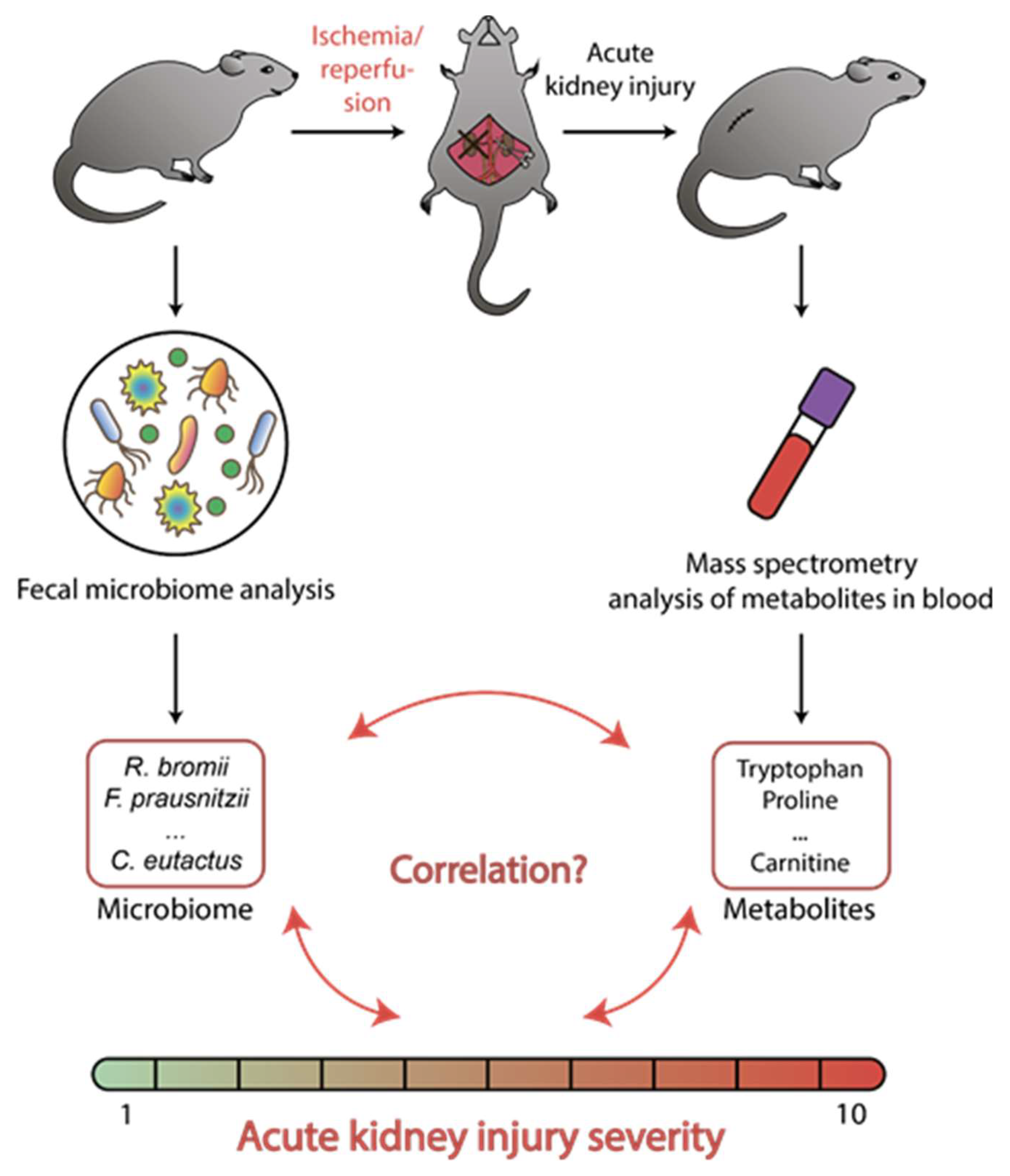
Microbiome-Metabolome Signature of Acute Kidney Injury
Abstract
1. Introduction
2. Results
2.1. AKI and Metabolome
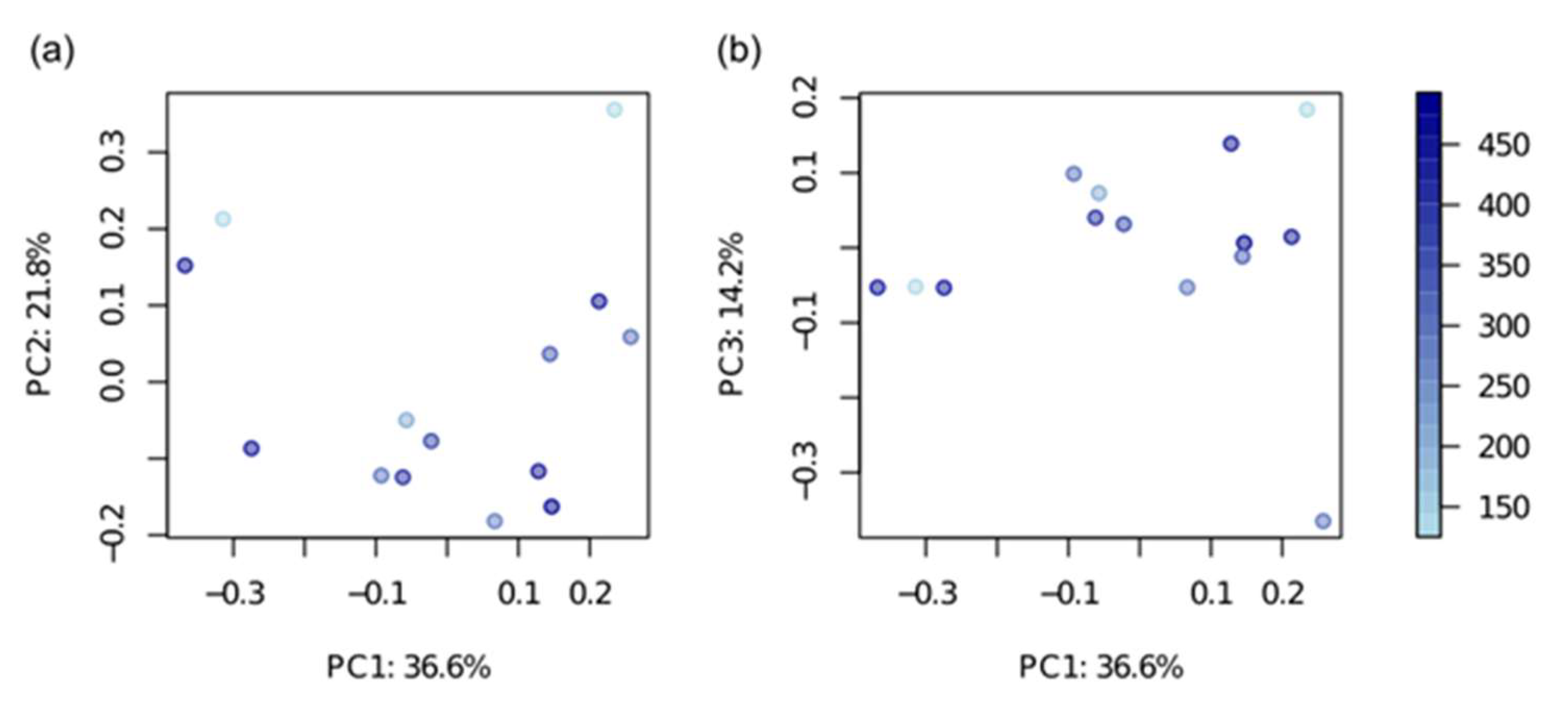
2.2. AKI and Microbiome
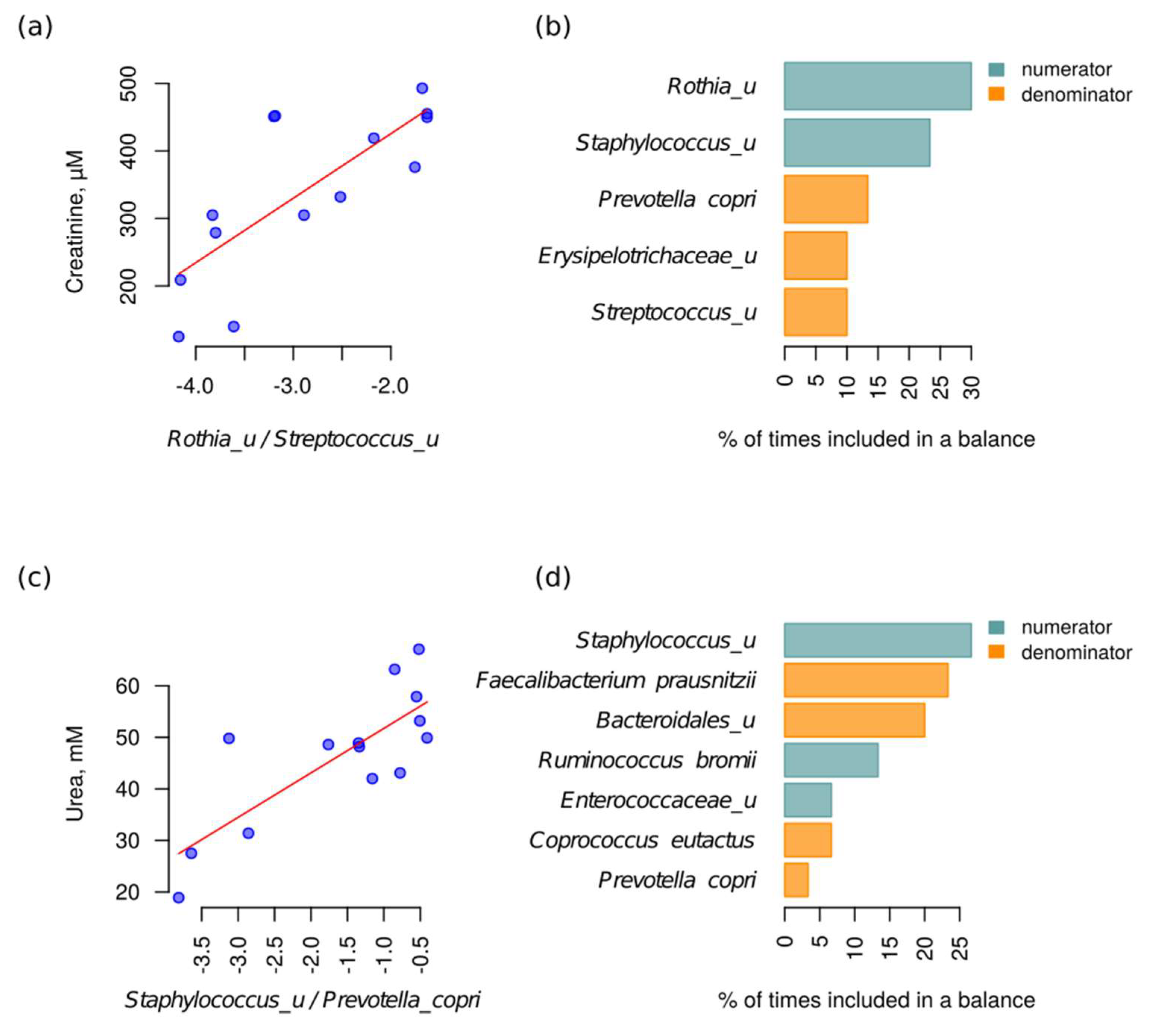
2.3. Metabolome and Microbiome during AKI
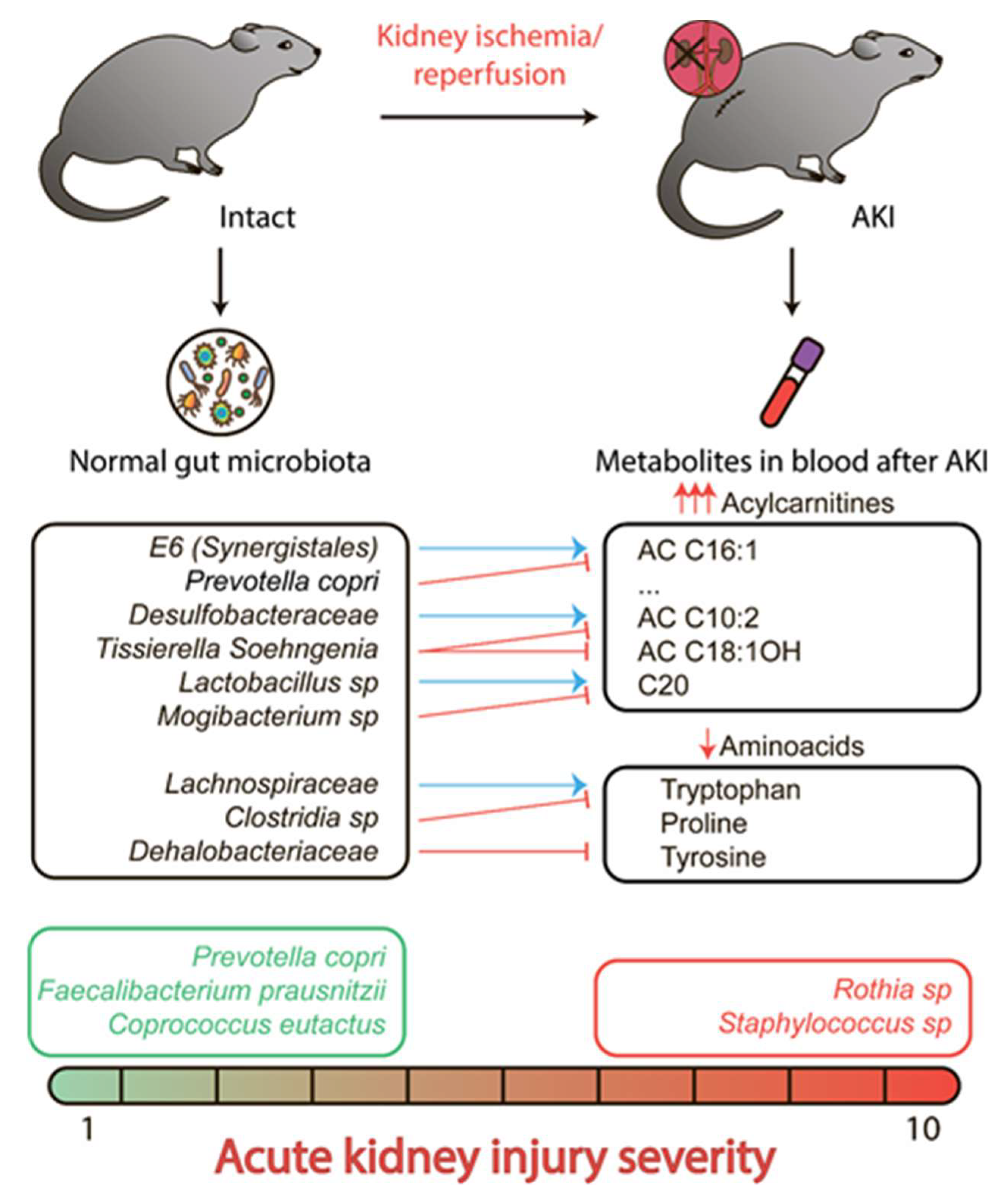
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. Kidney I/R Protocol
4.3. Tandem Mass Spectrometry
4.4. DNA Extraction and Sequencing
4.5. Bioinformatic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Rooks, M.G.; Garrett, W.S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 2016, 16, 341–352. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Devaraj, S. Gut microbiome in obesity, metabolic syndrome, and diabetes. Curr. Diabetes Rep. 2018, 18, 129. [Google Scholar] [CrossRef] [PubMed]
- Bouter, K.E.; van Raalte, D.H.; Groen, A.K.; Nieuwdorp, M. Role of the gut microbiome in the pathogenesis of obesity and obesity-related metabolic dysfunction. Gastroenterology 2017, 152, 1671–1678. [Google Scholar] [CrossRef] [PubMed]
- Goodman, B.; Gardner, H. The microbiome and cancer. J. Pathol. 2018, 244, 667–676. [Google Scholar] [CrossRef] [PubMed]
- Gorkiewicz, G.; Moschen, A. Gut microbiome: A new player in gastrointestinal disease. Virchows Arch. 2018, 472, 159–172. [Google Scholar] [CrossRef]
- Nishida, A.; Inoue, R.; Inatomi, O.; Bamba, S.; Naito, Y.; Andoh, A. Gut microbiota in the pathogenesis of inflammatory bowel disease. Clin. J. Gastroenterol. 2018, 11, 1–10. [Google Scholar] [CrossRef]
- Ahmadmehrabi, S.; Tang, W.H.W. Gut microbiome and its role in cardiovascular diseases. Curr. Opin. Cardiol. 2017, 32, 761–766. [Google Scholar] [CrossRef]
- Al Khodor, S.; Shatat, I.F. Gut microbiome and kidney disease: A bidirectional relationship. Pediatr. Nephrol. 2017, 32, 921–931. [Google Scholar] [CrossRef]
- Ji, W.; Zhu, Y.; Kan, P.; Cai, Y.; Wang, Z.; Wu, Z.; Yang, P. Analysis of intestinal microbial communities of cerebral infarction and ischemia patients based on high throughput sequencing technology and glucose and lipid metabolism. Mol. Med. Rep. 2017, 16, 5413–5417. [Google Scholar] [CrossRef]
- Felizardo, R.J.F.; Castoldi, A.; Andrade-Oliveira, V.; Câmara, N.O.S. The microbiota and chronic kidney diseases: A double-edged sword. Clin. Transl. Immunol. 2016, 5, e86. [Google Scholar] [CrossRef]
- Delzenne, N.M.; Neyrinck, A.M.; Bäckhed, F.; Cani, P.D. Targeting gut microbiota in obesity: Effects of prebiotics and probiotics. Nat. Rev. Endocrinol. 2011, 7, 639–646. [Google Scholar] [CrossRef] [PubMed]
- Van Spaendonk, H.; Ceuleers, H.; Witters, L.; Patteet, E.; Joossens, J.; Augustyns, K.; Lambeir, A.-M.; De Meester, I.; De Man, J.G.; De Winter, B.Y. Regulation of intestinal permeability: The role of proteases. World J. Gastroenterol. 2017, 23, 2106. [Google Scholar] [CrossRef] [PubMed]
- Vancamelbeke, M.; Vermeire, S. The intestinal barrier: A fundamental role in health and disease. Expert Rev. Gastroenterol. Hepatol. 2017, 11, 821–834. [Google Scholar] [CrossRef] [PubMed]
