Norpa Signalling and the Seasonal Circadian Locomotor Phenotype in Drosophila
Abstract
1. Introduction
2. Methods
3. Results
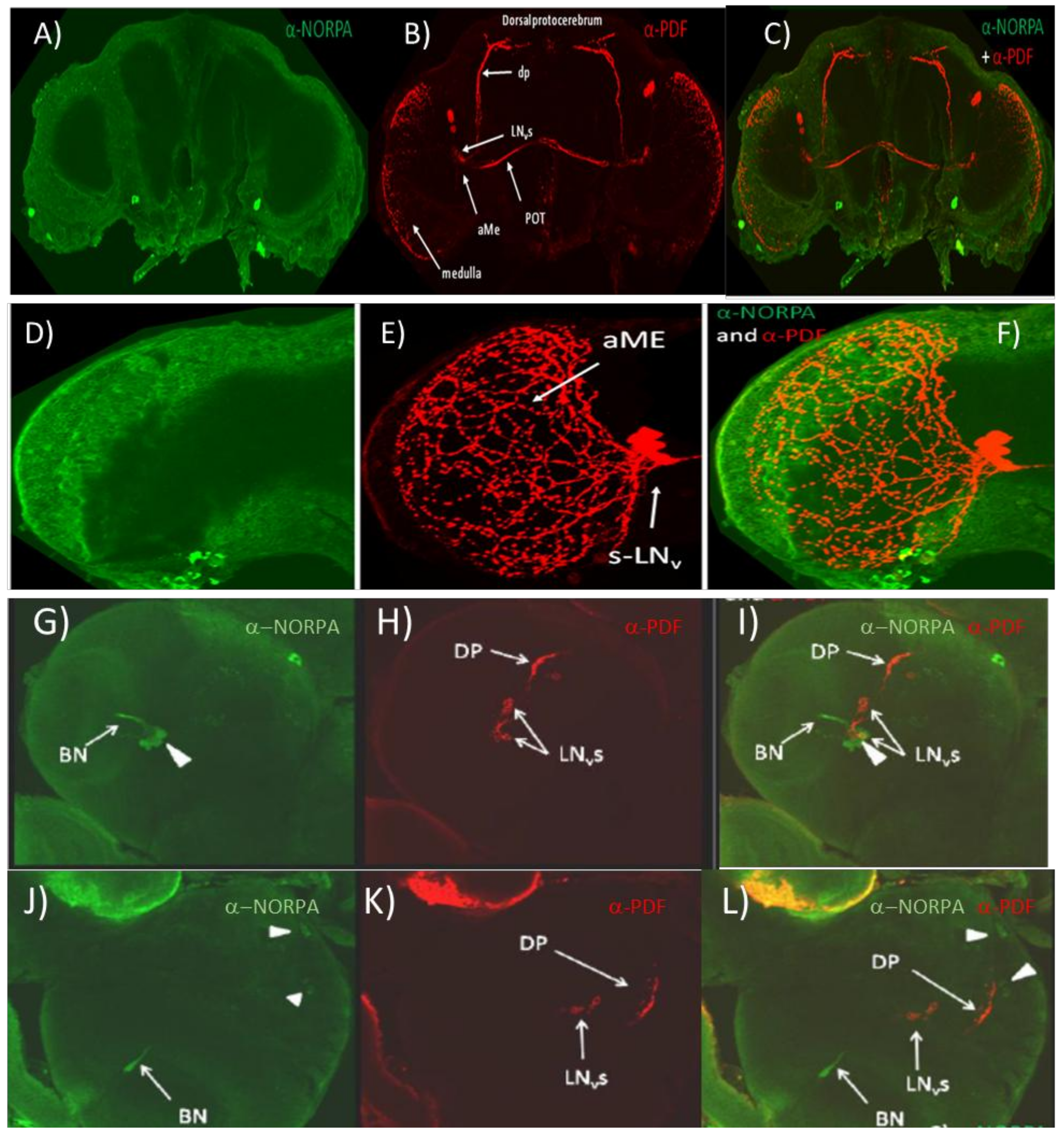
3.1. α-NORPA Antibody Does Not Penetrate into the Deeper Layers of the Adult CNS but Labels Neurons in the Larval Brain
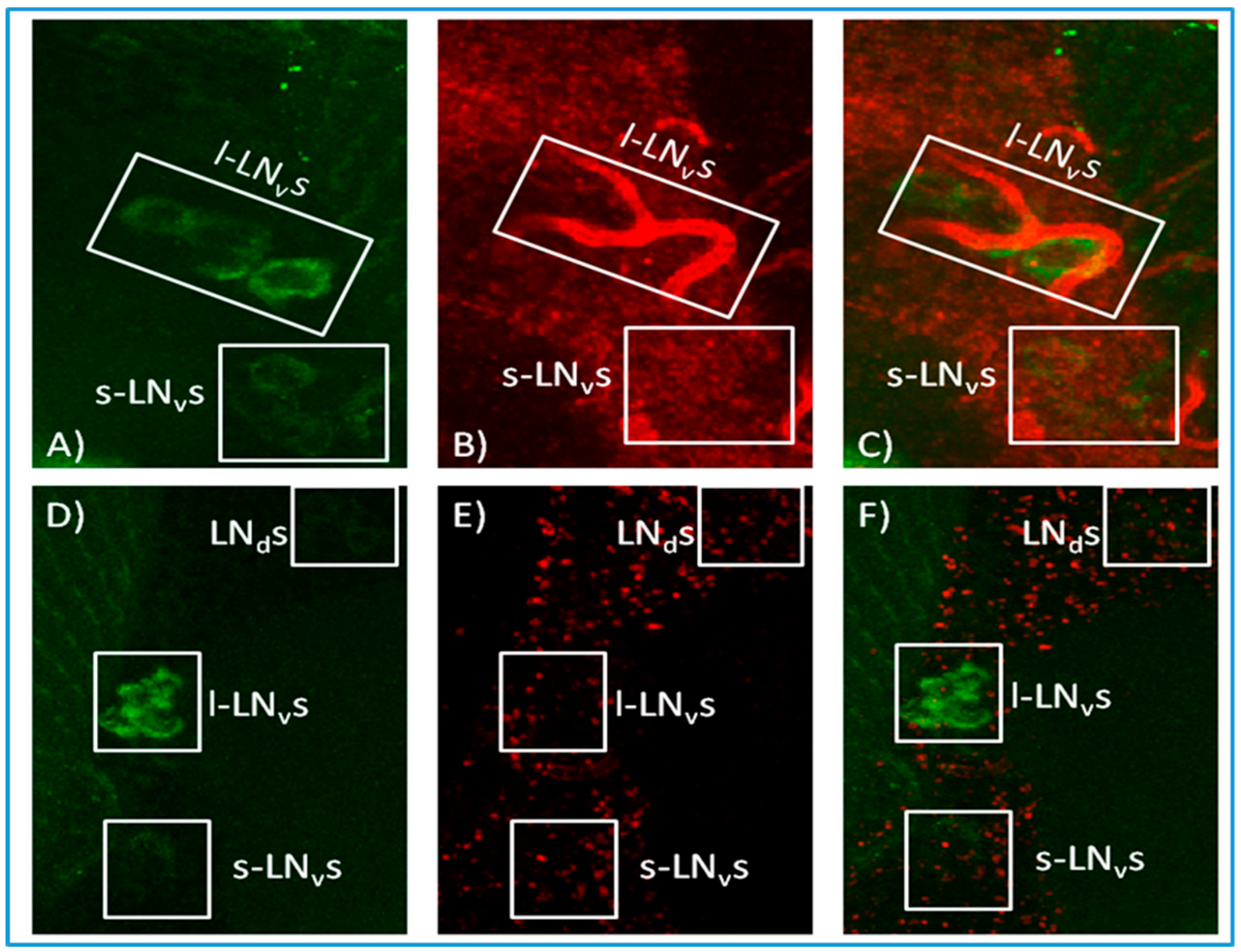
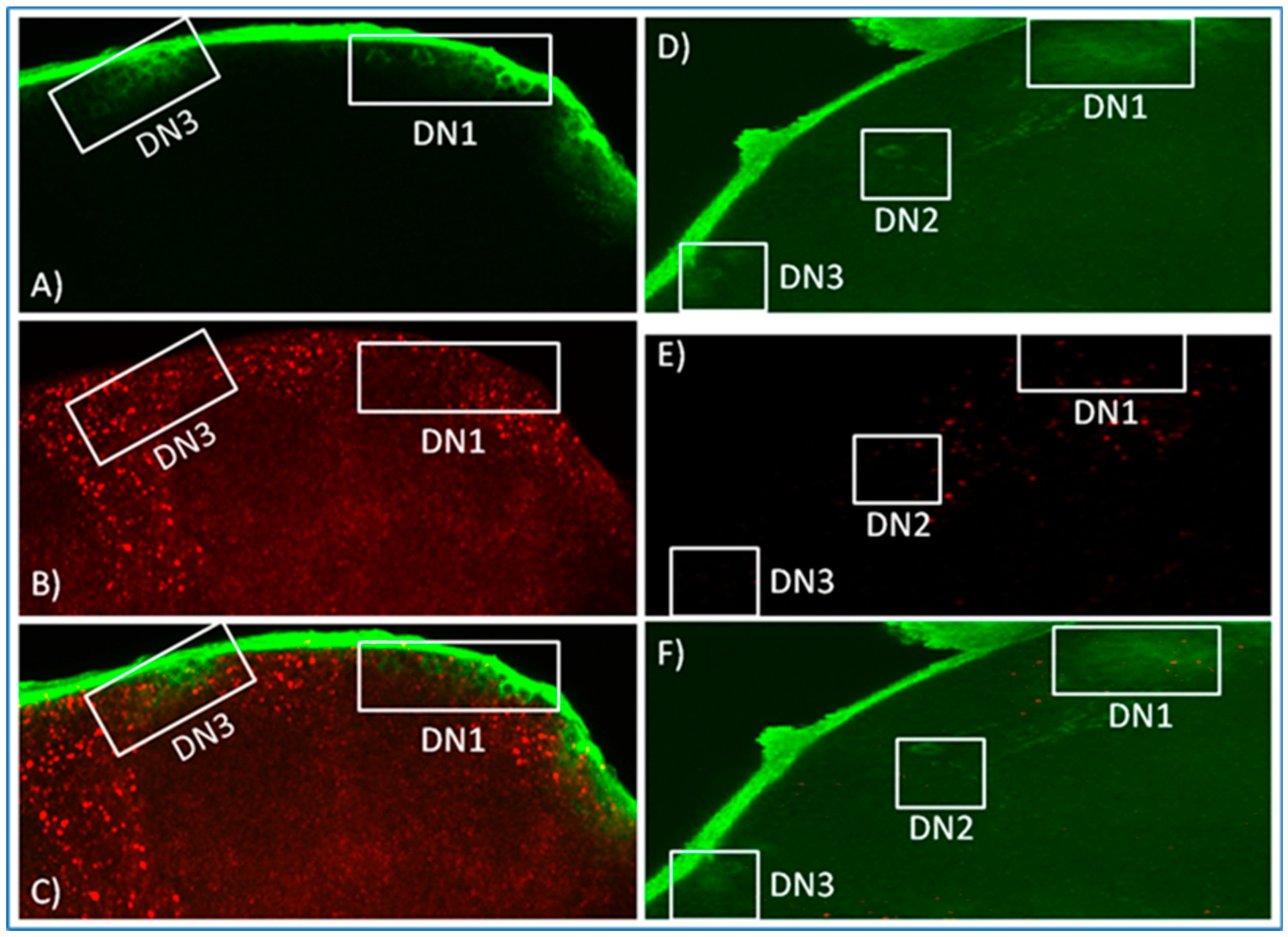
3.2. In Situ Hybridisation Reveals Expression of norpA in Clock Neurons
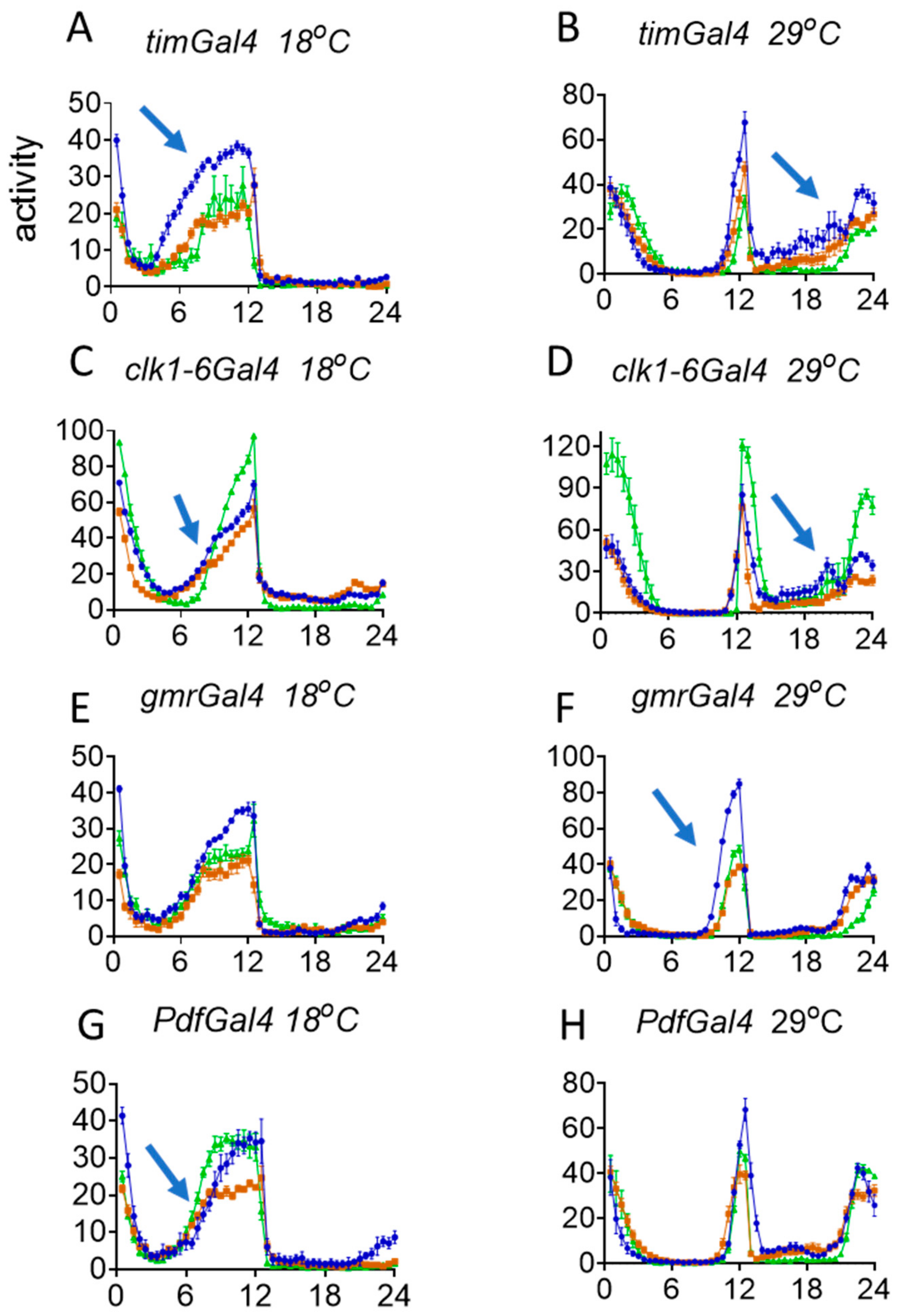
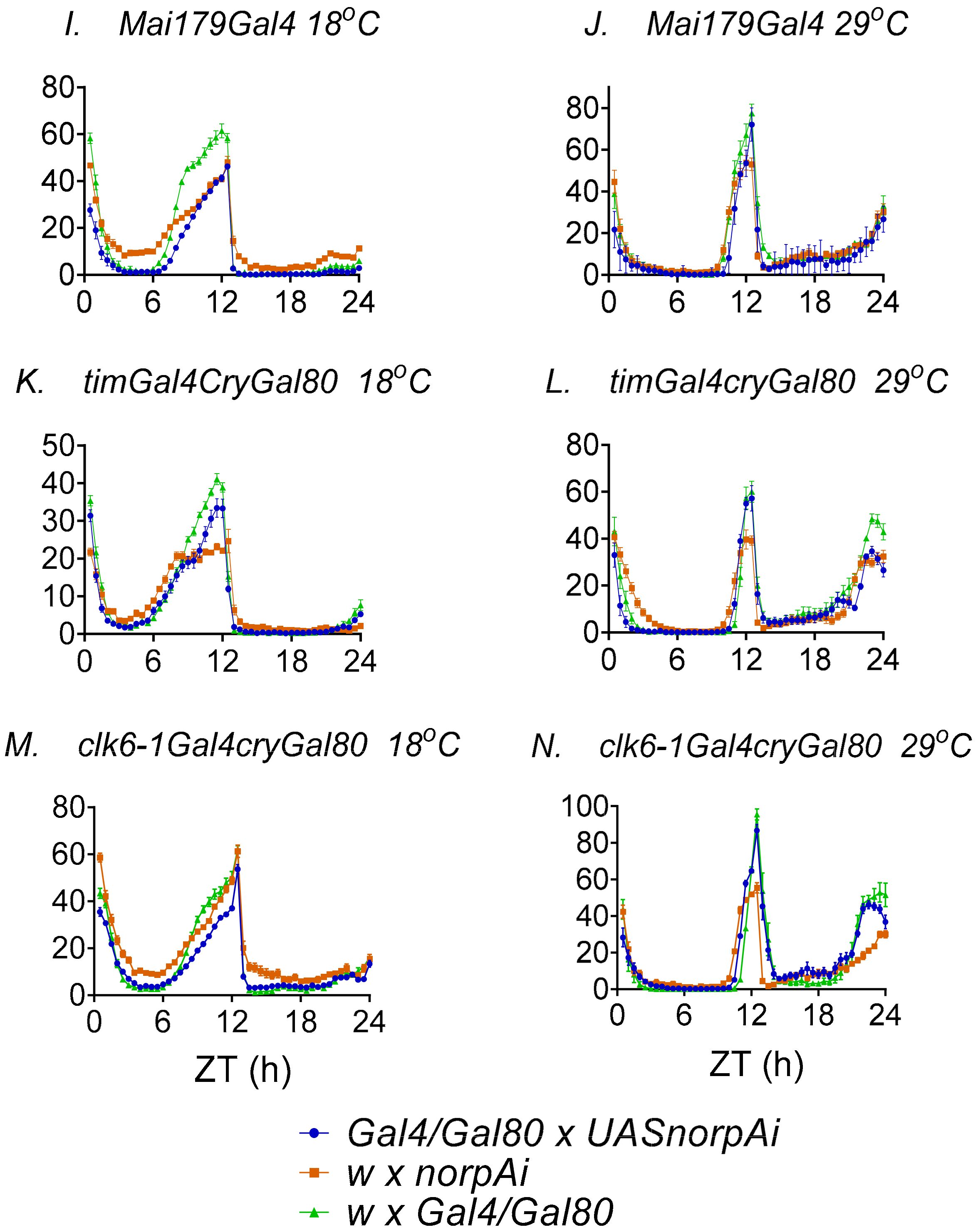
3.3. Behavioural Effects of Knockdown of norpA in Clock Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Pilorz, V.; Helfrich-Foerster, C.; Oster, H. The role of the circadian clock system in physiology. Pflügers Arch. Eur. J. Physiol. 2018, 470, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, Y.; Andersson, C.R.; Kondo, T.; Golden, S.S.; Johnson, C.H. Resonating circadian clocks enhance fitness in cyanobacteria. Proc. Natl. Acad. Sci. USA 1998, 95, 8660–8664. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, L.A.; Hennessy, J.M.; Peixoto, A.A.; Rosato, E.; Parkinson, H.; Costa, R.; Kyriacou, C.P. Natural Variation in a Drosophila Clock Gene and Temperature Compensation. Science 1997, 278, 2117–2120. [Google Scholar] [CrossRef] [PubMed]
- Tauber, E.; Zordan, M.; Sandrelli, F.; Pegoraro, M.; Osterwalder, N.; Breda, C.; Daga, A.; Selmin, A.; Monger, K.; Benna, C.; et al. Natural Selection Favors a Newly Derived timeless Allele in Drosophila melanogaster. Science 2007, 316, 1895–1898. [Google Scholar] [CrossRef]
- Zonato, V.; Vanin, S.; Costa, R.; Tauber, E.; Kyriacou, C.P. Inverse European Latitudinal Cline at the timeless Locus of Drosophila melanogaster Reveals Selection on a Clock Gene: Population Genetics of ls-tim. J. Biol. Rhythm. 2017, 33, 15–23. [Google Scholar] [CrossRef]
- Konopka, R.J.; Benzer, S. Clock Mutants of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1971, 68, 2112–2116. [Google Scholar] [CrossRef]
