Diagnosis of Bloodstream Infections: An Evolution of Technologies towards Accurate and Rapid Identification and Antibiotic Susceptibility Testing
Abstract
1. Challenges in Bloodstream Infection Diagnosis
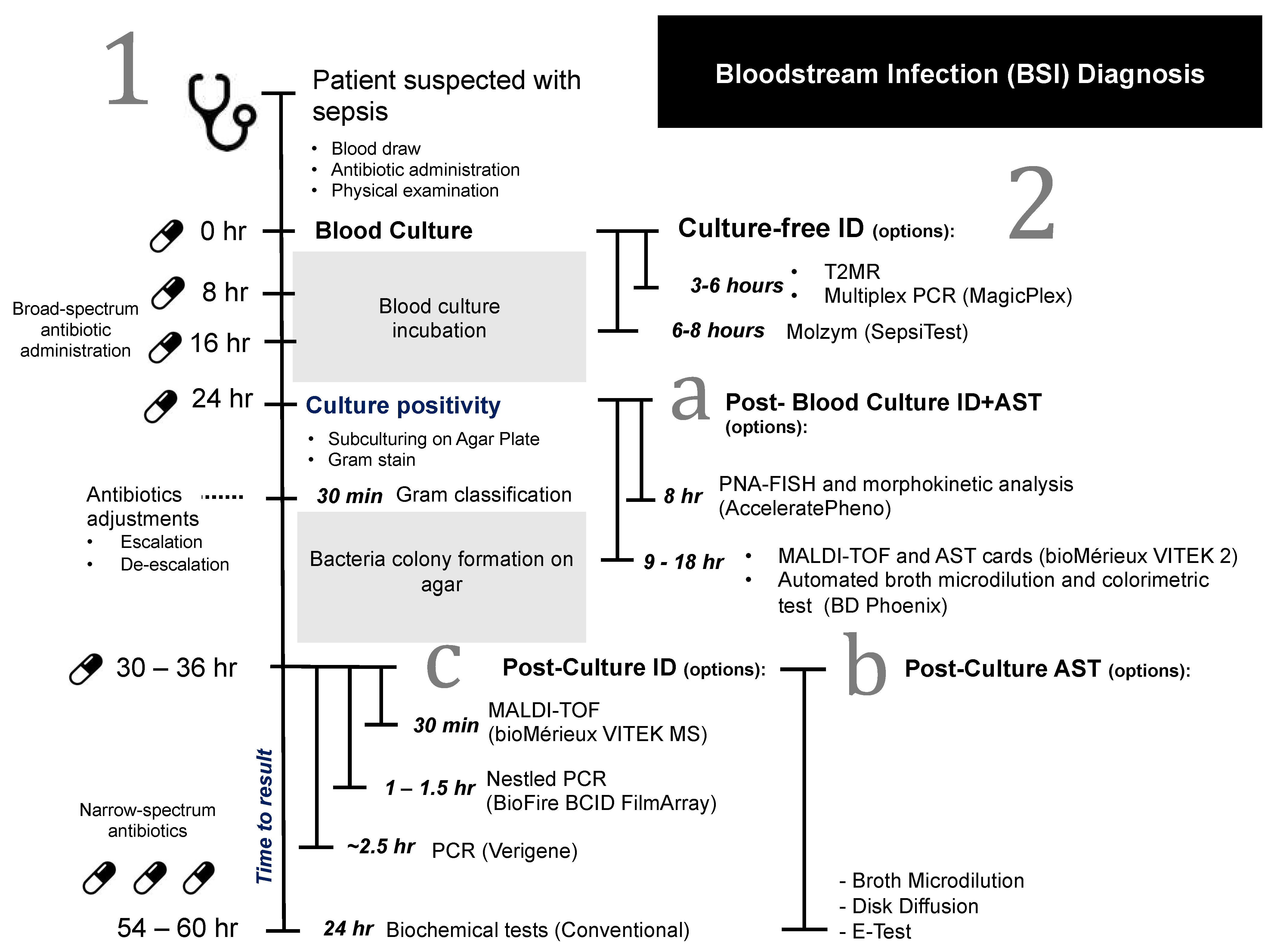
2. Overview of Current Blood Culture Diagnosis Workflow
3. Challenges of BSI Diagnosis and Clinical Significance for a Novel Technique
- (1)
- Sample preparation directly from whole blood that requires minimal handling and can process sufficient blood volume to ensure the capture of pathogens in low abundance;
- (2)
- Workflow that effectively separates, enriches, and concentrates pathogens or target analytes from background interferences;
- (3)
- Sensitive, quantitative, and accurate detection and species-level identification that differentiates pathogens from contaminants or commensals;
- (4)
- Timely and universal antibiotic susceptibility profiling independent of resistant mechanisms with MIC, reporting matches with critical clinical decision timepoints.
