Abstract
Textile hemp (Cannabis sativa L.) is a multipurpose crop producing biomass with uses in e.g., the textile, biocomposite, and construction sectors. It was previously shown that the hypocotyl of hemp is useful to study the kinetics of secondary tissue development, where primary and secondary growths are temporally uncoupled. We here sought to demonstrate that the stem of adult hemp plants is an additional suitable model to study the heterogeneous lignification of the tissues and the mechanisms underlying secondary cell wall formation in bast fibres. A targeted quantitative PCR analysis carried out on a set of twenty genes involved in cell wall biosynthesis clearly showed differences in expression in the core and cortical tissues along four stem regions spanning from elongation to cell wall thickening. Genes involved in phenylpropanoid biosynthesis and secondary cell wall cellulose synthases were expressed at higher levels in core tissues at the bottom, while specific genes, notably a class III peroxidase and a gene partaking in lignan biosynthesis, were highly expressed in the cortex of elongating internodes. The two systems, the hypocotyl and the adult stem of textile hemp, are equally valid and complementary to address questions related to lignification and secondary cell wall deposition.
1. Introduction
Industrial hemp (Cannabis sativa L.), which has a tetrahydrocannabinol (THC) content <0.3%, is a fibre crop historically used for textiles [1], and is currently considered a renewable resource for the provision of fibres substituting synthetic ones in composites [2,3]. Besides the application-oriented aspects of this crop, hemp is an interesting model to study questions related to lignification and the development of cellulose-rich (i.e., gelatinous) secondary cell walls (SCWs). Its stem comprises indeed a hollow lignified core (also known as hurd/shiv) and a cortex harbouring cellulosic phloem-supporting bast fibres. The crop can therefore produce inner lignified and peripheral cellulosic fibres. This feature is very interesting, as it enables researcher to investigate two aspects, lignification and cellulose-rich cell wall formation, in the same model [4,5].
It was already previously demonstrated that the hemp hypocotyl can provide valuable information relative to the transition from primary to secondary growth [6] and, more recently, lignification [7]. Additionally, the adult hemp stem was shown to provide detailed molecular data relative to the sequential developmental stages of bast fibres [8]. In particular, it was demonstrated that the snap point is a key region in the transition from elongation to bast fibre thickening, since the cell wall-related genes show major changes in expression [3,9].
With the goal of confirming the validity of the adult hemp stem as a model for cell wall studies, we have here undertaken a targeted qPCR analysis focused on twenty cell wall-related genes. We provide evidence for the differential expression of the genes in the inner/outer stem tissues along four stem regions and we propose a role in bast fibre development for some of the analysed genes.
2. Materials and Methods
2.1. Plant Material and Growth Conditions
C. sativa cv. Santhica 27 was used in this experiment. Plants were grown for six weeks in controlled conditions according to [8]. Eight different tissues sampled from four different heights were sampled. The heights were determined relatively to the snap point, which marks the transition between elongation and thickening of primary bast fibres [10]: above the snap point (ASP), internode containing the snap point (SP), below the snap point (BSP), and two internodes below the snap point (BBSP). The snap point was determined as previously described [9]. Each segment was separated in cortical tissues (harbouring the bast fibres, annotated as OUT) and core tissues (containing the xylem with its associated fibres and the pith, annotated as IN), according to [9,11]. To avoid excessive variation in gene expression, a segment of ca. 2 cm was collected in the middle of each internode. The samples were directly frozen in liquid nitrogen and stored at −80 °C until RNA extraction. Four biological replicates, each consisting of five plants, were used in this experiment.
2.2. RNA Extraction and RT-qPCR
Total RNA was extracted using the RNeasy Plant Mini Kit (Qiagen, Leusden, The Netherlands), with the on-column DNase treatment, following the instructions of the manufacturer with a modification. The 500 µL of RLC buffer were replaced by 450 µL of this buffer with 50 µL of 20% PEG (MW 20,000, Sigma, St. Louis, MO, USA) to maximise the extraction of total RNA [12]. The quantity and quality of the total RNAs were assessed spectrophotometrically and with a BioAnalyzer (all RINs > 7.5). Reverse transcription and RT-qPCR analysis were performed as described in [9]. The gene expression was normalised using eTIF3H and eTIF4, whose stability was determined with respect to previously reported reference genes (eTIF3E and Cyclophilin; [13]). A melt curve was performed at the end of each run to check the specificity of the PCR products. The characteristics of the primers are listed in Table S1. The target genes originate from several databases: our previously published hemp transcriptomes [6,8], a microarray-based experiment [14], or through BLAST search of orthologous Arabidopsis thaliana genes at the Medicinal Plant Genomics Resource [15]. The primers were validated via qPCR using a standard curve with a serial five-fold dilution of cDNA (20, 4, 0.8, 0.16, 0.032, and 0.0064 ng/µL). The normalised expression values were calculated in qBasePLUS, and the hierarchical clustering of expression values obtained with the software Cluster 3.0 [16].
