitotic tissues in higher organisms. Yeast 2001, 18, 499–509. [Google Scholar] [CrossRef] [PubMed]
- Longo, V.D.; Kennedy, B.K. Sirtuins in aging and age-related disease. Cell 2006, 126, 257–268. [Google Scholar] [CrossRef]
- Longo, V.D.; Shadel, G.S.; Kaeberlein, M.; Kennedy, B. Replicative and chronological aging in Saccharomyces cerevisiae. Cell Metab. 2012, 16, 18–31. [Google Scholar] [CrossRef] [PubMed]
- Gossmann, T.I.; Ziegler, M.; Puntervoll, P.; De Figueiredo, L.F.; Schuster, S.; Heiland, I. NAD+ biosynthesis and salvage—A phylogenetic perspective. FEBS J. 2012, 279, 3355–3363. [Google Scholar] [CrossRef] [PubMed]
- Dolle, C.; Skoge, R.; VanLinden, M.; Ziegler, M. NAD biosynthesis in humans—Enzymes, metabolites and therapeutic aspects. Curr. Top. Med. Chem. 2015, 13, 2907–2917. [Google Scholar] [CrossRef] [PubMed]
- Nikiforov, A.; Kulikova, V.; Ziegler, M. The human NAD metabolome: Functions, metabolism and compartmentalization. Crit. Rev. Biochem. Mol. Biol. 2015, 50, 284–297. [Google Scholar] [CrossRef]
- Stromland, O.; Niere, M.; Nikiforov, A.A.; VanLinden, M.R.; Heiland, I.; Ziegler, M. Keeping the balance in NAD metabolism. Biochem. Soc. Trans. 2019, 47, 119–130. [Google Scholar] [CrossRef]
- Croft, T.; Venkatakrishnan, P.; Lin, S.J. NAD+ metabolism and regulation: Lessons from yeast. Biomolecules 2020, 10, e330. [Google Scholar] [CrossRef]
- Suda, Y.; Tachikawa, H.; Yokota, A.; Nakanishi, H.; Yamashita, N.; Miura, Y.; Takahashi, N. Saccharomyces cerevisiae QNS1 codes for NAD(+) synthetase that is functionally conserved in mammals. Yeast 2003, 20, 995–1005. [Google Scholar] [CrossRef]
- Preiss, J.; Handler, P. Biosynthesis of diphosphopyridine nucleotide. J. Biol. Chem. 1958, 233, 488–492. [Google Scholar]
- Llorente, B.; Dujon, B. Transcriptional regulation of the Saccharomyces cerevisiae DAL5 gene family and identification of the high affinity nicotinic acid permease TNA1 (YGR260W). FEBS Lett. 2000, 475, 237–241. [Google Scholar] [CrossRef]
- Ghislain, M.; Talla, E.; François, J.M. Identification and functional analysis of the Saccharomyces cerevisiae nicotinamidase gene. PNCI Yeast 2002, 19, 215–224. [Google Scholar] [CrossRef] [PubMed]
- Tanny, J.C. Coupling of histone deacetylation to NAD breakdown by the yeast silencing protein Sir2: Evidence for acetyl transfer from substrate to an NAD breakdown product. Proc. Natl. Acad. Sci. USA 2001, 98, 415–420. [Google Scholar] [CrossRef] [PubMed]
- Sauve, A.A.; Wolberger, C.; Schramm, V.L.; Boeke, J.D. The biochemistry of Sirtuins. Annu. Rev. Biochem. 2006, 75, 435–465. [Google Scholar] [CrossRef] [PubMed]
- Sporty, J.; Lin, S.-J.; Kato, M.; Ognibene, T.; Stewart, B.; Turteltaub, K.; Bench, G. Quantitation of NAD+ biosynthesis from the salvage pathway in Saccharomyces cerevisiae. Yeast 2009, 26, 363–369. [Google Scholar] [CrossRef] [PubMed]
- Belenky, P.; Racette, F.G.; Bogan, K.L.; McClure, J.M.; Smith, J.S.; Brenner, C. Nicotinamide riboside promotes Sir2 silencing and extends lifespan via Nrk and Urh1/Pnp1/Meu1 pathways to NAD+. Cell 2007, 129, 473–484. [Google Scholar] [CrossRef] [PubMed]
