1. Introduction
Graph neural networks (GNNs) extend deep learning to graph-structured data by propagating and aggregating information along edges, enabling representation learning on non-Euclidean domains [
1,
2,
3]. They have achieved strong performance in diverse applications, including social networks [
4], bioinformatics [
5], and recommender systems [
6]. Existing GNNs are commonly categorized into spatial and spectral approaches. Spatial methods define neighborhood-wise aggregation rules [
4,
7,
8,
9,
10], whereas spectral methods derive convolution operators from graph signal processing and Laplacian spectra [
3,
11,
12,
13]. Both paradigms can be unified under the message-passing neural network (MPNN) framework [
14], where each layer updates node representations through permutation-invariant aggregation of neighborhood information.
Semi-supervised node classification is one of the most representative and practically important tasks for GNNs, where only a small subset of nodes is labeled and the goal is to infer labels for the remaining nodes by jointly exploiting node attributes and relational structure. In this setting, the graph provides an inductive bias for learning: neighborhood aggregation serves as a mechanism to propagate supervisory signals from labeled nodes to unlabeled ones, which can significantly improve generalization when labels are scarce. However, this advantage is inherently conditional on the quality of the relational structure: if observed edges are noisy, missing, or weakly aligned with class semantics, message passing may propagate misleading information and degrade class separability, especially under limited supervision [
15,
16]. Therefore, semi-supervised node classification offers a stringent testbed for studying the robustness of message passing under diverse graph structural regimes.
However, current semi-supervised node classification methods still face two major core limitations. First, many graph learning networks rely heavily on a given topology, or only reweight an existing graph, and the learned edge weights may inherit biases from the input adjacency [
9,
10,
17,
18,
19,
20,
21]. This issue becomes pronounced on heterophilous graphs, where connected nodes often carry different labels and observed edges are a weak proxy for semantic similarity. Second, local shared filters provide limited access to long-range yet semantically related nodes [
22,
23]. Although deeper and wider architectures have been explored [
24,
25,
26,
27], their effectiveness is often constrained by oversmoothing and by the difficulty of separating heterogeneous information as depth increases. Recent advances in graph homophily enhancement and non-local aggregation [
28,
29,
30,
31] improve long-range dependency modeling, but they typically depend on predefined structures or attention mechanisms under restricted spatial assumptions, leaving the geometry of node signals and the design space of relation operators insufficiently explored.
These two limitations are tightly coupled in practice. When the observed adjacency is unreliable (a common situation on heterophilous graphs), local propagation can introduce label-inconsistent messages; in such cases, merely reweighting observed edges may still be constrained by the same biased candidate set. Conversely, when one attempts to remedy locality by enlarging receptive fields, naive depth expansion may amplify oversmoothing and oversquashing effects, making it difficult to preserve discriminative information while aggregating broader contexts. This suggests that robust semi-supervised learning requires not only better aggregation operators, but also a principled way to control which node pairs communicate and how strongly they do so, under both homophilous and heterophilous regimes.
Motivated by the view that node signals can be regarded as samples from a latent geometric space, we ask whether message-passing probabilities can be learned from geometric relations among node representations rather than being tightly coupled to the observed adjacency. In particular, can mapping node signals into appropriate spaces mitigate correlation distortions induced by local graph convolution approximations, and can such relations induce a more informative topology for aggregation?
From this perspective, the observed graph can be treated as an imperfect measurement of latent relations, while node representations provide an alternative source of evidence for constructing communication patterns. If node signals admit an underlying geometric organization, then distances or inner products in an appropriate space can serve as a proxy for dependency strength and yield a topology that is more aligned with task semantics. Moreover, allowing multiple geometries provides additional flexibility: Euclidean spaces are naturally suited for locally smooth structures, while hyperbolic spaces can better represent uneven, hierarchical, or tree-like organizations that may arise in relational data. This motivates a unified framework that learns a geometry-aware topology from representations, and explicitly regulates the influence of the given adjacency rather than assuming it is fully reliable.