3. Results
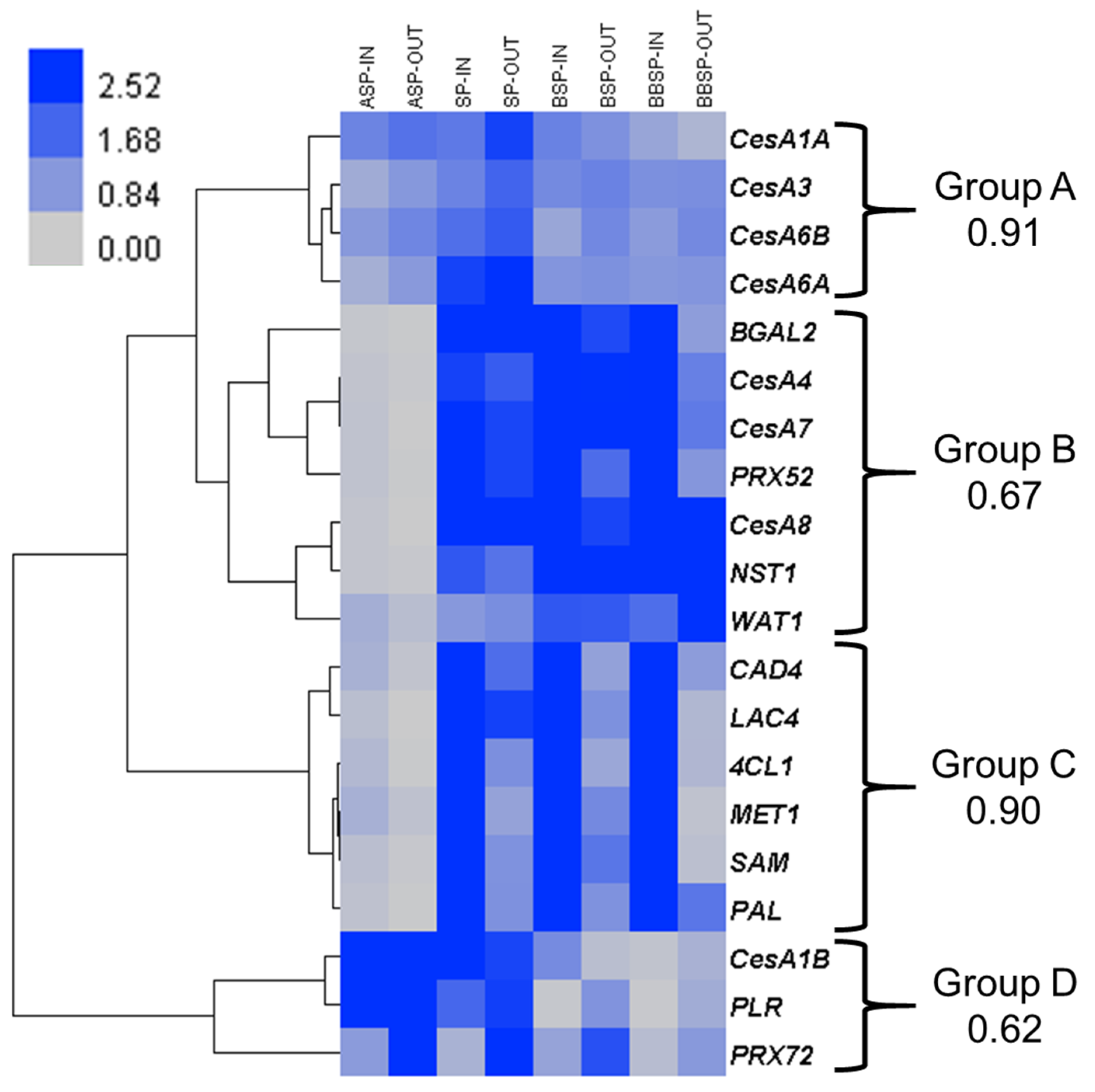
The hierarchical clustering of the expression values shows four main patterns (Figure 1). In group A, the genes associated with primary cell wall cellulose deposition are found (CesA1A, CesA3, CesA6A and CesA6B). These genes are ubiquitously expressed within the targeted tissues.