- Zehring, W.A.; Wheeler, D.A.; Reddy, P.; Konopka, R.J.; Kyriacou, C.P.; Rosbash, M.; Hall, J.C. P-element transformation with period locus DNA restores rhythmicity to mutant, arrhythmic Drosophila melanogaster. Cell 1984, 39, 369–376. [Google Scholar] [CrossRef]
- Bargiello, T.A.; Jackson, F.R.; Young, M.W. Restoration of circadian behavioural rhythms by gene transfer in Drosophila. Nature 1984, 312, 752–754. [Google Scholar] [CrossRef]
- E Hardin, P.; Hall, J.C.; Rosbash, M. Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature 1990, 343, 536–540. [Google Scholar] [CrossRef]
- Takahashi, J.S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 2016, 18, 164–179. [Google Scholar] [CrossRef]
- Callaway, E.; Ledford, H. Medicine Nobel awarded for work on circadian clocks. Nature 2017, 550, 18. [Google Scholar] [CrossRef] [PubMed]
- Dubowy, C.; Sehgal, A. Circadian Rhythms and Sleep in Drosophila melanogaster. Genetics 2017, 205, 1373–1397. [Google Scholar] [CrossRef] [PubMed]
- Koh, K.; Zheng, X.; Sehgal, A. JETLAG Resets the Drosophila Circadian Clock by Promoting Light-Induced Degradation of TIMELESS. Science 2006, 312, 1809–1812. [Google Scholar] [CrossRef] [PubMed]
- Peschel, N.; Veleri, S.; Stanewsky, R. Veela defines a molecular link between Cryptochrome and Timeless in the light-input pathway to Drosophila’s circadian clock. Proc. Natl. Acad. Sci. USA 2006, 103, 17313–17318. [Google Scholar] [CrossRef] [PubMed]
- Peschel, N.; Chen, K.-F.; Szabo, G.; Stanewsky, R. Light-Dependent Interactions between the Drosophila Circadian Clock Factors Cryptochrome, Jetlag, and Timeless. Curr. Biol. 2009, 19, 241–247. [Google Scholar] [CrossRef] [PubMed]
- Ozturk, N.; VanVickle-Chavez, S.J.; Akileswaran, L.; Van Gelder, R.N.; Sancar, A. Ramshackle (Brwd3) promotes light-induced ubiquitylation of Drosophila Cryptochrome by DDB1-CUL4-ROC1 E3 ligase complex. Proc. Natl. Acad. Sci. USA 2013, 110, 4980–4985. [Google Scholar] [CrossRef]
- Ceriani, M.F.; Darlington, T.K.; Staknis, D.; Más, P.; Petti, A.A.; Weitz, C.J.; Kay, S.A. Light-Dependent Sequestration of TIMELESS by CRYPTOCHROME. Science 1999, 285, 553–556. [Google Scholar] [CrossRef]