To address these questions, we propose a geometric graph learning paradigm that takes pairs of node signals, potentially after composite mappings across spaces, as input and outputs learnable communication probabilities. The paradigm supports three neighborhood regimes to compare local and non-local attention under both homophily and heterophily: (i) graph-free neighborhoods learned without relying on the input graph, (ii) local neighborhoods restricted to observed edges, and (iii) non-local neighborhoods that extend beyond the observed adjacency.
Based on this paradigm, we develop Geometric Graph Learning Network (G2LNet), a graph learning architecture compatible with MPNNs. G2LNet learns permutation-invariant node representations and infers geometric topologies via relation operators defined in Euclidean and hyperbolic spaces, together with aggregation schemes and constraint functions tailored to each neighborhood regime. We evaluate G2LNet on nine public benchmarks under multiple standard splits, including three homophilous citation networks and six heterophilous web and actor graphs. Experimental results demonstrate that the controlled variant of G2LNet consistently achieves significantly higher node classification accuracy compared to representative local and non-local benchmark models, confirming that geometric topology inference provides an effective pathway for robust message passing across graph mechanisms. Our main contributions include:
We propose G2LNet, a geometric graph learning framework that infers message-passing probabilities from geometric relations, reducing reliance on the observed adjacency.
We unify three neighborhood regimes within one architecture–graph-free, local, and non-local–enabling a controlled comparison of aggregation behaviors under homophily and heterophily.
We design Euclidean and hyperbolic relation operators (distance and inner-product) with a perceptual connectivity mechanism to regulate the influence of the input graph.
We introduce an end-to-end constraint objective to achieve stable structural learning. Experimental results demonstrate that the proposed architecture achieves superior classification performance to strong baseline models on most datasets.
3. Materials and Methods
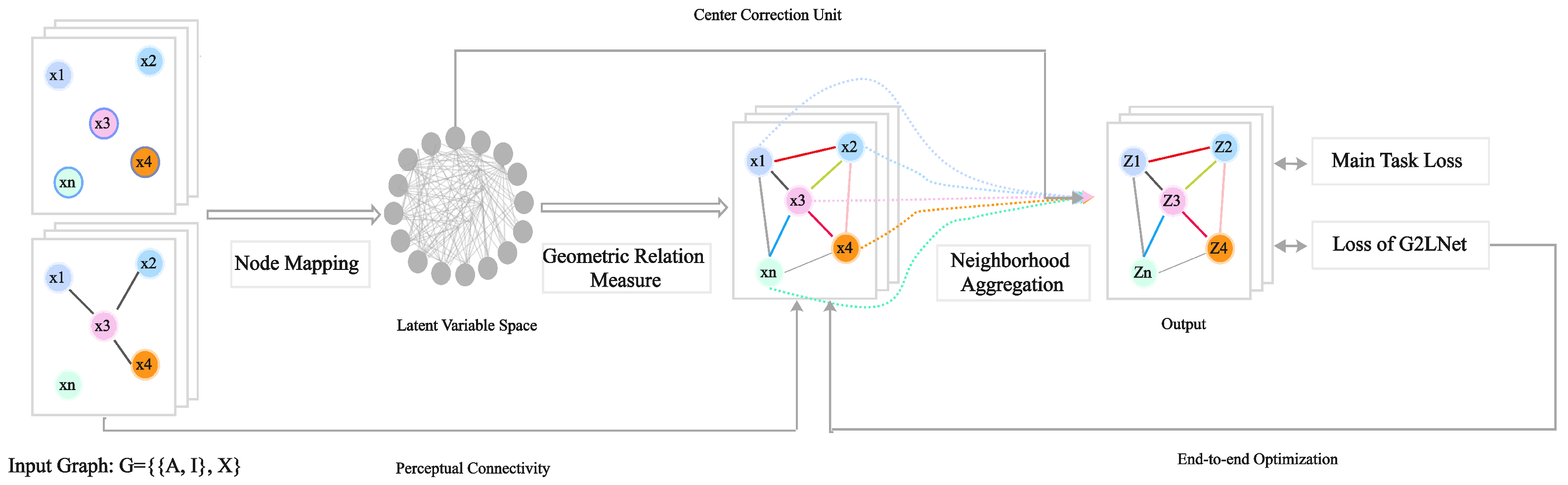
This section presents the proposed Geometric Graph Learning Network (G2LNet). As illustrated in
Figure 1, each layer consists of four components: (i) node mapping, (ii) geometric relation measure, (iii) neighborhood aggregation, and (iv) constraint regularization. The key idea is to infer a geometric topology–represented by learnable communication probabilities–from explicit relations between node representations in a chosen geometric space, and then perform message passing on the inferred neighborhoods. We denote the geometric space by the
and the network layer by
ℓ. Given node features
at layer
ℓ, G2LNet updates features in two coupled steps:
(1) Graph learning. In the graph learning phase, the graph learning function
P calculates the probability of information transfer
in the layer
-th, under the geometric space
based on the characteristic signals of the central node
and the neighboring node
. This probability is primarily determined by the metric function
and the constraint function
, defined as follows:
Extending the above equation to the entire graph constitutes a geometrically structured neighborhood
, where
, with the initial condition that
, and
S can be either the unitary matrix
I or the neighboring matrix of the graph
A. When
, it implies that the learning function
