3D Imaging and Quantitative Characterization of Mouse Capillary Coronary Network Architecture
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Model
2.2. Arterial Pressure Measurements
2.3. Mice Preparation
2.4. Optical Clearing
2.5. Shrinkage Measurement
2.6. Image Acquisition
2.6.1. Light Sheet Microscopy
2.6.2. Confocal Microscopy
2.7. Image Processing
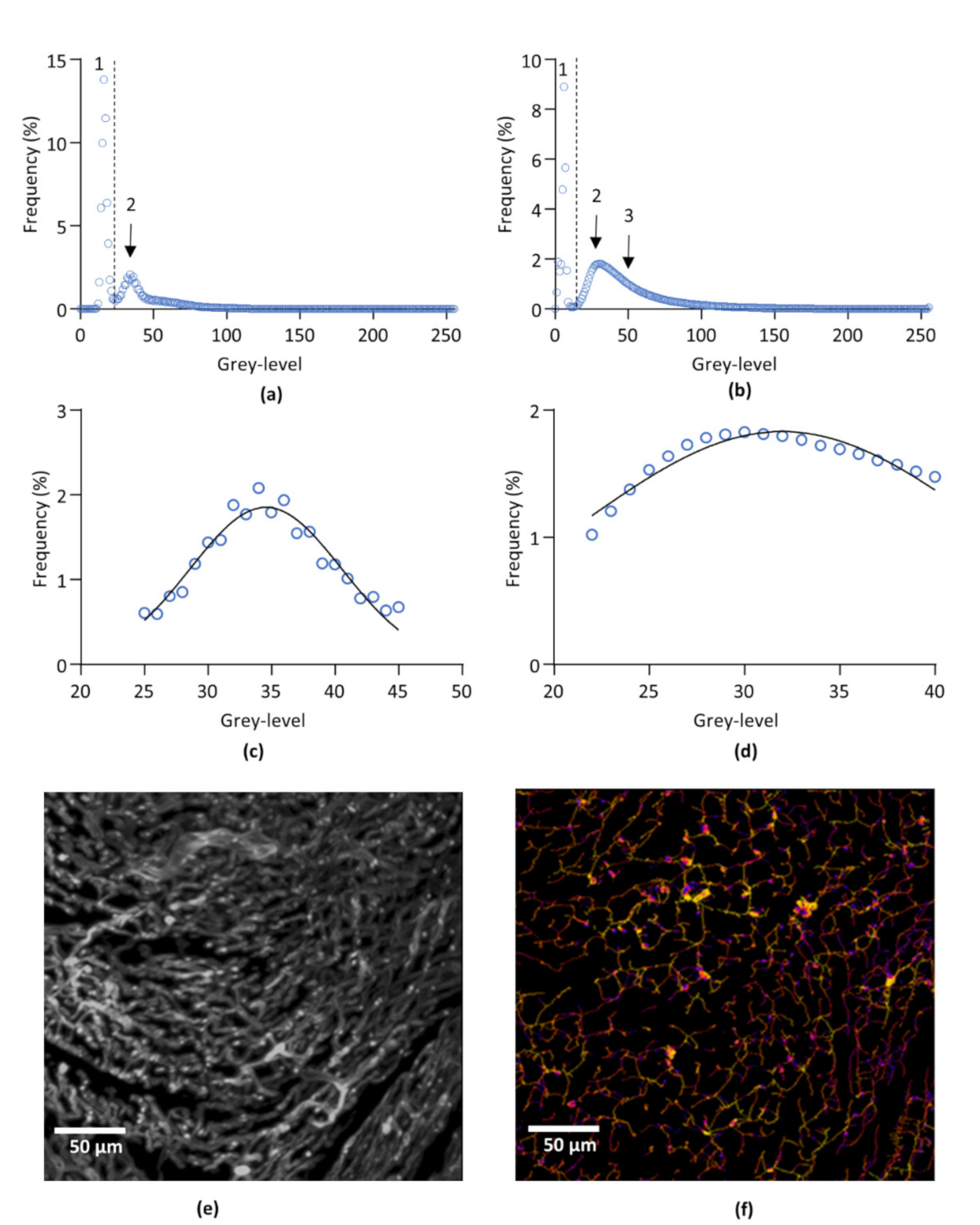
2.7.1. Segmentations
2.7.2. Filtering
2.7.3. Skeletonization and Distance Mapping
2.8. Data Analysis
2.8.1. Cardiac Volumes and Vascular Density
2.8.2. Fractal Dimension
2.8.3. Normalized Number of Segments
2.8.4. Normalized Total Capillary Length
2.8.5. Normalized Number and Percentage of Nodes
2.8.6. Segment Diameter
2.8.7. Tortuosity
2.9. Statistical Analysis
2.9.1. Global Parameters
2.9.2. Topological Parameters
3. Results
3.1. Cardiovascular Parameters
3.2. Shrinkage
3.3. Global Parameters
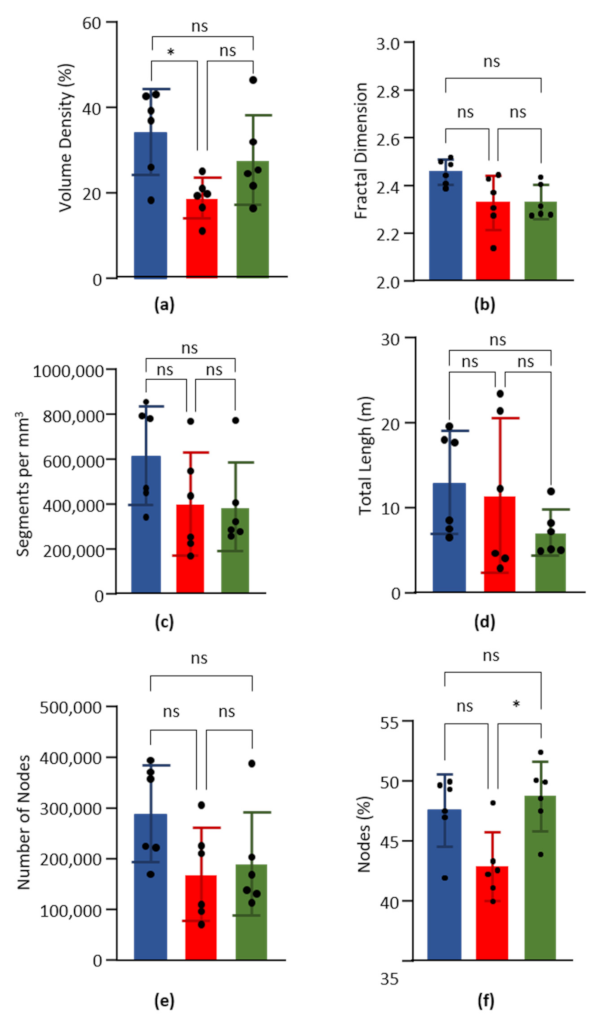
3.3.1. Vascular Density
3.3.2. Fractal Dimension
3.3.3. Normalized Number of Segments
3.3.4. Total Capillary Length
3.3.5. Number and Percentage of Nodes
3.4. Topological Parameters
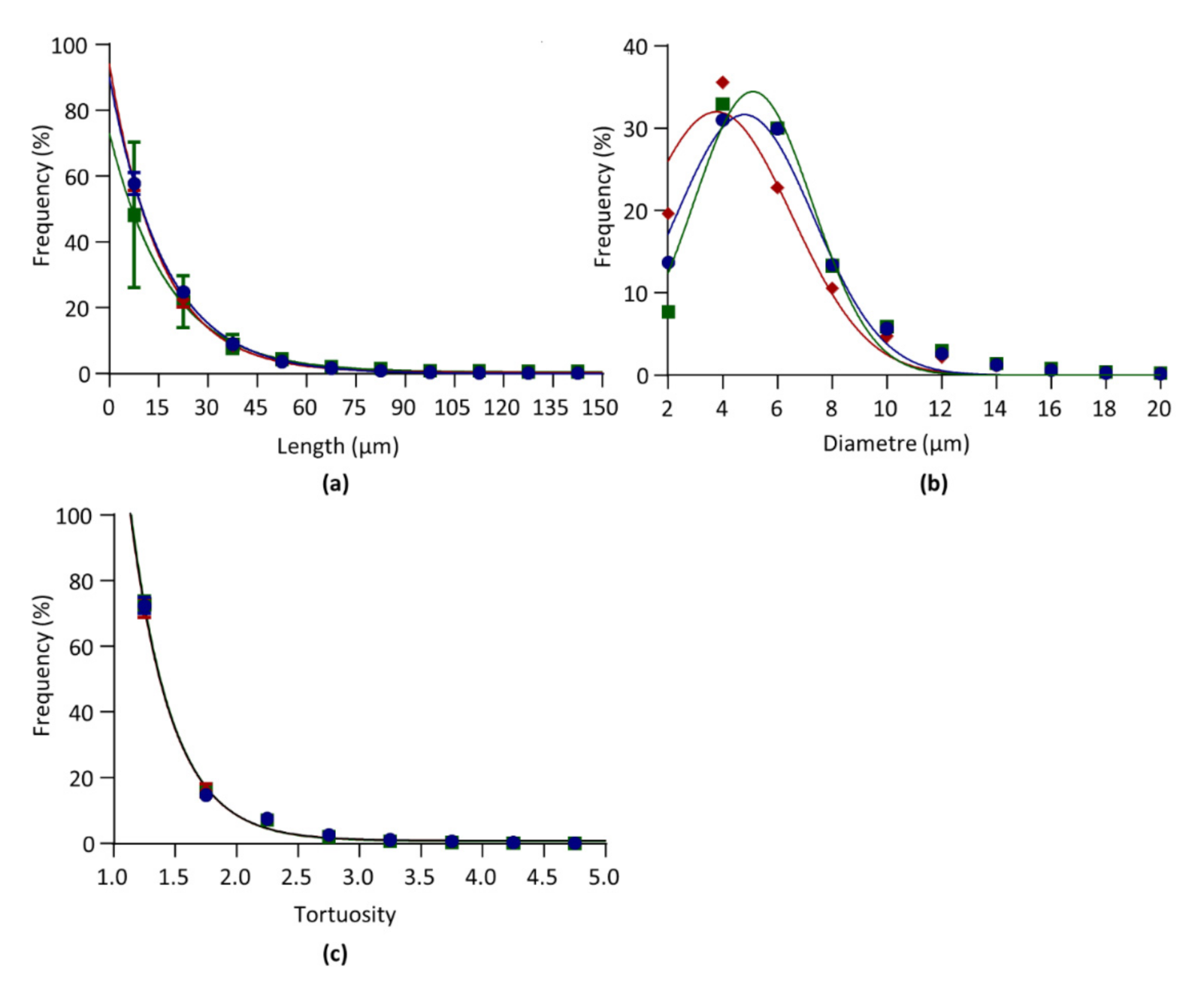
3.4.1. Segment Length
3.4.2. Diameter
3.4.3. Tortuosity
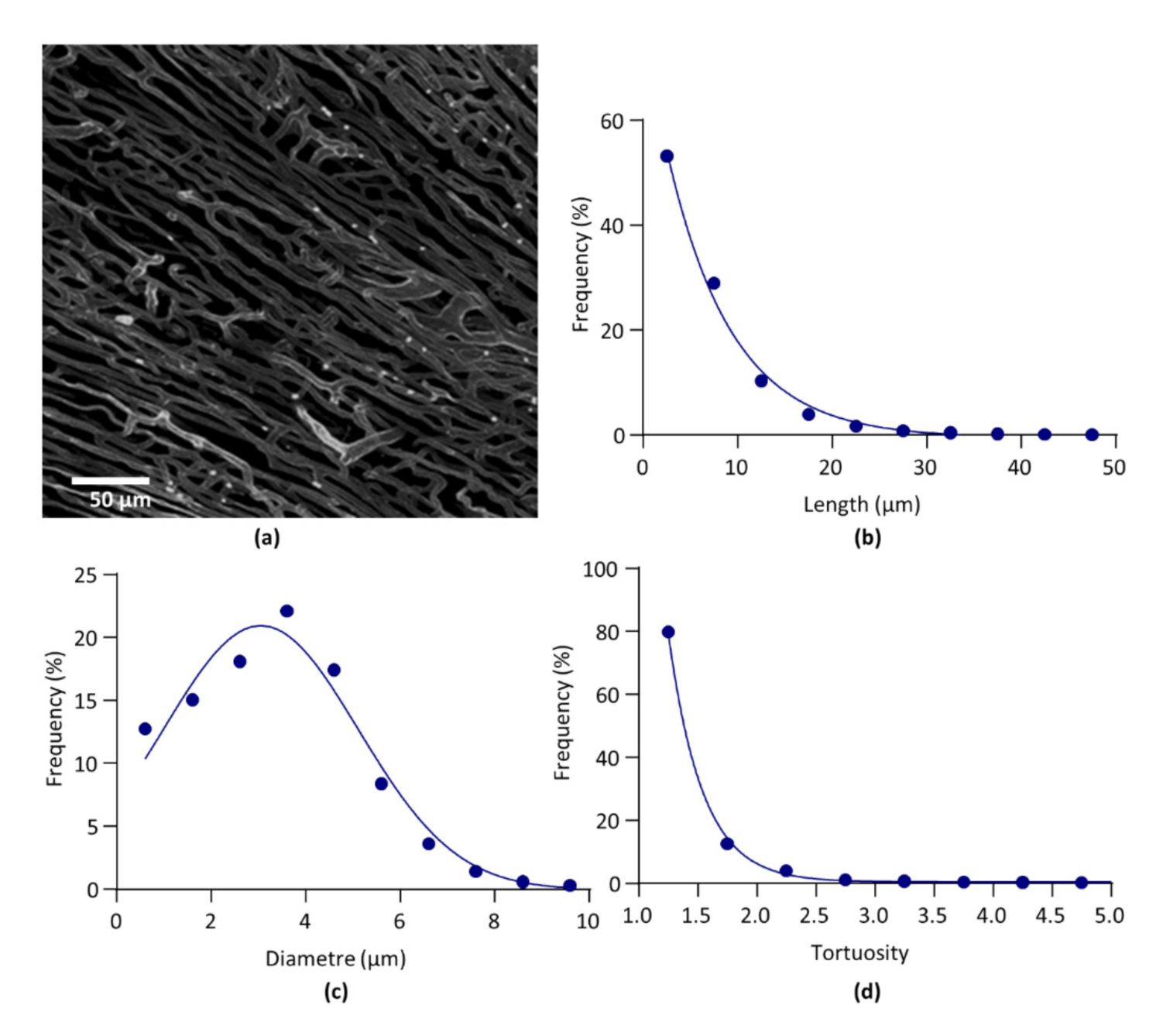
3.5. Confocal Microscopy
4. Discussion
4.1. 3D Imaging and Image Processing
4.2. Global and Archhitectural Parameters
4.3. Light Sheet Microscopy Versus Confocal Microscopy
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vancheri, F.; Longo, G.; Vancheri, S.; Henein, M. Coronary Microvascular Dysfunction. JCM 2020, 9, 2880. [Google Scholar] [CrossRef] [PubMed]
