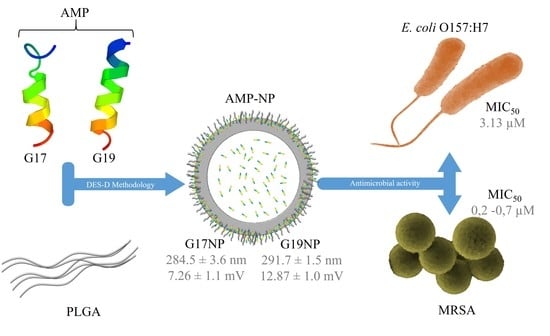
Potent and Specific Antibacterial Activity against Escherichia coli O157:H7 and Methicillin Resistant Staphylococcus aureus (MRSA) of G17 and G19 Peptides Encapsulated into Poly-Lactic-Co-Glycolic Acid (PLGA) Nanoparticles
Abstract
1. Introduction
2. Results and Discussion
2.1. Peptide Synthesis and Characterization
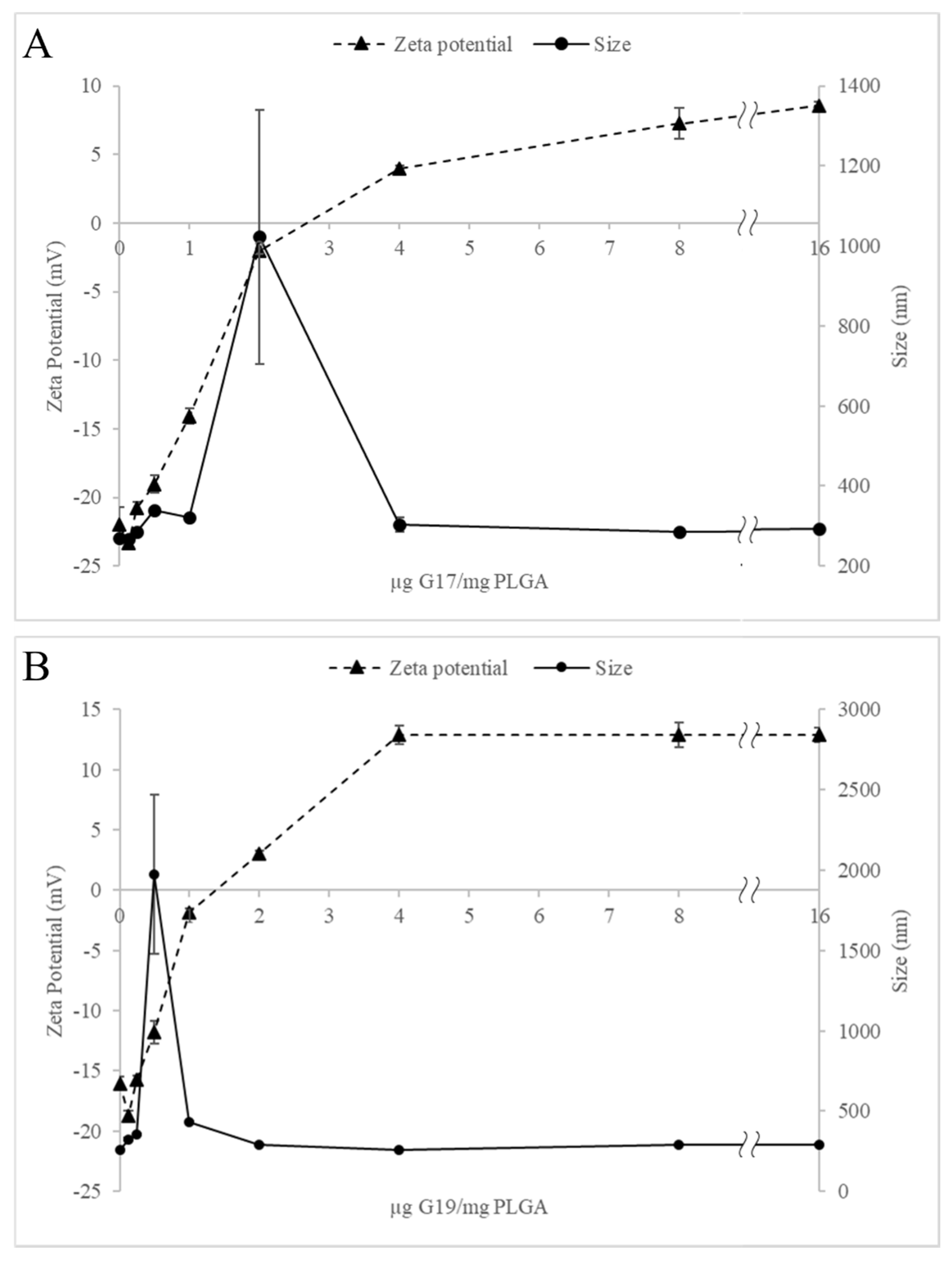
2.2. Physicochemical Properties of AMP-NPs
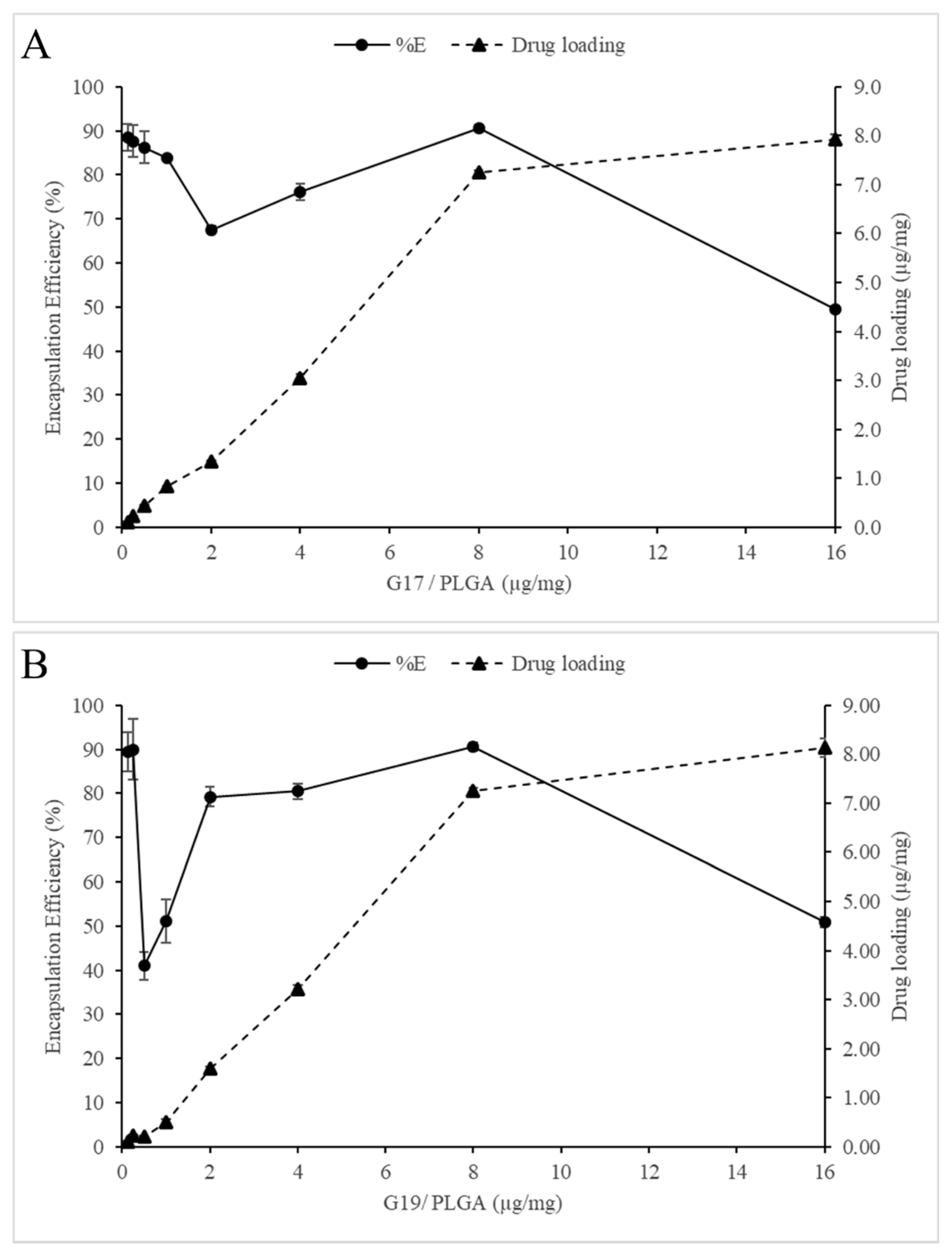
2.3. Encapsulation Efficiency and Load Capacity of the G17NP and G19NP
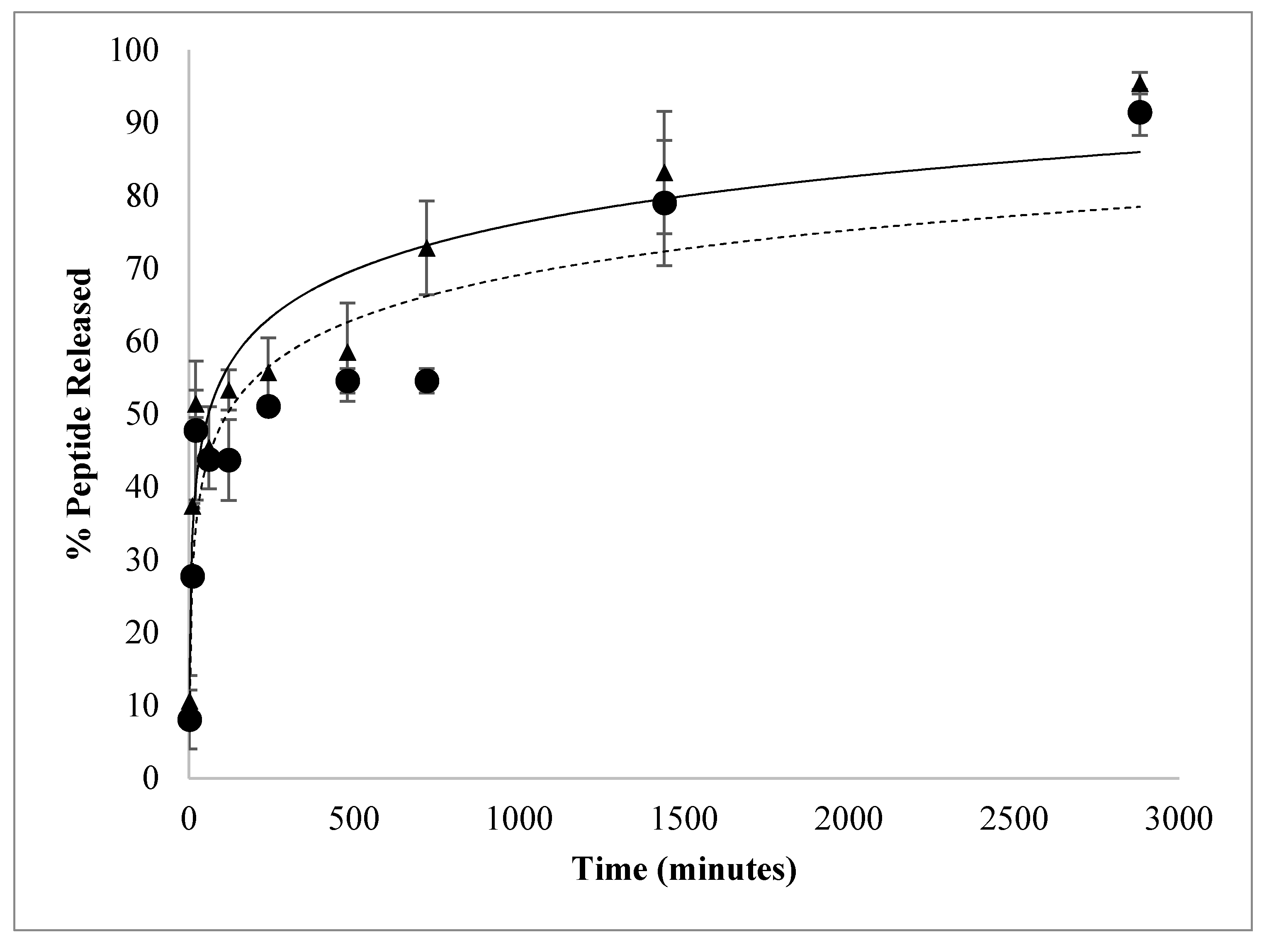
2.4. In Vitro Release of Antimicrobial Peptides from PLGA-NP
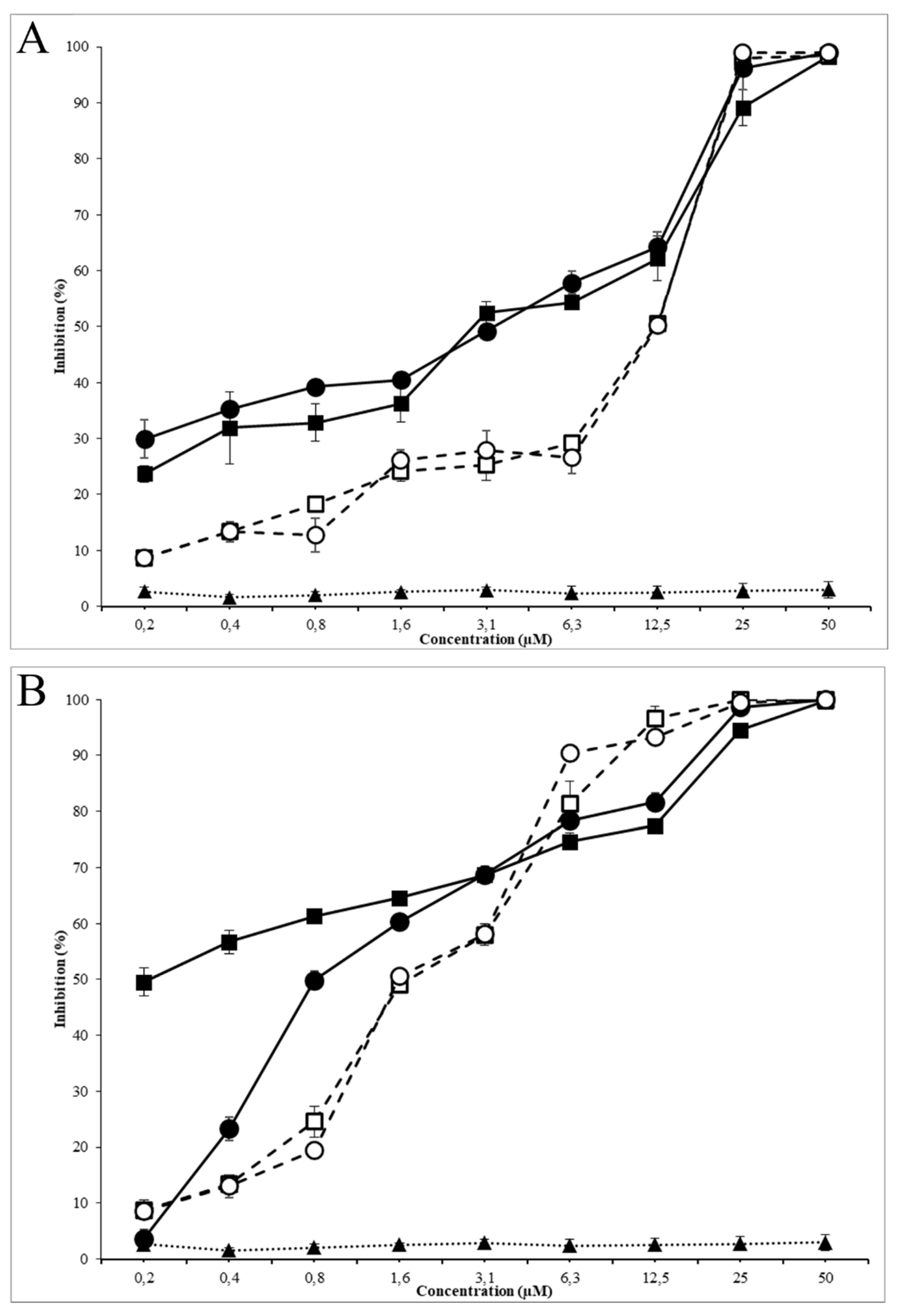
2.5. Determination of Minimum Inhibitory Concentration (MIC) and Minimum Bactericidal Concentration (MBC)
3. Materials and Methods
3.1. Materials
3.2. Peptide Synthesis and Characterization
3.3. Preparation of Polymeric NPs
3.4. Physicochemical Properties of Encapsulated Antimicrobial Peptides (AMP-Nps)
3.5. Determination of Peptide Loading and Encapsulation Efficiency of AMP-Nps
3.6. In Vitro Release of Peptide from Nps
3.7. Determination of Minimum Inhibitory Concentration (MIC) and Minimum Bactericidal Concentration (MBC)
3.8. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. Available online: https://www.who.int/medicines/publications/WHOPPL-Short_Summary_25Feb-ET_NM_WHO.pdf (accessed on 25 November 2019).
- World Health Organization. Antimicrobial Resistance: Global Report on Surveillance. Available online: https://www.who.int/drugresistance/documents/surveillancereport/en/ (accessed on 25 November 2019).
- Center for Disease Control and Prevention. Antibiotic Resistance Threats in the United States. 2019. Available online: https://www.cdc.gov/drugresistance/pdf/threats-report/2019-ar-threats-report-508.pdf (accessed on 9 November 2019).
- Marder, E.P.; Griffin, P.M.; Cieslak, P.R.; Dunn, J.; Hurd, S.; Jervis, R.; Lathrop, S.; Muse, A.; Ryan, P.; Smith, K.; et al. Preliminary Incidence and Trends of Infections with Pathogens Transmitted Commonly through Food—Foodborne Diseases Active Surveillance Network, 10 U.S. Sites, 2006–2017. MMWR. Morb. Mortal. Wkly. Rep. 2018, 67, 324–328. [Google Scholar] [CrossRef] [PubMed]