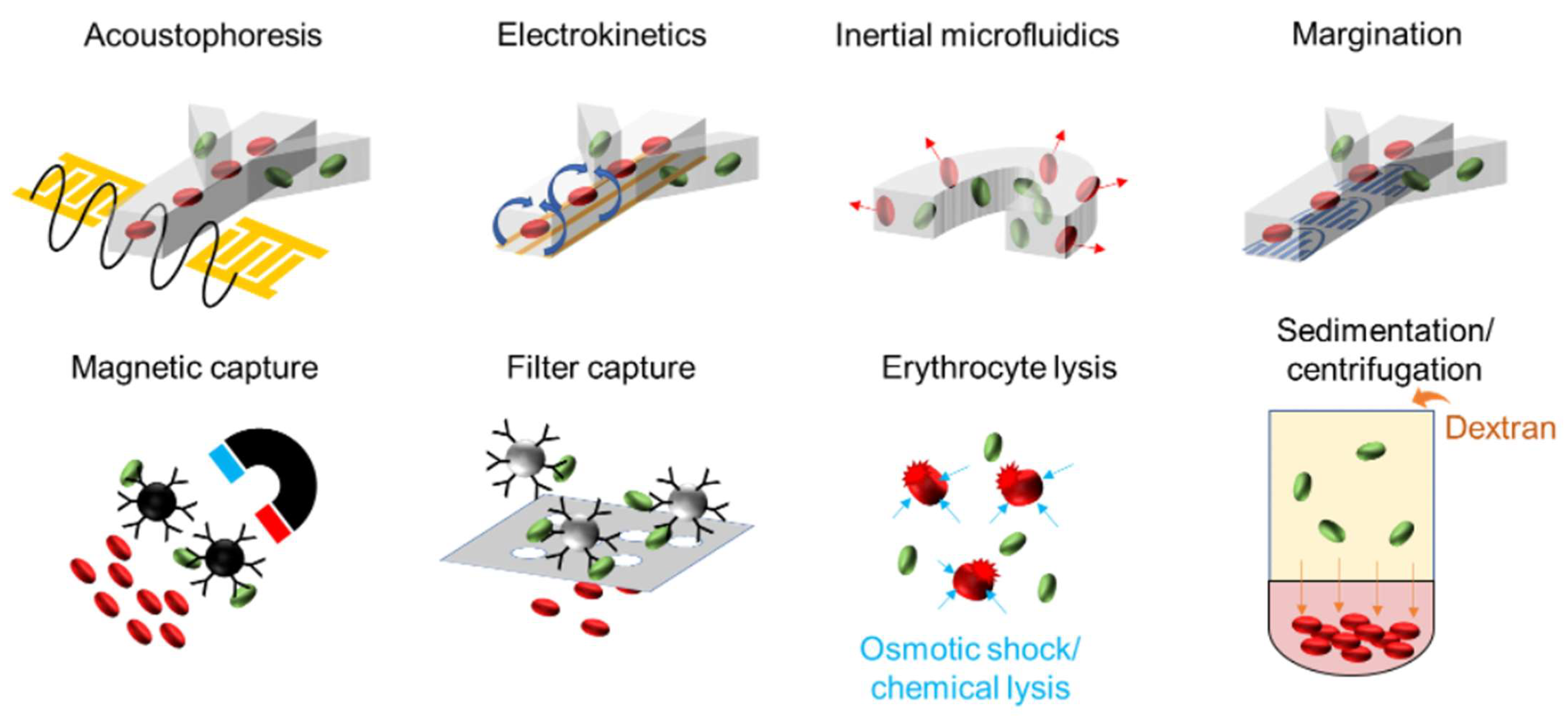
3.1. Sample Preparation and Assay Workflow for Capturing, Separating, and Enriching Low-Abundance Pathogens
3.1.1. Affinity Capture
3.1.2. Erythrocyte Depletion
3.1.3. Acoustophoresis
3.1.4. Electrokinetics
3.1.5. Inertial Focusing
3.2. Sensitive and Quantitative Pathogen Detection and ID with Timely AST
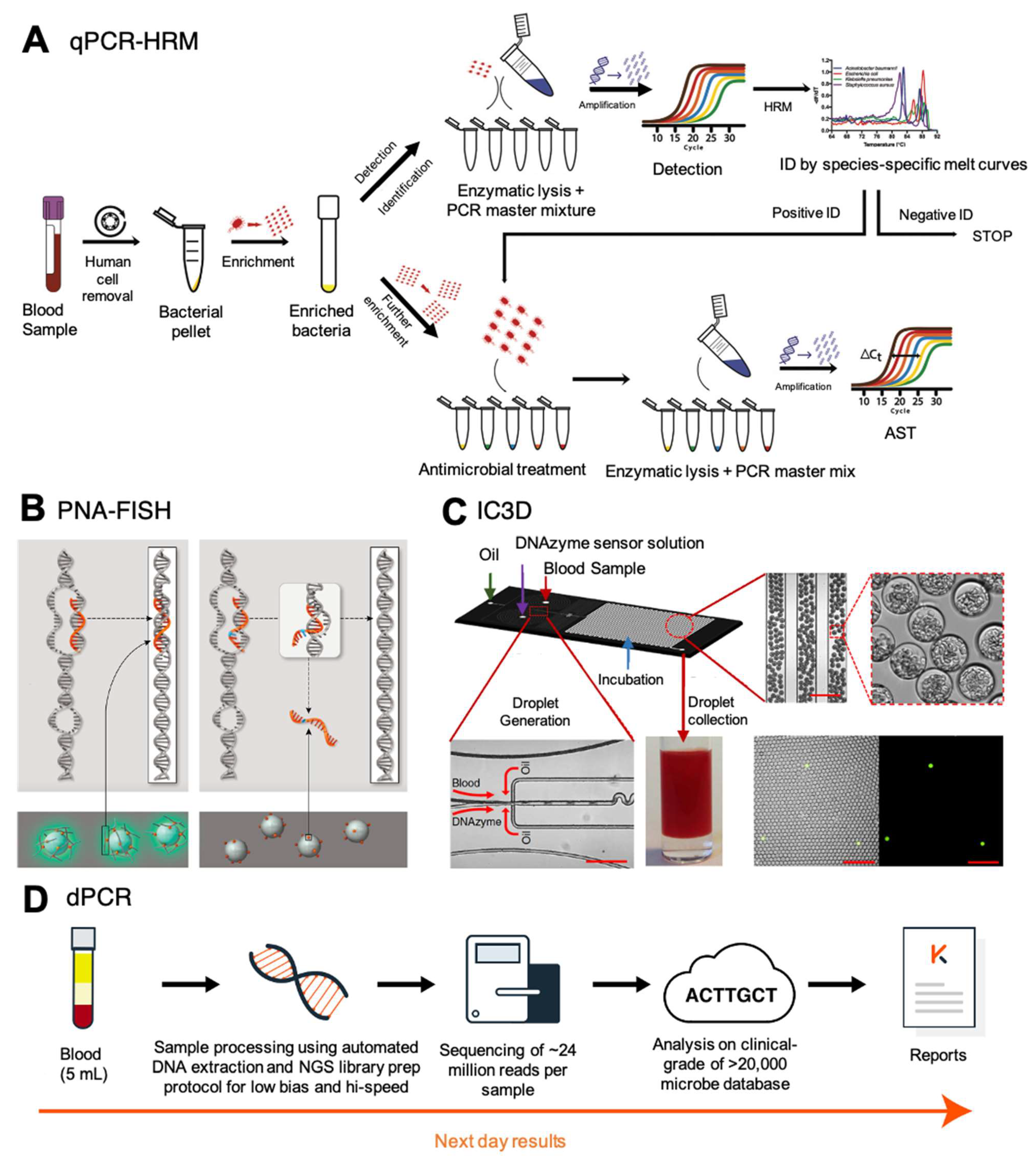
3.2.1. Digital PCR (dPCR)
3.2.2. Universal High-Resolution Melt (U-HRM) with Pheno-Molecular AST
3.2.3. Gamma Peptide Nucleic Acid (γPNA)
3.2.4. Next-Generation Sequencing (NGS)
3.2.5. Surface-Enhanced Raman Spectroscopy
3.2.6. Flow Cytometry
4. Summary and Future Outlook (Conclusions)
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Miller, J.M.; Binnicker, M.J.; Campbell, S.; Carroll, K.C.; Chapin, K.C.; Gilligan, P.H.; Gonzalez, M.D.; Jerris, R.C.; Kehl, S.C.; Patel, R.; et al. A guide to utilization of the microbiology laboratory for diagnosis of infectious diseases: 2018 update by the Infectious Diseases Society of America and the American Society for Microbiology. Clin. Infect. Dis. 2018, 67, e1–e94. [Google Scholar] [CrossRef] [PubMed]
- Castillo, D.J.; Rifkin, R.F.; Cowan, D.A.; Potgieter, M. The healthy human blood microbiome: Fact or fiction? Front. Cell. Infect. Microbiol. 2019, 9, 148. [Google Scholar] [CrossRef] [PubMed]
- Molitor, E. Susceptibility categories should be agreed upon. J. Clin. Microbiol. 2019, 57. [Google Scholar] [CrossRef] [PubMed]
- Opota, O.; Jaton, K.; Greub, G. Microbial diagnosis of bloodstream infection: Towards molecular diagnosis directly from blood. Clin. Microbiol. Infect. 2015, 21, 323–331. [Google Scholar] [CrossRef]
- Falconer, K.; Hammond, R.; Gillespie, S.H. Improving the recovery and detection of bloodstream pathogens from blood culture. J. Med. Microbiol. 2020, 69, 806–811. [Google Scholar] [CrossRef]
- Seymour, C.W.; Gesten, F.; Prescott, H.C.; Friedrich, M.E.; Iwashyna, T.J.; Phillips, G.S.; Lemeshow, S.; Osborn, T.; Terry, K.M.; Levy, M.M. Time to Treatment and Mortality during Mandated Emergency Care for Sepsis. N. Engl. J. Med. 2017, 376, 2235–2244. [Google Scholar] [CrossRef]
- CDC. Antibiotic Use in the United States, 2021 Update: Progress and Opportunities; U.S. Department of Health and Human Services: Washington, DC, USA, 2021; pp. 1–24. Available online: https://www.cdc.gov/antibiotic-use/index.html (accessed on 7 October 2021).
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Ling, T.K.W.; Liu, Z.K.; Cheng, A.F.B. Evaluation of the VITEK 2 system for rapid direct identification and susceptibility testing of gram-negative bacilli from positive blood cultures. J. Clin. Microbiol. 2003, 41, 4705–4707. [Google Scholar] [CrossRef]
- Barman, P.; Chopra, S.; Thukral, T. Direct testing by VITEK® 2: A dependable method to reduce turnaround time in Gram-negative bloodstream infections. J. Lab. Physicians 2018, 10, 260–264. [Google Scholar] [CrossRef]
- Munson, E.L.; Diekema, D.J.; Beekmann, S.E.; Chapin, K.C.; Doern, G.V. Detection and treatment of bloodstream infection: Laboratory reporting and antimicrobial management. J. Clin. Microbiol. 2003, 41, 495–497. [Google Scholar] [CrossRef][Green Version]
- Idelevich, E.A.; Becker, K. How to accelerate antimicrobial susceptibility testing. Clin. Microbiol. Infect. 2019, 25, 1347–1355. [Google Scholar] [CrossRef] [PubMed]
- van Belkum, A.; Burnham, C.A.D.; Rossen, J.W.A.; Mallard, F.; Rochas, O.; Dunne, W.M. Innovative and rapid antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 2020, 18, 299–311. [Google Scholar] [CrossRef] [PubMed]
- Vasala, A.; Hytönen, V.P.; Laitinen, O.H. Modern tools for rapid diagnostics of antimicrobial resistance. Front. Cell. Infect. Microbiol. 2020, 10, 308. [Google Scholar] [CrossRef] [PubMed]
- Datar, R.; Orenga, S.; Pogorelcnik, R.; Rochas, O.; Simner, P.J.; van Belkum, A. Recent Advances in Rapid Antimicrobial Susceptibility Testing. Clin. Chem. 2021, 68, 91–98. [Google Scholar] [CrossRef]
- Behera, B.; Vishnu, G.A.; Chatterjee, S.; Sreekumar, N.; Nagabhushan, A.; Rajendran, N.; Prathik, B.H.; Pandya, H.J. Emerging technologies for antibiotic susceptibility testing. Biosens. Bioelectron. 2019, 142, 111552. [Google Scholar] [CrossRef]
- Principi, N.; Silvestri, E.; Esposito, S. Advantages and limitations of bacteriophages for the treatment of bacterial infections. Front. Pharmacol. 2019, 10, 513. [Google Scholar] [CrossRef]
