Multi-Organ NMR Metabolomics to Assess In Vivo Overall Metabolic Impact of Cisplatin in Mice
Abstract
1. Introduction
2. Results
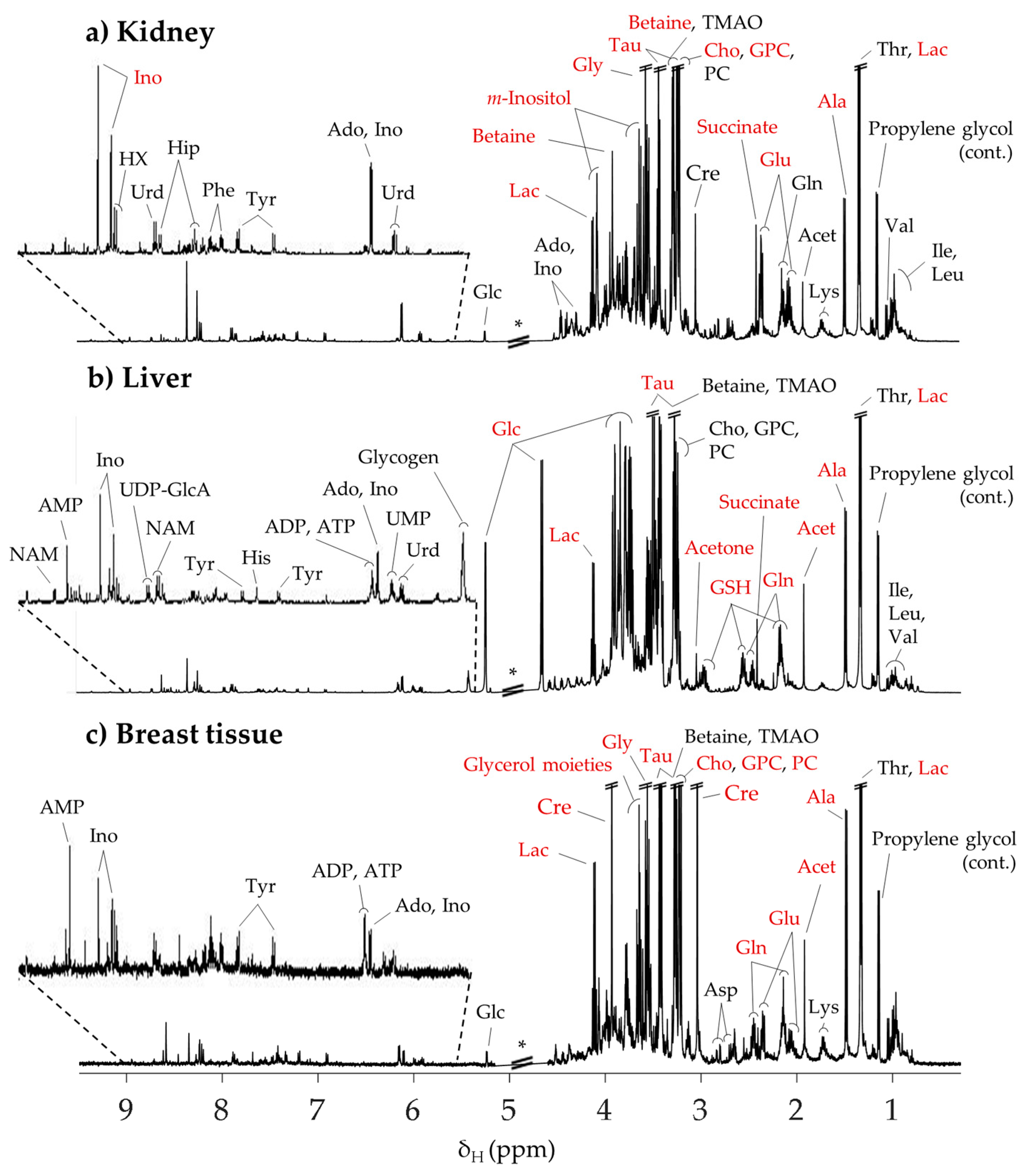
2.1. Typical 1H NMR Spectra of Aqueous Extracts of Mice Kidney, Liver and Breast Tissue
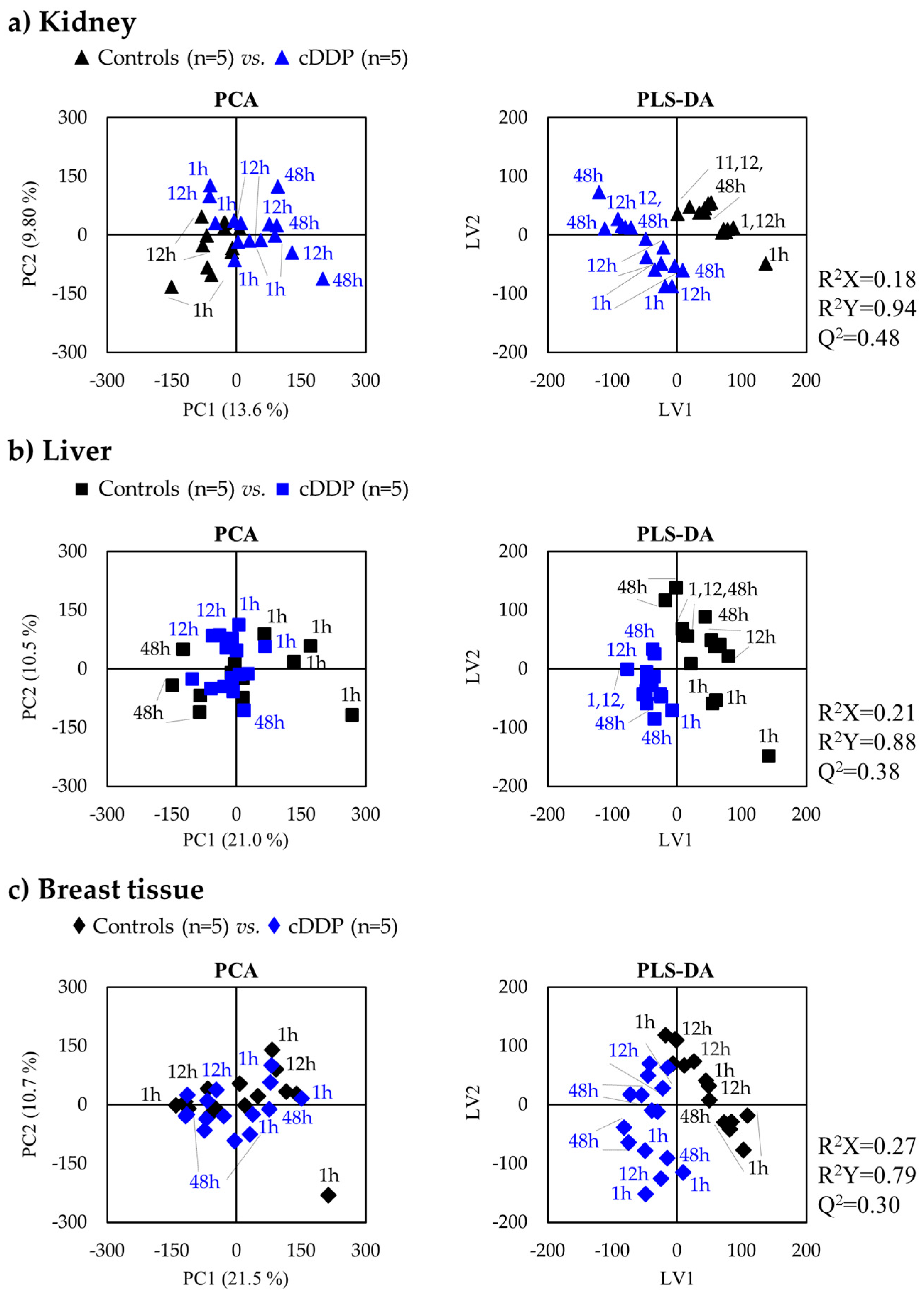
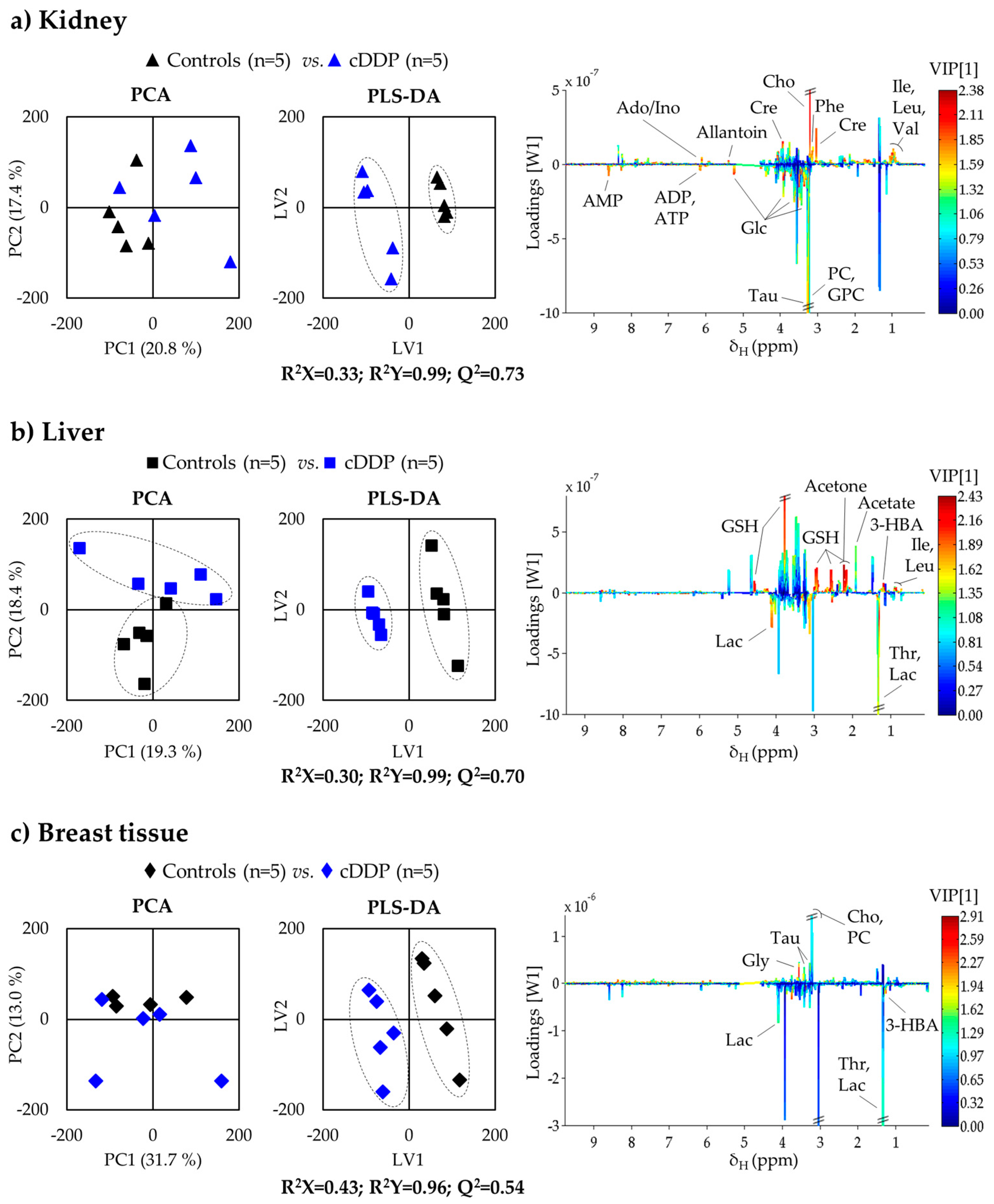
2.2. Impact of Mice Exposure to Cisplatin
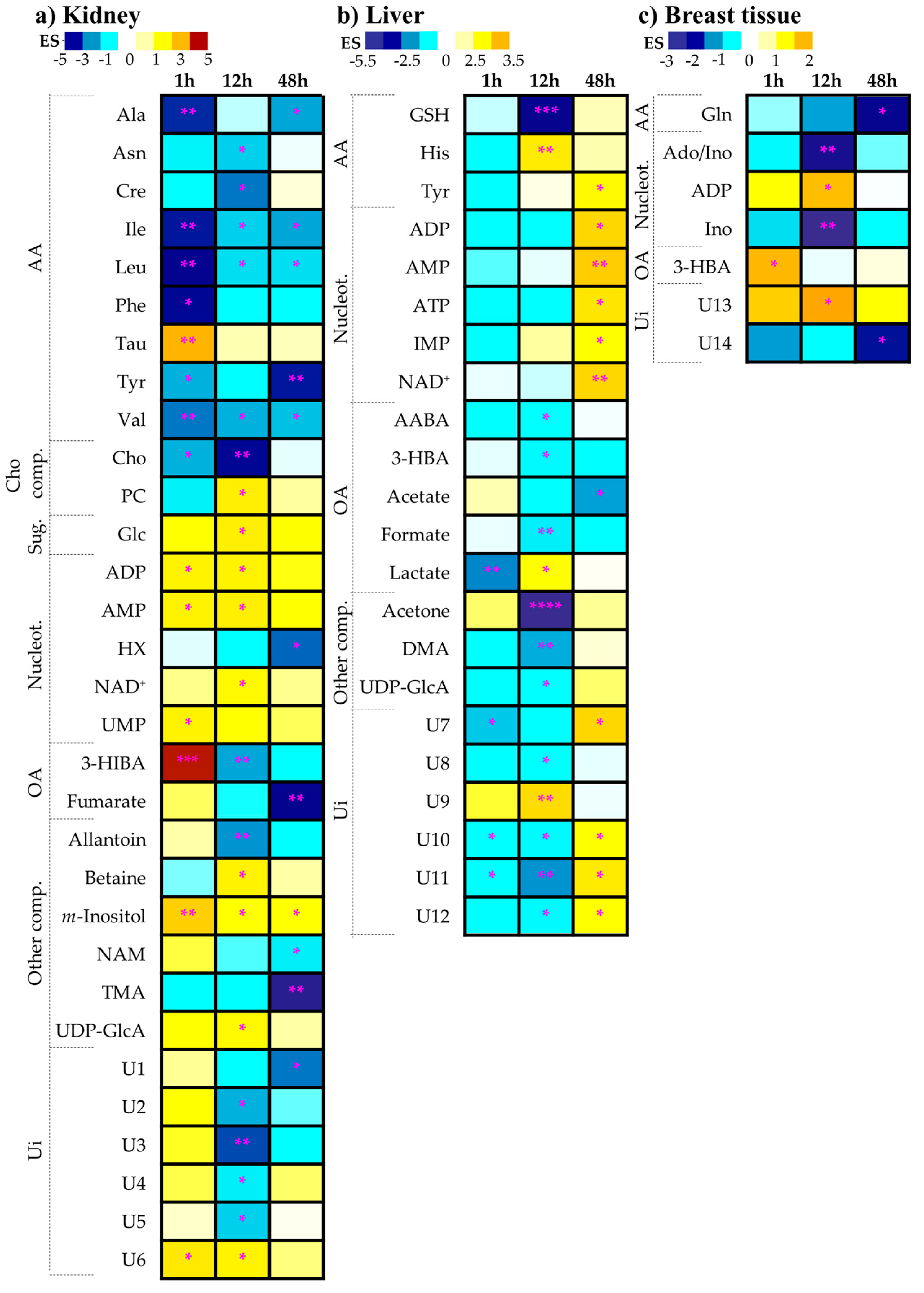
2.1.1. Kidney Metabolic Profiling
2.1.2. Liver Metabolic Profiling
2.1.3. Breast Tissue Metabolic Profiling
3. Discussion
4. Materials and Methods
4.1. Chemicals
4.2. Ethical Considerations
4.3. Animals
4.4. In Vivo Experimental Procedures
4.5. Sample Preparation for NMR
4.6. NMR Measurements
4.7. Data Processing and STATISTICS
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Dasari, S.; Tchounwou, P.B. Cisplatin in cancer therapy: Molecular mechanisms of action. Eur. J. Pharmacol. 2014, 740, 364–378. [Google Scholar] [CrossRef] [PubMed]
- Manohar, S.; Leung, N. Cisplatin nephrotoxicity: A review of the literature. J. Nephrol. 2018, 31, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Irie, M.; Hayakawa, E.; Fujimura, Y.; Honda, Y.; Setoyama, D.; Wariishi, H.; Hyodo, F.; Miura, D. Analysis of spatiotemporal metabolomic dynamics for sensitively monitoring biological alterations in cisplatin-induced acute kidney injury. Biochem. Biophys. Res. Commun. 2018, 496, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Achkar, I.W.; Abdulrahman, N.; Al-Sulaiti, H.; Joseph, J.M.; Uddin, S.; Mraiche, F. Cisplatin based therapy: The role of the mitogen activated protein kinase signaling pathway. J. Transl. Med. 2018, 16. [Google Scholar] [CrossRef]
- Perše, M.; Veceric-Haler, C. Cisplatin-Induced Rodent Model of Kidney Injury: Characteristics and Challenges. BioMed Res. Int. 2018, 2018, 1462802. [Google Scholar] [CrossRef]
- Khoo, S.H.G.; Al-Rubeai, M. Metabolomics as a Complementary Tool in Cell Culture. Biotechnol. Appl. Biochem. 2007, 47, 71–84. [Google Scholar] [CrossRef]
- Cuperlovic-Culf, M.; Barnett, A.D.; Culf, A.S.; Chute, I. Cell Culture Metabolomics: Applications and Future Directions. Drug Discov. Today 2010, 15, 610–621. [Google Scholar] [CrossRef]
- Coen, M.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. NMR-Based Metabolic Profiling and Metabonomic Approaches to Problems in Molecular Toxicology. Chem. Res. Toxicol. 2008, 21, 9–27. [Google Scholar] [CrossRef]
- Robertson, D.G.; Watkins, P.B.; Reily, M.D. Metabolomics in Toxicology: Preclinical and Clinical Applications. Toxicol. Sci. 2011, 120, S146–S170. [Google Scholar] [CrossRef]
- Lee, S.; Kang, S.; Park, S. Comparison of Metabolic Profiles of Normal and Cancer Cells in Response to Cytotoxic Agents. J. Korean Magn. Reson. Soc. 2017, 21, 31–43. [Google Scholar] [CrossRef]