- Fatima, R.; Aziz, M. Enterohemorrhagic Escherichia coli (EHEC); StatPearls Publishing: Petersburg FL, USA, 2020; pp. 1–10. [Google Scholar]
- Costa, F.; Teixeira, C.; Gomes, P.; Martins, M.C.L. Clinical Application of AMPs. Adv. Exp. Med. Biol. 2019, 1117, 281–298. [Google Scholar] [CrossRef] [PubMed]
- Gordon, Y.J.; Romanowski, E.G.; McDermott, A.M. A Review of Antimicrobial Peptides and Their Therapeutic Potential as Anti-Infective Drugs. Curr. Eye Res. 2005, 30, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Huang, J.; Chen, Y. Alpha-helical cationic antimicrobial peptides: Relationships of structure and function. Protein Cell 2010, 1, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Jenssen, H.; Hamill, P.; Hancock, R.E.W. Peptide Antimicrobial Agents. Clin. Microbiol. Rev. 2006, 19, 491–511. [Google Scholar] [CrossRef]
- Moghaddam, M.M.; Aghamollaei, H.; Kooshki, H.; Barjini, K.A.; Mirnejad, R.; Choopani, A. The development of antimicrobial peptides as an approach to prevention of antibiotic resistance. Rev. Med. Microbiol. 2015, 26, 98–110. [Google Scholar] [CrossRef]
- Bahar, A.; Ren, D. Antimicrobial Peptides. Pharmaceuticals 2013, 6, 1543–1575. [Google Scholar] [CrossRef]
- Jenssen, H.; Aspmo, S.I. Serum Stability of Peptides. In Methods in Molecular Biology; Otvos, L., Ed.; Humana Press: Totowa, NJ, USA, 2008; Volume 494, pp. 177–186. [Google Scholar]
- Oddo, A.; Hansen, P.R. Hemolytic Activity of Antimicrobial Peptides. In Antimicrobial Peptides: Methods and Protocols, Methods in Molecular Biology; Hansen, P.R., Ed.; Methods in Molecular Biology; Humana Press: Totowa, NY, USA, 2017; Volume 1548, pp. 427–435. [Google Scholar]
- Hemeg, H.A. Nanomaterials for alternative antibacterial therapy. Int. J. Nanomed. 2017, 12, 8211–8225. [Google Scholar] [CrossRef]
- Pelgrift, R.Y.; Friedman, A.J. Nanotechnology as a therapeutic tool to combat microbial resistance. Adv. Drug Deliv. Rev. 2013, 65, 1803–1815. [Google Scholar] [CrossRef] [PubMed]