P captures geometric dependency relationships from the fully graph-free initialization to capture geometric dependencies. The mapping function
is used to project the signal from the nodes into the new feature space. The geometric relation function
combines the projected features and the previous message passing probabilities to update the passing relations between the nodes, while
preserves the selectivity of the prior topology (Perceptual Connectivity in
Figure 1). The constraint function
is responsible for optimizing the update process of the neighborhood of the geometric structure.
(2) Geometric neighborhood aggregation. In the geometric neighborhood aggregation phase, the central node aggregates in a weighted manner neighbor information from the geometric structured neighborhood
quantified by the graph learning function
P via the neighborhood aggregation function
to update its own feature representation:
where,
,
is the permutation-invariant aggregation function, such as summation, mean and maximum.
is a layer of MLP.
is Center Correction Unit in
Figure 1.
3.1. Node Mapping
Classical neural networks have shown significant advantages in dealing with smooth, localized and combinatorial problems in continuous spaces [
37]. Through local sensory fields, parameter sharing and hierarchical feature extraction, these networks are able to efficiently learn discriminant functions for abstract features. However, given the local smoothness of the graph signal, its non-Euclidean properties and its topological complexity, and the frequency domain approximation of the graph convolution operator in Euclidean space, this may lead to distortions in the local correlations and global patterns of the graph signal. Therefore, in order to better capture the underlying local structure and patterns of the node signals
on the graph, we construct the permutation-equivariant geometric mapping function
, which is composite of the projection function
and the embedding function
, given by the following equations:
Space Projection. For the projection function
, it maps the graph signal to the vector space
, i.e.:
When a nodal signal on the graph is projected into hyperbolic space, i.e.,
, we use the exponential mapping to project the feature
that is in Euclidean space to the Poincaré ball model
:
where
is the curvature of the
ℓ-th layer,
is the the origin, and
interpreted as a node located in the tangent space at the origin.
Feature Embedding. Next, we then use the embedding function
to reduce the dimensionality of the node signals
in this space and to improve the model’s ability to adapt to nonlinear structures. This embedding function is denoted as:
Usually the embedding function
is a layer of simple MLP networks [
10,
18], which can be simply expressed as:
, where
, and the MLP is shared among all nodes in the graph, and only the nodes themselves are considered without aggregating information from the neighborhood. In particular, when
, the
function should be carried out in hyperbolic space, i.e., a layer of HMLP embeddings is denoted as:
Here,
and
, the point-to-point computation of nonlinear activations can be simply realized in the Poincaré ball model using the tangential method:
Compared with the traditional MLP, the advantages of HMLP mainly come from the unique geometric properties of the hyperbolic space.
3.2. Geometric Relation Measure
We first empirically define the “potential geometric dependency between nodes” as the dependency that increases with the similarity between nodes. For in-depth analysis, the geometric metric function is then classified into two types based on the distance as well as based on the inner product using Gaussian and linear kernels.