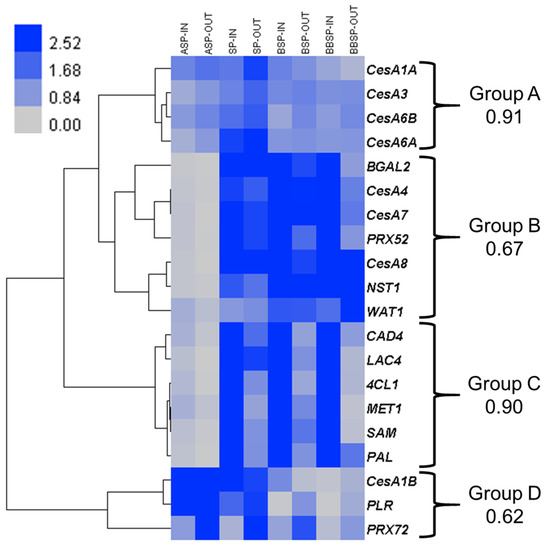
Figure 1.
Gene expression analysis targeting processes related to cell wall deposition. Heat map hierarchical clustering of the expression profiles of twenty genes at four stem regions, in inner (-IN) and outer (-OUT) tissues. For each group, the Pearson correlation coefficient is provided. Abbreviations are as in the text.
In group B, the orthologous genes known to be involved in SCW biosynthesis are found: the master transcriptional regulator of SCW deposition NAC secondary wall thickening promoting factor 1 NST1 [17], the three cellulose synthases associated with SCW biogenesis CesA4, CesA7, and CesA8 [18], as well as the class III peroxidase orthologous to Arabidopsis PRX52 [19]. These genes are more expressed at the SP and in the bottom internodes and slightly upregulated in the inner tissues as compared to outer tissues (Table S1). β-galactosidase 2 (BGAL2) is also found in this group, together with the gene Walls are thin 1 (WAT1). The expression pattern of WAT1 is somehow different from the other genes of this group, as it is slightly more expressed in the outer tissues from the SP downwards.
Group C gathers some major genes involved in lignification: Phenylalanine ammonia lyase (PAL), Cinnamyl alcohol dehydrogenase 4 (CAD4) and 4-hydroxycinnamoyl-CoA ligase 1 (4CL1) are part of the monolignol pathway, Methionine synthase 1 (MET1) and S-adenosylmethionine synthetase (SAM) are involved in monolignol methylation [20], and Laccase 4 (LAC4) is one of the enzymes partaking in lignin polymerisation [21]. These genes are in general more expressed in the core than in the cortical tissues.
Finally, the similar expression patterns of CesA1B, Pinoresinol lariciresinol reductase (PLR) and PRX72 form group D. These genes are more expressed above the SP. By contrast with the three other groups, PLR and PRX72 are upregulated in the outer tissues (Table S1), which may point to an important role in bast fibre development.
The expression of CesA1B strongly drops in the internodes below the snap point.
4. Discussion
In order to explain the molecular regulation leading to the different cell wall composition between core and cortical tissues, a targeted gene expression analysis was performed (Figure 1). It shows that the higher lignin content reported in xylem tissues (15% vs. 4% in the bast fibres [14]) is correlated with an upregulation of the genes of the phenylpropanoid/monolignol pathway, in accordance with previous results [14]. Those genes are found in clusters C (PAL, 4CL1, CAD4, LAC4, MET1, and SAM) and B (PRX52).
The CesAs associated with the cellulose synthase complex of the SCW (CesA4, CesA7 and CesA8; [18]) are grouped in cluster B. They are slightly more expressed in the inner tissues and weakly in the elongating stem (Figure 1). In the elongating internode (ASP), the deposition of SCW is restricted to the metaxylem and protoxylem [11]. In the secondary xylem, the SCW is deposited in a reticulated or pitted pattern, characterised by a massive deposition of cellulose [22]. Fibres and tracheary elements of the xylem have a xylan-type SCW (i.e., organised in S1, S2, and S3 sublayers), while bast fibres have a gelatinous-type SCW (S1, S2, and G-layer) [23]. Generally, there are, if any, only slight differences in the CesAs expression patterns between xylan-type and gelatinous-type SCWs [24]. For instance, CesaA4 and CesA7 are upregulated in the xylan-type bast fibres of jute [25], as also observed in hemp in our results (Figure 1). However, a recent study performed on flax has highlighted a higher expression of both primary and SCW-related CesAs in phloem fibres depositing their G-layer [26]. In addition, a strong bast fibre phenotype (reduced number and irregular cell shape associated with altered cell wall composition) has been observed in flax plants with virus-induced gene silencing of CesA genes (CesA1 or CesA6), usually acting in primary cell wall biogenesis [27]. The data presented in Figure 1 do not allow us to confirm nor refute this hypothesis, however, it is noteworthy to mention that several CesA isoforms may be missing from our analysis.
