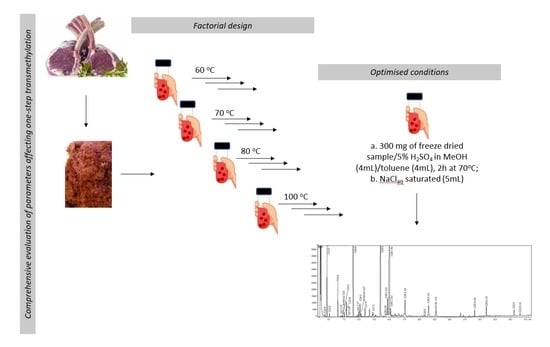
Comprehensive Evaluation of Parameters Affecting One-Step Method for Quantitative Analysis of Fatty Acids in Meat
Abstract
:1. Introduction
2. Results and Discussion
2.1. Influence of Reaction Temperature
2.2. Influence of Acid Concentration
2.3. Influence of Moisture Content
2.4. Influence of Solvent Volume and Sample Weight
2.5. Statistical Significance of the Influence of the Different Parameters Investigated
2.6. Influence of the Salt Type
2.7. Repeatability of the Optimised Direct Method for Quantitative Determination of Fatty Acid in Meat from Lamb, Beef and Venison
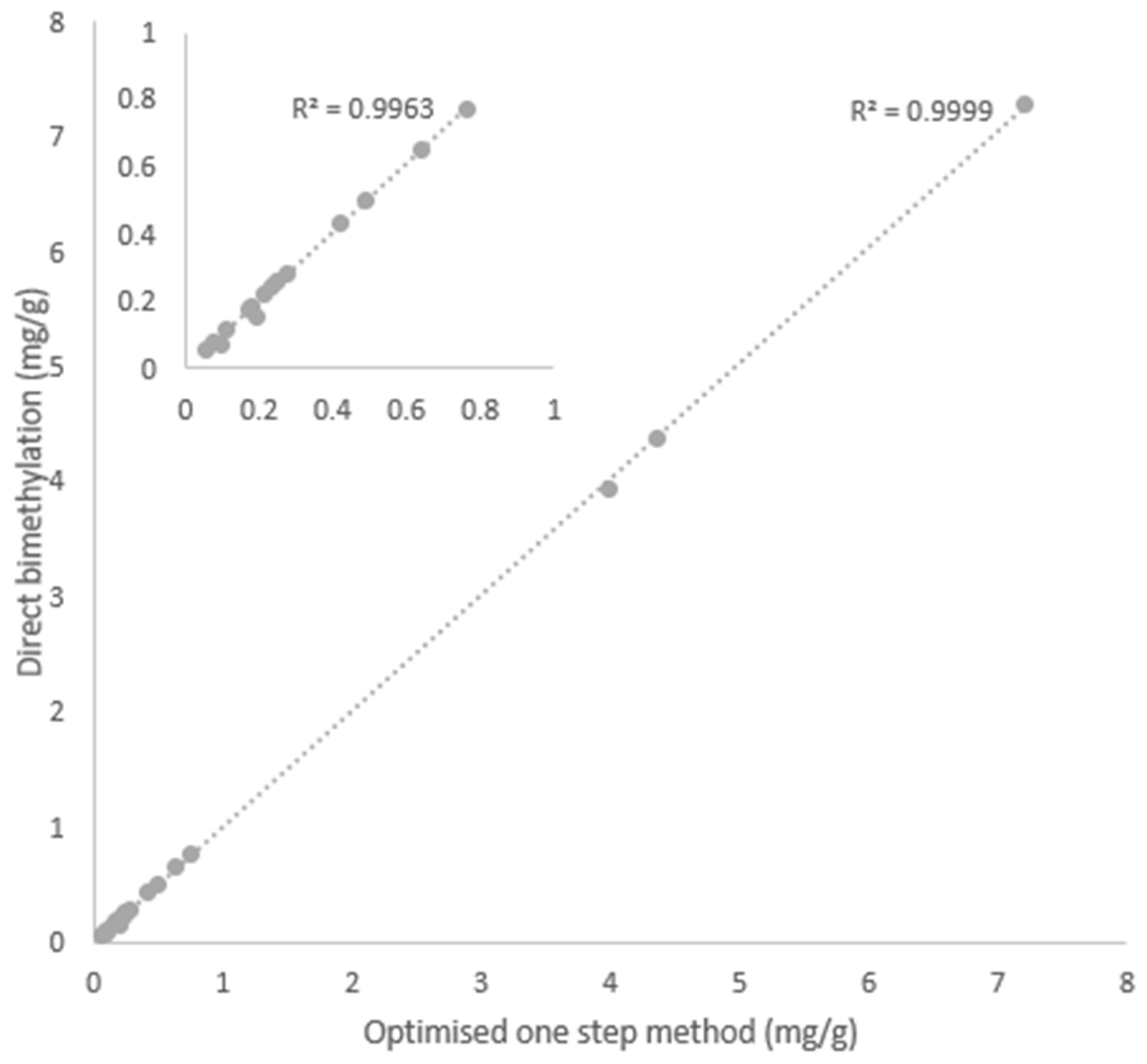
2.8. Comparative Evaluation Between the Optimised One-Step and Direct Bimethylation Procedures
3. Conclusions
4. Materials and Methods
4.1. Reagents
4.2. Samples
4.3. Experimental Design
4.4. FA Derivatisation using Bimethylation Procedure
4.5. FA Derivatisation using Optimised One-Step Procedure
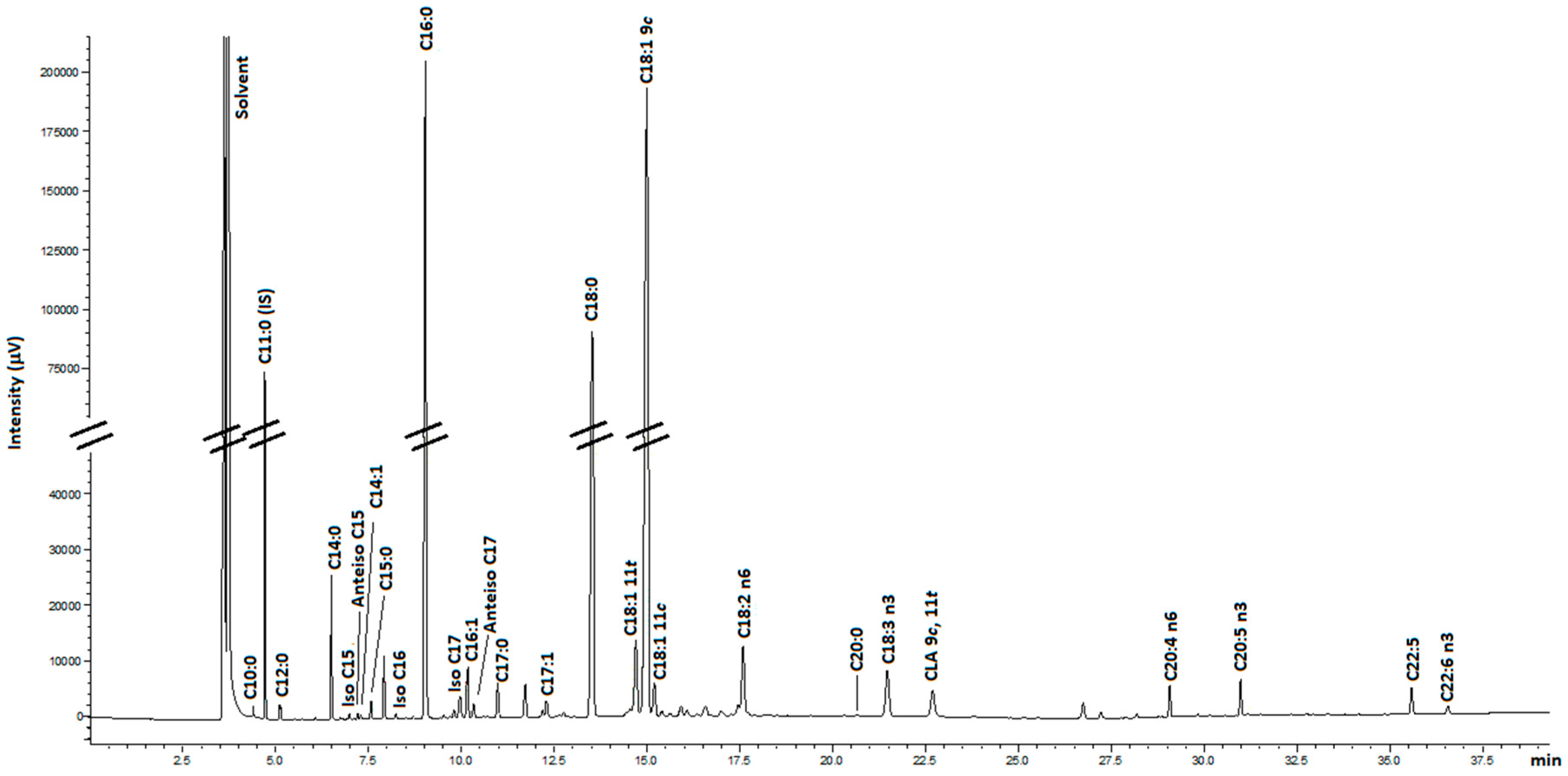
4.6. FAME Analysis
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jenkins, T.C. Technical note: Common analytical errors yielding inaccurate results during analysis of fatty acids in feed and digesta samples. J. Dairy Sci. 2010, 93, 1170–1174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pena, R.N.; Noguera, J.L.; García-Santana, M.J.; González, E.; Tejeda, J.F.; Ros-Freixedes, R.; Ibáñez-Escriche, N. Five genomic regions have a major impact on fat composition in Iberian pigs. Sci. Rep. 2019, 9, 2031. [Google Scholar] [CrossRef] [PubMed]
- Dugan, M.E.R.; Wood, J. Letter to the editor. Meat Sci. 2018, 143, 268. [Google Scholar] [CrossRef] [PubMed]
- De Paola, E.L.; Montevecchi, G.; Masino, F.; Antonelli, A.; Fiego, D.P.L. Single step extraction and derivatization of intramuscular lipids for fatty acid Ultra Fast GC analysis: Application on pig thigh. J. Food Sci. Technol. 2017, 54, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Wood, J.; Enser, M.; Fisher, A.; Nute, G.; Sheard, P.; Richardson, R.; Hughes, S.; Whittington, F. Fat deposition, fatty acid composition and meat quality: A review. Meat Sci. 2008, 78, 343–358. [Google Scholar] [CrossRef] [PubMed]
- Bhuiyan, M.S.A.; Lee, D.H.; Kim, H.J.; Lee, S.H.; Cho, S.H.; Yang, B.S.; Kim, S.D. Estimates of genetic parameters for fatty acid compositions in the longissimus dorsi muscle of Hanwoo cattle. Animal 2018, 12, 675–683. [Google Scholar] [CrossRef]