- Plautz, J.D.; Kaneko, M.; Hall, J.C.; Kay, S.A. Independent Photoreceptive Circadian Clocks Throughout Drosophila. Science 1997, 278, 1632–1635. [Google Scholar] [CrossRef]
- Ni, J.D.; Gurav, A.S.; Liu, W.; Ogunmowo, T.H.; Hackbart, H.; Elsheikh, A.; A Verdegaal, A.; Montell, C. Differential regulation of the Drosophila sleep homeostat by circadian and arousal inputs. eLife 2019, 8. [Google Scholar] [CrossRef]
- Grima, B.; Chélot, E.; Xia, R.; Rouyer, F. Morning and evening peaks of activity rely on different clock neurons of the Drosophila brain. Nature 2004, 431, 869–873. [Google Scholar] [CrossRef]
- Stoleru, D.; Peng, Y.; Agosto, J.; Rosbash, M. Coupled oscillators control morning and evening locomotor behaviour of Drosophila. Nature 2004, 431, 862–868. [Google Scholar] [CrossRef] [PubMed]
- Yao, Z.; Shafer, O.T. The Drosophila Circadian Clock Is a Variably Coupled Network of Multiple Peptidergic Units. Science 2014, 343, 1516–1520. [Google Scholar] [CrossRef] [PubMed]
- Dissel, S.; Hansen, C.N.; Özkaya, Ö.; Hemsley, M.; Kyriacou, C.P.; Rosato, E. The logic of circadian organization in Drosophila. Curr. Biol. 2014, 24, 2257–2266. [Google Scholar] [CrossRef] [PubMed]
- Yao, Z.; Bennett, A.J.; Clem, J.L.; Shafer, O.T. The Drosophila Clock Neuron Network Features Diverse Coupling Modes and Requires Network-wide Coherence for Robust Circadian Rhythms. Cell Rep. 2016, 17, 2873–2881. [Google Scholar] [CrossRef]
- Liang, X.; Holy, T.E.; Taghert, P.H. A Series of Suppressive Signals within the Drosophila Circadian Neural Circuit Generates Sequential Daily Outputs. Neuron 2017, 94, 1173–1189.e4. [Google Scholar] [CrossRef]
- King, A.N.; Sehgal, A. Molecular and circuit mechanisms mediating circadian clock output in the Drosophila brain. Eur. J. Neurosci. 2018, 51, 268–281. [Google Scholar] [CrossRef]
- Yoshii, T.; Todo, T.; Wülbeck, C.; Stanewsky, R.; Helfrich-Foerster, C. Cryptochrome is present in the compound eyes and a subset of Drosophila’s clock neurons. J. Comp. Neurol. 2008, 508, 952–966. [Google Scholar] [CrossRef]