The CesA genes analysed in this article were obtained by mining two resources: the Medicinal Plant Genomic Resource database (MPGR) [15] and our in-house hemp transcriptome assembly (originating from [6,8]). In this assembly, the contigs annotated as CesAs or cellulose synthase-like (Csl) were individually checked and used for a BLAST analysis in the hemp genome deposited in MPGR. Using this method, eight different CesA genes were retrieved. In the flax genome, between fifteen and sixteen predicted CesAs were found [27,28]. We may, thus, anticipate additional CesA isoforms in hemp, whose identifications rely on a robust annotation of the genomic resources available so far [29]. In agreement with the data from [27,28], CesA4, CesA7 and CesA8 were more expressed in tissues undergoing SCW formation, and may thus be considered as functional orthologous genes of AtCesA4, AtCesA7 and AtCesA8.
The master transcription factor of SCW deposition NST1 shows a trend similar to CesAs. As it is highly expressed both in inner and outer tissues undergoing SCW formation, this transcription factor may be involved in the development of xylem and bast fibres. This is in contrast with the data obtained in flax [30], where NST1 is less expressed in thickening bast fibres (which in this study correspond to BSP and BBSP samples), as compared to the top region above the SP (corresponding here to the ASP sample). The presence of secondary bast fibres originating from high cambial activity in hemp may explain this difference. Indeed, several transcription factors from the NST family (such as PtrWND1B) are suggested to contribute to the formation of bast fibres in poplar, based on their expression in this tissue [31]. We may thus invoke two scenarios, which are not mutually exclusive, to explain this particular gene expression. In the first one, NST1 is linked to the differentiation of secondary bast fibres, while in the second its expression leads to the regulation of the genes involved in the formation of the SCW (cellulose, xylan, and lignin biosyntheses; [32]). Assuming that hemp bast fibres are hypolignified, a molecular mechanism specifically downregulating the biosynthesis of monolignols and/or lignin should be present in hemp bast fibres. This control may take place at the transcriptional (through negative regulation of the expression of these genes) and/or at the posttranscriptional level. In flax bast fibres (which are also hypolignified), this regulation may be (partially) achieved through the degradation of transcripts of laccases involved in lignin polymerisation by microRNA397 [33].
The regulation of the biosynthesis of the non-cellulosic polysaccharides in gelatinous fibres, such as rhamnogalacturonan-I (RG-I), is still not well understood, but several NAC transcription factors are upregulated in poplar tension wood developing gelatinous fibres [34]. We may speculate that hemp NST1 may also regulate the biosynthesis of such polysaccharides, however, this remains to be confirmed.
Auxin homeostasis is a central element for SCW formation in fibres [35,36]. The tonoplast-localised auxin efflux protein WAT1 plays a key role in this respect. The Arabidopsis mutant line wat1-1 accumulates indole acetic acid (IAA), the main form of bioactive auxin, in the tonoplast, preventing its binding to nuclear targets (auxin-regulated genes), or endoplasmic reticulum-based auxin receptors [36]. This mutation has important transcriptomic consequences: NST1 is significantly downregulated, leading to lower expression of CesA4, CesA7, CesA8 and most of the genes of the lignin biosynthetic pathway [35]. The expression profile of the hemp ortholog of WAT1 points to an important role of auxin in the development and maturation of xylem and bast fibres. It suggests that the expression of NST1 may be driven by the pool of auxin present in the nucleus in a WAT1-dependent manner. Similarly, it was suggested that the auxin-induced expression of NST genes for the promotion of SCW formation in Arabidopsis fibres is mediated by the gene REVOLUTA [32], whose null mutant displays a significantly decreased expression of two putative auxin efflux carriers, PIN3 and PIN4 [37]. The expression profile of WAT1 in the present study is similar to the observations previously made in two other biological systems, namely, the developing hemp hypocotyl [6] and the developing hemp bast fibres [8]. This gene was highly expressed in hypocotyls undergoing secondary growth and cell wall thickening, as well as in bast fibres depositing their gelatinous layer. All these data suggest that WAT1 and auxin play a significant role in SCW deposition in hemp bast fibres.
The Arabidopsis BGAL2 specifically hydrolyses β-(1-3) and β-(1-4) linkages in galacto-oligosaccharides and β-(1-4) linkage in lupin galactan [38]. These two linkages are found in RG-I. This complex pectin is present with different structures in elongating tissues, in dividing regions, such as the cambium, and in the pectic matrix enrobing cellulose in flax and hemp bast fibres [39]. The expression profile of BGAL2 in hemp suggests a role in these events. According to previous microscopic observations [8], secondary growth occurs from the SP downwards (i.e., in the samples SP, BSP, and BBSP). Later in the development, secondary tissues originating from the cambium undergo intrusive growth [24,40]. Finally, the pectic matrix of the gelatinous layer of the bast fibres is enzymatically modified [41]. These three distinct processes require extensive modifications of the extracellular matrix, among them the BGAL2-driven RG-I degradation. The role of a specific BGAL in flax bast fibre maturation was already demonstrated [42].
