Complex Mixture Analysis of Organic Compounds in Yogurt by NMR Spectroscopy
Abstract
:1. Introduction
2. Results and Discussion
2.1. 1H NMR Spectra of Yogurt
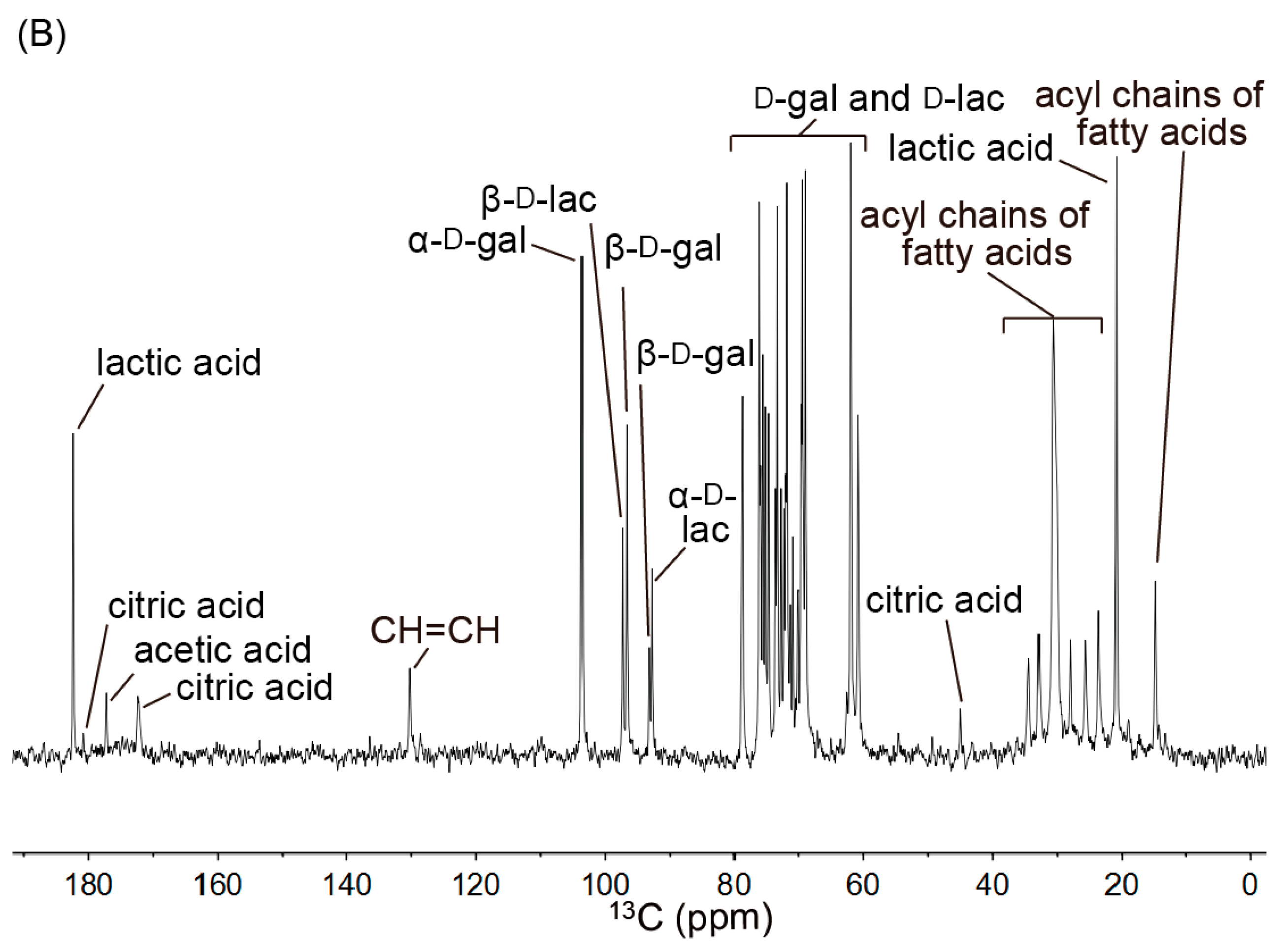
2.2. 13C NMR Spectra of Yogurt
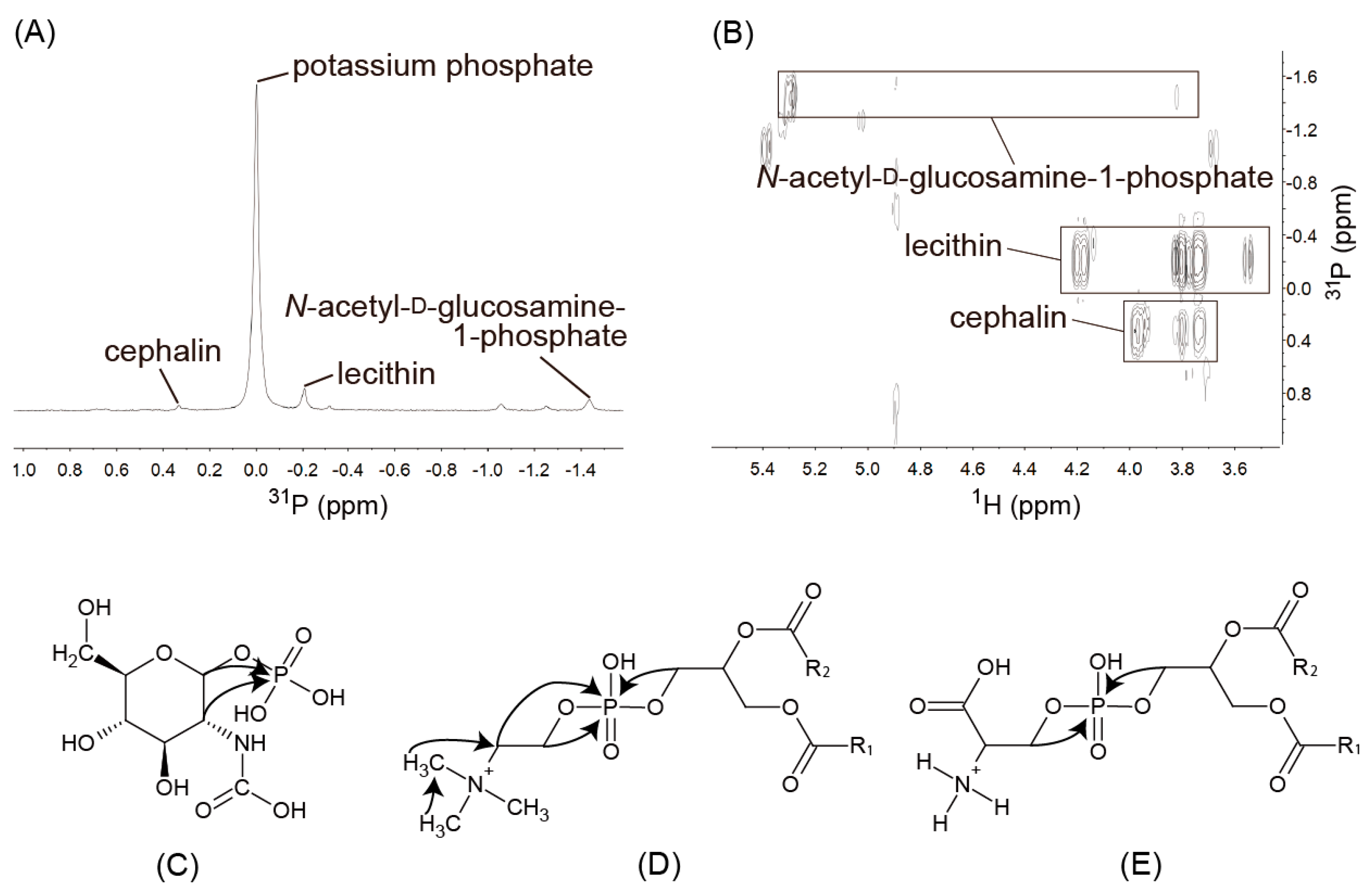
2.3. Identification of Several Components with 2D NMR Spectra
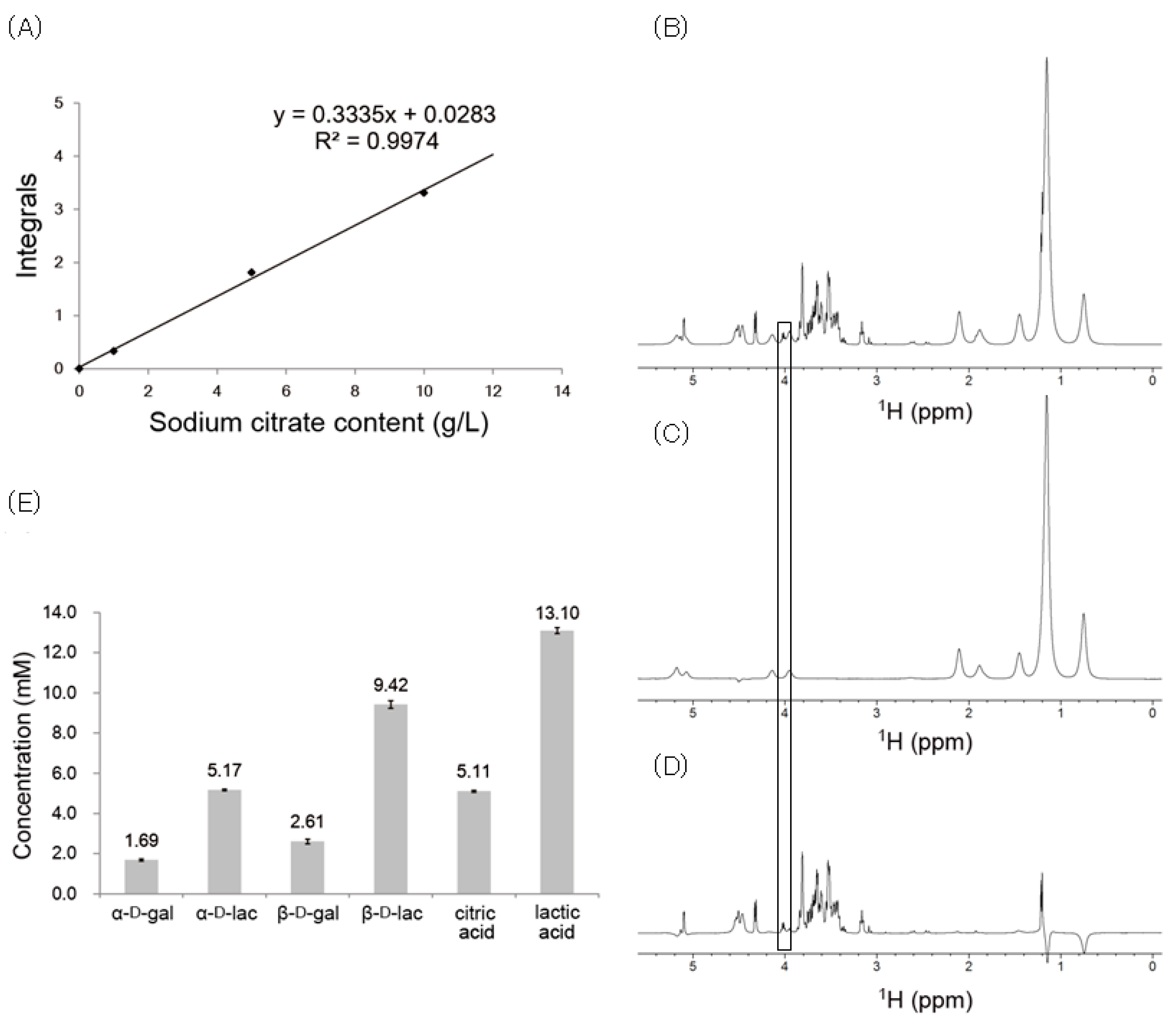
2.4. Concentrations of Yogurt Components
3. Experimental Section
3.1. Materials and Sample Preparation
3.2. NMR Spectroscopy
3.3. NMR Signal Assignments
3.4. Quantification of Yogurt Components
3.5. Diffusion Ordered Spectroscopy (DOSY)
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Gowda, G.N.; Raftery, D. Can NMR solve some significant challenges in metabolomics? J. Magn. Reson. 2015, 260, 144–160. [Google Scholar] [CrossRef] [PubMed]
- Monakhova, Y.B.; Ruge, W.; Kuballa, T.; Ilse, M.; Winkelmann, O.; Diehl, B.; Lachenmeier, D.W. Rapid approach to identify the presence of Arabica and Robusta species in coffee using 1H NMR spectroscopy. Food Chem. 2015, 182, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Hohmann, M.; Koospal, V.; Bauer-Christoph, C.; Christoph, N.; Wachter, H.; Diehl, B.; Holzgrabe, U. Quantitative 1H NMR analysis of egg yolk, alcohol, and total sugar content in egg liqueurs. J. Agric. Food Chem. 2015, 63, 4112–4119. [Google Scholar] [CrossRef] [PubMed]
- Hu, F.; Furihata, K.; Kato, Y.; Tanokura, M. Nondestructive quantification of organic compounds in whole milk without pretreatment by two-dimensional NMR spectroscopy. J. Agric. Food Chem. 2007, 55, 4307. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Teng, Z.; Parkin, K.L.; Wang, Q.; Zhang, Q.; Luo, W.; Zhao, M. Identification of bioactive metabolites dihydrocanadensolide, kojic acid, and vanillic acid in soy sauce using GC-MS, NMR spectroscopy, and single-crystal X-ray diffraction. J. Agric. Food Chem. 2014, 62, 8392–8401. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.; Furihata, K.; Miyakawa, T.; Tanokura, M. A pilot study of NMR-based sensory prediction of roasted coffee bean extracts. Food Chem. 2014, 152, 363–369. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Fresno, R.; Llorach, R.; Urpi-Sarda, M.; Khymenets, O.; Bulló, M.; Corella, D.; Andres-Lacueva, C. An NMR metabolomics approach reveals a combined-biomarkers model in a wine interventional trial with validation in free-living individuals of the PREDIMED study. Metabolomics 2015, 11, 797–806. [Google Scholar] [CrossRef]
- Koda, M.; Furihata, K.; Wei, F.; Miyakawa, T.; Tanokura, M. Metabolic discrimination of mango juice from various cultivars by band-selective NMR spectroscopy. J. Agric. Food Chem. 2012, 60, 1158–1166. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Song, Y.; Jing, W.; Wang, Y.; Yang, X.; Liu, D. Simultaneous determination of caffeine, gallic acid, theanine, (–)-epigallocatechin and (–)-epigallocatechin-3-gallate in green tea using quantitative 1H-NMR spectroscopy. Anal. Methods 2014, 6, 907–914. [Google Scholar] [CrossRef]