- Belaunzaran, X.; Lavín, P.; Mantecón, A.R.; Kramer, J.K.G.; Aldai, N. Effect of slaughter age and feeding system on the neutral and polar lipid composition of horse meat. Animal 2018, 12, 417–425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vahmani, P.; Rolland, D.; McAllister, T.; Block, H.; Proctor, S.; Guan, L.; Prieto, N.; López-Campos, Ó.; Aalhus, J.; Dugan, M.; et al. Effects of feeding steers extruded flaxseed on its own before hay or mixed with hay on animal performance, carcass quality, and meat and hamburger fatty acid composition. Meat Sci. 2017, 131, 9–17. [Google Scholar] [CrossRef]
- Schiavon, S.; Bergamaschi, M.; Pellattiero, E.; Simonetto, A.; Tagliapietra, F. Fatty Acid Composition of Lamb Liver, Muscle, And Adipose Tissues in Response to Rumen-Protected Conjugated Linoleic Acid (CLA) Supplementation Is Tissue Dependent. J. Agric. Food Chem. 2017, 65, 10604–10614. [Google Scholar] [CrossRef]
- Jaturasitha, S.; Chaiwang, N.; Kayan, A.; Kreuzer, M. Nutritional strategies to improve the lipid composition of meat, with emphasis on Thailand and Asia. Meat Sci. 2016, 120, 157–166. [Google Scholar] [CrossRef]
- Wood, J.; Richardson, R.; Nute, G.; Fisher, A.; Campo, M.M.; Kasapidou, E.; Sheard, P.; Enser, M. Effects of fatty acids on meat quality: A review. Meat Sci. 2004, 66, 21–32. [Google Scholar] [CrossRef]
- Raes, K. Meat quality, fatty acid composition and flavour analysis in Belgian retail beef. Meat Sci. 2003, 65, 1237–1246. [Google Scholar] [CrossRef]
- Arshad, M.S.; Sohaib, M.; Ahmad, R.S.; Nadeem, M.T.; Imran, A.; Arshad, M.U.; Kwon, J.-H.; Amjad, Z. Ruminant meat flavor influenced by different factors with special reference to fatty acids. Lipids Heal. Dis. 2018, 17, 223. [Google Scholar] [CrossRef] [PubMed]
- Flakemore, A.R.; Malau-Aduli, B.S.; Nichols, P.D.; Malau-Aduli, A.E.O. Omega-3 fatty acids, nutrient retention values, and sensory meat eating quality in cooked and raw Australian lamb. Meat Sci. 2017, 123, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Scollan, N.; Hocquette, J.-F.; Nuernberg, K.; Dannenberger, D.; Richardson, I.; Moloney, A. Innovations in beef production systems that enhance the nutritional and health value of beef lipids and their relationship with meat quality. Meat Sci. 2006, 74, 17–33. [Google Scholar] [CrossRef] [PubMed]