Assuming that the node signals
and
have been embedded into the corresponding geometric space by the geometric mapping function
in Equation (
7), and then the graphs are computed using the geometric relation function
. In order to avoid the over-independence of the graphs at each layer as well as the influence of the heterogeneous graphs, we establish perceptual connectivity for the graphs at each layer:
where
,
,
is the perception factor, which serves as a hyperparameter in the network. Note that while the embedding function can still be a GCN network [
29], it is not suitable for deeper GCN networks–its limitation lies in the inability to learn affine functions capable of distinguishing features of nodes belonging to different categories (see Proof A2 in
Appendix B for details).
Distance metrics. Distance metrics are crucial in quantifying the similarity of data points. The kernel function, as an efficient similarity metric, possesses smoothness and locality, and its value decreases with increasing distance between nodes, reflecting the tendency of similarity to decrease with increasing distance. Specifically, the distance metric based on the kernel function can be defined as:
where
, it should be noted that
when
.
determines the rate at which the distance decays, with smaller making the effect of distance on similarity more significant and larger making the effect of distance relatively weaker. In this work, we take
in order to contrast it with the shortest path in hyperbolic space.
Inner Product Measure. Compared to distance metrics, inner product metrics focus more on how well the node feature vectors are aligned in the same space. To quantify this similarity, this paper uses Pearson’s correlation coefficient to explain the strength of the connection between nodes:
where,
and
are the mean and standard deviation of the eigenvectors
of node
, respectively, and
is a very small number that avoids dividing by 0. Since
, we perform the squaring operation. When
, dot product operation should be performed in tangent space, i.e.,:
.
3.3. Neighborhood Aggregation
In Graph Neural Networks (GNNs), the aggregation function, as a core component of information integration, is designed to efficiently extract the features of nodes and their local neighborhoods through an iterative process at each layer. The design of the aggregation function must follow Permutation Invariance (PI), which means that the aggregation result should remain unchanged no matter how the order of the nodes is changed. Given an input graph
, the graph learning function P constructs a geometrically structured neighborhood
under the joint action of the geometric metric function
in Equation (
13) and the constraint function
in Equation (
16), which constitutes a metric space connecting potentially similar nodes in the graph. We can directly use this neighborhood in the aggregation function to aggregate and update node features. In order to enhance the sensitivity of the aggregation function to different structural features, especially the local structural information of the nodes, we introduce a trick called the center correction unit
to adjust the features of the center node in the aggregation process. Ultimately, we designed an alignment-invariant aggregation function in Equation (
6) (see Proof A5 in
Appendix B).
3.4. Constraint Function
Learning a topology from features is inherently ill-posed: without explicit regularization, the inferred connectivity can easily collapse to (i) a nearly dense graph (trivial high-connectivity solution) or (ii) a highly smooth topology that accelerates oversmoothing in deep message passing [
36]. We therefore train G2LNet with a joint objective that combines the node classification loss and an end-to-end topology regularizer:
where
is the cross-entropy on labeled nodes, and
regularizes the learned communication probabilities
.
Firstly, to make the regularization well-defined across different embedding spaces, we explicitly specify the
spatial state in which feature discrepancies are measured. Let
denote the mapped node embedding used for topology inference. When
, we measure differences directly in Euclidean space; when
, we first project hyperbolic embeddings to the tangent space (at
) via the logarithmic map, so that standard vector norms are applicable. Concretely,
and we impose a Laplacian-style smoothness penalty that assigns large communication probability only when the mapped features are close:
where
stacks
,
is the (possibly normalized) Laplacian of
, and
controls the strength of smoothing. Since
is inferred from geometric relations measure, Equation (
18) regularizes how topology is formed, discouraging high weights between geometrically dissimilar nodes while still permitting non-local edges when they are geometrically supported.