- Peker, N.; Couto, N.; Sinha, B.; Rossen, J.W. Diagnosis of bloodstream infections from positive blood cultures and directly from blood samples: Recent developments in molecular approaches. Clin. Microbiol. Infect. 2018, 24, 944–955. [Google Scholar] [CrossRef]
- Maugeri, G.; Lychko, I.; Sobral, R.; Roque, A.C.A. Identification and antibiotic-susceptibility profiling of infectious bacterial agents: A review of current and future trends. Biotechnol. J. 2019, 14, 1700750. [Google Scholar] [CrossRef]
- Shin, D.J.; Andini, N.; Hsieh, K.; Yang, S.; Wang, T.-H. Emerging analytical techniques for rapid pathogen identification and susceptibility testing. Annu. Rev. Anal. Chem. 2019, 12, 41–67. [Google Scholar] [CrossRef]
- Fredborg, M.; Andersen, K.R.; Jørgensen, E.; Droce, A.; Olesen, T.; Jensen, B.B.; Rosenvinge, F.S.; Sondergaard, T.E. Real-time optical antimicrobial susceptibility testing. J. Clin. Microbiol. 2013, 51, 2047–2053. [Google Scholar] [CrossRef]
- Zhu, N.; Wong, P.K. Advances in Viral Diagnostic Technologies for Combating COVID-19 and Future Pandemics. SLAS Technol. Transl. Life Sci. Innov. 2020, 25, 513–521. [Google Scholar] [CrossRef] [PubMed]
- Mariella, R. Sample preparation: The weak link in microfluidics-based biodetection. Biomed. Microdevices 2008, 10, 777–784. [Google Scholar] [CrossRef] [PubMed]
- Xia, N.; Hunt, T.P.; Mayers, B.T.; Alsberg, E.; Whitesides, G.M.; Westervelt, R.M.; Ingber, D.E. Combined microfluidic-micromagnetic separation of living cells in continuous flow. Biomed. Microdevices 2006, 8, 299–308. [Google Scholar] [CrossRef] [PubMed]
- Yung, C.W.; Fiering, J.; Mueller, A.J.; Ingber, D.E. Micromagnetic-microfluidic blood cleansing device. Lab Chip 2009, 9, 1171–1177. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.H.; Super, M.; Yung, C.W.; Cooper, R.M.; Domansky, K.; Graveline, A.R.; Mammoto, T.; Berthet, J.B.; Tobin, H.; Cartwright, M.J.; et al. An extracorporeal blood-cleansing device for sepsis therapy. Nat. Med. 2014, 20, 1211–1216. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.L.; Wang, C.H.; Chen, Y.S.; Chien, C.C.; Kuo, F.C.; You, H.L.; Lee, M.S.; Lee, G.B. An integrated microfluidic system for early detection of sepsis-inducing bacteria. Lab Chip 2021, 21, 113–121. [Google Scholar] [CrossRef]
- Lee, J.J.; Jeong, K.J.; Hashimoto, M.; Kwon, A.H.; Rwei, A.; Shankarappa, S.A.; Tsui, J.H.; Kohane, D.S. Synthetic ligand-coated magnetic nanoparticles for microfluidic bacterial separation from blood. Nano Lett. 2014, 14, 1–5. [Google Scholar] [CrossRef]
- Zelenin, S.; Hansson, J.; Ardabili, S.; Ramachandraiah, H.; Brismar, H.; Russom, A. Microfluidic-based isolation of bacteria from whole blood for sepsis diagnostics. Biotechnol. Lett. 2015, 37, 825–830. [Google Scholar] [CrossRef]
- Li, H.; Torab, P.; Mach, K.E.; Surrette, C.; England, M.R.; Craft, D.W.; Thomas, N.J.; Liao, J.C.; Puleo, C.; Wong, P.K. Adaptable microfluidic system for single-cell pathogen classification and antimicrobial susceptibility testing. Proc. Natl. Acad. Sci. USA 2019, 116, 10270–10279. [Google Scholar] [CrossRef]
- Forsyth, B.; Torab, P.; Lee, J.H.; Malcom, T.; Wang, T.H.; Liao, J.C.; Yang, S.; Kvam, E.; Puleo, C.; Wong, P.K. A Rapid Single-Cell Antimicrobial Susceptibility Testing Workflow for Bloodstream Infections. Biosensors 2021, 11, 288. [Google Scholar] [CrossRef]
- Ai, Y.; Sanders, C.K.; Marrone, B.L. Separation of Escherichia coli Bacteria from Peripheral Blood Mononuclear Cells Using Standing Surface Acoustic Waves. Anal. Chem. 2013, 85, 9126–9134. [Google Scholar] [CrossRef] [PubMed]
- Ohlsson, P.; Evander, M.; Petersson, K.; Mellhammar, L.; Lehmusvuori, A.; Karhunen, U.; Soikkeli, M.; Seppä, T.; Tuunainen, E.; Spangar, A.; et al. Integrated Acoustic Separation, Enrichment, and Microchip Polymerase Chain Reaction Detection of Bacteria from Blood for Rapid Sepsis Diagnostics. Anal. Chem. 2016, 88, 9403–9411. [Google Scholar] [CrossRef] [PubMed]
- Van Assche, D.; Reithuber, E.; Qiu, W.; Laurell, T.; Henriques-Normark, B.; Mellroth, P.; Ohlsson, P.; Augustsson, P. Gradient acoustic focusing of sub-micron particles for separation of bacteria from blood lysate. Sci. Rep. 2020, 10, 3670. [Google Scholar] [CrossRef]