- Casciaro, B.; D’Angelo, I.; Zhang, X.; Loffredo, M.R.; Conte, G.; Cappiello, F.; Quaglia, F.; Di, Y.-P.P.; Ungaro, F.; Mangoni, M.L. Poly(lactide-co-glycolide) Nanoparticles for Prolonged Therapeutic Efficacy of Esculentin-1a-Derived Antimicrobial Peptides against Pseudomonas aeruginosa Lung Infection: In Vitro and in Vivo Studies. Biomacromolecules 2019. [Google Scholar] [CrossRef]
- Cruz, J.; Rondon-Villarreal, P.; Torres, R.G.; Urquiza, M.; Guzman, F.; Alvarez, C.; Abengozar, M.A.; Sierra, D.A.; Rivas, L.; Fernandez-Lafuente, R.; et al. Design of Bactericidal Peptides Against Escherichia coli O157:H7, Pseudomonas aeruginosa and methicillin-resistant Staphylococcus aureus. Med. Chem. (Los. Angeles) 2018, 14, 741–752. [Google Scholar] [CrossRef]
- Cruz, J.; Flórez, J.; Torres, R.; Urquiza, M.; Gutiérrez, J.A.; Guzmán, F.; Ortiz, C.C. Antimicrobial activity of a new synthetic peptide loaded in polylactic acid or poly(lactic-co-glycolic) acid nanoparticles against Pseudomonas aeruginosa, Escherichia coli O157:H7 and methicillin-resistant Staphylococcus aureus (MRSA). Nanotechnology 2017, 28, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Waghu, F.H.; Barai, R.S.; Gurung, P.; Idicula-Thomas, S. CAMPR3: A database on sequences, structures and signatures of antimicrobial peptides. Nucleic Acids Res. 2016, 44, D1094–D1097. [Google Scholar] [CrossRef] [PubMed]
- Guruprasad, K.; Reddy, B.V.B.; Pandit, M.W. Correlation between stability of a protein and its dipeptide composition: A novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng. Des. Sel. 1990, 4, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Krishnakumari, V.; Nagaraj, R. Antimicrobial and hemolytic activities of crabrolin, a 13-residue peptide from the venom of the European hornet, Vespa crabro, and its analogs. J. Pept. Res. 2009, 50, 88–93. [Google Scholar] [CrossRef] [PubMed]
- Umerska, A.; Cassisa, V.; Bastiat, G.; Matougui, N.; Nehme, H.; Manero, F.; Eveillard, M.; Saulnier, P. Synergistic interactions between antimicrobial peptides derived from plectasin and lipid nanocapsules containing monolaurin as a cosurfactant against Staphylococcus aureus. Int. J. Nanomed. 2017, 12, 5687–5699. [Google Scholar] [CrossRef]