- Castillo, D.J.; Rifkin, R.F.; Cowan, D.A.; Potgieter, M. The healthy human blood microbiome: Fact or fiction? Front. Cell. Infect. Microbiol. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Possemiers, S.; Van De Wiele, T.; Guiot, Y.; Everard, A.; Rottier, O.; Geurts, L.; Naslain, D.; Neyrinck, A.; Lambert, D.M.; et al. Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut 2009, 58, 1091–1103. [Google Scholar] [CrossRef] [PubMed]
- Bonavia, A.; Singbartl, K. A review of the role of immune cells in acute kidney injury. Pediatr. Nephrol. 2018, 33, 1629–1639. [Google Scholar] [CrossRef]
- Anders, H.-J.; Andersen, K.; Stecher, B. The intestinal microbiota, a leaky gut, and abnormal immunity in kidney disease. Kidney Int. 2013, 83, 1010–1016. [Google Scholar] [CrossRef]
- Emal, D.; Rampanelli, E.; Stroo, I.; Butter, L.M.; Teske, G.J.; Claessen, N.; Stokman, G.; Florquin, S.; Leemans, J.C.; Dessing, M.C. Depletion of gut microbiota protects against renal ischemia-reperfusion injury. J. Am. Soc. Nephrol. 2017, 28, 1450–1461. [Google Scholar] [CrossRef]
- Rabb, H.; Pluznick, J.; Noel, S. The microbiome and acute kidney injury. Nephron 2018, 140, 120–123. [Google Scholar] [CrossRef]
- Olszak, T.; An, D.; Zeissig, S.; Vera, M.P.; Richter, J.; Franke, A.; Glickman, J.N.; Siebert, R.; Baron, R.M.; Kasper, D.L.; et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 2012, 336, 489–493. [Google Scholar] [CrossRef]
- Jang, H.R.; Gandolfo, M.T.; Ko, G.J.; Satpute, S.; Racusen, L.; Rabb, H. Early exposure to germs modifies kidney damage and inflammation after experimental ischemia-reperfusion injury. Am. J. Physiol. Ren. Physiol. 2009, 297. [Google Scholar] [CrossRef] [PubMed]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ankawi, G.; Sun, J.; Digvijay, K.; Yin, Y.; Rosner, M.H.; Ronco, C. Gut-kidney crosstalk in septic acute kidney injury. Crit. Care 2018, 22, 117. [Google Scholar] [CrossRef] [PubMed]
- Jakobsson, H.E.; Rodríguez-Piñeiro, A.M.; Schütte, A.; Ermund, A.; Boysen, P.; Bemark, M.; Sommer, F.; Bäckhed, F.; Hansson, G.C.; Johansson, M.E.V. The composition of the gut microbiota shapes the colon mucus barrier. EMBO Rep. 2015, 16, 164–177. [Google Scholar] [CrossRef]
- Hamer, H.M.; Jonkers, D.; Venema, K.; Vanhoutvin, S.; Troost, F.J.; Brummer, R.J. Review article: The role of butyrate on colonic function. Aliment. Pharmacol. Ther. 2008, 27, 104–119. [Google Scholar] [CrossRef]
- Lameire, N.H.; Bagga, A.; Cruz, D.; De Maeseneer, J.; Endre, Z.; Kellum, J.A.; Liu, K.D.; Mehta, R.L.; Pannu, N.; Van Biesen, W.; et al. Acute kidney injury: An increasing global concern. Lancet 2013, 382, 170–179. [Google Scholar] [CrossRef]
- Plotnikov, E.Y.; Kazachenko, A.V.; Vyssokikh, M.Y.; Vasileva, A.K.; Tcvirkun, D.V.; Isaev, N.K.; Kirpatovsky, V.I.; Zorov, D.B. The role of mitochondria in oxidative and nitrosative stress during ischemia/reperfusion in the rat kidney. Kidney Int. 2007, 72, 1493–1502. [Google Scholar] [CrossRef]
- Rivera-Pinto, J.; Egozcue, J.J.; Pawlowsky-Glahn, V.; Paredes, R.; Noguera-Julian, M.; Calle, M.L. Balances: A new perspective for microbiome analysis. mSystems 2018, 3. [Google Scholar] [CrossRef]