- Abdulkadir, S.; Tsuchiya, M. One-step method for quantitative and qualitative analysis of fatty acids in marine animal samples. J. Exp. Mar. Boil. Ecol. 2008, 354, 1–8. [Google Scholar] [CrossRef]
- O’Fallon, J.V.; Busboom, J.R.; Nelson, M.L.; Gaskins, C.T. A direct method for fatty acid methyl ester synthesis: Application to wet meat tissues, oils, and feedstuffs. J. Anim. Sci. 2007, 85, 1511–1521. [Google Scholar] [CrossRef] [Green Version]
- Murrieta, C.; Hess, B.; Rule, D. Comparison of acidic and alkaline catalysts for preparation of fatty acid methyl esters from ovine muscle with emphasis on conjugated linoleic acid. Meat Sci. 2003, 65, 523–529. [Google Scholar] [CrossRef]
- Sukhija, P.S.; Palmquist, D.L. Rapid method for determination of total fatty acid content and composition of feedstuffs and feces. J. Agric. Food Chem. 1988, 36, 1202–1206. [Google Scholar] [CrossRef]
- Araujo, P.; Nguyen, T.-T.; Frøyland, L.; Wang, J.; Kang, J.X. Evaluation of a Rapid Method for the Quantitative Analysis of Fatty Acids in Various Matrices. J. Chromatogr. A 2008, 1212, 106–113. [Google Scholar] [CrossRef]
- Griffiths, M.J.; Van Hille, R.P.; Harrison, S.T.L.; Griffiths, M.; Hille, R.P. Selection of Direct Transesterification as the Preferred Method for Assay of Fatty Acid Content of Microalgae. Lipids 2010, 45, 1053–1060. [Google Scholar] [CrossRef] [PubMed]
- Castro-Gómez, P.; Fontecha, J.; Rodríguez-Alcalá, L.M. A high-performance direct transmethylation method for total fatty acids assessment in biological and foodstuff samples. Talanta 2014, 128, 518–523. [Google Scholar] [CrossRef] [PubMed]
- Hidalgo, P.; Ciudad, G.; Schober, S.; Mittelbach, M.; Navia, R. Improving the FAME Yield of in Situ Transesterification from Microalgal Biomass through Particle Size Reduction and Cosolvent Incorporation. Energy Fuels 2015, 29, 823–832. [Google Scholar] [CrossRef]
- Carrapiso, A.I.; Timón, M.L.; Petrón, M.J.; Tejeda, J.F.; García, C. In situ transesterification of fatty acids from Iberian pig subcutaneous adipose tissue. Meat Sci. 2000, 56, 159–164. [Google Scholar] [CrossRef]