- Rogueda, P.G.; Traini, D. The nanoscale in pulmonary delivery. Part 1: Deposition, fate, toxicology and effects. Expert Opin. Drug Deliv. 2007, 4, 595–606. [Google Scholar] [CrossRef]
- Wang, L.; Hu, C.; Shao, L. The antimicrobial activity of nanoparticles: Present situation and prospects for the future. Int. J. Nanomed. 2017, 12, 1227–1249. [Google Scholar] [CrossRef]
- Yin Win, K.; Feng, S.-S. Effects of particle size and surface coating on cellular uptake of polymeric nanoparticles for oral delivery of anticancer drugs. Biomaterials 2005, 26, 2713–2722. [Google Scholar] [CrossRef]
- He, C.; Hu, Y.; Yin, L.; Tang, C.; Yin, C. Effects of particle size and surface charge on cellular uptake and biodistribution of polymeric nanoparticles. Biomaterials 2010, 31, 3657–3666. [Google Scholar] [CrossRef]
- Tiyaboonchai, W.; Woiszwillo, J.; Middaugh, C.R. Formulation and characterization of amphotericin B–polyethylenimine–dextran sulfate nanoparticles. J. Pharm. Sci. 2001, 90, 902–914. [Google Scholar] [CrossRef] [PubMed]
- Water, J.J.; Smart, S.; Franzyk, H.; Foged, C.; Nielsen, H.M. Nanoparticle-mediated delivery of the antimicrobial peptide plectasin against Staphylococcus aureus in infected epithelial cells. Eur. J. Pharm. Biopharm. 2015, 92, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Na, R.; Wang, X.; Liu, H.; Zhao, L.; Sun, X.; Ma, G.; Cui, F. Fabrication of antimicrobial peptide-loaded PLGA/Chitosan composite microspheres for long-Acting bacterial resistance. Molecules 2017, 22, 1637. [Google Scholar] [CrossRef] [PubMed]
- Khayata, N.; Abdelwahed, W.; Chehna, M.F.; Charcosset, C.; Fessi, H. Preparation of vitamin E loaded nanocapsules by the nanoprecipitation method: From laboratory scale to large scale using a membrane contactor. Int. J. Pharm. 2012, 423, 419–427. [Google Scholar] [CrossRef]
- Huang, M.; Khor, E.; Lim, L.-Y. Uptake and Cytotoxicity of Chitosan Molecules and Nanoparticles: Effects of Molecular Weight and Degree of Deacetylation. Pharm. Res. 2004, 21, 344–353. [Google Scholar] [CrossRef]