- Khwaja, A. KDIGO clinical practice guidelines for acute kidney injury. Nephron Clin. Pract. 2012, 120, 179–184. [Google Scholar] [CrossRef]
- Schrezenmeier, E.V.; Barasch, J.; Budde, K.; Westhoff, T.; Schmidt-Ott, K.M. Biomarkers in acute kidney injury—Pathophysiological basis and clinical performance. Acta Physiol. 2017, 219, 554–572. [Google Scholar] [CrossRef]
- Teo, S.H.; Endre, Z.H. Biomarkers in acute kidney injury (AKI). Best Pract. Res. Clin. Anaesthesiol. 2017, 31, 331–344. [Google Scholar] [CrossRef] [PubMed]
- Evenepoel, P.; Poesen, R.; Meijers, B. The gut–kidney axis. Pediatr. Nephrol. 2017, 32, 2005–2014. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Noel, S.; Pluznick, J.L.; Hamad, A.R.A.; Rabb, H. Gut microbiota-kidney cross-talk in acute kidney injury. Semin. Nephrol. 2019, 39, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Thursby, E.; Juge, N. Introduction to the human gut microbiota. Biochem. J. 2017, 474, 1823–1836. [Google Scholar] [CrossRef] [PubMed]
- Neuman, H.; Debelius, J.W.; Knight, R.; Koren, O. Microbial endocrinology: The interplay between the microbiota and the endocrine system. FEMS Microbiol. Rev. 2015, 39, 509–521. [Google Scholar] [CrossRef] [PubMed]
- Sommer, F.; Bäckhed, F. The gut microbiota-masters of host development and physiology. Nat. Rev. Microbiol. 2013, 11, 227–238. [Google Scholar] [CrossRef]
- Lee, G.R. The balance of th17 versus treg cells in autoimmunity. Int. J. Mol. Sci. 2018, 19, 730. [Google Scholar] [CrossRef]
- Cerf-Bensussan, N.; Gaboriau-Routhiau, V. The immune system and the gut microbiota: Friends or foes? Nat. Rev. Immunol. 2010, 10, 735–744. [Google Scholar] [CrossRef]
- Vatanen, T.; Kostic, A.D.; D’Hennezel, E.; Siljander, H.; Franzosa, E.A.; Yassour, M.; Kolde, R.; Vlamakis, H.; Arthur, T.D.; Hämäläinen, A.M.; et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 2016, 165, 842–853. [Google Scholar] [CrossRef]
- Salguero, M.; Al-Obaide, M.; Singh, R.; Siepmann, T.; Vasylyeva, T. Dysbiosis of Gram-negative gut microbiota and the associated serum lipopolysaccharide exacerbates inflammation in type2 diabetic patients with chronic kidney disease. Exp. Ther. Med. 2019, 18. [Google Scholar] [CrossRef]
- Andrade-Oliveira, V.; Amano, M.T.; Correa-Costa, M.; Castoldi, A.; Felizardo, R.J.F.; de Almeida, D.C.; Bassi, E.J.; Moraes-Vieira, P.M.; Hiyane, M.I.; Rodas, A.C.D.; et al. Gut bacteria products prevent AKI induced by ischemia-reperfusion. J. Am. Soc. Nephrol. 2015, 26, 1877–1888. [Google Scholar] [CrossRef] [PubMed]
- Yoshifuji, A.; Wakino, S.; Irie, J.; Tajima, T.; Hasegawa, K.; Kanda, T.; Tokuyama, H.; Hayashi, K.; Itoh, H. Gut Lactobacillus protects against the progression of renal damage by modulating the gut environment in rats. Nephrol. Dial. Transplant. 2016, 31, 401–412. [Google Scholar] [CrossRef] [PubMed]
- Tao, S.; Li, L.; Li, L.; Liu, Y.; Ren, Q.; Shi, M.; Liu, J.; Jiang, J.; Ma, H.; Huang, Z.; et al. Understanding the gut–kidney axis among biopsy-proven diabetic nephropathy, type 2 diabetes mellitus and healthy controls: An analysis of the gut microbiota composition. Acta Diabetol. 2019, 56. [Google Scholar] [CrossRef] [PubMed]