- Lee, M.R.F.; Tweed, J.K.S. Isomerisation of cis-9 trans-11 conjugated linoleic acid (CLA) to trans-9 trans-11 CLA during acidic methylation can be avoided by a rapid base catalysed methylation of milk fat. J. Dairy Res. 2008, 75, 354–356. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Juarez, M.; Polvillo, O.; Contò, M.; Ficco, A.; Ballico, S.; Failla, S. Comparison of four extraction/methylation analytical methods to measure fatty acid composition by gas chromatography in meat. J. Chromatogr. A 2008, 1190, 327–332. [Google Scholar] [CrossRef] [PubMed]
- Hoerr, C.W.; Harwood, H.J. Solubilities of high molecular weight aliphatic compounds in n-hexane. J. Org. Chem. 1951, 16, 779–791. [Google Scholar] [CrossRef]
- Crompton, M.J.; Dunstan, R.H. Evaluation of in-situ fatty acid extraction protocols for the analysis of staphylococcal cell membrane associated fatty acids by gas chromatography. J. Chromatogr. B 2018, 1084, 80–88. [Google Scholar] [CrossRef]
- Yamasaki, M.; Kishihara, K.; Ikeda, I.; Sugano, M.; Yamada, K. A recommended esterification method for gas chromatographic measurement of conjugated linoleic acid. J. Am. Oil Chem. Soc. 1999, 76, 933–938. [Google Scholar] [CrossRef]
- Liu, Z.; Ezernieks, V.; Rochfort, S.; Cocks, B. Comparison of methylation methods for fatty acid analysis of milk fat. Food Chem. 2018, 261, 210–215. [Google Scholar] [CrossRef]
- Kramer, J.K.G.; Fellner, V.; Dugan, M.E.R.; Sauer, F.D.; Mossoba, M.M.; Yurawecz, M.P. Evaluating acid and base catalysts in the methylation of milk and rumen fatty acids with special emphasis on conjugated dienes and total trans fatty acids. Lipids 1997, 32, 1219–1228. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Moate, P.; Rochfort, S. A simplified protocol for fatty acid profiling of milk fat without lipid extraction. Int. Dairy J. 2019, 90, 68–71. [Google Scholar] [CrossRef]
- MacGee, J.; Williams, M.G. Prepartion of sphingolipid fatty acid methyl esters for determination by gas—liquid chromatography. J. Chromatogr. A 1981, 205, 281–288. [Google Scholar] [CrossRef]
- Dong, T.; Yu, L.; Gao, D.; Yu, X.; Miao, C.; Zheng, Y.; Lian, J.; Li, T.; Chen, S. Direct quantification of fatty acids in wet microalgal and yeast biomass via a rapid in situ fatty acid methyl ester derivatization approach. Appl. Microbiol. Biotechnol. 2015, 99, 10237–10247. [Google Scholar] [CrossRef] [PubMed]