- Kuczenski, R.; Chang, H.-C.; Revzin, A. Dielectrophoretic microfluidic device for the continuous sorting of Escherichia coli from blood cells. Biomicrofluidics 2011, 5, 32005–3200515. [Google Scholar] [CrossRef] [PubMed]
- Bisceglia, E.; Cubizolles, M.; Trainito, C.; Berthier, J.; Pudda, C.; Français, O.; Mallard, F.; Le Pioufle, B. A generic and label free method based on dielectrophoresis for the continuous separation of microorganism from whole blood samples. Sensors Actuators B Chem. 2015, 212, 335–343. [Google Scholar] [CrossRef]
- Gao, J.; Sin, M.L.Y.; Liu, T.; Gau, V.; Liao, J.C.; Wong, P.K. Hybrid Electrokinetic Manipulation in High-Conductivity Media. Lab Chip 2011, 11, 1770–1775. [Google Scholar] [CrossRef] [PubMed]
- Mach, A.J.; Di Carlo, D. Continuous scalable blood filtration device using inertial microfluidics. Biotechnol. Bioeng. 2010, 107, 302–311. [Google Scholar] [CrossRef]
- Hou, H.W.; Bhattacharyya, R.P.; Hung, D.T.; Han, J. Direct detection and drug-resistance profiling of bacteremias using inertial microfluidics. Lab Chip 2015, 15, 2297–2307. [Google Scholar] [CrossRef]
- Faridi, M.A.; Ramachandraiah, H.; Banerjee, I.; Ardabili, S.; Zelenin, S.; Russom, A. Elasto-inertial microfluidics for bacteria separation from whole blood for sepsis diagnostics. J. Nanobiotechnol. 2017, 15, 3. [Google Scholar] [CrossRef]
- Narayana Iyengar, S.; Kumar, T.; Martensson, G.; Russom, A. High resolution and rapid separation of bacteria from blood using elasto-inertial microfluidics. Electrophoresis 2021, 42, 2538–2551. [Google Scholar] [CrossRef]
- Lu, X.; Chow, J.J.M.; Koo, S.H.; Jiang, B.; Tan, T.Y.; Yang, D.; Ai, Y. Sheathless and high-throughput elasto-inertial bacterial sorting for enhancing molecular diagnosis of bloodstream infection. Lab Chip 2021, 21, 2163–2177. [Google Scholar] [CrossRef] [PubMed]
- Wei Hou, H.; Gan, H.Y.; Bhagat, A.A.; Li, L.D.; Lim, C.T.; Han, J. A microfluidics approach towards high-throughput pathogen removal from blood using margination. Biomicrofluidics 2012, 6, 24115–2411513. [Google Scholar] [CrossRef] [PubMed]
- Chu, Y.W.; Engebretson, D.A.; Carey, J.R. Bioconjugated magnetic nanoparticles for the detection of bacteria. J. Biomed. Nanotechnol. 2013, 9, 1951–1961. [Google Scholar] [CrossRef] [PubMed]
- Pitt, W.G.; Alizadeh, M.; Husseini, G.A.; McClellan, D.S.; Buchanan, C.M.; Bledsoe, C.G.; Robison, R.A.; Blanco, R.; Roeder, B.L.; Melville, M.; et al. Rapid separation of bacteria from blood-review and outlook. Biotechnol. Prog. 2016, 32, 823–839. [Google Scholar] [CrossRef]
- Burklund, A.; Zhang, J.X.J. Microfluidics-Based Organism Isolation from Whole Blood: An Emerging Tool for Bloodstream Infection Diagnosis. Ann. Biomed. Eng. 2019, 47, 1657–1674. [Google Scholar] [CrossRef]
- Wang, S.; Inci, F.; Chaunzwa, T.L.; Ramanujam, A.; Vasudevan, A.; Subramanian, S.; Chi Fai Ip, A.; Sridharan, B.; Gurkan, U.A.; Demirci, U. Portable microfluidic chip for detection of Escherichia coli in produce and blood. Int. J. Nanomed. 2012, 7, 2591–2600. [Google Scholar] [CrossRef][Green Version]
- Hammes, W.P.; Neuhaus, F.C. On the mechanism of action of vancomycin: Inhibition of peptidoglycan synthesis in Gaffkya homari. Antimicrob. Agents Chemother. 1974, 6, 722–728. [Google Scholar] [CrossRef]
- Alborn, W.E., Jr.; Allen, N.E.; Preston, D.A. Daptomycin disrupts membrane potential in growing Staphylococcus aureus. Antimicrob. Agents Chemother. 1991, 35, 2282–2287. [Google Scholar] [CrossRef]
- Lin, Y.S.; Tsai, P.J.; Weng, M.F.; Chen, Y.C. Affinity capture using vancomycin-bound magnetic nanoparticles for the MALDI-MS analysis of bacteria. Anal. Chem. 2005, 77, 1753–1760. [Google Scholar] [CrossRef]
- Wang, M.; Fan, E.; Wu, Y.; Fu, Z. Daptomycin-modified magnetic beads integrated with lysostaphin for selective analysis of Staphylococcus. J. Pharm. Biomed. Anal. 2019, 175, 112785. [Google Scholar] [CrossRef]
- Wadstrom, T.; Ljungh, A. Glycosaminoglycan-binding microbial proteins in tissue adhesion and invasion: Key events in microbial pathogenicity. J. Med. Microbiol. 1999, 48, 223–233. [Google Scholar] [CrossRef] [PubMed]
- Mattsby-Baltzer, I.; Bergstrom, T.; McCrea, K.; Ward, R.; Adolfsson, L.; Larm, O. Affinity apheresis for treatment of bacteremia caused by Staphylococcus aureus and/or methicillin-resistant S. aureus (MRSA). J. Microbiol. Biotechnol. 2011, 21, 659–664. [Google Scholar] [CrossRef] [PubMed]