- Belenky, P.A.; Moga, T.G.; Brenner, C. Saccharomyces cerevisiae YOR071C encodes the high affinity nicotinamide riboside transporter Nrt1. J. Biol. Chem. 2008, 283, 8075–8079. [Google Scholar] [CrossRef] [PubMed]
- Emanuelli, M.; Carnevali, F.; Lorenzi, M.; Raffaelli, N.; Amici, A.; Ruggieri, S.; Magni, G. Identification and characterization of YLR328W, the Saccharomyces cerevisiae structural gene encoding NMN adenylyltransferase. Expression and characterization of the recombinant enzyme. FEBS Lett. 1999, 455, 13–17. [Google Scholar] [CrossRef]
- Lu, S.P.; Kato, M.; Lin, S.J. Assimilation of endogenous nicotinamide riboside is essential for calorie restriction-mediated life span extension in Saccharomyces cerevisiae. J. Biol. Chem. 2009, 284, 17110–17119. [Google Scholar] [CrossRef]
- Belenky, P.; Stebbins, R.; Bogan, K.L.; Evans, C.R.; Brenner, C. Nrt1 and Tna1-independent export of NAD+ precursor vitamins promotes NAD+ homeostasis and allows engineering of vitamin production. PLoS ONE 2011, 6, e19710. [Google Scholar] [CrossRef]
- Tempel, W.; Rabeh, W.M.; Bogan, K.L.; Belenky, P.; Wojcik, M.; Seidle, H.F.; Nedyalkova, L.; Yang, T.; Sauve, A.A.; Park, H.-W.; et al. Nicotinamide riboside kinase structures reveal new pathways to NAD+. PLoS Biol. 2007, 5, e263. [Google Scholar] [CrossRef]
- Belenky, P.; Christensen, K.C.; Gazzaniga, F.; Pletnev, A.A.; Brenner, C. Nicotinamide riboside and nicotinic acid riboside salvage in fungi and mammals quantitative basis for Urh1 and purine nucleoside phosphorylase function in NAD+ metabolism. J. Biol. Chem. 2009, 284, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Bogan, K.L.; Evans, C.; Belenky, P.; Song, P.; Burant, C.F.; Kennedy, R.; Brenner, C. Identification of Isn1 and Sdt1 as glucose- and vitamin-regulated nicotinamide mononucleotide and nicotinic acid adenine dinucleotide 5′-nucleotidases responsible for production of nicotinamide riboside and nicotinic acid riboside. J. Biol. Chem. 2009, 284, 34861–34869. [Google Scholar] [CrossRef] [PubMed]
- Steinkraus, K.A.; Kaeberlein, M.K.B. Replicative aging in yeast: A means to an end. Annu. Rev. Cell Dev. Biol. 2009, 24, 29–54. [Google Scholar] [CrossRef] [PubMed]
- Kaeberlein, M.; McVey, M.; Guarente, L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 1999, 13, 2570–2580. [Google Scholar] [CrossRef]
- Imai, S.I.; Armstrong, C.M.; Kaeberlein, M.; Guarente, L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 2000, 403, 795–800. [Google Scholar] [CrossRef]
- Gartenberg, M.R.; Smith, J.S. The nuts and bolts of transcriptionally silent chromatin in Saccharomyces cerevisiae. Genetics 2016, 203, 1563–1599. [Google Scholar] [CrossRef]
- Bitterman, K.J.; Anderson, R.M.; Cohen, H.Y.; Latorre-Esteves, M.; Sinclair, D.A. Inhibition of silencing and accelerated aging by nicotinamide, a putative negative regulator of yeast Sir2 and human SIRT1. J. Biol. Chem. 2002, 277, 45099–45107. [Google Scholar] [CrossRef]
- Sinclair, D.A.; Guarente, L. Extrachromosomal rDNA circles—A cause of aging in yeast. Cell 1997, 91, 1033–1042. [Google Scholar] [CrossRef]