The expression profiles of CesA1B, PLR, and PRX72 is in sharp contrast with the genes from clusters B and C. PLR and PRX72 are more expressed in the elongating internode (ASP) and in the outer tissues. PLR is an entry enzyme for the biosynthesis of lignans. Lignans are formed by enantioselective coupling of two monolignol units [43]. This family of molecules is involved in plant growth [44], lignin distribution during SCW biosynthesis [45], and redox homeostasis during lignification [46]. From the data here reported, it is possible to propose that PLR regulates stem elongation via the biosynthesis of specific lignans [47]. The expression of PLR was also higher in the elongating hypocotyls (6 and 9 days after sowing; [7]). A functional analysis of this gene, as well as a detailed chemical characterisation of lignans present in elongating- and non-elongating tissues, will validate this hypothesis. The expression of PLR and PRX72 is higher in the cortical tissue, and may thus be important for the development of bast fibres. The Arabidopsis ortholog of PRX72 is involved in lignin biosynthesis [48]. A mutant defective in AtPRX72 shows thinner SCWs only in interfascicular fibres and a lower lignin S/G ratio. Based on its expression profile in hemp (Figure 1), a role in bast fibre lignification is, however, unlikely. Indeed, bast fibres lignify mostly after they reach their final size and S-lignin is deposited at the latest stage of lignification [48].
CesA1B is strongly expressed in the internode ASP, both in inner and outer tissues, consistent with its role in the deposition of cellulose in the primary cell wall. In this region of the stem, the bast and xylem fibres grow intrusively, while the cells of the other tissues may eventually end their symplastic elongation [49]. The upregulation of CesA1B in the ASP region suggests a role in intrusive growth. The expression pattern of CesA1B shows a clustering that is not coinciding with that of the other primary CesAs; maybe its function is different and associated with the early stages of xylem cell development. It would be interesting to determine whether this gene is specifically induced upon gravistimulation in the xylem cells of the stem.
5. Conclusions
In this article, a gene expression analysis was performed, which aimed at analysing key actors involved in the biosynthesis of the cell wall in hemp stem tissues. The differential cell wall composition of inner and outer stem tissues is at least partially regulated at the gene expression level, especially for lignin. The differential expression of genes controlling the cell wall composition of several types of tissues is a promising result for new research lines.
Supplementary Materials
The following are available online at http://www.mdpi.com/2079-6439/6/2/27/s1, Table S1: Primers and raw gene expression data.
Acknowledgments
The authors acknowledge the Fonds National de la Recherche, Luxembourg, (Project CANCAN C13/SR/5774202) for partial financial support. The authors thank Aude Corvisy and Laurent Solinhac for their technical support.
Author Contributions
Marc Behr and Gea Guerriero designed the experiment and collected the samples. Marc Behr performed the RT-qPCR analysis and analysed the data. Marc Behr and Gea Guerriero wrote the manuscript. Jean-Francois Hausman and Stanley Lutts critically revised the manuscript. Jean-Francois Hausman led the Project CANCAN. All the authors have read and approved the final version of this manuscript.
Conflicts of Interest
The authors declare no conflict of interest. The founding sponsors had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
References
- Skoglund, G.; Nockert, M.; Holst, B. Viking and early middle ages northern Scandinavian textiles proven to be made with hemp. Sci. Rep. 2013, 3, 2686. [Google Scholar] [CrossRef] [PubMed]