- Weis, W.I.; Drickamer, K.; Hendrickson, W.A. Structure of a C-type mannose-binding protein complexed with an oligosaccharide. Nature 1992, 360, 127–134. [Google Scholar] [CrossRef]
- Jung, S.H.; Hahn, Y.K.; Oh, S.; Kwon, S.; Um, E.; Choi, S.; Kang, J.H. Advection Flows-Enhanced Magnetic Separation for High-Throughput Bacteria Separation from Undiluted Whole Blood. Small 2018, 14, e1801731. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Torab, P.; Wong, P.K. Detection of bacterial infection via a fidget spinner. Nat. Biomed. Eng. 2020, 4, 577–578. [Google Scholar] [CrossRef] [PubMed]
- Martel, J.M.; Smith, K.C.; Dlamini, M.; Pletcher, K.; Yang, J.; Karabacak, M.; Haber, D.A.; Kapur, R.; Toner, M. Continuous Flow Microfluidic Bioparticle Concentrator. Sci. Rep. 2015, 5, 11300. [Google Scholar] [CrossRef]
- Ferroni, A.; Suarez, S.; Beretti, J.L.; Dauphin, B.; Bille, E.; Meyer, J.; Bougnoux, M.E.; Alanio, A.; Berche, P.; Nassif, X. Real-time identification of bacteria and Candida species in positive blood culture broths by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J. Clin. Microbiol. 2010, 48, 1542–1548. [Google Scholar] [CrossRef]
- Prod’hom, G.; Bizzini, A.; Durussel, C.; Bille, J.; Greub, G. Matrix-assisted laser desorption ionization-time of flight mass spectrometry for direct bacterial identification from positive blood culture pellets. J. Clin. Microbiol. 2010, 48, 1481–1483. [Google Scholar] [CrossRef]
- Saffert, R.T.; Cunningham, S.A.; Mandrekar, J.; Patel, R. Comparison of three preparatory methods for detection of bacteremia by MALDI-TOF mass spectrometry. Diagn. Microbiol. Infect. Dis. 2012, 73, 21–26. [Google Scholar] [CrossRef]
- Hariu, M.; Watanabe, Y.; Oikawa, N.; Manaka, T.; Seki, M. Evaluation of blood culture broths with lysis buffer to directly identify specific pathogens by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry methods. Infect. Drug Resist. 2018, 11, 1573–1579. [Google Scholar] [CrossRef]
- Kok, J.; Thomas, L.C.; Olma, T.; Chen, S.C.; Iredell, J.R. Identification of bacteria in blood culture broths using matrix-assisted laser desorption-ionization Sepsityper and time of flight mass spectrometry. PLoS ONE 2011, 6, e23285. [Google Scholar] [CrossRef] [PubMed]
- Meex, C.; Neuville, F.; Descy, J.; Huynen, P.; Hayette, M.P.; De Mol, P.; Melin, P. Direct identification of bacteria from BacT/ALERT anaerobic positive blood cultures by MALDI-TOF MS: MALDI Sepsityper kit versus an in-house saponin method for bacterial extraction. J. Med. Microbiol. 2012, 61, 1511–1516. [Google Scholar] [CrossRef]
- Tanner, H.; Evans, J.T.; Gossain, S.; Hussain, A. Evaluation of three sample preparation methods for the direct identification of bacteria in positive blood cultures by MALDI-TOF. BMC Res. Notes 2017, 10, 48. [Google Scholar] [CrossRef] [PubMed]
- Quach, A.; Ferrante, A. The Application of Dextran Sedimentation as an Initial Step in Neutrophil Purification Promotes Their Stimulation, due to the Presence of Monocytes. J. Immunol. Res. 2017, 2017, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Tanavde, V.M.; Desai, S.S.; Rao, S.G.A. Erythrocyte depletion of human umbilical cord blood using dextran sedimentation. Indian J. Med. Res. 1997, 106, 16–19. [Google Scholar]
- Wu, M.; Ozcelik, A.; Rufo, J.; Wang, Z.; Fang, R.; Jun Huang, T. Acoustofluidic separation of cells and particles. Microsyst Nanoeng 2019, 5, 32. [Google Scholar] [CrossRef]
- Bruus, H. Acoustofluidics 10: Scaling laws in acoustophoresis. Lab Chip 2012, 12, 1578–1586. [Google Scholar] [CrossRef]
- Wu, M.; Ouyang, Y.; Wang, Z.; Zhang, R.; Huang, P.H.; Chen, C.; Li, H.; Li, P.; Quinn, D.; Dao, M.; et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc. Natl. Acad. Sci. USA 2017, 114, 10584–10589. [Google Scholar] [CrossRef]
- Wu, M.; Huang, P.H.; Zhang, R.; Mao, Z.; Chen, C.; Kemeny, G.; Li, P.; Lee, A.V.; Gyanchandani, R.; Armstrong, A.J.; et al. Circulating Tumor Cell Phenotyping via High-Throughput Acoustic Separation. Small 2018, 14, e1801131. [Google Scholar] [CrossRef]
- Ohlsson, P.; Petersson, K.; Augustsson, P.; Laurell, T. Acoustic impedance matched buffers enable separation of bacteria from blood cells at high cell concentrations. Sci. Rep. 2018, 8, 9156. [Google Scholar] [CrossRef]
- Lu, Y.; Liu, T.; Lamanda, A.C.; Sin, M.L.; Gau, V.; Liao, J.C.; Wong, P.K. AC Electrokinetics of Physiological Fluids for Biomedical Applications. J. Lab. Autom. 2015, 20, 611–620. [Google Scholar] [CrossRef] [PubMed]