- Ulberth, F.; Henninger, M. One-step extraction/methylation method for determining the fatty acid composition of processed foods. J. Am. Oil Chem. Soc. 1992, 69, 174–177. [Google Scholar] [CrossRef]
- Ken’ichi, I.; Yumeto, F. Preparation of fatty acid methyl esters for gas-liquid chromatography. J. Lipid Res. 2010, 51, 635–640. [Google Scholar] [Green Version]
- Lee, M.; Tweed, J.; Kim, E.; Scollan, N. Beef, chicken and lamb fatty acid analysis—A simplified direct bimethylation procedure using freeze-dried material. Meat Sci. 2012, 92, 863–866. [Google Scholar] [CrossRef]



| Fatty Acids | P-values | R2-adj Values | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| - | Individual Effect | Interaction | |||||||||
| - | Temperature | Time | Acid Concentration | Volume | Temp * Time | Temp * Acid Concentration | Temp * Vol | Time * Acid Concentration | Time * Vol | Acid Concentration * Vol | |
| Total Fatty Acids | <0.001 | <0.001 | <0.001 | 0.005 | <0.001 | 0.052 | 0.002 | 0.237 | 0.052 | 0.212 | 45.4% |
| C10:0 | <0.001 | <0.001 | 0.002 | <0.001 | <0.001 | 0.525 | 0.015 | 0.417 | 0.105 | 0.299 | 33.5% |
| C12:0 | <0.001 | <0.001 | 0.005 | 0.061 | <0.001 | 0.679 | 0.086 | 0.488 | 0.184 | 0.546 | 26.5% |
| C14:0 | <0.001 | <0.001 | <0.001 | 0.022 | <0.001 | 0.067 | <0.001 | 0.284 | 0.042 | 0.199 | 45.4% |
| Iso C15 | <0.001 | <0.001 | 0.004 | <0.001 | <0.001 | 0.071 | 0.239 | 0.503 | 0.348 | 0.400 | 42.6% |
| Anteiso C15 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | 0.066 | 0.036 | 0.346 | 0.031 | 0.558 | 43.5% |
| C15:0 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | 0.130 | 0.003 | 0.222 | 0.026 | 0.275 | 46.3% |
| Iso C16 | <0.001 | 0.129 | 0.008 | 0.366 | 0.049 | 0.408 | 0.125 | 0.754 | 0.101 | 0.914 | 36.5% |
| C16:0 | <0.001 | <0.001 | <0.001 | 0.005 | <0.001 | 0.062 | 0.001 | 0.334 | 0.047 | 0.235 | 46.0% |
| Iso c17 | 0.041 | <0.001 | <0.001 | <0.001 | <0.001 | 0.019 | <0.001 | 0.328 | 0.059 | 0.223 | 41.2% |
| C16:1 | <0.001 | <0.001 | <0.001 | 0.002 | <0.001 | 0.037 | 0.002 | 0.269 | 0.056 | 0.238 | 43.3% |
| Anteiso C17 | <0.001 | <0.001 | 0.001 | 0.010 | <0.001 | 0.149 | 0.017 | 0.416 | 0.048 | 0.301 | 38.5% |
| C17:0 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | 0.063 | 0.001 | 0.310 | 0.050 | 0.219 | 45.9% |
| C17:1 | <0.001 | 0.001 | <0.001 | 0.001 | <0.001 | 0.046 | 0.006 | 0.250 | 0.056 | 0.232 | 39.3% |