- Taqueti, V.R.; Di Carli, M.F. Coronary Microvascular Disease Pathogenic Mechanisms and Therapeutic Options. J. Am. Coll. Cardiol. 2018, 72, 2625–2641. [Google Scholar] [CrossRef] [PubMed]
- de Feyter, P.J.; Serruys, P.W.; Davies, M.J.; Richardson, P.; Lubsen, J.; Oliver, M.F. Quantitative Coronary Angiography to Measure Progression and Regression of Coronary Atherosclerosis. Value, Limitations, and Implications for Clinical Trials. Circulation 1991, 84, 412–423. [Google Scholar] [CrossRef]
- Müller, B.; Lang, S.; Dominietto, M.; Rudin, M.; Schulz, G.; Deyhle, H.; Germann, M.; Pfeiffer, F.; David, C.; Weitkamp, T. High-Resolution Tomographic Imaging of Microvessels. Proc. SPIE 2008, 7078, 70780B. [Google Scholar] [CrossRef]
- Wu, Y.; Ip, J.E.; Huang, J.; Zhang, L.; Matsushita, K.; Liew, C.-C.; Pratt, R.E.; Dzau, V.J. Essential Role of ICAM-1/CD18 in Mediating EPC Recruitment, Angiogenesis, and Repair to the Infarcted Myocardium. Circ. Res. 2006, 99, 315–322. [Google Scholar] [CrossRef]
- Markovič, R.; Peltan, J.; Gosak, M.; Horvat, D.; Žalik, B.; Seguy, B.; Chauvel, R.; Malandain, G.; Couffinhal, T.; Duplàa, C.; et al. Planar Cell Polarity Genes Frizzled4 and Frizzled6 Exert Patterning Influence on Arterial Vessel Morphogenesis. PLoS ONE 2017, 12, e0171033. [Google Scholar] [CrossRef]
- Lugo-Hernandez, E.; Squire, A.; Hagemann, N.; Brenzel, A.; Sardari, M.; Schlechter, J.; Sanchez-Mendoza, E.H.; Gunzer, M.; Faissner, A.; Hermann, D.M. 3D Visualization and Quantification of Microvessels in the Whole Ischemic Mouse Brain Using Solvent-Based Clearing and Light Sheet Microscopy. J. Cereb. Blood Flow Metab. 2017, 37, 3355–3367. [Google Scholar] [CrossRef]