- Stanewsky, R.; Kaneko, M.; Emery, P.; Beretta, B.; Wager-Smith, K.; A Kay, S.; Rosbash, M.; Hall, J.C. The cryb mutation identifies cryptochrome as a circadian photoreceptor in Drosophila. Cell 1998, 95, 681–692. [Google Scholar] [CrossRef]
- Senthilan, P.; Grebler, R.; Reinhard, N.; Rieger, D.; Helfrich-Foerster, C. Role of Rhodopsins as Circadian Photoreceptors in the Drosophila melanogaster. Biology 2019, 8, 6. [Google Scholar] [CrossRef]
- Montell, C. Drosophila visual transduction. Trends Neurosci. 2012, 35, 356–363. [Google Scholar] [CrossRef]
- Ni, J.D.; Baik, L.S.; Holmes, T.C.; Montell, C. A rhodopsin in the brain functions in circadian photoentrainment in Drosophila. Nature 2017, 545, 340–344. [Google Scholar] [CrossRef] [PubMed]
- Davis, F.P.; Nern, A.; Picard, S.; Reiser, M.B.; Rubin, G.M.; Eddy, S.R.; Henry, G.L. A genetic, genomic, and computational resource for exploring neural circuit function. eLife 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Ogueta, M.; Hardie, R.C.; Stanewsky, R. Non-canonical Phototransduction Mediates Synchronization of the Drosophila melanogaster Circadian Clock and Retinal Light Responses. Curr. Biol. 2018, 28, 1725–1735.e3. [Google Scholar] [CrossRef] [PubMed]
- Lachaise, D.; Cariou, M.-L.; David, J.R.; Lemeunier, F.; Tsacas, L.; Ashburner, M. Historical Biogeography of the Drosophila melanogaster Species Subgroup. Evol. Biol. 1988, 22, 159–225. [Google Scholar] [CrossRef]
- Vanin, S.; Bhutani, S.; Montelli, S.; Menegazzi, P.; Green, E.W.; Pegoraro, M.; Sandrelli, F.; Costa, R.; Kyriacou, C.P. Unexpected features of Drosophila circadian behavioural rhythms under natural conditions. Nature 2012, 484, 371–375. [Google Scholar] [CrossRef]
- Green, E.W.; O’Callaghan, E.K.; Hansen, C.N.; Bastianello, S.; Bhutani, S.; Vanin, S.; Armstrong, D.; Costa, R.; Kyriacou, C.P. Drosophila circadian rhythms in seminatural environments: Summer afternoon component is not an artifact and requires TrpA1 channels. Proc. Natl. Acad. Sci. USA 2015, 112, 8702–8707. [Google Scholar] [CrossRef]