- Wong, P.K.; Wang, T.H.; Deval, J.H.; Ho, C.M. Electrokinetics in micro devices for biotechnology applications. IEEE-ASME Trans. Mechatron. 2004, 9, 366–376. [Google Scholar] [CrossRef]
- Pohl, H.A.; Hawk, I. Separation of Living and Dead Cells by Dielectrophoresis. Science 1966, 152, 647. [Google Scholar] [CrossRef]
- Ramos, A.; Morgan, H.; Green, N.G.; Castellanos, A. Ac electrokinetics: A review of forces in microelectrode structures. J. Phys. D-Appl. Phys. 1998, 31, 2338–2353. [Google Scholar] [CrossRef]
- Lu, Y.; Ren, Q.; Liu, T.; Leung, S.L.; Gau, V.; Liao, J.C.; Chan, C.L.; Wong, P.K. Long-range electrothermal fluid motion in microfluidic systems. Int. J. Heath Mass Transf. 2016, 98, 341–349. [Google Scholar] [CrossRef]
- Liu, T.; Lu, Y.; Gau, V.; Liao, J.C.; Wong, P.K. Rapid antimicrobial susceptibility testing with electrokinetics enhanced biosensors for diagnosis of acute bacterial infections. Ann. Biomed. Eng. 2014, 42, 2314–2321. [Google Scholar] [CrossRef]
- Gao, J.; Jeffries, L.; Mach, K.E.; Craft, D.W.; Thomas, N.J.; Gau, V.; Liao, J.C.; Wong, P.K. A Multiplex Electrochemical Biosensor for Bloodstream Infection Diagnosis. SLAS Technol. 2017, 22, 466–474. [Google Scholar] [CrossRef]
- Huge, B.J.; Champion, M.M.; Dovichi, N.J. Capillary Zone Electrophoresis with Fraction Collection for Separation, Culturing, and Identification of Bacteria from an Environmental Microbiome. Anal. Chem. 2019, 91, 4649–4655. [Google Scholar] [CrossRef]
- Jiang, X.; Liu, S.; Zhang, Y.; Ji, Y.; Sohail, A.; Cao, C.; Wang, P.; Xiao, H. Free-Flow Isoelectric Focusing for Comprehensive Separation and Analysis of Human Salivary Microbiome for Lung Cancer. Anal. Chem. 2020, 92, 12017–12025. [Google Scholar] [CrossRef]
- Di Carlo, D. Inertial microfluidics. Lab Chip 2009, 9, 3038–3046. [Google Scholar] [CrossRef]
- Di Carlo, D.; Irimia, D.; Tompkins, R.G.; Toner, M. Continuous inertial focusing, ordering, and separation of particles in microchannels. Proc. Natl. Acad. Sci. USA 2007, 104, 18892–18897. [Google Scholar] [CrossRef] [PubMed]
- Martel, J.M.; Toner, M. Inertial focusing in microfluidics. Annu. Rev. Biomed. Eng. 2014, 16, 371–396. [Google Scholar] [CrossRef]
- Zhang, J.; Yan, S.; Yuan, D.; Alici, G.; Nguyen, N.T.; Ebrahimi Warkiani, M.; Li, W. Fundamentals and applications of inertial microfluidics: A review. Lab Chip 2016, 16, 10–34. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Graham, M.D. Margination and segregation in confined flows of blood and other multicomponent suspensions. Soft Matter 2012, 8, 10536–10548. [Google Scholar] [CrossRef]
- Carboni, E.; Tschudi, K.; Nam, J.; Lu, X.; Ma, A.W.K. Particle margination and its implications on intravenous anticancer drug delivery. AAPS PharmSciTech 2014, 15, 762–771. [Google Scholar] [CrossRef]
- Lee, A.; Mirrett, S.; Reller, L.B.; Weinstein, M.P. Detection of bloodstream infections in adults: How many blood cultures are needed? J. Clin. Microbiol. 2007, 45, 3546–3548. [Google Scholar] [CrossRef] [PubMed]
- Henning, C.; Aygül, N.; Dinnétz, P.; Wallgren, K.; Özenci, V. Detailed analysis of the characteristics of sample volume in blood culture bottles. J. Clin. Microbiol. 2019, 57, 1–9. [Google Scholar] [CrossRef]
- Peretz, A.; Isakovich, N.; Pastukh, N.; Koifman, A.; Glyatman, T.; Brodsky, D. Performance of Gram staining on blood cultures flagged negative by an automated blood culture system. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 1539–1541. [Google Scholar] [CrossRef]
- Dubourg, G.; Raoult, D. Emerging methodologies for pathogen identification in positive blood culture testing. Expert Rev. Mol. Diagn. 2016, 16, 97–111. [Google Scholar] [CrossRef]
- Gajdács, M.; Baráth, Z.; Kárpáti, K.; Szabó, D.; Usai, D.; Zanetti, S.; Donadu, M.G. No correlation between biofilm formation, virulence factors, and antibiotic resistance in pseudomonas aeruginosa: Results from a laboratory-based in vitro study. Antibiotics 2021, 10, 1134. [Google Scholar] [CrossRef]
- Di Domenico, E.G.; Marchesi, F.; Cavallo, I.; Toma, L.; Sivori, F.; Papa, E.; Spadea, A.; Cafarella, G.; Terrenato, I.; Prignano, G.; et al. The Impact of Bacterial Biofilms on End-Organ Disease and Mortality in Patients with Hematologic Malignancies Developing a Bloodstream Infection. Microbiol. Spectr. 2021, 9, e0055021. [Google Scholar] [CrossRef] [PubMed]