- Smith, A.F.; Doyeux, V.; Berg, M.; Peyrounette, M.; Haft-Javaherian, M.; Larue, A.-E.; Slater, J.H.; Lauwers, F.; Blinder, P.; Tsai, P.; et al. Brain Capillary Networks Across Species: A Few Simple Organizational Requirements Are Sufficient to Reproduce Both Structure and Function. Front. Physiol. 2019, 10, 233. [Google Scholar] [CrossRef]
- Valero-Muñoz, M.; Backman, W.; Sam, F. Murine Models of Heart Failure With Preserved Ejection Fraction. JACC Basic Transl. Sci. 2017, 2, 770–789. [Google Scholar] [CrossRef] [PubMed]
- Costa, E.C.; Silva, D.N.; Moreira, A.F.; Correia, I.J. Optical Clearing Methods: An Overview of the Techniques Used for the Imaging of 3D Spheroids. Biotechnol. Bioeng. 2019, 116, 2742–2763. [Google Scholar] [CrossRef]
- Renier, N.; Adams, E.L.; Kirst, C.; Wu, Z.; Azevedo, R.; Kohl, J.; Autry, A.E.; Kadiri, L.; Umadevi Venkataraju, K.; Zhou, Y.; et al. Mapping of Brain Activity by Automated Volume Analysis of Immediate Early Genes. Cell 2016, 165, 1789–1802. [Google Scholar] [CrossRef] [PubMed]
- Robertson, R.T.; Levine, S.T.; Haynes, S.M.; Gutierrez, P.; Baratta, J.L.; Tan, Z.; Longmuir, K.J. Use of Labeled Tomato Lectin for Imaging Vasculature Structures. Histochem. Cell Biol. 2015, 143, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Breisch, E.A.; Houser, S.R.; Carey, R.A.; Spann, J.F.; Bove, A.A. Myocardial Blood Flow and Capillary Density in Chronic Pressure Overload of the Feline Left Ventricle. Cardiovasc. Res. 1980, 14, 469–475. [Google Scholar] [CrossRef] [PubMed]