- Ferreira-Halder, C.V.; de Sousa Faria, A.V.; Andrade, S.S. Action and function of Faecalibacterium prausnitzii in health and disease. Best Pract. Res. Clin. Gastroenterol. 2017, 31, 643–648. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Duncan, S.H.; Garcia-Gil, L.J.; Martinez-Medina, M. Faecalibacterium prausnitzii: From microbiology to diagnostics and prognostics. ISME J. 2017, 11, 841–852. [Google Scholar] [CrossRef] [PubMed]
- Nylund, L.; Nermes, M.; Isolauri, E.; Salminen, S.; De Vos, W.M.; Satokari, R. Severity of atopic disease inversely correlates with intestinal microbiota diversity and butyrate-producing bacteria. Allergy 2015, 70, 241–244. [Google Scholar] [CrossRef]
- Rodríguez-Piñeiro, A.M.; Johansson, M.E.V. The colonic mucus protection depends on the microbiota. Gut Microbes 2015, 6, 326–330. [Google Scholar] [CrossRef]
- Barrows, I.R.; Ramezani, A.; Raj, D.S. Gut feeling in AKI: The long arm of short-chain fatty acids. J. Am. Soc. Nephrol. 2015, 26, 1755–1757. [Google Scholar] [CrossRef][Green Version]
- Jiang, S.; Xie, S.; Lv, D.; Wang, P.; He, H.; Zhang, T.; Zhou, Y.; Lin, Q.; Zhou, H.; Jiang, J.; et al. Alteration of the gut microbiota in Chinese population with chronic kidney disease. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Jiang, S.; Xie, S.; Lv, D.; Zhang, Y.; Deng, J.; Zeng, L.; Chen, Y. A reduction in the butyrate producing species Roseburia spp. and Faecalibacterium prausnitzii is associated with chronic kidney disease progression. Antonie Van Leeuwenhoek 2016, 109, 1389–1396. [Google Scholar] [CrossRef]
- Zybailov, B.L.; Glazko, G.V.; Rahmatallah, Y.; Andreyev, D.S.; Taylor McElroy, D.; Karaduta, O.; Byrum, S.D.; Orr, L.; Tackett, A.J.; Mackintosh, S.G.; et al. Metaproteomics reveals potential mechanisms by which dietary resistant starch supplementation attenuates chronic kidney disease progression in rats. PLoS ONE 2019, 14. [Google Scholar] [CrossRef] [PubMed]
- Navab-Moghadam, F.; Sedighi, M.; Khamseh, M.E.; Alaei-Shahmiri, F.; Talebi, M.; Razavi, S.; Amirmozafari, N. The association of type II diabetes with gut microbiota composition. Microb. Pathog. 2017, 110, 630–636. [Google Scholar] [CrossRef] [PubMed]
- Ganesan, K.; Chung, S.K.; Vanamala, J.; Xu, B. Causal relationship between diet-induced gut microbiota changes and diabetes: A novel strategy to transplant Faecalibacterium prausnitzii in preventing diabetes. Int. J. Mol. Sci. 2018, 19, 720. [Google Scholar] [CrossRef] [PubMed]
- Machiels, K.; Joossens, M.; Sabino, J.; De Preter, V.; Arijs, I.; Eeckhaut, V.; Ballet, V.; Claes, K.; Van Immerseel, F.; Verbeke, K.; et al. A decrease of the butyrate-producing species roseburia hominis and faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 2014, 63, 1275–1283. [Google Scholar] [CrossRef]
- Lopez-Siles, M.; Enrich-Capó, N.; Aldeguer, X.; Sabat-Mir, M.; Duncan, S.H.; Garcia-Gil, L.J.; Martinez-Medina, M. Alterations in the abundance and Co-occurrence of Akkermansia muciniphila and Faecalibacterium prausnitzii in the colonic mucosa of inflammatory bowel disease subjects. Front. Cell. Infect. Microbiol. 2018, 8. [Google Scholar] [CrossRef]