- Heidari, R.; Sheikh, A.F.; Hashemzadeh, M.; Farshadzadeh, Z.; Salmanzadeh, S.; Saki, M. Antibiotic resistance, biofilm production ability and genetic diversity of carbapenem-resistant Pseudomonas aeruginosa strains isolated from nosocomial infections in southwestern Iran. Mol. Biol. Rep. 2022, 1, 3. [Google Scholar] [CrossRef] [PubMed]
- Devonshire, A.S.; Honeyborne, I.; Gutteridge, A.; Whale, A.S.; Nixon, G.; Wilson, P.; Jones, G.; McHugh, T.D.; Foy, C.A.; Huggett, J.F. Highly reproducible absolute quantification of mycobacterium tuberculosis complex by digital PCR. Anal. Chem. 2015, 87, 3706–3713. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Bai, R.; Zhao, Z.; Tao, L.; Ma, M.; Ji, Z.; Jian, M.; Ding, Z.; Dai, X.; Bao, F.; et al. Application of droplet digital PCR to detect the pathogens of infectious diseases. Biosci. Rep. 2018, 38, BSR20181170. [Google Scholar] [CrossRef]
- Kang, D.K.; Ali, M.M.; Zhang, K.; Huang, S.S.; Peterson, E.; Digman, M.A.; Gratton, E.; Zhao, W. Rapid detection of single bacteria in unprocessed blood using Integrated Comprehensive Droplet Digital Detection. Nat. Commun. 2014, 5, 5427. [Google Scholar] [CrossRef] [PubMed]
- Wouters, Y.; Dalloyaux, D.; Christenhusz, A.; Roelofs, H.M.J.; Wertheim, H.F.; Bleeker-Rovers, C.P.; te Morsche, R.H.; Wanten, G.J.A. Droplet digital polymerase chain reaction for rapid broad-spectrum detection of bloodstream infections. Microb. Biotechnol. 2020, 13, 657–668. [Google Scholar] [CrossRef]
- Abram, T.J.; Cherukury, H.; Ou, C.-Y.; Vu, T.; Toledano, M.; Li, Y.; Grunwald, J.T.; Toosky, M.N.; Tifrea, D.F.; Slepenkin, A.; et al. Rapid bacterial detection and antibiotic susceptibility testing in whole blood using one-step, high throughput blood digital PCR. Lab Chip 2020, 20, 477–489. [Google Scholar] [CrossRef]
- Montgomery, J.L.; Sanford, L.N.; Wittwer, C.T. High-resolution DNA melting analysis in clinical research and diagnostics. Expert Rev. Mol. Diagn. 2010, 10, 219–240. [Google Scholar] [CrossRef]
- Wittwer, C.T.; Reed, G.H.; Gundry, C.N.; Vandersteen, J.G.; Pryor, R.J. High-resolution genotyping by amplicon melting analysis using LCGreen. Clin. Chem. 2003, 49, 853–860. [Google Scholar] [CrossRef]
- Yang, S.; Ramachandran, P.; Rothman, R.; Hsieh, Y.H.; Hardick, A.; Won, H.; Kecojevic, A.; Jackman, J.; Gaydos, C. Rapid identification of biothreat and other clinically relevant bacterial species by use of universal PCR coupled with high-resolution melting analysis. J. Clin. Microbiol. 2009, 47, 2252–2255. [Google Scholar] [CrossRef]
- Rothman, R.; Ramachandran, P.; Yang, S.; Hardick, A.; Won, H.; Kecojevic, A.; Quianzon, C.; Hsieh, Y.-H.; Gaydos, C. Use of quantitative broad-based polymerase chain reaction for detection and identification of common bacterial pathogens in cerebrospinal fluid. Acad. Emerg. Med. 2010, 17, 741–747. [Google Scholar] [CrossRef] [PubMed]
- Won, H.; Rothman, R.; Ramachandran, P.; Hsieh, Y.H.; Kecojevic, A.; Carroll, K.C.; Aird, D.; Gaydos, C.; Yang, S. Rapid identification of bacterial pathogens in positive blood culture bottles by use of a broad-based PCR assay coupled with high-resolution melt analysis. J. Clin. Microbiol. 2010, 48, 3410–3413. [Google Scholar] [CrossRef] [PubMed]
- Hardick, J.; Won, H.; Jeng, K.; Hsieh, Y.H.; Gaydos, C.A.; Rothman, R.E.; Yang, S. Identification of bacterial pathogens in ascitic fluids from patients with suspected spontaneous bacterial peritonitis by use of broad- range PCR (16S PCR) coupled with high-resolution melt analysis. J. Clin. Microbiol. 2012, 50, 2428–2432. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, Y.; Park, S.; Yang, S.; Wang, T.-H. An all-in-one microfluidic device for parallel DNA extraction and gene analysis. Biomed. Microdevices 2010, 12, 1043–1049. [Google Scholar] [CrossRef] [PubMed]
- Jeng, K.; Yang, S.; Won, H.; Gaydos, C.A.; Hsieh, Y.H.; Kecojevic, A.; Carroll, K.C.; Hardick, J.; Rothmana, R.E. Application of a 16S rRNA PCR-high-resolution melt analysis assay for rapid detection of Salmonella bacteremia. J. Clin. Microbiol. 2012, 50, 1122–1124. [Google Scholar] [CrossRef]
- Masek, B.J.; Hardick, J.; Won, H.; Yang, S.; Hsieh, Y.H.; Rothman, R.E.; Gaydos, C.A. Sensitive detection and serovar differentiation of typhoidal and nontyphoidal salmonella enterica species using 16S rRNA gene PCR coupled with high-resolution melt analysis. J. Mol. Diagn. 2014, 16, 261–266. [Google Scholar] [CrossRef]