| C18:0 | <0.001 | <0.001 | <0.001 | 0.011 | <0.001 | 0.063 | 0.001 | 0.322 | 0.039 | 0.216 | 47.1% |
| C18:1 9t | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | 0.847 | 0.004 | 0.871 | 0.091 | 0.558 | 65.1% |
| C18:1 11t | <0.001 | <0.001 | <0.001 | 0.003 | <0.001 | 0.093 | <0.001 | 0.258 | 0.038 | 0.232 | 45.1% |
| C18:1 9c | <0.001 | 0.01 | 0.001 | <0.001 | <0.001 | 0.156 | 0.005 | 0.443 | 0.176 | 0.681 | 37.7% |
| C18:1 11c | <0.001 | 0.010 | 0.001 | <0.001 | <0.001 | 0.156 | 0.005 | 0.443 | 0.176 | 0.681 | 33.6% |
| C18:2 n6 | <0.001 | 0.077 | <0.001 | 0.111 | <0.001 | 0.005 | 0.003 | 0.260 | 0.053 | 0.177 | 29.2% |
| C20:0 | 0.407 | 0.061 | 0.001 | 0.308 | <0.001 | 0.254 | 0.055 | 0.541 | 0.735 | 0.636 | 4.7% |
| C18:3 n3 | <0.001 | 0.014 | <0.001 | 0.013 | <0.001 | 0.013 | 0.002 | 0.269 | 0.041 | 0.198 | 33.5% |
| CLA 9c, 11t | <0.001 | 0.784 | 0.301 | 0.274 | <0.001 | <0.001 | <0.001 | 0.282 | 0.066 | 0.333 | 38.3% |
| C20:4 n6 | 0.695 | 0.821 | 0.001 | 0.031 | <0.001 | 0.003 | 0.039 | 0.123 | 0.271 | 0.084 | 17.5% |
| C20:5 n3 | <0.001 | 0.554 | 0.856 | 0.934 | 0.906 | 0.819 | 0.443 | 0.660 | 0.861 | 0.729 | 24.7% |
| C22:5 | 0.543 | 0.739 | <0.001 | 0.002 | <0.001 | 0.001 | 0.006 | 0.385 | 0.011 | 0.112 | 23.0% |
| C22:6 n3 | 0.053 | 0.143 | <0.001 | 0.067 | <0.001 | 0.028 | 0.056 | 0.412 | 0.196 | 0.093 | 21.8% |
| FAs | Na2CO3 | NaCl | ||
|---|---|---|---|---|
| Mean ± STD | RSD | Mean ± STD | RSD | |
| C10:0 | 0.21 ± 0.00 | 1.5 | 0.20 ± 0.00 | 2.4 |
| C12:0 | 0.44 ± 0.01 | 1.6 | 0.44 ± 0.01 | 1.9 |
| C14:0 | 3.81 ± 0.07 | 1.9 | 3.76 ± 0.07 | 1.9 |
| iso C15 | 0.13 ± 0.00 | 2.8 | 0.13 ± 0.00 | 2.7 |
| anteiso C15 | 0.19 ± 0.00 | 2.3 | 0.19 ± 0.01 | 3.6 |
| C14:1 | 0.10 ± 0.00 | 1.7 | 0.10 ± 0.00 | 2.0 |
| C15:0 | 0.45 ± 0.01 | 1.2 | 0.43 ± 0.01 | 2.2 |
| iso C16 | 0.16 ± 0.01 | 3.8 | 0.16 ± 0.01 | 4.0 |
| C16:0 | 34.26 ± 0.37 | 1.1 | 34.14 ± 0.38 | 1.1 |
| iso C17 | 0.54 ± 0.01 | 1.5 | 0.53 ± 0.01 | 1.7 |
| C16:1 | 1.60 ± 0.02 | 1.1 | 1.57 ± 0.02 | 1.5 |
| anteiso C17 | 0.61 ± 0.01 | 2.2 | 0.59 ± 0.01 | 1.8 |
| C17:0 | 1.44 ± 0.01 | 0.8 | 1.40 ± 0.02 | 1.5 |
| C17:1 | 0.75 ± 0.00 | 0.6 | 0.73 ± 0.01 | 0.9 |
| C18:0 | 24.17 ± 0.26 | 1.1 | 24.05 ± 0.27 | 1.1 |
| C18:1 9t | 0.31 ± 0.01 | 1.8 | 0.30 ± 0.01 | 3.1 |
| C18:1 11t | 3.77 ± 0.04 | 1.0 | 3.69 ± 0.06 | 1.8 |