- Menegazzi, P.; Yoshii, T.; Helfrich-Foerster, C. Laboratory versus nature: The two sides of the Drosophila circadian clock. J. Biol. Rhythm. 2012, 27, 433–442. [Google Scholar] [CrossRef]
- Wheeler, D.A.; Hamblen-Coyle, M.J.; Dushay, M.S.; Hall, J.C. Behavior in Light-Dark Cycles of Drosophila Mutants That Are Arrhythmic, Blind, or Both. J. Biol. Rhythm. 1993, 8, 67–94. [Google Scholar] [CrossRef]
- Glaser, F.T.; Stanewsky, R. Temperature Synchronization of the Drosophila Circadian Clock. Curr. Biol. 2005, 15, 1352–1363. [Google Scholar] [CrossRef]
- Azevedo, R.V.; Hansen, C.N.; Chen, K.; Rosato, E.; Kyraicou, C.P. Disrupted gultamate signalling in Drosophila generates rhythms in constant light. Front. Physiol. 2020, 11, 145. [Google Scholar] [CrossRef]
- Sehadova, H.; Glaser, F.T.; Gentile, C.; Simoni, A.; Giesecke, A.; Albert, J.; Stanewsky, R. Temperature Entrainment of Drosophila’s Circadian Clock Involves the Gene nocte and Signaling from Peripheral Sensory Tissues to the Brain. Neuron 2009, 64, 251–266. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Xu, M.; Anantaprakorn, Y.; Rosing, M.; Stanewsky, R. nocte Is Required for Integrating Light and Temperature Inputs in Circadian Clock Neurons of Drosophila. Curr. Biol. 2018, 28, 1595–1605.e3. [Google Scholar] [CrossRef] [PubMed]
- Simoni, A.; Wolfgang, W.; Topping, M.P.; Kavlie, R.G.; Stanewsky, R.; Albert, J.T. A Mechanosensory Pathway to the Drosophila Circadian Clock. Science 2014, 343, 525–528. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Buhl, E.; Xu, M.; Croset, V.; Rees, J.S.; Lilley, K.S.; Benton, R.; Hodge, J.J.L.; Stanewsky, R. Drosophila Ionotropic Receptor 25a mediates circadian clock resetting by temperature. Nature 2015, 527, 516–520. [Google Scholar] [CrossRef]
- Majercak, J.; Sidote, D.; E Hardin, P.; Edery, I. How a Circadian Clock Adapts to Seasonal Decreases in Temperature and Day Length. Neuron 1999, 24, 219–230. [Google Scholar] [CrossRef]