- Lindstrom, D.L.; Leverich, C.K.; Henderson, K.A.; Gottschling, D.E. Replicative age induces mitotic recombination in the ribosomal RNA gene cluster of Saccharomyces cerevisiae. PLoS Genet. 2011, 7, e1002015. [Google Scholar] [CrossRef]
- Ganley, A.R.D.; Kobayashi, T. Ribosomal DNA and cellular senescence: New evidence supporting the connection between rDNA and aging. FEMS Yeast Res. 2014, 14, 49–59. [Google Scholar] [CrossRef]
- Dang, W.; Steffen, K.K.; Perry, R.; Dorsey, J.A.; Johnson, F.B.; Shilatifard, A.; Kaeberlein, M.; Kennedy, B.K.; Berger, S.L. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 2009, 459, 802–807. [Google Scholar] [CrossRef] [PubMed]
- Borra, M.T.; Langer, M.R.; Slama, J.T.; Denu, J.M. Substrate specificity and kinetic mechanism of the Sir2 family of NAD+-dependent histone/protein deacetylases. Biochemistry 2004, 43, 9877–9887. [Google Scholar] [CrossRef] [PubMed]
- Kato, M.; Lin, S.J. Regulation of NAD+ metabolism, signaling and compartmentalization in the yeast Saccharomyces cerevisiae. DNA Repair 2014, 23, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Denu, J.M. Vitamins and aging: Pathways to NAD+ synthesis. Cell 2007, 129, 453–454. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.S.; Brachmann, C.B.; Celic, I.; Kenna, M.A.; Muhammad, S.; Starai, V.J.; Avalos, J.L.; Escalante-Semerena, J.C.; Grubmeyer, C.; Wolberger, C.; et al. A phylogenetically conserved NAD+-dependent protein deacetylase activity in the Sir2 protein family. Proc. Natl. Acad. Sci. USA 2000, 97, 6658–6663. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.M.; Bitterman, K.J.; Wood, J.G.; Medvedik, O.; Cohen, H.; Lin, S.S.; Manchester, J.K.; Gordon, J.I.; Sinclair, D.A. Manipulation of a nuclear NAD+ salvage pathway delays aging without altering steady-state NAD+ levels. J. Biol. Chem. 2002, 277, 18881–18890. [Google Scholar] [CrossRef]
- Sandmeier, J.J.; Celic, I.; Boeke, J.D.; Smith, J.S. Telomeric and rDNA silencing in Saccharomyces cerevisiae are dependent on a nuclear NAD(+) salvage pathway. Genetics 2002, 160, 877–889. [Google Scholar]
- Bedalov, A.; Hirao, M.; Posakony, J.; Nelson, M.; Simon, J.A. NAD+-dependent deacetylase Hst1p controls biosynthesis and cellular NAD+ levels in Saccharomyces cerevisiae. Mol. Cell. Biol. 2003, 23, 7044–7054. [Google Scholar] [CrossRef]
- Lu, S.P.; Lin, S.J. Phosphate-responsive signaling pathway is a novel component of NAD+ metabolism in Saccharomyces cerevisiae. J. Biol. Chem. 2011, 286, 14271–14281. [Google Scholar] [CrossRef]
- Kato, M.; Lin, S.J. YCL047C/POF1 is a novel nicotinamide mononucleotide adenylyltransferase (NMNAT) in Saccharomyces cerevisiae. J. Biol. Chem. 2014, 289, 15577–15587. [Google Scholar] [CrossRef]
- McClure, J.M.; Wierman, M.B.; Maqani, N.; Smith, J.S. Isonicotinamide enhances Sir2 protein-mediated silencing and longevity in yeast by raising intracellular NAD+ concentration. J. Biol. Chem. 2012, 287, 20957–20966. [Google Scholar] [CrossRef] [PubMed]