- Andre, C.M.; Hausman, J.F.; Guerriero, G. Cannabis sativa: The plant of the thousand and one molecules. Front. Plant Sci. 2016, 7, 19. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Behr, M.; Backes, A.; Faleri, C.; Hausman, J.-F.; Lutts, S.; Cai, G. Bast fibre formation: Insights from next-generation sequencing. Procedia Eng. 2017, 200, 229–235. [Google Scholar] [CrossRef]
- Guerriero, G.; Sergeant, K.; Hausman, J.-F. Integrated -omics: A powerful approach to understanding the heterogeneous lignification of fibre crops. Int. J. Mol. Sci. 2013, 14, 10958–10978. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Sergeant, K.; Hausman, J.F. Wood biosynthesis and typologies: A molecular rhapsody. Tree Physiol. 2014, 34, 839–855. [Google Scholar] [CrossRef] [PubMed]
- Behr, M.; Legay, S.; Zizková, E.; Motyka, V.; Dobrev, P.I.; Hausman, J.F.; Lutts, S.; Guerriero, G. Studying secondary growth and bast fiber development: The hemp hypocotyl peeks behind the wall. Front. Plant Sci. 2016, 7, 1733. [Google Scholar] [CrossRef] [PubMed]
- Behr, M.; Sergeant, K.; Leclercq, C.C.; Planchon, S.; Guignard, C.; Lenouvel, A.; Renaut, J.; Hausman, J.-F.; Lutts, S.; Guerriero, G. Insights into the molecular regulation of monolignol-derived product biosynthesis in the growing hemp hypocotyl. BMC Plant Biol. 2018, 18, 1. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Behr, M.; Legay, S.; Mangeot-Peter, L.; Zorzan, S.; Ghoniem, M.; Hausman, J.-F. Transcriptomic profiling of hemp bast fibres at different developmental stages. Sci. Rep. 2017, 7, 4961. [Google Scholar] [CrossRef] [PubMed]
- Behr, M.; Legay, S.; Hausman, J.-F.; Lutts, S.; Guerriero, G. Molecular investigation of the stem snap point in textile hemp. Genes 2017, 8, 363. [Google Scholar] [CrossRef] [PubMed]
- Gorshkova, T.A.; Sal’nikov, V.V.; Chemikosova, S.B.; Ageeva, M.V.; Pavlencheva, N.V.; van Dam, J.E.G. The snap point: A transition point in Linum usitatissimum bast fiber development. Ind. Crops Prod. 2003, 18, 213–221. [Google Scholar] [CrossRef]
- Guerriero, G.; Mangeot-Peter, L.; Legay, S.; Behr, M.; Lutts, S.; Siddiqui, K.S.; Hausman, J.-F. Identification of fasciclin-like arabinogalactan proteins in textile hemp (Cannabis sativa L.): In silico analyses and gene expression patterns in different tissues. BMC Genom. 2017, 18, 741. [Google Scholar] [CrossRef] [PubMed]
- Gehrig, H.H.; Winter, K.; Cushman, J.; Borland, A.; Taybi, T. An improved RNA isolation method for succulent plant species rich in polyphenols and polysaccharides. Plant Mol. Biol. Rep. 2000, 18, 369–376. [Google Scholar] [CrossRef]
- Mangeot-Peter, L.; Legay, S.; Hausman, J.F.; Esposito, S.; Guerriero, G. Identification of reference genes for RT-qPCR data normalization in Cannabis sativa stem tissues. Int. J. Mol. Sci. 2016, 17, 1556. [Google Scholar] [CrossRef] [PubMed]
- Van den Broeck, H.C.; Maliepaard, C.; Ebskamp, M.J.M.; Toonen, M.A.J.; Koops, A.J. Differential expression of genes involved in C1 metabolism and lignin biosynthesis in wooden core and bast tissues of fibre hemp (Cannabis sativa L.). Plant Sci. 2008, 174, 205–220. [Google Scholar] [CrossRef]
- Medicinal Plant Genomics Resource. Available online: http://medicinalplantgenomics.msu.edu/index.shtml (accessed on 1 July 2017).
- Eisen, M.B.; Spellman, P.T.; Brown, P.O.; Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 1998, 95, 14863–14868. [Google Scholar] [CrossRef] [PubMed]
- Cassan-Wang, H.; Goué, N.; Saidi, M.N.; Legay, S.; Sivadon, P.; Goffner, D.; Grima-Pettenati, J. Identification of novel transcription factors regulating secondary cell wall formation in Arabidopsis. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Hill, J.L.; Hammudi, M.B.; Tien, M. The Arabidopsis cellulose synthase complex: A proposed hexamer of CESA trimers in an equimolar stoichiometry. Plant Cell 2014, 26, 4834–4842. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Pérez, F.; Pomar, F.; Pedreno, M.A.; Novo-Uzal, E. The suppression of atprx52 affects fibers but not xylem lignification in Arabidopsis by altering the proportion of syringyl units. Physiol. Plant. 2014, 154, 395–406. [Google Scholar] [CrossRef] [PubMed]