- Hu, F.; Furihata, K.; Ito-Ishida, M.; Kaminogawa, S.; Tanokura, M. Nondestructive observation of bovine milk by NMR spectroscopy: Analysis of existing states of compounds and detection of new compounds. J. Agric. Food Chem. 2004, 52, 4969–4974. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.; Furihata, K.; Hu, F.; Miyakawa, T.; Tanokura, M. Complex mixture analysis of organic compounds in green coffee bean extract by two-dimensional NMR spectroscopy. Magn. Reson. Chem. 2010, 48, 857–865. [Google Scholar] [CrossRef] [PubMed]
- Da Costa, M.P.; da Silva Frasao, B.; da Costa Lima, B.R.C.; Rodrigues, B.L.; Junior, C.A.C. Simultaneous analysis of carbohydrates and organic acids by HPLC-DAD-RI for monitoring goat’s milk yogurts fermentation. Talanta 2016, 152, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Weerathilake, W.A.; Rasika, D.M.; Ruwanmali, J.K.; Munasinghe, M.A. The evolution, processing, varieties and health benefits of yogurt. Int. J. Sci. Res. Publ. 2014, 4, 1–10. [Google Scholar]
- Siefarth, C.; Tran, T.B.T.; Mittermaier, P.; Pfeiffer, T.; Buettner, A. Effect of radio frequency heating on yoghurt, I: Technological applicability, shelf-life and sensorial quality. Foods 2014, 3, 318–335. [Google Scholar] [CrossRef]
- Lobato-Calleros, C.; Ramírez-Santiago, C.; Vernon-Carter, E.J.; Álvarez-Ramírez, J. Impact of native and chemically modified starches addition as fat replacers in the viscoelasticity of reduceD-fat stirred yogurt. J. Food Eng. 2014, 131, 110–115. [Google Scholar] [CrossRef]
- Pan, X.D.; Wu, P.G.; Yang, D.J.; Wang, L.Y.; Shen, X.H.; Zhu, C.Y. Simultaneous determination of melamine and cyanuric acid in dairy products by mixeD-mode solid phase extraction and GC–MS. Food Control. 2013, 30, 545–548. [Google Scholar] [CrossRef]