- Low, K.H.; Lim, C.; Ko, H.W.; Edery, I. Natural Variation in the Splice Site Strength of a Clock Gene and Species-Specific Thermal Adaptation. Neuron 2008, 60, 1054–1067. [Google Scholar] [CrossRef]
- Cao, W.; Edery, I. A Novel Pathway for Sensory-Mediated Arousal Involves Splicing of an Intron in the period Clock Gene. Sleep 2015, 38, 41–51. [Google Scholar] [CrossRef]
- Low, K.H.; Chen, W.-F.; Yildirim, E.; Edery, I. Natural Variation in the Drosophila melanogaster Clock Gene Period Modulates Splicing of Its 3′-Terminal Intron and Mid-Day Siesta. PLoS ONE 2012, 7, e49536. [Google Scholar] [CrossRef]
- Cao, W.; Edery, I. Mid-day siesta in natural populations of D. melanogaster from Africa exhibits an altitudinal cline and is regulated by splicing of a thermosensitive intron in the period clock gene. BMC Evol. Biol. 2017, 17, 32. [Google Scholar] [CrossRef]
- Yang, Y.; Edery, I. Parallel clinal variation in the mid-day siesta of Drosophila melanogaster implicates continent-specific targets of natural selection. PLoS Genet. 2018, 14, e1007612. [Google Scholar] [CrossRef]
- Collins, B.H.; Rosato, E.; Kyriacou, C.P. Seasonal behavior in Drosophila melanogaster requires the photoreceptors, the circadian clock, and phospholipase C. Proc. Natl. Acad. Sci. USA 2004, 101, 1945–1950. [Google Scholar] [CrossRef] [PubMed]
- Majercak, J.; Chen, W.-F.; Edery, I. Splicing of the period Gene 3′-Terminal Intron Is Regulated by Light, Circadian Clock Factors, and Phospholipase C. Mol. Cell. Biol. 2004, 24, 3359–3372. [Google Scholar] [CrossRef] [PubMed]
- Bloomquist, B.; Shortridge, R.; Schneuwly, S.; Perdew, M.; Montell, C.; Steller, H.; Rubin, G.M.; Pak, W. Isolation of a putative phospholipase c gene of Drosophila, norpA, and its role in phototransduction. Cell 1988, 54, 723–733. [Google Scholar] [CrossRef]
- Zhu, L.; McKay, R.R.; Shortridge, R.D. Tissue-specific expression of phospholipase C encoded by the norpA gene of Drosophila melanogaster. J. Biol. Chem. 1993, 268, 15994–16001. [Google Scholar]