- Orlandi, I.; Pellegrino Coppola, D.; Strippoli, M.; Ronzulli, R.; Vai, M. Nicotinamide supplementation phenocopies SIR2 inactivation by modulating carbon metabolism and respiration during yeast chronological aging. Mech. Ageing Dev. 2017, 161, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Gallo, C.M.; Smith, D.L.; Smith, J.S. Nicotinamide clearance by Pnc1 directly regulates Sir2-mediated silencing and longevity. Mol. Cell. Biol. 2004, 24, 1301–1312. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.M.; Bitterman, K.J.; Wood, J.G.; Medvedik, O.; Sinclair, D.A. Nicotinamide and PNC1 govern lifespan extension by calorie restriction in Saccharomyces cerevisiae. Nature 2003, 423, 181–185. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.J.; Defossez, P.A.; Guarente, L. Requirement of NAD and SIR2 for life-span extension by calorie restriction in Saccharomyces cerevisiae. Science 2000, 289, 2126–2128. [Google Scholar] [CrossRef]
- Kaeberlein, M.; Kirkland, K.T.; Fields, S.; Kennedy, B.K. Sir2-independent life span extension by calorie restriction in yeast. PLoS Biol. 2004, 2, e296. [Google Scholar] [CrossRef] [PubMed]
- Tsuchiya, M.; Dang, N.; Kerr, E.O.; Hu, D.; Steffen, K.K.; Oakes, J.A.; Kennedy, B.K.; Kaeberlein, M. Sirtuin-independent effects of nicotinamide on lifespan extension from calorie restriction in yeast. Aging Cell 2006, 5, 505–514. [Google Scholar] [CrossRef] [PubMed]
- Katewa, S.D.; Kapahi, P. Dietary restriction and aging, 2009. Aging Cell 2010, 9, 105–112. [Google Scholar] [CrossRef]
- Anderson, R.M.; Weindruch, R. The caloric restriction paradigm: Implications for healthy human aging. Am. J. Hum. Biol. 2012, 24, 101–106. [Google Scholar] [CrossRef]
- Kapahi, P.; Kaeberlein, M.; Hansen, M. Dietary restriction and lifespan: Lessons from invertebrate models. Ageing Res. Rev. 2017, 39, 3–14. [Google Scholar] [CrossRef]
- Lee, Y.-L.; Lee, C.-K. Transcriptional response according to strength of calorie restriction in Saccharomyces cerevisiae. Mol. Cells 2008, 26, 299–307. [Google Scholar] [PubMed]
- Maslanka, R.; Kwolek-Mirek, M.; Zadrag-Tecza, R. Consequences of calorie restriction and calorie excess for the physiological parameters of the yeast Saccharomyces cerevisiae cells. FEMS Yeast Res. 2017, 17, e087. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.J.; Ford, E.; Haigis, M.; Liszt, G.; Guarente, L. Calorie restriction extends yeast life span by lowering the level of NADH. Genes Dev. 2004, 18, 12–16. [Google Scholar] [CrossRef]
- Evans, C.; Bogan, K.L.; Song, P.; Burant, C.F.; Kennedy, R.T.; Brenner, C. NAD+ metabolite levels as a function of vitamins and calorie restriction: Evidence for different mechanisms of longevity. BMC Chem. Biol. 2010, 10, e2. [Google Scholar] [CrossRef] [PubMed]
- Mei, S.C.; Brenner, C. Quantification of protein copy number in yeast: The NAD+ metabolome. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Sauve, A.A.; Moir, R.D.; Schramm, V.L.; Willis, I.M. Chemical activation of Sir2-dependent silencing by relief of nicotinamide inhibition. Mol. Cell 2005, 17, 595–601. [Google Scholar] [CrossRef]
- Fabrizio, P.; Longo, V.D. The chronological life span of Saccharomyces cerevisiae. Methods Mol. Biol. 2007, 371, 89–95. [Google Scholar]