- Duarte, I.F.; Ladeirinha, A.F.; Lamego, I.; Gil, A.M.; Carvalho, L.; Carreira, I.M.; Melo, J.B. Potential Markers of Cisplatin Treatment Response Unveiled by NMR Metabolomics of Human Lung Cells. Mol. Pharm. 2013, 10, 4242–4251. [Google Scholar] [CrossRef] [PubMed]
- Maria, R.M.; Altei, W.F.; Selistre-de-Araujo, H.S.; Colnago, L.A. Impact of chemotherapy on metabolic reprogramming: Characterization of the metabolic profile of breast cancer MDA-MB-231 cells using H-1 HR-MAS NMR spectroscopy. J. Pharm. Biomed Anal. 2017, 146, 324–328. [Google Scholar] [CrossRef] [PubMed]
- Mandujano-Tinoco, E.A.; Gallardo-Perez, J.C.; Marin-Hernandez, A.; Moreno-Sanchez, R.; Rodriguez-Enriquez, S. Anti-mitochondrial therapy in human breast cancer multi-cellular spheroids. Biochim. Biophys. Acta 2013, 1833, 541–551. [Google Scholar] [CrossRef] [PubMed]
- Lamego, I.; Marques, M.P.M.; Duarte, I.F.; Martins, A.S.; Oliveira, H.; Gil, A.M. Impact of the Pd(2)Spermine Chelate on Osteosarcoma Metabolism: An NMR Metabolomics Study. J. Proteome Res. 2017, 16, 1773–1783. [Google Scholar] [CrossRef] [PubMed]
- Duarte, I.F.; Lamego, I.; Marques, J.; Marques, M.P.M.; Blaise, B.J.; Gil, A.M. Nuclear Magnetic Resonance (NMR) Study of the Effect of Cisplatin on the Metabolic Profile of MG-63 Osteosarcoma Cells. J. Proteome Res. 2010, 9, 5877–5886. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.Y.; Wilson, M.; Mirbahai, L.; McConville, C.; Arvanitis, T.N.; Griffin, J.L.; Kauppinen, R.A.; Peet, A.C. In Vitro Metabonomic Study Detects Increases in UDP-GlcNAc and UDP-GalNAc, as Early Phase Markers of Cisplatin Treatment Response in Brain Tumor Cells. J. Proteome Res. 2011, 10, 3493–3500. [Google Scholar] [CrossRef]
- Alonezi, S.; Tusiimire, J.; Wallace, J.; Dufton, M.J.; Parkinson, J.A.; Young, L.C.; Clements, C.J.; Park, J.K.; Jeon, J.W.; Ferro, V.A.; et al. Metabolomic Profiling of the Synergistic Effects of Melittin in Combination with Cisplatin on Ovarian Cancer Cells. Metabolites 2017, 7, 14. [Google Scholar] [CrossRef]
- Poisson, L.M.; Munkarah, A.; Madi, H.; Datta, I.; Hensley-Alford, S.; Tebbe, C.; Buekers, T.; Giri, S.; Rattan, R. A metabolomic approach to identifying platinum resistance in ovarian cancer. J. Ovarian Res. 2015, 8, 13. [Google Scholar] [CrossRef]
- De Castro, F.; Benedetti, M.; Antonaci, G.; Del Coco, L.; De Pascali, S.A.; Muscella, A.; Marsigliante, S.; Fanizzi, F.P. Response of Cisplatin Resistant Skov-3 Cells to [Pt(O,O ‘-Acac)(gamma-Acac)(DMS)] Treatment Revealed by a Metabolomic H-1-NMR Study. Molecules 2018, 23, 2301. [Google Scholar] [CrossRef]
- Obrist, F.; Michels, J.; Durand, S.; Chery, A.; Pol, J.; Levesque, S.; Joseph, A.; Astesana, V.; Pietrocola, F.; Wu, G.S.; et al. Metabolic vulnerability of cisplatin-resistant cancers. EMBO J. 2018, 37. [Google Scholar] [CrossRef]