- Tyakht, A.V.; Manolov, A.I.; Kanygina, A.V.; Ischenko, D.S.; Kovarsky, B.A.; Popenko, A.S.; Pavlenko, A.V.; Elizarova, A.V.; Rakitina, D.V.; Baikova, J.P.; et al. Genetic diversity of Escherichia coli in gut microbiota of patients with Crohn’s disease discovered using metagenomic and genomic analyses. BMC Genom. 2018, 19. [Google Scholar] [CrossRef]
- Petrov, V.A.; Saltykova, I.V.; Zhukova, I.A.; Alifirova, V.M.; Zhukova, N.G.; Dorofeeva, Y.B.; Tyakht, A.V.; Kovarsky, B.A.; Alekseev, D.G.; Kostryukova, E.S.; et al. Analysis of gut microbiota in patients with parkinson’s disease. Bull. Exp. Biol. Med. 2017, 162, 734–737. [Google Scholar] [CrossRef]
- Nagpal, R.; Wang, S.; Solberg Woods, L.C.; Seshie, O.; Chung, S.T.; Shively, C.A.; Register, T.C.; Craft, S.; McClain, D.A.; Yadav, H. Comparative microbiome signatures and short-chain fatty acids in mouse, rat, non-human primate, and human feces. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Vanhatalo, A.; Blackwell, J.R.; L’Heureux, J.E.; Williams, D.W.; Smith, A.; van der Giezen, M.; Winyard, P.G.; Kelly, J.; Jones, A.M. Nitrate-responsive oral microbiome modulates nitric oxide homeostasis and blood pressure in humans. Free Radic. Biol. Med. 2018, 124, 21–30. [Google Scholar] [CrossRef]
- Manrique, P.; Freire, M.O.; Chen, C.; Zadeh, H.H.; Young, M.; Suci, P. Perturbation of the indigenous rat oral microbiome by ciprofloxacin dosing. Mol. Oral Microbiol. 2013, 28, 404–414. [Google Scholar] [CrossRef]
- Zhang, S.L.; Bai, L.; Goel, N.; Bailey, A.; Jang, C.J.; Bushman, F.D.; Meerlo, P.; Dinges, D.F.; Sehgal, A. Human and rat gut microbiome composition is maintained following sleep restriction. Proc. Natl. Acad. Sci. USA 2017, 114, E1564–E1571. [Google Scholar] [CrossRef] [PubMed]
- Imhann, F.; Jan Bonder, M.; Vich Vila, A.; Fu, J.; Mujagic, Z.; Vork, L.; Tigchelaar, E.F.; Jankipersadsing, S.A.; Cenit, M.C.; Harmsen, H.J.M.; et al. Proton pump inhibitors affect the gut microbiome. Gut 2016. [Google Scholar] [CrossRef] [PubMed]
- Dubinkina, V.B.; Tyakht, A.V.; Odintsova, V.Y.; Yarygin, K.S.; Kovarsky, B.A.; Pavlenko, A.V.; Ischenko, D.S.; Popenko, A.S.; Alexeev, D.G.; Taraskina, A.Y.; et al. Links of gut microbiota composition with alcohol dependence syndrome and alcoholic liver disease. Microbiome 2017, 5, 141. [Google Scholar] [CrossRef] [PubMed]
- Lecomte, V.; Kaakoush, N.O.; Maloney, C.A.; Raipuria, M.; Huinao, K.D.; Mitchell, H.M.; Morris, M.J. Changes in gut microbiota in rats fed a high fat diet correlate with obesity-associated metabolic parameters. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Dewhirst, F.E.; Chen, T.; Izard, J.; Paster, B.J.; Tanner, A.C.R.; Yu, W.H.; Lakshmanan, A.; Wade, W.G. The human oral microbiome. J. Bacteriol. 2010, 192, 5002–5017. [Google Scholar] [CrossRef] [PubMed]
- Hyde, E.R.; Luk, B.; Cron, S.; Kusic, L.; McCue, T.; Bauch, T.; Kaplan, H.; Tribble, G.; Petrosino, J.F.; Bryan, N.S. Characterization of the rat oral microbiome and the effects of dietary nitrate. Free Radic. Biol. Med. 2014, 77, 249–257. [Google Scholar] [CrossRef]