- Kunda, P.B.; Benavente, F.; Catalá-Clariana, S.; Giménez, E.; Barbosa, J.; Sanz-Nebot, V. Identification of bioactive peptides in a functional yogurt by micro liquid chromatography time-of-flight mass spectrometry assisted by retention time prediction. J. Chromatogr. A 2012, 1229, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Coscia, A.; Orrù, S.; Di Nicola, P.; Giuliani, F.; Rovelli, I.; Peila, C.; Bertino, E. Cow’s milk proteins in human milk. J. Biol. Regul. Homeost. Agents 2012, 26, 39–42. [Google Scholar] [PubMed]
- Ahrens, B.; Lopes de Oliveira, L.C.; Grabenhenrich, L.; Schulz, G.; Niggemann, B.; Wahn, U.; Beyer, K. Individual cow’s milk allergens as prognostic markers for tolerance development? Clin. Exp. Allergy 2012, 42, 1630–1637. [Google Scholar] [CrossRef] [PubMed]
- Mekmene, O.; Gaucheron, F. Determination of calcium-binding constants of caseins, phosphoserine, citrate and pyrophosphate: A modelling approach using free calcium measurement. Food Chem. 2011, 127, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Milewski, S.; Gabriel, I.; Olchowy, J. Enzymes of UDP-GlcNAc biosynthesis in yeast. Yeast 2006, 23, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Ardawatia, H.; Liberles, D.A. A systematic analysis of lineage-specific evolution in metabolic pathways. Gene 2007, 387, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Shiby, V.K.; Mishra, H.N. Fermented milks and milk products as functional foods—A review. Crit. Rev. Food Sci. Nutr. 2013, 53, 482–496. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.S. Diffusion ordered nuclear magnetic resonance spectroscopy: Principles and applications. Prog. Nucl. Magn. Reson. Spectrosc. 1999, 34, 203–256. [Google Scholar] [CrossRef]