- Zong, L.; Pi, Z.F.; Liu, S.; Liu, Z.Q.; Song, F.R. Metabolomics analysis of multidrug-resistant breast cancer cells in vitro using methyl-tert-butyl ether method. RSC Adv. 2018, 8, 15831–15841. [Google Scholar] [CrossRef]
- Chao, W.-W.; Cheng, Y.-W.; Chen, Y.-R.; Lee, S.-H.; Cjou, C.-Y.; Shyur, L.-F. Phyto-sesquiterpene lactone deoxyelephantopin and cisplatin synergistically suppress lung metastasis of B16 melanoma in mice with reduced nephrotoxicity. Phytomedicine 2019, 56, 194–206. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.H.; Wang, Z.J.; Liu, X.H.; Duan, J.C.; Feng, G.S.; Yin, Y.X.; Gu, J.; Chen, Z.L.; Gao, S.G.; Bai, H.; et al. Prediction of Chemotherapeutic Efficacy in Non-Small Cell Lung Cancer by Serum Metabolomic Profiling. Clin. Cancer Res. 2018, 24, 2100–2109. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Zhou, Y.P.; Geng, H.; Song, D.D.; Tang, J.; Zhu, X.M.; Yu, D.; Hu, S.; Cui, Y.F. Serum Metabolic Profile Alteration Reveals Response to Platinum-Based Combination Chemotherapy for Lung Cancer: Sensitive Patients Distinguished from Insensitive ones. Sci. Rep. 2017, 7, 17524. [Google Scholar] [CrossRef]
- Tandberg, D.J.; Holt, T.; Kelsey, C.R. Plasma Metabolites and Risk of Radiation-induced Esophagitis: A Secondary Analysis from a Prospective Study. Anticancer Res. 2017, 37, 719–725. [Google Scholar] [CrossRef]
- Doskocz, M.; Marchewka, Z.; Jez, M.; Passowicz-Muszynska, E.; Dlugosz, A. Preliminary Study on J-Resolved NMR Method Usability for Toxic Kidney’s Injury Assessment. Adv. Clin. Exp. Med. 2015, 24, 629–635. [Google Scholar] [CrossRef]
- Liu, S.; Wang, W.; Zhou, X.; Gu, R.; Ding, Z. Dose responsive effects of cisplatin in L02 cells using NMR-based metabolomics. Environ. Toxicol. Pharmacol. 2014, 37, 150–157. [Google Scholar] [CrossRef]
- Wilmes, A.; Bielow, C.; Ranninger, C.; Bellwon, P.; Aschauer, L.; Limonciel, A.; Chassaigne, H.; Kristl, T.; Aiche, S.; Huber, C.G.; et al. Mechanism of cisplatin proximal tubule toxicity revealed by integrating transcriptomics, proteomics, metabolomics and biokinetics. Toxicol. In Vitro 2015, 30, 117–127. [Google Scholar] [CrossRef]
- Heyman, S.N.; Rosen, S.; Rosenberger, C. Animal models of renal dysfunction: Acute kidney injury. Expert Opin. Drug Discov. 2009, 4, 629–641. [Google Scholar] [CrossRef]
- Portilla, D.; Li, S.; Nagothu, K.K.; Megyesi, J.; Kaissling, B.; Schnackenberg, L.; Safirstein, R.L.; Beger, R.D. Metabolomic study of cisplatin-induced nephrotoxicity. Kidney Int. 2006, 69, 2194–2204. [Google Scholar] [CrossRef]