- Potter, R.F.; Groom, A.C. Capillary Diameter and Geometry in Cardiac and Skeletal Muscle Studied by Means of Corrosion Casts. Microvasc. Res. 1983, 25, 68–84. [Google Scholar] [CrossRef]
- Doube, M.; Kłosowski, M.M.; Arganda-Carreras, I.; Cordelières, F.P.; Dougherty, R.P.; Jackson, J.S.; Schmid, B.; Hutchinson, J.R.; Shefelbine, S.J. BoneJ: Free and Extensible Bone Image Analysis in ImageJ. Bone 2010, 47, 1076–1079. [Google Scholar] [CrossRef]
- Ollion, J.; Cochennec, J.; Loll, F.; Escudé, C.; Boudier, T. TANGO: A Generic Tool for High-Throughput 3D Image Analysis for Studying Nuclear Organization. Bioinformatics 2013, 29, 1840–1841. [Google Scholar] [CrossRef] [PubMed]
- Luppe, M. Fractal Dimension Based on Minkowski-Bouligand Method Using Exponential Dilations. Electron. Lett. 2015, 51, 475–477. [Google Scholar] [CrossRef]
- Merz, S.F.; Korste, S.; Bornemann, L.; Michel, L.; Stock, P.; Squire, A.; Soun, C.; Engel, D.R.; Detzer, J.; Lörchner, H.; et al. Contemporaneous 3D Characterization of Acute and Chronic Myocardial I/R Injury and Response. Nat. Commun. 2019, 10, 2312. [Google Scholar] [CrossRef]
- Schmidt-Nielsen, K.; Pennycuik, P. Capillary Density in Mammals in Relation to Body Size and Oxygen Consumption. Am. J. Physiol. Leg. Content 1961, 200, 746–750. [Google Scholar] [CrossRef]
- Amann, K.; Wiest, G.; Zimmer, G.; Gretz, N.; Ritz, E.; Mall, G. Reduced Capillary Density in the Myocardium of Uremic Rats—A Stereological Study. Kidney Int. 1992, 42, 1079–1085. [Google Scholar] [CrossRef]