- Bonawitz, N.D.; Rodeheffer, M.S.; Shadel, G.S. Defective mitochondrial gene expression results in reactive oxygen species-mediated inhibition of respiration and reduction of yeast life span. Mol. Cell. Biol. 2006, 26, 4818–4829. [Google Scholar] [CrossRef]
- Ocampo, A.; Liu, J.; Schroeder, E.A.; Shadel, G.S.; Barrientos, A. Mitochondrial respiratory thresholds regulate yeast chronological life span and its extension by caloric restriction. Cell Metab. 2012, 16, 55–67. [Google Scholar] [CrossRef]
- Baccolo, G.; Stamerra, G.; Damiano Pellegrino, C.; Orlandi, I.; Vai, M. Mitochondrial Metabolism and Aging in Yeast. In International Review of Cell and Molecular Biology; Academic Press: Cambridge, MA, USA, 2018; Volume 340, pp. 1–33. [Google Scholar]
- Casatta, N.; Porro, A.; Orlandi, I.; Brambilla, L.; Vai, M. Lack of Sir2 increases acetate consumption and decreases extracellular pro-aging factors. Biochim. Biophys. Acta Mol. Cell Res. 2013, 1833, 593–601. [Google Scholar] [CrossRef]
- Lin, Y.Y.; Lu, J.Y.; Zhang, J.; Walter, W.; Dang, W.; Wan, J.; Tao, S.C.; Qian, J.; Zhao, Y.; Boeke, J.D.; et al. Protein acetylation microarray reveals that NuA4 controls key metabolic target regulating gluconeogenesis. Cell 2009, 136, 1073–1084. [Google Scholar] [CrossRef] [PubMed]
- Orlandi, I.; Stamerra, G.; Strippoli, M.; Vai, M. During yeast chronological aging resveratrol supplementation results in a short-lived phenotype Sir2-dependent. Redox Biol. 2017, 12, 745–754. [Google Scholar] [CrossRef] [PubMed]
- Fabrizio, P.; Gattazzo, C.; Battistella, L.; Wei, M.; Cheng, C.; McGrew, K.; Longo, V.D. Sir2 blocks extreme life-span extension. Cell 2005, 123, 655–667. [Google Scholar] [CrossRef]
- Smith, D.L.; McClure, J.M.; Matecic, M.; Smith, J.S. Calorie restriction extends the chronological lifespan of Saccharomyces cerevisiae independently of the Sirtuins. Aging Cell 2007, 6, 649–662. [Google Scholar] [CrossRef] [PubMed]
- Orlandi, I.; Stamerra, G.; Vai, M. Altered expression of mitochondrial NAD+ carriers influences yeast chronological lifespan by modulating cytosolic and mitochondrial metabolism. Front. Genet. 2018, 9, e676. [Google Scholar] [CrossRef] [PubMed]
- Wei, M.; Fabrizio, P.; Hu, J.; Ge, H.; Cheng, C.; Li, L.; Longo, V.D. Life span extension by calorie restriction depends on Rim15 and transcription factors downstream of Ras/PKA, Tor, and Sch9. PLoS Genet. 2008, 4, 139–149. [Google Scholar] [CrossRef]
- Wei, M.; Fabrizio, P.; Madia, F.; Hu, J.; Ge, H.; Li, L.M.; Longo, V.D. Tor1/Sch9-regulated carbon source substitution is as effective as calorie restriction in life span extension. PLoS Genet. 2009, 5, e1000467. [Google Scholar] [CrossRef]
- Turck, D.; Castenmiller, J.; de Henauw, S.; Hirsch-Ernst, K.I.; Kearney, J.; Maciuk, A.; Mangelsdorf, I.; McArdle, H.J.; Naska, A.; Pelaez, C.; et al. Safety of nicotinamide riboside chloride as a novel food pursuant to Regulation (EU) 2015/2283 and bioavailability of nicotinamide from this source, in the context of Directive 2002/46/EC. EFSA J. 2019, 17, e5775. [Google Scholar]
- Yoshino, J.; Baur, J.A.; Imai, S.I. NAD+ intermediates: The biology and therapeutic potential of NMN and NR. Cell Metab. 2018, 27, 513–528. [Google Scholar] [CrossRef]