- Zhang, P.; Chen, J.; Wang, Y.; Huang, Y.; Tian, Y.; Zhang, Z.; Xu, F. Discovery of Potential Biomarkers with Dose- and Time-Dependence in Cisplatin Induced Nephrotoxicity Using Metabolomics Integrated with a Principal Component-Based Area Calculation Strategy. Chem. Res. Toxicol. 2016, 29, 776–783. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, X.; Zhou, H.; Fan, S.; Wang, Y.; Zhang, L.; Ju, L.; Wu, X.; Wu, H.; Zhang, Y. Metabonomics study on nephrotoxicity induced by intraperitoneal and intravenous cisplatin administration using rapid resolution liquid chromatography coupled with quadrupole-time-of- flight mass spectrometry (RRLC-Q-TOF-MS). RSC Adv. 2014, 4, 8260. [Google Scholar] [CrossRef]
- Zhang, P.; Li, W.; Chen, J.; Li, R.; Zhang, Z.; Huang, Y.; Xu, F. Branched-Chain Amino Acids as Predictors for Individual Differences of Cisplatin Nephrotoxicity in Rats: A Pharmacometabonomics Study. J. Proteome Res. 2017, 16, 1753–1762. [Google Scholar] [CrossRef] [PubMed]
- Uehara, T.; Horinouchid, A.; Morikawaa, Y.; Tonomuraa, Y.; Minamie, K.; Onob, A.; Yamatec, J.; Yamadab, H.; Ohnof, Y.; Urushidanib, T. Identification of metabolomic biomarkers for drug-induced acute kidney injury in rats. J. Appl. Toxicol. 2014, 34, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- Ezaki, T.; Nishiumi, S.; Azumaa, T.; Yoshida, M. Metabolomics for the early detection of cisplatin induced nephrotoxicity. Toxicol. Res. 2017, 6, 843. [Google Scholar] [CrossRef] [PubMed]
- Xu, E.Y.; Perlina, A.; Vu, H.; Troth, S.P.; Brennan, R.J.; Aslamkhan, A.G.; Xu, Q. Integrated Pathway Analysis of Rat Urine Metabolic Profiles and Kidney Transcriptomic Profiles To Elucidate the Systems Toxicology of Model Nephrotoxicants. Chem. Res. Toxicol. 2008, 21, 1548–1561. [Google Scholar] [CrossRef]
- Ryu, S.H.; Lee, J.D.; Kim, J.W.; Kim, S.; Kim, K.B. 1H NMR toxicometabolomics following cisplatin-induced nephrotoxicity in male rats. J. Toxicol. Sci. 2019, 44, 57–71. [Google Scholar] [CrossRef]
- Espandiari, P.; Rosenzweig, B.; Zhang, J.; Zhou, Y.; Schnackenberg, L.; Vaidya, V.S.; Goering, P.L.; Brown, R.P.; Bonventre, V.; Mahjoob, K.; et al. Age-related differences in susceptibility to cisplatin-induced renal toxicity. J. Appl. Toxicol. 2009, 30, 172–182. [Google Scholar] [CrossRef]
- Won, A.J.; Kim, S.; Kim, Y.G.; Kim, K.-B.; Choi, W.S.; Kacew, S.; Kim, K.S.; Jung, J.H.; Lee, B.M.; Kimb, S.; et al. Discovery of urinary metabolomic biomarkers for early detection of acute kidney injury. Mol. Biosyst. 2016, 12, 133. [Google Scholar] [CrossRef]
- Portilla, D.; Schnackenberg, L.; Beger, R.D. Metabolomics as an Extension of Proteomic Analysis: Study of acute kidney injury. Semin. Nephrol. 2007, 27, 609–620. [Google Scholar] [CrossRef]