- Shen, B.; Li, C.; Tarczynski, M.C. High free-methionine and decreased lignin content result from a mutation in the Arabidopsis S-adenosyl-l-methionine synthetase 3 gene. Plant J. 2002, 29, 371–380. [Google Scholar] [CrossRef] [PubMed]
- Berthet, S.; Demont-Caulet, N.; Pollet, B.; Bidzinski, P.; Cézard, L.; Le Bris, P.; Borrega, N.; Hervé, J.; Blondet, E.; Balzergue, S.; et al. Disruption of LACCASE4 and 17 results in tissue-specific alterations to lignification of Arabidopsis thaliana stems. Plant Cell 2011, 23, 1124–1137. [Google Scholar] [CrossRef] [PubMed]
- Schuetz, M.; Benske, A.; Smith, R.A.; Watanabe, Y.; Tobimatsu, Y.; Ralph, J.; Demura, T.; Ellis, B.; Samuels, A.L. Laccases direct lignification in the discrete secondary cell wall domains of protoxylem. Plant Physiol. 2014, 166, 798–807. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Hausman, J.-F.; Strauss, J.; Ertan, H.; Siddiqui, K.S. Lignocellulosic biomass: Biosynthesis, degradation, and industrial utilization. Eng. Life Sci. 2016, 16, 1–16. [Google Scholar] [CrossRef]
- Gorshkova, T.; Brutch, N.; Chabbert, B.; Deyholos, M.; Hayashi, T.; Lev-Yadun, S.; Mellerowicz, E.J.; Morvan, C.; Neutelings, G.; Pilate, G. Plant fiber formation: State of the art, recent and expected progress, and open questions. Crit. Rev. Plant Sci. 2012, 31, 201–228. [Google Scholar] [CrossRef]
- Islam, M.S.; Saito, J.A.; Emdad, E.M.; Ahmed, B.; Islam, M.M.; Halim, A.; Hossen, Q.M.M.; Hossain, M.Z.; Ahmed, R.; Hossain, M.S.; et al. Comparative genomics of two jute species and insight into fibre biogenesis. Nat. Plants 2017, 3, 16223. [Google Scholar] [CrossRef] [PubMed]
- Mokshina, N.; Gorshkov, O.; Ibragimova, N.; Chernova, T.; Gorshkova, T. Cellulosic fibres of flax recruit both primary and secondary cell wall cellulose synthases during deposition of thick tertiary cell walls and in the course of graviresponse. Funct. Plant Biol. 2017, 44, 820–831. [Google Scholar] [CrossRef]
- Chantreau, M.; Chabbert, B.; Billiard, S.; Hawkins, S.; Neutelings, G. Functional analyses of cellulose synthase genes in flax (Linum usitatissimum) by virus-induced gene silencing. Plant Biotechnol. J. 2015, 1312–1324. [Google Scholar] [CrossRef] [PubMed]
- Mokshina, N.; Gorshkova, T.; Deyholos, M.K. Chitinase-like and cellulose synthase gene expression in gelatinous-type cellulosic walls of flax (Linum usitatissimum L.) bast fibers. PLoS ONE 2014, 9, e97949. [Google Scholar] [CrossRef] [PubMed]
- Van Bakel, H.; Stout, J.M.; Cote, A.G.; Tallon, C.M.; Sharpe, A.G.; Hughes, T.R.; Page, J.E. The draft genome and transcriptome of Cannabis sativa. Genome Biol. 2011, 12, R102. [Google Scholar] [CrossRef] [PubMed]
- Gorshkov, O.; Mokshina, N.; Gorshkov, V.; Chemikosova, S.; Gogolev, Y.; Gorshkova, T. Transcriptome portrait of cellulose-enriched flax fibres at advanced stage of specialization. Plant Mol. Biol. 2017, 93, 431–449. [Google Scholar] [CrossRef] [PubMed]
- Ohtani, M.; Nishikubo, N.; Xu, B.; Yamaguchi, M.; Mitsuda, N.; Goué, N.; Shi, F.; Ohme-Takagi, M.; Demura, T. A NAC domain protein family contributing to the regulation of wood formation in poplar. Plant J. 2011, 67, 499–512. [Google Scholar] [CrossRef] [PubMed]
- Mitsuda, N.; Iwase, A.; Yamamoto, H.; Yoshida, M.; Seki, M.; Shinozaki, K.; Ohme-Takagi, M. NAC transcription factors, NST1 and NST3, are key regulators of the formation of secondary walls in woody tissues of Arabidopsis. Plant Cell 2007, 19, 270–280. [Google Scholar] [CrossRef] [PubMed]
- Le Roy, J.; Blervacq, A.-S.; Créach, A.; Huss, B.; Hawkins, S.; Neutelings, G. Spatial regulation of monolignol biosynthesis and laccase genes control developmental and stress-related lignin in flax. BMC Plant Biol. 2017, 17, 124. [Google Scholar] [CrossRef] [PubMed]
- Grant, E.H.; Fujino, T.; Beers, E.P.; Brunner, A.M. Characterization of NAC domain transcription factors implicated in control of vascular cell differentiation in Arabidopsis and Populus. Planta 2010, 232, 337–352. [Google Scholar] [CrossRef] [PubMed]