- Kim, S.; McKay, R.R.; Miller, K.; Shortridge, R.D. Multiple Subtypes of Phospholipase C Are Encoded by the norpA Gene of Drosophila melanogaster. J. Biol. Chem. 1995, 270, 14376–14382. [Google Scholar] [CrossRef]
- McKay, R.R.; Chen, D.-M.; Miller, K.; Kim, S.; Stark, W.S.; Shortridge, R.D. Phospholipase C Rescues Visual Defect in norpA Mutant of Drosophila melanogaster. J. Biol. Chem. 1995, 270, 13271–13276. [Google Scholar] [CrossRef]
- Gummadova, J.O.; Coutts, G.A.; Glossop, N.R.J. Analysis of the Drosophila Clock Promoter Reveals Heterogeneity in Expression between Subgroups of Central Oscillator Cells and Identifies a Novel Enhancer Region. J. Biol. Rhythm. 2009, 24, 353–367. [Google Scholar] [CrossRef]
- Collins, B.H.; Dissel, S.; Gaten, E.; Rosato, E.; Kyriacou, C.P. Disruption of Cryptochrome partially restores circadian rhythmicity to the arrhythmic period mutant of Drosophila. Proc. Natl. Acad. Sci. USA 2005, 102, 19021–19026. [Google Scholar] [CrossRef]
- Wülbeck, C.; Helfrich-Foerster, C. RNA In Situ Hybridizations on Drosophila Whole Mounts. Adv. Struct. Saf. Stud. 2007, 362, 495–511. [Google Scholar] [CrossRef]
- Malpel, S.; Klarsfeld, A.; Rouyer, F. Larval optic nerve and adult extra-retinal photoreceptors sequentially associate with clock neurons during Drosophila brain development. Development 2002, 129, 1443–1453. [Google Scholar]
- Shortridge, R.D.; Yoon, J.; Lending, C.R.; Bloomquist, B.T.; Perdew, M.H.; Pak, W.L. A Drosophila phospholipase C gene that is expressed in the central nervous system. J. Biol. Chem. 1991, 266, 12474–12480. [Google Scholar] [PubMed]
- Boothroyd, C.E.; Wijnen, H.; Naef, F.; Saez, L.; Young, M.W. Integration of Light and Temperature in the Regulation of Circadian Gene Expression in Drosophila. PLoS Genet. 2007, 3, e54. [Google Scholar] [CrossRef] [PubMed]
- Montelli, S.; Mazzotta, G.M.; Vanin, S.; Caccin, L.; Corrà, S.; De Pitta, C.; Boothroyd, C.; Green, E.W.; Kyriacou, C.P.; Costa, R. period and timeless mRNA Splicing Profiles under Natural Conditions in Drosophila melanogaster. J. Biol. Rhythm. 2015, 30, 217–227. [Google Scholar] [CrossRef]