- Kwon, H.K.; Kim, M.; Wen, H.; Kang, S.; Yang, H.-J.; Choi, M.-J.; Lee, H.S.; Choi, D.; Park, I.S.; Suh, I.J.; et al. Predicting idiopathic toxicity of cisplatin by a pharmacometabonomic approach. Kidney Int. 2011, 79, 529–537. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Boudonck, K.J.; Mitechell, M.W.; Német, L.; Keresztes, L.; Nyska, A.; Shinar, D.; Rosenstock, M. Discovery of Metabolomics Biomarkers for Early Detection of Nephrotoxicity. Toxicol. Pathol. 2009, 37, 280–292. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Chen, J.-Q.; Huang, W.-Q.; Li, W.; Huang, Y.; Zhang, Z.-J.; Xu, F.-G. Renal Medulla is More Sensitive to Cisplatin than Cortex Revealed by Untargeted Mass Spectrometry-Based Metabolomics in Rats. Sci. Rep. 2017, 7, 44804. [Google Scholar] [CrossRef] [PubMed]
- Zhong, F.; Liu, X.; Zhou, Q.; Hao, X.; Lu, Y.; Guo, S.; Wang, W.; Lin, D.; Chen, N. 1H NMR spectroscopy analysis of metabolites in the kidneys provides new insight into pathophysiological mechanisms: Applications for treatment with Cordyceps sinensis. Nephrol. Dial. Transplant. 2012, 27, 556–565. [Google Scholar] [CrossRef]
- Ling, Y.S.; Liang, H.-J.; Chung, M.-H.; Lin, M.-H.; Lin, C.-Y. NMR- and MS-based metabolomics: Various organ responses following naphthalene intervention. Mol. BioSyst. 2014, 10, 1918–1931. [Google Scholar] [CrossRef]
- Zheng, H.; Ni, Z.; Cai, A.; Zhang, X.; Chen, J.; Shu, Q.; Gao, H. Balancing metabolome coverage and reproducibility for untargeted NMR-based metabolic profiling in tissue samples through mixture design methods. Anal. Bioanal. Chem. 2018, 410, 7783–7792. [Google Scholar] [CrossRef]
- Jiang, L.; Si, Z.-H.; Li, M.-H.; Zhao, H.; Fu, Y.-H.; Xing, Y.-X.; Hong, W.; Ruan, L.Y.; Li, P.-M.; Wang, J.-S. 1H NMR-based metabolomics study of liver damage induced by ginkgolic acid (15:1) in mice. J. Pharm. Biomed. Anal. 2017, 136, 44–54. [Google Scholar] [CrossRef]
- Chen, M.; Zheng, H.; Xu, M.; Zhao, L.; Zhang, Q.; Song, J.; Zhao, Z.; Lu, S.; Weng, Q.; Wu, X.; et al. Changes in hepatic metabolic profile during the evolution of STZ-induced diabetic rats via an 1H NMR-based metabonomic investigation. Biosci. Rep. 2019, 39, 12. [Google Scholar] [CrossRef]
- Feng, X.; Xu, H.; Chen, J.-F.; Ruan, L.-Y.; Zhao, W.-L.; Meng, H.-H.; Liu, W.-Y.; Zhao, W.-L.; Zheng, Q.; Liu, Z.-C.; et al. Potential hepatoxicity risk of the shell of Herpetospermum caudigerum Wall in rats based on 1H-NMR metabonomicsXin. J. Pharm. Biomed. Anal. 2019, 176, 112800. [Google Scholar] [CrossRef]
- Berben, L.; Sereika, S.M.; Engberg, S. Effect size estimation: Methods and examples. Int. J. Nurs. Stud. 2012, 49, 1039–1047. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Guldris, S.C.; Parra, E.G.; Amenós, A.C. Gut microbiota in chronic kidney disease. Nefrologia 2017, 37, 9–19. [Google Scholar] [CrossRef]