Secondly, to prevent the inferred topology from degenerating into a dense graph and to enforce a selective communication pattern, we penalize the magnitude of
:
which is particularly important in the non-local and graph-free regimes where the candidate node pairs grow rapidly. If an undirected topology is required, we further add a symmetry regularizer:
where
controls the strength of symmetry enforcement. We treat this term as optional because directed communication can be beneficial when
P is interpreted as asymmetric information flow.
Finally, the topology regularizer at layer
is
Together, these constraints (i) couple topology inference to geometric feature consistency, (ii) avoid trivial dense solutions by enforcing a communication budget, and (iii) stabilize training by regulating propagation strength, thereby supporting topology inference under local, non-local, and graph-free regimes.
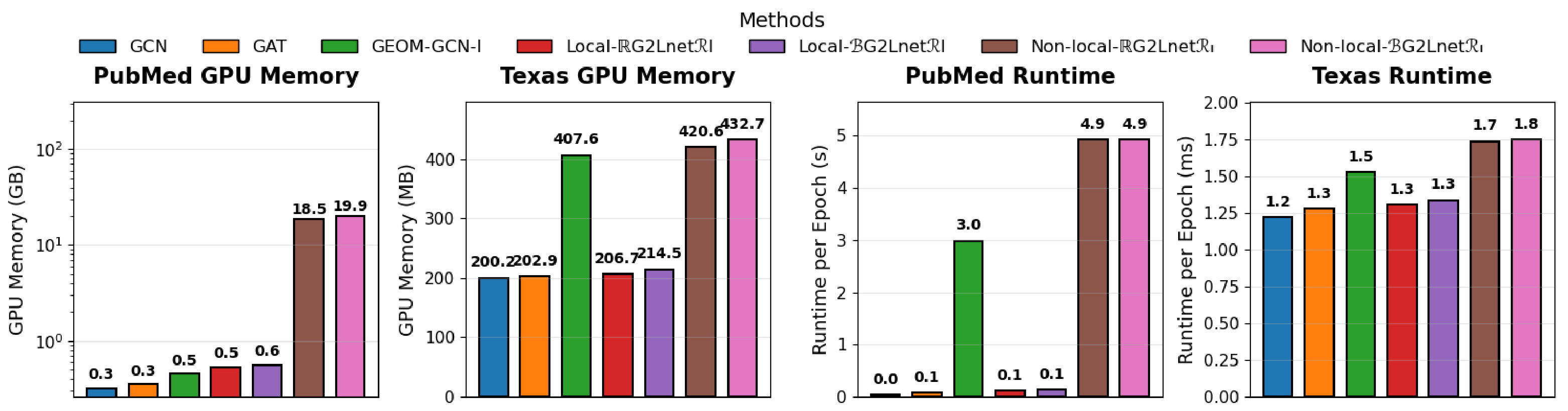
3.5. Computational Complexity Analysis
We compare G2LNet with representative local and non-local methods, focusing on the additional cost of relation computation. Let be the number of nodes, the number of observed edges, and f the embedding dimension.
Local Aggregation. When candidates are restricted to (i.e., pairs), relation computation scales as for inner-product relations and for distance relations (after embedding).
Non-local/graph-free Aggregation. When candidates include all pairs (or a large expanded set), relation computation scales as (inner-product) or (distance after embedding).
5. Conclusions
This paper proposes the Geometric Graph Learning Network (G2LNet), which treats the attention mechanism as a learnable communication probability induced by explicit geometric relationships between node representations, thereby enabling topological inference beyond observed adjacency relations. G2LNet unifies local, non-local, and agraph-based neighborhoods within a single architecture, supports relational operators in both Euclidean and hyperbolic spaces, and introduces an end-to-end constrained objective function to regularize inferred connections. Experiments across nine benchmarks demonstrate that G2LNet’s controlled variant achieves optimal classification performance.
Future work will prioritize scaling non-local and graph-free inference, developing principled criteria for selecting relation operators and neighborhood regimes, and extending the framework to dynamic and multi-relational graphs. Beyond semi-supervised node classification, the proposed geometric topology inference is applicable to settings with noisy or missing edges, including web-scale information networks, recommendation, and other heterogeneous relational learning problems.