- Gómez-Sequeda, N.S.; Torres, R.; Ortiz López, C.C. Synthesis, characterization, and in vitro activity against Candida spp. of fluconazole encapsulated on cationic and conventional nanoparticles of poly(lactic-co-glycolic acid). Nanotechnol. Sci. Appl. 2017, 10, 95–104. [Google Scholar] [CrossRef]
- Stoll, S. The Importance of Zeta Potential Measurements & Role of Ionic Strength in Flocculation Processes. Water Technol. 2013, 4, 1–5. [Google Scholar]
- Malvern Panalytical. Pharmaceutical Formulations and the Importance of Zeta Potential to Pharmaceutical Formulations with Supplier Data by Malvern. Available online: http://www.azonano.com/article.aspx?ArticleID=1234 (accessed on 12 November 2019).
- Cohen-Sela, E.; Chorny, M.; Koroukhov, N.; Danenberg, H.D.; Golomb, G. A new double emulsion solvent diffusion technique for encapsulating hydrophilic molecules in PLGA nanoparticles. J. Control. Release 2009, 133, 90–95. [Google Scholar] [CrossRef]
- Sun, Z.; Zhong, J.; Liang, X.; Liu, J.; Chen, X.; Huan, L. Novel mechanism for nisin resistance via proteolytic degradation of nisin by the nisin resistance protein NSR. Antimicrob. Agents Chemother. 2009, 53, 1964–1973. [Google Scholar] [CrossRef]
- Sakoulas, G.; Guram, K.; Reyes, K.; Nizet, V.; Zervos, M. Human cathelicidin LL-37 Resistance and increased daptomycin MIC in methicillin-resistant Staphylococcus aureus strain USA600 (ST45) are associated with increased mortality in a hospital setting. J. Clin. Microbiol. 2014, 52, 2172–2174. [Google Scholar] [CrossRef]
- Saxena, T.; Kaushik, P.; Krishna Mohan, M. Prevalence of E. coli O157:H7 in water sources: An overview on associated diseases, outbreaks and detection methods. Diagn. Microbiol. Infect. Dis. 2015, 82, 249–264. [Google Scholar] [CrossRef] [PubMed]
- Panyam, J.; William, D.; Dash, A.; Leslie-Pelecky, D.; Labhasetwar, V. Solid-state solubility influences encapsulation and release of hydrophobic drugs from PLGA/PLA nanoparticles. J. Pharm. Sci. 2004, 93, 1804–1814. [Google Scholar] [CrossRef] [PubMed]
- Xie, S.; Tao, Y.; Pan, Y.; Qu, W.; Cheng, G.; Huang, L.; Chen, D.; Wang, X.; Liu, Z.; Yuan, Z. Biodegradable nanoparticles for intracellular delivery of antimicrobial agents. J. Control. Release 2014, 187, 101–117. [Google Scholar] [CrossRef] [PubMed]