- Kikenny, C.; Browne, W.J.; Cuthill, I.C.; Emerson, M.; Altman, D.G. Improving Bioscience Research Reporting: The ARRIVE Guidelines for Reporting Animal Research. PLoS Biol. 2010, 8, e1000412. [Google Scholar] [CrossRef]
- Beckonert, O.; Keun, H.C.; Ebbels, T.M.; Bundy, J.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat. Protoc. 2007, 2, 2692–2703. [Google Scholar] [CrossRef]
- Lin, C.Y.; Wu, H.; Tjeerdema, R.S.; Viant, M.R. Evaluation of metabolite extraction strategies from tissue samples using NMR metabolomics. Metabolomics 2007, 3, 55–67. [Google Scholar] [CrossRef]
- Le Belle, J.E.; Harris, N.G.; Williams, S.R.; Bhakoo, K.K. A comparison of cell and tissue extraction techniques using high-resolution 1H-NMR spectroscopy. NMR Biomed. 2002, 15, 37–44. [Google Scholar] [CrossRef]
- Wu, H.; Southam, A.D.; Hines, A.; Viant, M.R. High-throughput tissue extraction protocol for NMR- and MS-based metabolomics. Anal. Biochem. 2008, 372, 204–212. [Google Scholar] [CrossRef]
- Wishart, D.S.; Feunang, Y.D.; Marcu, A.; Guo, A.C.; Liang, K.; Vázquez-Fresno, R.; Sajed, T.; Johnson, D.; Li, C.; Karu, N.; et al. HMDB 4.0—The Human Metabolome Database for 2018. Nucleic Acids Res. 2018, 46, D608–D617. [Google Scholar] [CrossRef]
| Metabolite | δH ppm (multiplicity) | 1 h | 12 h | 48 h | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ES | ± | Error | p-Value | ES | ± | Error | p-Value | ES | ± | Error | p-Value | ||
| 3-HIBA | 1.09 (d) | 4.7 | ± | 2.5 | 1.9 × 10−4 a | −2.2 | ± | 1.6 | 9.5 × 10−3 | - | - | - | - |
| ADP c | 8.28 (s) | 1.8 | ± | 1.6 | 2.9 × 10−2 | 1.8 | ± | 1.5 | 1.6 × 10−2 | - | - | - | - |
| Ala b | 1.48 (d) | −3.0 | ± | 1.9 | 9.5 × 10−3 a | - | - | - | - | −2.2 | ± | 1.7 | 1.1 × 10−2 |
| Allantoin | 5.39 (s) | - | - | - | - | −2.3 | ± | 1.6 | 9.3 × 10−3 | - | - | - | - |
| AMP | 8.61 (s) | 1.8 | ± | 1.5 | 3.3 × 10−2 | 1.9 | ± | 1.5 | 2.8 × 10−2 | - | - | - | - |
| Asn b, c | 2.96 (m) | - | - | - | - | −1.9 | ± | 1.5 | 2.3 × 10−2 | - | - | - | - |
| Betaine | 3.90 (s) | - | - | - | - | 1.8 | ± | 1.5 | 3.8 × 10−2 | - | - | - | - |
| Cho | 3.21 (s) | −2.1 | ± | 1.6 | 1.6 × 10−2 | −3.2 | ± | 1.9 | 2.7 × 10−3 | - | - | - | - |
| Cre c | 3.93 (s) | - | - | - | - | −2.5 | ± | 1.6 | 1.1 × 10−2 | - | - | - | - |
| Fumarate | 6.52 (s) | - | - | - | - | - | - | - | - | −3.5 | ± | 2.1 | 2.7 × 10−3 a |
| Glc b | 5.23 (d) | - | - | - | - | 1.9 | ± | 1.5 | 2.7 × 10−2 | - | - | - | - |
| Hypoxanthine b | 8.21 (s) | - | - | - | - | - | - | - | - | −2.6 | ± | 1.8 | 3.3 × 10−2 |
| Ile b, c | 1.01 (d) | −3.1 | ± | 1.9 | 4.1 × 10−3 a | −1.9 | ± | 1.5 | 3.3 × 10−2 | −2.2 | ± | 1.7 | 1.5 × 10−2 |
| Leu b, c | 0.96 (t) | −3.5 | ± | 2.1 | 1.2 × 10−3 a | −1.8 | ± | 1.5 | 3.0 × 10−2 | −1.8 | ± | 1.6 | 2.5 × 10−2 |
| m-Inositol | 3.62 (t) | 2.5 | ± | 1.7 | 6.8 × 10−3 a | 1.5 | ± | 1.4 | 4.4 × 10−2 | 1.6 | ± | 1.5 | 3.2 × 10−2 |