| Compound | 1H Multiplicity J (Hz) | Chemical Shift (ppm) | Assignment a | ||
|---|---|---|---|---|---|
| 1H | 13C | 31P | |||
| d-lactose | d(7.70) | 4.33 | 103.57 | CH-1′ | |
| 3.43 | 71.79 | CH-2′ | |||
| 3.54 | 73.31 | CH-3′ | |||
| 3.80 | 69.38 | CH-4′ | |||
| 3.61 | 76.15 | CH-5′ | |||
| 3.64 | 61.94 | CH2OH-6′ | |||
| d(3.05) | 5.11 | 92.62 | CH-α1 | ||
| t(8.85) | 3.47 | 71.98 | CH-α2 | ||
| 3.71 | 72.24 | CH-α3 | |||
| 3.54 | 78.89 | CH-α4 | |||
| 3.82 | 70.86 | CH-α5 | |||
| 3.76 | 60.66 | CH2OH-α6 | |||
| d(7.90) | 4.55 | 96.55 | CH-β1 | ||
| t(7.83) | 3.16 | 74.66 | CH-β2 | ||
| 3.53 | 75.17 | CH-β3 | |||
| 3.54 | 78.75 | CH-β4 | |||
| 3.52 | 75.56 | CH-β5 | |||
| 3.82 | 60.80 | CH2OH-β6 | |||
| d-galactose | d(3.35) | 5.14 | 93.06 | CH-α1 | |
| 3.69 | 69.18 | CH-α2 | |||
| 3.72 | 69.99 | CH-α3 | |||
| 3.85 | 70.11 | CH-α4 | |||
| 3.96 | 71.25 | CH-α5 | |||
| 3.66 | 62.02 | CH2OH-α6 | |||
| d(7.80) | 4.46 | 97.23 | CH-β1 | ||
| 3.36 | 72.70 | CH-β2 | |||
| 3.53 | 75.17 | CH-β3 | |||
| 3.82 | 69.56 | CH-β4 | |||
| 3.60 | 75.94 | CH-β5 | |||
| 3.63 | 61.84 | CH2OH-β6 | |||
| acyl chains of fatty acids | 0.76 | 14.74 | CH3-ω1 | ||
| 1.17 | 23.54 | CH2-ω2 | |||
| 1.45 | 25.60 | CH2-Δ3 | |||
| 1.89 | 27.93 | oleate CH2-ω8,11 | |||
| 1.17 | 29.78–30.90 | CH2(ω4-n) | |||
| 1.14 | 32.79 | CH2-ω3 | |||
| 2.11 | 34.37 | CH2-Δ2 | |||
| 5.19 | 130.18 | oleate HC=CH | |||
| glycerol backbone of fats | 3.98, 4.16 | 62.54 | glycerol-1,3 | ||
| 5.10 | 69.59 | glycerol-2 | |||
| acetic acid | s | 1.93 | 22.72 | CH3 | |