| C18:1 9c | 56.32 ± 0.63 | 1.1 | 56.28 ± 0.47 | 0.8 |
| C18:1 11c | 1.39 ± 0.01 | 0.7 | 1.35 ± 0.01 | 1.0 |
| C18:2 n6 | 4.70 ± 0.03 | 0.7 | 4.58 ± 0.04 | 0.9 |
| C20:0 | 0.10 ± 0.00 | 1.5 | 0.10 ± 0.00 | 3.4 |
| C18:3 n3 | 3.04 ± 0.02 | 0.5 | 2.98 ± 0.06 | 2.0 |
| CLA 9c, 11t | 1.72 ± 0.02 | 0.9 | 1.71 ± 0.08 | 4.6 |
| C22 | 0.16 ± 0.00 | 1.4 | 0.15 ± 0.04 | 23.0 |
| C20:4 n6 | 1.20 ± 0.02 | 1.7 | 1.17 ± 0.02 | 1.6 |
| C22:1 | 0.10 ± 0.04 | 4.2 | 0.08 ± 0.01 | 15.9 |
| C20:5 n3 | 1.17 ± 0.01 | 1.2 | 1.14 ± 0.01 | 1.2 |
| C24:0 | 0.07 ± 0.01 | 11.5 | 0.07 ± 0.01 | 11.1 |
| C24:1 | 0.06 ± 0.00 | 7.4 | 0.09 ± 0.02 | 27.3 |
| C22:5 | 1.08 ± 0.01 | 1.3 | 1.06 ± 0.01 | 1.3 |
| C22:6 n3 | 0.30 ± 0.01 | 2.5 | 0.29 ± 0.01 | 3.5 |
| Unreported FA | 10.48 ± 0.88 | 8.4 | 10.28 ± 0.85 | 8.3 |
| Sum FA | 154.85 ± 1.83 | 1.2 | 153.55 ± 1.87 | 1.2 |
| FA | Lamb Chumps | Lamb Loin | Lamb Shoulder | Beef | Venison | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Mean ± STD | RSD | Mean ± STD | RSD | Mean ± STD | RSD | Mean ± STD | RSD | Mean ± STD | RSD | |
| C14:0 | 0.84 ± 0.01 | 1.25 | 0.70 ± 0.01 | 1.47 | 8.36 ± 0.27 | 3.18 | 0.64 ± 0.01 | 1.34 | 0.21 ± 0.01 | 4.38 |
| C14:1 | 0.03 ± 0.00 | 4.08 | 0.02 ± 0.00 | 5.09 | 0.22 ± 0.02 | 8.77 | 0.18 ± 0.00 | 2.53 | 0.06 ± 0.00 | 3.74 |
| C15:0 | 0.13 ± 0.00 | 1.59 | 0.10 ± 0.00 | 2.72 | 1.29 ± 0.04 | 3.39 | 0.13 ± 0.00 | 2.31 | 0.05 ± 0.00 | 5.06 |
| C16:0 | 5.61 ± 0.06 | 1.14 | 5.12 ± 0.04 | 0.69 | 42.21 ± 1.22 | 2.89 | 7.50 ± 0.11 | 1.46 | 1.67 ± 0.04 | 2.69 |
| C16:1 | 0.28 ± 0.01 | 2.40 | 0.24 ± 0.01 | 2.87 | 2.01 ± 0.06 | 3.03 | 0.90 ± 0.01 | 1.17 | 0.32 ± 0.01 | 3.16 |
| C17:0 | 0.30 ± 0.01 | 1.77 | 0.25 ± 0.00 | 1.41 | 2.91 ± 0.09 | 3.07 | 0.31 ± 0.01 | 1.86 | 0.06 ± 0.00 | 5.52 |
| C17:1 | 0.14 ± 0.00 | 1.96 | 0.12 ± 0.00 | 2.28 | 1.01 ± 0.03 | 3.10 | 0.25 ± 0.00 | 1.62 | 0.04 ± 0.00 | 7.28 |
| C18:0 | 4.44 ± 0.05 | 1.12 | 3.88 ± 0.02 | 0.63 | 35.39 ± 0.92 | 2.61 | 4.47 ± 0.06 | 1.33 | 1.54 ± 0.03 | 1.86 |
| C18:1 11t | 0.80 ± 0.02 | 1.89 | 0.69 ± 0.02 | 3.23 | 7.86 ± 0.30 | 3.79 | 0.37 ± 0.01 | 2.09 | 0.12±0.01 | 5.93 |
| C18:1 9c | 7.68 ± 0.09 | 1.21 | 7.34 ± 0.06 | 0.81 | 53.06 ± 1.37 | 2.58 | 12.07 ± 0.15 | 1.26 | 1.61 ± 0.04 | 2.46 |
| C18:1 11c | 0.34 ± 0.00 | 1.34 | 0.29 ± 0.01 | 4.68 | 1.90 ± 0.09 | 4.61 | 0.55 ± 0.01 | 2.71 | 0.37 ± 0.01 | 1.85 |