| Niacinamide b | 7.60 (dd) | - | - | - | - | - | - | - | - | −1.7 | ± | 1.5 | 3.6 × 10−2 |
| PC | 3.22 (s) | - | - | - | - | 1.9 | ± | 1.5 | 1.7 × 10−2 | - | - | - | - |
| Phe b,c | 7.33 (d) | −3.2 | ± | 2.0 | 1.6 × 10−2 | - | - | - | - | - | - | - | - |
| Tau | 3.42 (t) | 3.0 | ± | 1.9 | 3.2 × 10−3 a | - | - | - | - | - | - | - | - |
| TMA | 2.89 (s) | - | - | - | - | - | - | - | - | -4.3 | ± | 2.4 | 1.1 × 10−3 a |
| Tyr b, c | 6.90 (d) | −2.1 | ± | 1.6 | 1.6 × 10−2 | - | - | - | - | -3.1 | ± | 1.9 | 2.5 × 10−3 a |
| NAD+ c | 8.43 (s) | - | - | - | - | 1.7 | ± | 1.4 | 1.6 × 10−2 | - | - | - | - |
| UDP-GlcA | 7.95 (d) | - | - | - | - | 1.7 | ± | 1.4 | 3.0 × 10−2 | - | - | - | - |
| UMP c | 5.99 (m) | 1.8 | ± | 1.6 | 2.8 × 10−2 | - | - | - | - | - | - | - | - |
| Val b, c | 1.05 (d) | −2.5 | ± | 1.7 | 8.9 × 10−3 a | −2.1 | ± | 1.5 | 2.0 × 10−2 | −2.0 | ± | 1.6 | 2.1 × 10−2 |
| U1 | 0.89 (t) | - | - | - | - | - | - | - | - | −2.5 | ± | 1.8 | 1.6 × 10−2 |
| U2 | 0.93 (s) | - | - | - | - | −2.1 | ± | 1.6 | 1.2 × 10−2 | - | - | - | - |
| U3 | 1.62 (d) | - | - | - | - | −2.8 | ± | 1.7 | 5.0 × 10−3 | - | - | - | - |
| U4 | 2.92 (s) | - | - | - | - | −1.7 | ± | 1.4 | 3.5 × 10−2 | - | - | - | - |
| U5 | 3.15 (s) | - | - | - | - | −1.9 | ± | 1.5 | 3.2 × 10−2 | - | - | - | - |
| U6 | 3.35 (s) | 2.0 | ± | 1.6 | 1.8 × 10−2 | 1.8 | ± | 1.5 | 2.2 × 10−2 | - | - | - | - |
| Metabolite | δH ppm (multiplicity) | 1 h | 12 h | 48 h | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ES | ± | Error | p-Value | ES | ± | Error | p-Value | ES | ± | Error | p-Value | ||
| 2-aminobutyrate † | 0.80 (t) | - | - | - | - | −1.6 | ± | 1.4 | 4.4 × 10−2 | - | - | - | - |
| 3-HBA | 1.20 (d) | - | - | - | - | −1.8 | ± | 1.5 | 3.1 × 10−2 | - | - | - | - |
| Acetate b | 1.92 (s) | - | - | - | - | - | - | - | - | −2.5 | ± | 1.7 | 2.0 × 10−2 |
| Acetone | 2.24 (s) | - | - | - | - | −5.1 | ± | 2.6 | 5.2 × 10−5a | - | - | - | - |
| ADP | 8.54 (s) | - | - | - | - | - | - | - | - | 2.6 | ± | 1.8 | 1.6 × 10−2 |
| AMP b | 4.51 (dd) | - | - | - | - | - | - | - | - | 2.8 | ± | 1.8 | 3.8 × 10−3 |
| ATP b | 8.52 (s) | - | - | - | - | - | - | - | - | 2.3 | ± | 1.7 | 1.6 × 10−2 |
| DMA | 2.73 (s) | - | - | - | - | −2.4 | ± | 1.6 | 6.0 × 10−3 a | - | - | - | - |
| Formate | 8.46 (s) | - | - | - | - | −1.9 | ± | 1.5 | 7.9 × 10−3 a | - | - | - | - |
| GSH | 2.55 (m) | - | - | - | - | −4.0 | ± | 2.1 | 2.7 × 10−4 a | - | - | - | - |
| His | 7.08 (s) | - | - | - | - | 2.2 | ± | 1.6 | 9.110−3 | - | - | - | - |
| IMP | 8.58 (s) | - | - | - | - | - | - | - | - | 1.7 | ± | 1.5 | 3.2 × 10−2 |
| Lactate b | 4.10 (q) | −2.7 | ± | 1.7 | 6.9 × 10−3 | 1.7 | ± | 1.5 | 3.0 × 10−2 | - | - | - | - |
| NAD+ | 8.43 (s) | - | - | - | - | - | - | - | - | 2.6 | ± | 1.8 | 9.9 × 10−3 |