- Houghten, R.A. General method for the rapid solid-phase synthesis of large numbers of peptides: Specificity of antigen-antibody interaction at the level of individual amino acids. Proc. Natl. Acad. Sci. USA 1985, 82, 5131–5135. [Google Scholar] [CrossRef]
- Fields, G.B.; Noble, R.L. Solid phase peptide synthesis utilizing 9-fluorenylmethoxycarbonyl amino acids. Int. J. Pept. Protein Res. 1990, 35, 161–214. [Google Scholar] [CrossRef]
- Jofré, C.; Guzmán, F.; Cárdenas, C.; Albericio, F.; Marshall, S.H. A natural peptide and its variants derived from the processing of infectious pancreatic necrosis virus (IPNV) displaying enhanced antimicrobial activity: A novel alternative for the control of bacterial diseases. Peptides 2011, 32, 852–858. [Google Scholar] [CrossRef]
- Wadhwani, P.; Epand, R.F.; Heidenreich, N.; Bürck, J.; Ulrich, A.S.; Epand, R.M. Membrane-active peptides and the clustering of anionic lipids. Biophys. J. 2012, 103, 265–274. [Google Scholar] [CrossRef]
- Santoveña, A.; Oliva, A.; Guzman, F.; Patarroyo, M.E.; Llabrés, M.; Fariña, J.B. Chromatographic characterization of synthetic peptides: SPf66 malaria vaccine. J. Chromatogr. B 2002, 766, 3–12. [Google Scholar] [CrossRef]
- Rivera, P.A.; Martinez-Oharriz, M.C.; Rubio, M.; Irache, J.M.; Espuelas, S. Fluconazole encapsulation in PLGA microspheres by spray-drying. J. Microencapsul. 2004, 21, 203–211. [Google Scholar] [CrossRef]
- Aguilar, M. HPLC of Peptides and Proteins: Basic Theory and Methodology. In HPLC of Peptides and Proteins; Aguilar, M., Ed.; Humana Press: Totowa, NJ, USA, 2004; Volume 251, pp. 3–8. [Google Scholar]
- Thomasin, C.; Corradin, G.; Men, Y.; Merkle, H.P.; Gander, B. Tetanus toxoid and synthetic malaria antigen containing poly(lactide)/poly(lactide-co-glycolide) microspheres: Importance of polymer degradation and antigen release for immune response. J. Control. Release 1996, 41, 131–145. [Google Scholar] [CrossRef]
- Paredes, D.; Ortiz, C.; Torres, R. Synthesis, characterization, and evaluation of antibacterial effect of Ag nanoparticles against Escherichia coli O157:H7 and methicillin-resistant Staphylococcus aureus (MRSA). Int. J. Nanomed. 2014, 9, 1717–1729. [Google Scholar] [CrossRef]