- Braidy, N.; Liu, Y. NAD+ therapy in age-related degenerative disorders: A benefit/risk analysis. Exp. Gerontol. 2020, 132, e110831. [Google Scholar] [CrossRef]
- Trammell, S.A.J.; Schmidt, M.S.; Weidemann, B.J.; Redpath, P.; Jaksch, F.; Dellinger, R.W.; Li, Z.; Abel, E.D.; Migaud, M.E.; Brenner, C. Nicotinamide riboside is uniquely and orally bioavailable in mice and humans. Nat. Commun. 2016, 7, e12948. [Google Scholar] [CrossRef] [PubMed]
- Airhart, S.E.; Shireman, L.M.; Risler, L.J.; Anderson, G.D.; Nagana Gowda, G.A.; Raftery, D.; Tian, R.; Shen, D.D.; O’Brien, K.D. An open-label, non-randomized study of the pharmacokinetics of the nutritional supplement nicotinamide riboside (NR) and its effects on blood NAD+ levels in healthy volunteers. PLoS ONE 2017, 12, e0186459. [Google Scholar] [CrossRef] [PubMed]
- Martens, C.R.; Denman, B.A.; Mazzo, M.R.; Armstrong, M.L.; Reisdorph, N.; McQueen, M.B.; Chonchol, M.; Seals, D.R. Chronic nicotinamide riboside supplementation is well-tolerated and elevates NAD+ in healthy middle-aged and older adults. Nat. Commun. 2018, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Dollerup, O.L.; Christensen, B.; Svart, M.; Schmidt, M.S.; Sulek, K.; Ringgaard, S.; Stodkilde-Jorgensen, H.; Moller, N.; Brenner, C.; Treebak, J.T.; et al. A randomized placebo-controlled clinical trial of nicotinamide riboside in obese men: Safety, insulin-sensitivity, and lipid-mobilizing effects. Am. J. Clin. Nutr. 2018, 108, 343–353. [Google Scholar] [CrossRef]
- Dollerup, O.L.; Trammell, S.A.J.; Hartmann, B.; Holst, J.J.; Christensen, B.; Moller, N.; Gillum, M.P.; Treebak, J.T.; Jessen, N. Effects of nicotinamide riboside on endocrine pancreatic function and incretin hormones in nondiabetic men with obesity. J. Clin. Endocrinol. Metab. 2019, 104, 5703–5714. [Google Scholar] [CrossRef]
- Dollerup, O.L.; Chubanava, S.; Agerholm, M.; Sondergard, S.D.; Altıntaş, A.; Moller, A.B.; Hoyer, K.F.; Ringgaard, S.; Stodkilde-Jorgensen, H.; Lavery, G.G.; et al. Nicotinamide riboside does not alter mitochondrial respiration, content or morphology in skeletal muscle from obese and insulin-resistant men. J. Physiol. 2020, 598, 731–754. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Orlandi, I.; Alberghina, L.; Vai, M. Nicotinamide, Nicotinamide Riboside and Nicotinic Acid—Emerging Roles in Replicative and Chronological Aging in Yeast. Biomolecules 2020, 10, 604. https://doi.org/10.3390/biom10040604
Orlandi I, Alberghina L, Vai M. Nicotinamide, Nicotinamide Riboside and Nicotinic Acid—Emerging Roles in Replicative and Chronological Aging in Yeast. Biomolecules. 2020; 10(4):604. https://doi.org/10.3390/biom10040604
Chicago/Turabian StyleOrlandi, Ivan, Lilia Alberghina, and Marina Vai. 2020. "Nicotinamide, Nicotinamide Riboside and Nicotinic Acid—Emerging Roles in Replicative and Chronological Aging in Yeast" Biomolecules 10, no. 4: 604. https://doi.org/10.3390/biom10040604
APA StyleOrlandi, I., Alberghina, L., & Vai, M. (2020). Nicotinamide, Nicotinamide Riboside and Nicotinic Acid—Emerging Roles in Replicative and Chronological Aging in Yeast. Biomolecules, 10(4), 604. https://doi.org/10.3390/biom10040604