| Tyr | 6.90 (d) | - | - | - | - | - | - | - | - | 1.9 | ± | 1.6 | 2.3 × 10−2 |
| UDP-GlcA | 7.95 (d) | - | - | - | - | -1.7 | ± | 1.4 | 3.0 × 10−2 | - | - | - | - |
| U7 | 0.73 (s) | −2.2 | ± | 1.6 | 1.6 × 10−2 | - | - | - | - | 2.6 | ± | 1.8 | 1.6 × 10−2 |
| U8 | 0.85 (t) | - | - | - | - | −1.6 | ± | 1.4 | 1.6 × 10−2 | - | - | - | - |
| U9 | 3.10 (d) | - | - | - | - | 2.5 | ± | 1.7 | 9.8 × 10−3 a | - | - | - | - |
| U10 | 4.31 (d) | −1.6 | ± | 1.4 | 4.7 × 10−2 | −1.8 | ± | 1.5 | 2.1 × 10−2 | 1.7 | ± | 1.5 | 3.3 × 10−2 |
| U11 | 6.18 (s) | −1.7 | ± | 1.5 | 3.6 × 10−2 | −2.6 | ± | 1.7 | 7.8 × 10−3 a | 2.2 | ± | 1.7 | 2.3 × 10−2 |
| U12 | 8.28 (br) | - | - | - | - | −1.8 | ± | 1.5 | 3.5 × 10−2 | 1.8 | ± | 1.5 | 4.2 × 10−2 |
| Metabolite | δH ppm (multiplicity) | 1 h | 12 h | 48 h | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ES | ± | Error | p-value | ES | ± | Error | p-value | ES | ± | Error | p-value | ||
| 3-HBA | 1.20 (d) | 1.8 | ± | 1.5 | 4.3 × 10−2 | - | - | - | - | - | - | - | - |
| Ado/Ino | 6.10 (d) | - | - | - | - | −2.4 | ± | 1.6 | 6.1 × 10−3a | - | - | - | - |
| ADP | 8.54 (s) | - | - | - | - | 1.7 | ± | 1.4 | 3.1 × 10−2 a | - | - | - | - |
| Gln | 2.45 (m) | - | - | - | - | - | - | - | - | −2.2 | ± | 1.6 | 1.2 × 10−2 |
| Ino | 8.35 (s) | - | - | - | - | −2.8 | ± | 1.7 | 2.4 × 10−3 a | - | - | - | - |
| U13 | 1.25 (s) | - | - | - | - | 2.0 | ± | 1.5 | 2.6 × 10−2 a | - | - | - | - |
| U14 | 7.68 (s) | - | - | - | - | - | - | - | - | −1.9 | ± | 1.6 | 3.1 × 10−2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carneiro, T.J.; Araújo, R.; Vojtek, M.; Gonçalves-Monteiro, S.; Diniz, C.; Batista de Carvalho, A.L.M.; Marques, M.P.M.; Gil, A.M. Multi-Organ NMR Metabolomics to Assess In Vivo Overall Metabolic Impact of Cisplatin in Mice. Metabolites 2019, 9, 279. https://doi.org/10.3390/metabo9110279
Carneiro TJ, Araújo R, Vojtek M, Gonçalves-Monteiro S, Diniz C, Batista de Carvalho ALM, Marques MPM, Gil AM. Multi-Organ NMR Metabolomics to Assess In Vivo Overall Metabolic Impact of Cisplatin in Mice. Metabolites. 2019; 9(11):279. https://doi.org/10.3390/metabo9110279
Chicago/Turabian StyleCarneiro, Tatiana J., Rita Araújo, Martin Vojtek, Salomé Gonçalves-Monteiro, Carmen Diniz, Ana L.M. Batista de Carvalho, Maria Paula M. Marques, and Ana M. Gil. 2019. "Multi-Organ NMR Metabolomics to Assess In Vivo Overall Metabolic Impact of Cisplatin in Mice" Metabolites 9, no. 11: 279. https://doi.org/10.3390/metabo9110279
APA StyleCarneiro, T. J., Araújo, R., Vojtek, M., Gonçalves-Monteiro, S., Diniz, C., Batista de Carvalho, A. L. M., Marques, M. P. M., & Gil, A. M. (2019). Multi-Organ NMR Metabolomics to Assess In Vivo Overall Metabolic Impact of Cisplatin in Mice. Metabolites, 9(11), 279. https://doi.org/10.3390/metabo9110279