- Fouque, D.; Holt, S.; Guebre-Egziabher, F.; Nakamura, K.; Vianey-Saban, C.; Hadj-Aïssa, A.; Hoppel, C.L.; Kopple, J.D. Relationship between serum carnitine, acylcarnitines, and renal function in patients with chronic renal disease. J. Ren. Nutr. 2006, 16, 125–131. [Google Scholar] [CrossRef]
- Hoppel, C. The role of carnitine in normal and altered fatty acid metabolism. Am. J. Kidney Dis. 2003, 41. [Google Scholar] [CrossRef]
- Reuter, S.E.; Evans, A.M. Carnitine and Acylcarnitines: Pharmacokinetic, pharmacological and clinical aspects. Clin. Pharmacokinet. 2012, 51, 553–572. [Google Scholar] [CrossRef]
- Liepinsh, E.; Makrecka-Kuka, M.; Volska, K.; Kuka, J.; Makarova, E.; Antone, U.; Sevostjanovs, E.; Vilskersts, R.; Strods, A.; Tars, K.; et al. Long-chain acylcarnitines determine ischaemia/reperfusion-induced damage in heart mitochondria. Biochem. J. 2016, 473, 1191–1202. [Google Scholar] [CrossRef]
- Færgeman, N.J.; Knudsen, J. Role of long-chain fatty acyl-CoA esters in the regulation of metabolism and in cell signalling. Biochem. J. 1997, 323, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Wanner, C.; Riegel, W.; Schaefer, R.M.; Hörl, W.H. Carnitine and carnitine esters in acute renal failure. Nephrol. Dial. Transplant. 1989, 4, 951–956. [Google Scholar] [CrossRef] [PubMed]
- Yamada, K.A.; Kanter, E.M.; Newatia, A. Long-chain acylcarnitine induces Ca2+ efflux from the sarcoplasmic reticulum. J. Cardiovasc. Pharmacol. 2000, 36, 14–21. [Google Scholar] [CrossRef] [PubMed]
- Adeva-Andany, M.M.; Calvo-Castro, I.; Fernández-Fernández, C.; Donapetry-García, C.; Pedre-Piñeiro, A.M. Significance of l-carnitine for human health. IUBMB Life 2017, 69, 578–594. [Google Scholar] [CrossRef] [PubMed]
- Pekala, J.; Patkowska-Sokola, B.; Bodkowski, R.; Jamroz, D.; Nowakowski, P.; Lochynski, S.; Librowski, T. L-Carnitine—Metabolic functions and meaning in humans life. Curr. Drug Metab. 2011, 12, 667–678. [Google Scholar] [CrossRef] [PubMed]
- Moghaddas, A.; Dashti-Khavidaki, S. L-Carnitine and potential protective effects against ischemia-reperfusion injury in noncardiac organs: From experimental data to potential clinical applications. J. Diet. Suppl. 2018, 15, 740–756. [Google Scholar] [CrossRef]
- Estaphan, S.; Eissa, H.; Elattar, S.; Rashed, L.; Farouk, M. A study on the effect of cimetidine and l-carnitine on myoglobinuric acute kidney injury in male rats. Injury 2015, 46, 1223–1230. [Google Scholar] [CrossRef]
- Jafari, A.; Khatami, M.R.; Dashti-Khavidaki, S.; Lessan-Pezeshki, M.; Abdollahi, A.; Moghaddas, A. Protective effects of L-Carnitine against delayed graft function in kidney transplant recipients: A pilot, randomized, double-blinded, placebo-controlled clinical trial. J. Ren. Nutr. 2017, 27, 113–126. [Google Scholar] [CrossRef]
- Mister, M.; Noris, M.; Szymczuk, J.; Azzollini, N.; Aiello, S.; Abbate, M.; Trochimowicz, L.; Gaguardini, E.; Arduini, A.; Perico, N.; et al. Propionyl-L-carnitine prevents renal function deterioration due to ischemia/reperfusion. Kidney Int. 2002, 61, 1064–1078. [Google Scholar] [CrossRef]
- Ferrari, R.; Merli, E.; Cicchitelli, G.; Mele, D.; Fucili, A.; Ceconi, C. Therapeutic effects of L-carnitine and propionyl-L-carnitine on cardiovascular diseases: A review. Ann. N. Y. Acad. Sci. 2004, 1033, 79–91. [Google Scholar] [CrossRef]