- Reishofer, G.; Koschutnig, K.; Enzinger, C.; Ebner, F.; Ahammer, H. Fractal Dimension and Vessel Complexity in Patients with Cerebral Arteriovenous Malformations. PLoS ONE 2012, 7, e41148. [Google Scholar] [CrossRef] [PubMed]
- Han, H.-C. Twisted Blood Vessels: Symptoms, Etiology and Biomechanical Mechanisms. J. Vasc. Res. 2012, 49, 185–197. [Google Scholar] [CrossRef] [PubMed]
| SAP (Mean ± SD) | DAP (Mean ± SD) | MAP (Mean ± SD) | BPM (Mean ± SD) |
|---|---|---|---|
| 123.1 ± 15 mm Hg | 89.7 ± 6.9 mm Hg | 100.5 ± 9.5 mm Hg | 303.6 ± 31 |
| L (Mean ± SD) | W (Mean ± SD) | T (Mean ± SD) | Volume (Mean ± SD) |
|---|---|---|---|
| 6.70 ± 4.2% | 10.7 ± 4.2% | 2.91 ± 8.2% | 19.18 ± 7.7% |
| Parameter | LV | S | RV |
|---|---|---|---|
| Vascular density | 34.4 ± 11% | 18.9 ± 4.7% | 27.8 ± 11% |
| Fractal dimension | 2.46 ± 0.05 | 2.32 ± 0.11 | 2.33 ± 0.07 |
| Number of segments/mm3 of cardiac tissue | 615,784 ± 220,000 | 399,922 ± 230,000 | 387,457 ± 200,000 |
| Total length/mm3 of cardiac tissue | 13.01 ± 6.1 m | 11.5 ± 9.2 m | 7.11 ± 2.7 m |
| Number of nodes/mm3 of cardiac tissue | 289,878 ± 96,000 | 169,886 ± 92,000 | 190,392 ± 100,000 |
| Number of nodes/number of segments | 47.56 ± 3.0% | 42.9 ± 2.9% | 48.72 ± 2.9% |
| Parameter | LV | S | RV |
|---|---|---|---|
| Length | |||
| λ (mean ± SEM) | 16.9 ± 0.6 µm | 15.6 ± 0.6 µm | 17.9 ± 4.4 µm |
| Diameter | |||
| μ (mean ± SEM) | 4.81 ± 0.24 µm | 3.78 ± 0.51 µm | 5.12 ± 0.25 µm |
| σ (mean ± SEM) | 2.52 ± 0.27 µm | 2.75 ± 0.44 µm | 2.17 ± 0.26 µm |
| Tortuosity | |||