- Zhang, L.; Chung, B.Y.; Lear, B.C.; Kilman, V.L.; Liu, Y.; Mahesh, G.; Meissner, R.-A.; E Hardin, P.; Allada, R. DN1p Circadian Neurons Coordinate Acute Light and PDF Inputs to Produce Robust Daily Behavior in Drosophila. Curr. Biol. 2010, 20, 591–599. [Google Scholar] [CrossRef][Green Version]
- Yadlapalli, S.; Jiang, C.; Bahle, A.; Reddy, P.; Meyhofer, E.; Shafer, O.T. Circadian clock neurons constantly monitor environmental temperature to set sleep timing. Nature 2018, 555, 98–102. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Yu, J.; Jung, H.J.; Abruzzi, K.C.; Luo, W.; Griffith, L.C.; Rosbash, M. Circadian neuron feedback controls the Drosophila sleep–activity profile. Nature 2016, 536, 292–297. [Google Scholar] [CrossRef] [PubMed]
- Abruzzi, K.; Zadina, A.; Luo, W.; Wiyanto, E.; Rahman, R.; Guo, F.; Shafer, O.; Rosbash, M. RNA-seq analysis of Drosophila clock and non-clock neurons reveals neuron-specific cycling and novel candidate neuropeptides. PLoS Genet. 2017, 13, e1006613. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Cao, W.; Edery, I. The SR protein B52/SRp55 regulates splicing of the period thermosensitive intron and mid-day siesta in Drosophila. Sci. Rep. 2018, 8, 1872. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Breda, C.; Rosato, E.; Kyriacou, C.P. Norpa Signalling and the Seasonal Circadian Locomotor Phenotype in Drosophila. Biology 2020, 9, 130. https://doi.org/10.3390/biology9060130
Breda C, Rosato E, Kyriacou CP. Norpa Signalling and the Seasonal Circadian Locomotor Phenotype in Drosophila. Biology. 2020; 9(6):130. https://doi.org/10.3390/biology9060130
Chicago/Turabian StyleBreda, Carlo, Ezio Rosato, and Charalambos P. Kyriacou. 2020. "Norpa Signalling and the Seasonal Circadian Locomotor Phenotype in Drosophila" Biology 9, no. 6: 130. https://doi.org/10.3390/biology9060130
APA StyleBreda, C., Rosato, E., & Kyriacou, C. P. (2020). Norpa Signalling and the Seasonal Circadian Locomotor Phenotype in Drosophila. Biology, 9(6), 130. https://doi.org/10.3390/biology9060130