- Bremer, J. Carnitine—Metabolism and functions. Physiol. Rev. 1983, 63, 1420–1480. [Google Scholar] [CrossRef]
- Uchino, S. Creatinine. Curr. Opin. Crit. Care 2010, 16, 562–567. [Google Scholar] [CrossRef]
- Bessman, S.P.; Carpenter, C.L. The creatine-creatine phosphate energy shuttle. Annu. Rev. Biochem. 1985, 54, 831–862. [Google Scholar] [CrossRef]
- Popkov, V.A.; Silachev, D.N.; Zalevsky, A.O.; Zorov, D.B.; Plotnikov, E.Y. Mitochondria as a source and a target for uremic toxins. Int. J. Mol. Sci. 2019, 20, 94. [Google Scholar] [CrossRef]
- Kopple, J.D. Phenylalanine and tyrosine metabolism in chronic kidney failure. J. Nutr. 2007, 137, 1586S–1590S. [Google Scholar] [CrossRef]
- Saito, A.; Niwa, T.; Maeda, K.; Kobayashi, K.; Yamamoto, Y.; Ohta, K. Tryptophan and indolic tryptophan metabolites in chronic renal failure. Am. J. Clin. Nutr. 1980, 33, 1402–1406. [Google Scholar] [CrossRef]
- Ganapathy, V.; Roesel, R.A.; Howard, J.C.; Leibach, F.H. Interaction of proline, 5-oxoproline, and pipecolic acid for renal transport in the rabbit. J. Biol. Chem. 1983, 258, 2266–2272. [Google Scholar]
- Tomasova, L.; Konopelski, P.; Ufnal, M. Gut bacteria and hydrogen sulfide: The new old players in circulatory system homeostasis. Molecules 2016, 21, 1558. [Google Scholar] [CrossRef]
- Feliers, D.; Lee, H.J.; Kasinath, B.S. Hydrogen sulfide in renal physiology and disease. Antioxid. Redox Signal. 2016, 25, 720–731. [Google Scholar] [CrossRef]
- Kasinath, B.S.; Feliers, D.; Lee, H.J. Hydrogen sulfide as a regulatory factor in kidney health and disease. Biochem. Pharmacol. 2018, 149, 29–41. [Google Scholar] [CrossRef]
- Weber, G.J.; Pushpakumar, S.B.; Sen, U. Hydrogen sulfide alleviates hypertensive kidney dysfunction through an epigenetic mechanism. Am. J. Physiol. Heart Circ. Physiol. 2017, 312, H874–H885. [Google Scholar] [CrossRef]
- Sun, H.J.; Wu, Z.Y.; Cao, L.; Zhu, M.Y.; Liu, T.T.; Guo, L.; Lin, Y.; Nie, X.W.; Bian, J.S. Hydrogen sulfide: Recent progression and perspectives for the treatment of diabetic nephropathy. Molecules 2019, 24, 2857. [Google Scholar] [CrossRef]
- Rivers, J.R.; Badiei, A.; Bhatia, M. Hydrogen sulfide as a therapeutic target for inflammation. Expert Opin. Ther. Targets 2012, 16, 439–449. [Google Scholar] [CrossRef]
- Volokh, O.; Klimenko, N.; Berezhnaya, Y.; Tyakht, A.; Nesterova, P.; Popenko, A.; Alexeev, D. Human gut microbiome response induced by fermented dairy product intake in healthy volunteers. Nutrients 2019, 11, 547. [Google Scholar] [CrossRef]
- Efimova, D.; Tyakht, A.; Popenko, A.; Vasilyev, A.; Altukhov, I.; Dovidchenko, N.; Odintsova, V.; Klimenko, N.; Loshkarev, R.; Pashkova, M.; et al. Knomics-Biota—A system for exploratory analysis of human gut microbiota data. BioData Min. 2018, 11. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.; Kindt, R.; Legendre, P.; O’hara, R.; Simpson, G.; Solymos, P.; Stevens, M.; Wagner, H. Vegan: Community Ecology Package, Version 2.0–7. Available online: https://cran.r-project.org/web/packages/vegan/ (accessed on 2 April 2020).