| Peptide | Sequence | Net Charge | MW (Da) | m/z | PAP (%) | pI | II | GRAVY |
|---|---|---|---|---|---|---|---|---|
| G17 | ATKKCGLFKILKGVGKI | 5 | 1804.31 | 1804.19 | 96 | 10.20 | 18.06 | 0.382 |
| G19 | ATKKCALWSILKAVAKI | 4 | 1844.33 | 1864.19 | 97.3 | 10.04 | 24.68 | 0.735 |
| NP | (µg AMP /mgPLGA) Ratio | Mean Zeta Potential (mV) | Mean Size (nm) | Mean PDI | Mean Encapsulation Efficiency (%) | Mean Peptide Loading (µg AMP/mg PLGA) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Empty NP | 0 | −21.97 | ± | 1.25 | 270.87 | ± | 2.96 | 0.11 | ± | 0.03 | - | - | - | - | ||
| G17NP | 16 | 8.56 | ± | 0.30 | 293.20 | ± | 1.61 | 0.24 | ± | 0.03 | 49.50 | ± | 0.70 | 7.92 | ± | 0.11 |
| 8 | 7.26 | ± | 1.11 | 284.50 | ± | 3.61 | 0.22 | ± | 0.02 | 90.59 | ± | 0.67 | 7.25 | ± | 0.05 | |
| 4 | 3.99 | ± | 0.23 | 303.33 | ± | 18.65 | 0.27 | ± | 0.07 | 76.10 | ± | 1.87 | 3.04 | ± | 0.07 | |
| 2 | −2.06 | ± | 0.21 | 1022.40 | ± | 317.18 | 0.83 | ± | 0.27 | 67.58 | ± | 0.83 | 1.35 | ± | 0.02 | |
| 1 | −14.10 | ± | 0.56 | 322.27 | ± | 4.25 | 0.22 | ± | 0.02 | 83.79 | ± | 0.14 | 0.84 | ± | 0.00 | |
| 0.50 | −19.03 | ± | 0.64 | 339.53 | ± | 1.86 | 0.12 | ± | 0.08 | 86.19 | ± | 3.59 | 0.43 | ± | 0.02 | |
| 0.25 | −20.80 | ± | 0.46 | 285.37 | ± | 1.61 | 0.08 | ± | 0.04 | 87.64 | ± | 3.66 | 0.22 | ± | 0.01 | |
| 0.12 | −23.37 | ± | 0.45 | 266.30 | ± | 4.35 | 0.12 | ± | 0.04 | 88.48 | ± | 3.01 | 0.11 | ± | 0.00 | |
| G19NP | 16 | 12.90 | ± | 0.60 | 291.67 | ± | 3.38 | 0.16 | ± | 0.11 | 50.85 | ± | 1.20 | 8.14 | ± | 0.19 |
| 8 | 12.87 | ± | 1.02 | 291.77 | ± | 1.59 | 0.26 | ± | 0.06 | 90.60 | ± | 0.94 | 7.25 | ± | 0.07 | |
| 4 | 12.90 | ± | 0.80 | 257.83 | ± | 2.84 | 0.13 | ± | 0.05 | 80.48 | ± | 1.76 | 3.22 | ± | 0.07 | |
| 2 | 3.03 | ± | 0.30 | 290.70 | ± | 3.67 | 0.08 | ± | 0.04 | 79.27 | ± | 2.26 | 1.59 | ± | 0.04 | |
| 1 | −1.86 | ± | 0.37 | 432.63 | ± | 6.66 | 0.16 | ± | 0.04 | 51.10 | ± | 4.98 | 0.51 | ± | 0.05 | |
| 0.50 | −11.80 | ± | 0.95 | 1976.00 | ± | 496.56 | 0.82 | ± | 0.12 | 41.05 | ± | 3.18 | 0.21 | ± | 0.02 | |