- Ranocha, P.; Denancé, N.; Vanholme, R.; Freydier, A.; Martinez, Y.; Hoffmann, L.; Köhler, L.; Pouzet, C.; Renou, J.-P.; Sundberg, B.; et al. Walls are thin 1 (WAT1), an Arabidopsis homolog of Medicago truncatula nodulin21, is a tonoplast-localized protein required for secondary wall formation in fibers. Plant J. 2010, 63, 469–483. [Google Scholar] [CrossRef] [PubMed]
- Ranocha, P.; Dima, O.; Nagy, R.; Felten, J.; Corratgé-Faillie, C.; Novák, O.; Morreel, K.; Lacombe, B.; Martinez, Y.; Pfrunder, S.; et al. Arabidopsis WAT1 is a vacuolar auxin transport facilitator required for auxin homoeostasis. Nat. Commun. 2013, 4, 2625. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Ye, Z.-H. Alteration of auxin polar transport in the Arabidopsis ifl1 mutants. Plant Physiol. 2001, 126, 549–563. [Google Scholar] [CrossRef] [PubMed]
- Gantulga, D.; Turan, Y.; Bevan, D.R.; Esen, A. The Arabidopsis at1g45130 and at3g52840 genes encode beta-galactosidases with activity toward cell wall polysaccharides. Phytochemistry 2008, 69, 1661–1670. [Google Scholar] [CrossRef] [PubMed]
- Mikshina, P.V.; Petrova, A.A.; Gorshkova, T.A. Functional diversity of rhamnogalacturonans I. Russ. Chem. Bull. 2015, 64, 1014–1023. [Google Scholar] [CrossRef]
- Lev-Yadun, S. Intrusive growth—The plant analog of dendrite and axon growth in animals. New Phytol. 2001, 150, 508–512. [Google Scholar] [CrossRef]
- Mikshina, P.; Chernova, T.; Chemikosova, S.B.; Ibragimova, N.; Mokshina, N.; Gorshkova, T. Cellulosic fibers: Role of matrix polysaccharides in structure and function. In Cellulose—Fundamental Aspects; van de Ven, T., Godbout, L., Eds.; InTech: Rijeka, Croatia, 2013; pp. 91–112. [Google Scholar]
- Roach, M.J.; Mokshina, N.Y.; Badhan, A.; Snegireva, A.V.; Hobson, N.; Deyholos, M.K.; Gorshkova, T.A. Development of cellulosic secondary walls in flax fibers requires ß-galactosidase. Plant Physiol. 2011, 156, 1351–1363. [Google Scholar] [CrossRef] [PubMed]
- Nakatsubo, T.; Mizutani, M.; Suzuki, S.; Hattori, T.; Umezawa, T. Characterization of Arabidopsis thaliana pinoresinol reductase, a new type of enzyme involved in lignan biosynthesis. J. Biol. Chem. 2008, 283, 15550–15557. [Google Scholar] [CrossRef] [PubMed]
- Cutillo, F.; D’Abrosca, B.; DellaGreca, M.; Fiorentino, A.; Zarrelli, A. Lignans and neolignans from Brassica fruticulosa: Effects on seed germination and plant growth. J. Agric. Food Chem. 2003, 51, 6165–6172. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Zeng, Y.; Yin, Y.; Pu, Y.; Jackson, L.A.; Engle, N.L.; Martin, M.Z.; Tschaplinski, T.J.; Ding, S.Y.; Ragauskas, A.J.; et al. Pinoresinol reductase 1 impacts lignin distribution during secondary cell wall biosynthesis in Arabidopsis. Phytochemistry 2015, 112, 170–178. [Google Scholar] [CrossRef] [PubMed]
- Niculaes, C.; Morreel, K.; Kim, H.; Lu, F.; McKee, L.S.; Ivens, B.; Haustraete, J.; Vanholme, B.; Rycke, R.D.; Hertzberg, M.; et al. Phenylcoumaran benzylic ether reductase prevents accumulation of compounds formed under oxidative conditions in poplar xylem. Plant Cell 2014, 26, 3775–3791. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, S.; Ichikawa, H.; Nishiwaki, H.; Shuto, Y. Evaluation of plant growth regulatory activity of furofuran lignan bearing a 7,9′:7′,9-diepoxy structure using optically pure (+)- and (−)-enantiomers. J. Agric. Food Chem. 2015, 63, 5224–5228. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Pérez, F.; Pomar, F.; Pedreño, M.A.; Novo-Uzal, E. Suppression of Arabidopsis peroxidase 72 alters cell wall and phenylpropanoid metabolism. Plant Sci. 2015, 239, 192–199. [Google Scholar] [CrossRef] [PubMed]
- Snegireva, A.; Chernova, T.; Ageeva, M.; Lev-Yadun, S.; Gorshkova, T. Intrusive growth of primary and secondary phloem fibres in hemp stem determines fibre-bundle formation and structure. AoB Plants 2015, 7, plv061. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).