| C18:2 n6 | 1.59 ± 0.02 | 1.00 | 1.12 ± 0.02 | 1.52 | 4.57 ± 0.18 | 3.96 | 0.68 ± 0.01 | 1.07 | 1.30 ± 0.02 | 1.51 |
| C18:3 n3 | 0.88 ± 0.01 | 1.05 | 0.64 ± 0.01 | 1.04 | 3.67 ± 0.11 | 2.95 | 0.42 ± 0.01 | 2.19 | 0.51 ± 0.01 | 2.19 |
| CLA 9c, 11t | 0.29 ± 0.01 | 2.09 | 0.28 ± 0.01 | 2.66 | 2.40 ± 0.11 | 4.39 | 0.19 ± 0.01 | 4.20 | 0.04 ± 0.00 | 9.07 |
| C20:4 n6 | 0.34 ± 0.00 | 1.00 | 0.27 ± 0.00 | 1.12 | 0.37 ± 0.02 | 4.91 | 0.25 ± 0.00 | 1.56 | 0.49 ± 0.00 | 0.98 |
| C20:5 n3 | 0.24 ± 0.00 | 0.76 | 0.22 ± 0.00 | 1.33 | 0.23 ± 0.02 | 7.01 | 0.18 ± 0.00 | 1.81 | 0.35 ± 0.01 | 1.62 |
| C22:5 | 0.22 ± 0.00 | 1.73 | 0.20 ± 0.00 | 2.15 | 0.39 ± 0.02 | 5.44 | 0.23 ± 0.00 | 1.92 | 0.32 ± 0.01 | 1.59 |
| C22:6 n3 | 0.07 ± 0.00 | 2.60 | 0.07 ± 0.00 | 4.38 | <0.15 | - | 0.04 ± 0.00 | 9.95 | 0.09 ± 0.00 | 5.06 |
| Unreported FA | 2.77 ± 0.12 | 4.34 | 2.26 ± 0.08 | 3.35 | 14.44 ± 0.55 | 3.81 | 2.52 ± 0.07 | 2.85 | 1.86 ± 0.05 | 2.45 |
| SUM FA | 26.97 ± 0.37 | 1.36 | 23.81 ± 0.20 | 0.82 | 182.37 ± 4.95 | 2.71 | 31.86 ± 0.39 | 1.21 | 11.03 ± 0.20 | 1.77 |
| Variables | Levels Used | Optimum Conditions | ||||
|---|---|---|---|---|---|---|
| Temperature (°C) | 60 | 70 | 80 | 100 | 70 | |
| Time (hr) | 1 | 2 | 3 | 2 | ||
| Concentration (% H2SO4 in methanol) | 3 | 5 | 7 | 9 | 5 | |
| Acid and toluene volume (mL) | 2 | 3 | 4 | 5 | 6 | 4 (each) |
| Sample weight (mg) | 100 | 200 | 300 | 400 | 500 | 300 |
| Moisture (%) | 0 | 2.5 | 5 | 7.5 | 0–7.5% | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Agnew, M.P.; Craigie, C.R.; Weralupitiya, G.; Reis, M.M.; Johnson, P.L.; Reis, M.G. Comprehensive Evaluation of Parameters Affecting One-Step Method for Quantitative Analysis of Fatty Acids in Meat. Metabolites 2019, 9, 189. https://doi.org/10.3390/metabo9090189
Agnew MP, Craigie CR, Weralupitiya G, Reis MM, Johnson PL, Reis MG. Comprehensive Evaluation of Parameters Affecting One-Step Method for Quantitative Analysis of Fatty Acids in Meat. Metabolites. 2019; 9(9):189. https://doi.org/10.3390/metabo9090189
Chicago/Turabian StyleAgnew, Michael P., Cameron R. Craigie, Gayani Weralupitiya, Marlon M. Reis, Patricia L. Johnson, and Mariza G. Reis. 2019. "Comprehensive Evaluation of Parameters Affecting One-Step Method for Quantitative Analysis of Fatty Acids in Meat" Metabolites 9, no. 9: 189. https://doi.org/10.3390/metabo9090189