| 0.25 | −15.77 | ± | 0.40 | 353.67 | ± | 5.93 | 0.17 | ± | 0.02 | 89.92 | ± | 6.88 | 0.22 | ± | 0.02 | |
| 0.12 | −18.73 | ± | 0.40 | 324.87 | ± | 7.27 | 0.12 | ± | 0.05 | 89.45 | ± | 4.39 | 0.11 | ± | 0.01 | |
| Microorganism | E. coli O157:H7 | MRSA | ||||
|---|---|---|---|---|---|---|
| Compound | MIC50 (µM) | MIC90 (µM) | MBC (µM) | MIC50 (µM) | MIC90 (µM) | MBC (µM) |
| G17 | 12.5 | 20 | 70 | 1.5 | 12.5 | 100 |
| G17NP | 3.13 | 25 | >100 | 0.2 | 12.5 | 25 |
| G19 | 12.5 | 20 | 70 | 1.5 | 6.5 | 70 |
| G19NP | 3.13 | 20 | 100 | 0.7 | 20 | 100 |
| NP | >100 | >100 | >100 | >100 | >100 | >100 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gómez-Sequeda, N.; Ruiz, J.; Ortiz, C.; Urquiza, M.; Torres, R. Potent and Specific Antibacterial Activity against Escherichia coli O157:H7 and Methicillin Resistant Staphylococcus aureus (MRSA) of G17 and G19 Peptides Encapsulated into Poly-Lactic-Co-Glycolic Acid (PLGA) Nanoparticles. Antibiotics 2020, 9, 384. https://doi.org/10.3390/antibiotics9070384
Gómez-Sequeda N, Ruiz J, Ortiz C, Urquiza M, Torres R. Potent and Specific Antibacterial Activity against Escherichia coli O157:H7 and Methicillin Resistant Staphylococcus aureus (MRSA) of G17 and G19 Peptides Encapsulated into Poly-Lactic-Co-Glycolic Acid (PLGA) Nanoparticles. Antibiotics. 2020; 9(7):384. https://doi.org/10.3390/antibiotics9070384
Chicago/Turabian StyleGómez-Sequeda, Nicolás, Jennifer Ruiz, Claudia Ortiz, Mauricio Urquiza, and Rodrigo Torres. 2020. "Potent and Specific Antibacterial Activity against Escherichia coli O157:H7 and Methicillin Resistant Staphylococcus aureus (MRSA) of G17 and G19 Peptides Encapsulated into Poly-Lactic-Co-Glycolic Acid (PLGA) Nanoparticles" Antibiotics 9, no. 7: 384. https://doi.org/10.3390/antibiotics9070384
APA StyleGómez-Sequeda, N., Ruiz, J., Ortiz, C., Urquiza, M., & Torres, R. (2020). Potent and Specific Antibacterial Activity against Escherichia coli O157:H7 and Methicillin Resistant Staphylococcus aureus (MRSA) of G17 and G19 Peptides Encapsulated into Poly-Lactic-Co-Glycolic Acid (PLGA) Nanoparticles. Antibiotics, 9(7), 384. https://doi.org/10.3390/antibiotics9070384