| 175.40 | COOH | ||||
| citric acid | AB | 2.53,2.69 | 44.93 | CH2 | |
| 75.80 | C | ||||
| 177.70 | CH2-COOH | ||||
| 181.40 | COOH | ||||
| lactic acid | d(6.60) | 1.23 | 20.84 | CH3 | |
| dd | 4.08 | 68.95 | CH | ||
| 182.50 | COOH | ||||
| alanine | d(7.15) | 1.36 | 17.09 | CH3 | |
| 3.66 | 51.49 | CH | |||
| 176.4 | COOH | ||||
| creatine | s | 2.90 | 37.40 | CH3 | |
| s | 3.81 | 54.78 | CH2 | ||
| 158.30 | C | ||||
| 175.40 | COOH | ||||
| lecithin | s | 3.10 | 54.28 | trimethylamine | |
| 3.74, 3.80 | −0.20 | CH2-3 | |||
| 4.18 | 60.22 | −0.20 | CH2-4 | ||
| 3.54 | 67.09 | −0.20 | CH2-5 | ||
| cephalin | 3.74, 3.80 | 0.33 | CH2-3 | ||
| 3.97 | 0.33 | CH2-4 | |||
| N-acetyl-d-glucosamine-1-phosphate | 5.30 | −1.44 | O-CH | ||
| 3.83 | −1.44 | N-CH | |||
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, Y.; Hu, F.; Miyakawa, T.; Tanokura, M. Complex Mixture Analysis of Organic Compounds in Yogurt by NMR Spectroscopy. Metabolites 2016, 6, 19. https://doi.org/10.3390/metabo6020019
Lu Y, Hu F, Miyakawa T, Tanokura M. Complex Mixture Analysis of Organic Compounds in Yogurt by NMR Spectroscopy. Metabolites. 2016; 6(2):19. https://doi.org/10.3390/metabo6020019
Chicago/Turabian StyleLu, Yi, Fangyu Hu, Takuya Miyakawa, and Masaru Tanokura. 2016. "Complex Mixture Analysis of Organic Compounds in Yogurt by NMR Spectroscopy" Metabolites 6, no. 2: 19. https://doi.org/10.3390/metabo6020019
APA StyleLu, Y., Hu, F., Miyakawa, T., & Tanokura, M. (2016). Complex Mixture Analysis of Organic Compounds in Yogurt by NMR Spectroscopy. Metabolites, 6(2), 19. https://doi.org/10.3390/metabo6020019