| Metabolite | AKI vs. Control | Correlation with SCr | |||
|---|---|---|---|---|---|
| AKI/Control | FDR Adjusted p-Value | Mean in AKI, µM | Mean in Control, µM | ||
| Malonylcarnitine (AC C3DC) | 6.71 | 0.00073 | 0.244 | 0.036 | 0.89 |
| Glutarylcarnitine (AC C5DC) | 4.55 | 0.00301 | 0.253 | 0.056 | 0.76 |
| Decadienoylcarnitine (AC C10:2) | 4.30 | 0.00073 | 0.016 | 0.004 | 0.72 |
| 3-hydroxybutyrylcarnitine (AC C4OH) | 3.85 | 0.00432 | 0.061 | 0.016 | 0.63 |
| Linoleylcarnitine (AC C18:2) | 3.76 | 0.00128 | 0.082 | 0.022 | 0.63 |
| Methylmalonylcarnitine (AC C4DC) | 3.51 | 0.00301 | 0.154 | 0.044 | 0.85 |
| Hexanoylcarnitine (AC C6) | 3.41 | 0.00073 | 0.046 | 0.014 | 0.75 |
| Acetylcarnitine (AC C2) | 3.15 | 0.00167 | 26.116 | 8.296 | 0.77 |
| Octanoylcarnitine (AC C8) | 3.13 | 0.00081 | 0.021 | 0.007 | 0.86 |
| Oleoylcarnitine (AC C18:1) | 3.08 | 0.00002 | 0.123 | 0.040 | 0.79 |
| 3-hydroxystearylcarnitine (AC C18OH) | 2.75 | 0.00891 | 0.023 | 0.008 | 0.73 |
| 3-hydroxyoleylcarnitine (AC C18:1OH) | 2.70 | 0.00891 | 0.020 | 0.007 | 0.51 |
| 3-hydroxypalmitoylcarnitine (AC C16OH) | 2.44 | 0.01884 | 0.021 | 0.009 | 0.74 |
| Hydroxyhexanoylcarnitine (AC C6OH) | 2.41 | 0.00573 | 0.026 | 0.011 | 0.71 |
| Arachidylcarnitine (C20) | 2.39 | 0.00228 | 0.035 | 0.015 | 0.32 |
| Palmitoylcarnitine (AC C16) | 2.32 | 0.00073 | 0.236 | 0.102 | 0.70 |
| Stearoylcarnitine (AC C18) | 2.30 | 0.00073 | 0.132 | 0.058 | 0.66 |
| 3-hydroxypalmitoleylcarnitine (AC C16:1OH) | 2.25 | 0.01022 | 0.023 | 0.010 | 0.54 |
| Tetradecadienoylcarnitine (AC C14:2) | 2.23 | 0.00108 | 0.045 | 0.020 | 0.72 |
| Tetradecenoylcarnitine (AC C14:1) | 2.16 | 0.00432 | 0.101 | 0.047 | 0.69 |
| Palmitoleylcarnitine (AC C16:1) | 2.14 | 0.00482 | 0.052 | 0.024 | 0.66 |
| 3-hydroxyisovalerylcarnitine (AC C5OH) | 2.12 | 0.00827 | 0.092 | 0.043 | 0.73 |
| Butyrylcarnitine (AC C4) | 2.10 | 0.01656 | 0.455 | 0.216 | 0.57 |
| Octenoylcarnitine (AC C8:1) | 2.09 | 0.00223 | 0.012 | 0.006 | 0.81 |
| Adipylcarnitine (AC C6DC) | 2.03 | 0.00991 | 0.096 | 0.047 | 0.77 |
| Myristylcarnitine (AC C14) | 2.00 | 0.00281 | 0.095 | 0.047 | 0.74 |
| Dodecanoylcarnitine (AC C12) | 2.00 | 0.00159 | 0.124 | 0.062 | 0.61 |
| Decanoylcarnitine (AC C10) | 1.96 | 0.00788 | 0.033 | 0.017 | 0.82 |
| Decenoylcarnitine (AC C10:1) | 1.81 | 0.01221 | 0.027 | 0.015 | 0.65 |
| 3-hydroxymyristylcarnitine (AC C14OH) | 1.76 | 0.03478 | 0.012 | 0.007 | 0.67 |
| Free carnitine (AC C0) | 1.72 | 0.01656 | 51.941 | 30.283 | 0.64 |
| Tyrosine (AA Tyr) | 0.73 | 0.00573 | 57.410 | 78.415 | -0.54 |
| Tryptophan (AA Trp) | 0.66 | 0.01200 | 11.244 | 16.991 | -0.61 |
| Proline (AA Pro) | 0.60 | 0.00223 | 114.723 | 192.781 | -0.70 |
| Metabolite | Taxon | p-Value, adj. | R2, adj. | Position in a Balance | % of Times Included in a Balance |
|---|---|---|---|---|---|
| Hexadecenoylcarnitine (AC C16:1) | unclassified E6 (Synergistales) | 0.00028 | 0.74793 | NUM | 50 |
| Prevotella copri | DEN | 56 | |||
| Tryptophan | unclassified Lachnospiraceae | 0.00028 | 0.72956 | NUM | 37 |
| unclassified Clostridia | DEN | 43 | |||
| Decadienoylcarnitine (AC C10:2) | unclassified Desulfobacteraceae | 0.00028 | 0.73393 | NUM | 33 |
| unclassified Tissierella/Soehngenia | DEN | 33 | |||
| Arachidylcarnitine (C20) | unclassified Lactobacillus | 0.00045 | 0.69365 | NUM | 33 |
| unclassified Mogibacterium | DEN | 30 | |||
| Tyrosine | unclassified Dehalobacteriaceae | 0.00135 | 0.6112 | DEN | 33 |
| Hydroxyoleoylcarnitine (AC C18:1OH) | unclassified Tissierella/Soehngenia | 0.00617 | 0.46336 | DEN | 26 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Andrianova, N.V.; Popkov, V.A.; Klimenko, N.S.; Tyakht, A.V.; Baydakova, G.V.; Frolova, O.Y.; Zorova, L.D.; Pevzner, I.B.; Zorov, D.B.; Plotnikov, E.Y. Microbiome-Metabolome Signature of Acute Kidney Injury. Metabolites 2020, 10, 142. https://doi.org/10.3390/metabo10040142
Andrianova NV, Popkov VA, Klimenko NS, Tyakht AV, Baydakova GV, Frolova OY, Zorova LD, Pevzner IB, Zorov DB, Plotnikov EY. Microbiome-Metabolome Signature of Acute Kidney Injury. Metabolites. 2020; 10(4):142. https://doi.org/10.3390/metabo10040142
Chicago/Turabian StyleAndrianova, Nadezda V., Vasily A. Popkov, Natalia S. Klimenko, Alexander V. Tyakht, Galina V. Baydakova, Olga Y. Frolova, Ljubava D. Zorova, Irina B. Pevzner, Dmitry B. Zorov, and Egor Y. Plotnikov. 2020. "Microbiome-Metabolome Signature of Acute Kidney Injury" Metabolites 10, no. 4: 142. https://doi.org/10.3390/metabo10040142
APA StyleAndrianova, N. V., Popkov, V. A., Klimenko, N. S., Tyakht, A. V., Baydakova, G. V., Frolova, O. Y., Zorova, L. D., Pevzner, I. B., Zorov, D. B., & Plotnikov, E. Y. (2020). Microbiome-Metabolome Signature of Acute Kidney Injury. Metabolites, 10(4), 142. https://doi.org/10.3390/metabo10040142