| τ (mean ± SEM) | 0.32 ± 0.02 | 0.35 ± 0.02 | 0.35 ± 0.02 |
| Parameters | LSM | CM | (LSM-CM) |
|---|---|---|---|
| Vascular density | 43.3% | 31.2% | 12.1% |
| Fractal dimension | 2.49 | 2.53 | 0.04 |
| Number of segments/mm3 of cardiac tissue | 794,032 | 4,297,330 | 3,503,298 |
| Number of nodes/mm3 of cardiac tissue | 394,296 | 1,992,201 | 1,597,905 |
| Number of nodes/number of segments | 49.7% | 46.4% | 3.3% |
| Total length/mm3 of cardiac tissue | 17.7 m | 27.1 m | 9.4 m |
| Length | |||
| λ (mean ± SEM) | 28.9 ± 9.8 µm | 6.89 ± 1.0 µm | 22.01 µm |
| Diameter | |||
| μ (mean ± SEM) | 4.61 ± 0.68 µm | 3.05 ± 0.29 µm | 1.56 µm |
| σ (mean ± SEM) | 2.25 ± 0.61 µm | 2.07 ± 0.26 µm | 0.18 µm |
| Tortuosity | |||
| τ (mean ± SEM) | 0.31 ± 0.07 | 0.27 ± 0.02 | 0.04 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nicolas, N.; Roux, E. 3D Imaging and Quantitative Characterization of Mouse Capillary Coronary Network Architecture. Biology 2021, 10, 306. https://doi.org/10.3390/biology10040306
Nicolas N, Roux E. 3D Imaging and Quantitative Characterization of Mouse Capillary Coronary Network Architecture. Biology. 2021; 10(4):306. https://doi.org/10.3390/biology10040306
Chicago/Turabian StyleNicolas, Nabil, and Etienne Roux. 2021. "3D Imaging and Quantitative Characterization of Mouse Capillary Coronary Network Architecture" Biology 10, no. 4: 306. https://doi.org/10.3390/biology10040306
APA StyleNicolas, N., & Roux, E. (2021). 3D Imaging and Quantitative Characterization of Mouse Capillary Coronary Network Architecture. Biology, 10(4), 306. https://doi.org/10.3390/biology10040306