- Fraley, S.I.; Hardick, J.; Jo Masek, B.; Athamanolap, P.; Rothman, R.E.; Gaydos, C.A.; Carroll, K.C.; Wakefield, T.; Wang, T.H.; Yang, S. Universal digital high-resolution melt: A novel approach to broad-based profiling of heterogeneous biological samples. Nucleic Acids Res. 2013, 41, e175. [Google Scholar] [CrossRef]
- Athamanolap, P.; Parekh, V.; Fraley, S.I.; Agarwal, V.; Shin, D.J.; Jacobs, M.A.; Wang, T.H.; Yang, S. Trainable high resolution melt curve machine learning classifier for large-scale reliable genotyping of sequence variants. PLoS ONE 2014, 9, e109094. [Google Scholar] [CrossRef]
- Fraley, S.I.; Athamanolap, P.; Masek, B.J.; Hardick, J.; Carroll, K.C.; Hsieh, Y.H.; Rothman, R.E.; Gaydos, C.A.; Wang, T.H.; Yang, S. Nested machine learning facilitates increased sequence content for large-scale automated high resolution melt genotyping. Sci. Rep. 2016, 6, 19218. [Google Scholar] [CrossRef]
- Athamanolap, P.; Hsieh, K.; O’Keefe, C.M.; Zhang, Y.; Yang, S.; Wang, T.H. Nanoarray digital PCR with high-resolution melt enables broad bacteria identification and pheno-molecular antimicrobial susceptibility test. Anal. Chem. 2019, 91, 12784–12792. [Google Scholar] [CrossRef]
- Chen, L.; Shin, D.J.; Zheng, S.; Melendez, J.H.; Gaydos, C.A.; Wang, T.H. Direct-qPCR assay for coupled identification and antimicrobial susceptibility testing of Neisseria gonorrhoeae. ACS Infect. Dis. 2018, 4, 1377–1384. [Google Scholar] [CrossRef] [PubMed]
- Schoepp, N.G.; Schlappi, T.S.; Curtis, M.S.; Butkovich, S.S.; Miller, S.; Humphries, R.M.; Ismagilov, R.F. Rapid pathogen-specific phenotypic antibiotic susceptibility testing using digital LAMP quantification in clinical samples. Sci. Transl. Med. 2017, 9, eaal3693. [Google Scholar] [CrossRef] [PubMed]
- Schoepp, N.G.; Khorosheva, E.M.; Schlappi, T.S.; Curtis, M.S.; Humphries, R.M.; Hindler, J.A.; Ismagilov, R.F. Digital quantification of DNA replication and chromosome segregation enables determination of antimicrobial susceptibility after only 15 minutes of antibiotic exposure. Angew. Chem. Int. Ed. 2016, 55, 9557–9561. [Google Scholar] [CrossRef] [PubMed]
- Andini, N.; Hu, A.; Zhou, L.; Cogill, S.; Wang, T.H.; Wittwer, C.T.; Yang, S. A “culture” shift: Broad bacterial detection, identification, and antimicrobial susceptibility testing directly from whole blood. Clin. Chem. 2018, 64, 1453–1462. [Google Scholar] [CrossRef] [PubMed]
- Athamanolap, P.; Hsieh, K.; Chen, L.; Yang, S.; Wang, T.H. Integrated bacterial identification and antimicrobial susceptibility testing using PCR and high-resolution melt. Anal. Chem. 2017, 89, 11529–11536. [Google Scholar] [CrossRef]
- Zhang, Y.; Hu, A.; Andini, N.; Yang, S. A ‘culture’ shift: Application of molecular techniques for diagnosing polymicrobial infections. Biotechnol. Adv. 2019, 37, 476–490. [Google Scholar] [CrossRef]
- Yang, X.; Hashemi, M.M.; Andini, N.; Li, M.M.; Kuang, S.; Carroll, K.C.; Wang, T.H.; Yang, S. RNA markers for ultra-rapid molecular antimicrobial susceptibility testing in fluoroquinolone-treated Klebsiella pneumoniae. J. Antimicrob. Chemother. 2020, 75, 1747–1755. [Google Scholar] [CrossRef]
- Gupta, A.; Mishra, A.; Puri, N. Peptide nucleic acids: Advanced tools for biomedical applications. J. Biotechnol. 2017, 259, 148–159. [Google Scholar] [CrossRef]
- Sacui, I.; Hsieh, W.C.; Manna, A.; Sahu, B.; Ly, D.H. Gamma peptide nucleic acids: As orthogonal nucleic acid recognition codes for organizing molecular self-assembly. J. Am. Chem. Soc. 2015, 137, 8603–8610. [Google Scholar] [CrossRef]
- Mach, K.E.; Kaushik, A.M.; Hsieh, K.; Wong, P.K.; Wang, T.H.; Liao, J.C. Optimizing peptide nucleic acid probes for hybridization-based detection and identification of bacterial pathogens. Analyst 2019, 144, 1565–1574. [Google Scholar] [CrossRef]
- Deck, M.K.; Anderson, E.S.; Buckner, R.J.; Colasante, G.; Coull, J.M.; Crystal, B.; Latta, P.D.; Fuchs, M.; Fuller, D.; Harris, W.; et al. Multicenter evaluation of the Staphylococcus QuickFISH method for simultaneous identification of Staphylococcus aureus and coagulase-negative staphylococci directly from blood culture bottles in less than 30 minutes. J. Clin. Microbiol. 2012, 50, 1994–1998. [Google Scholar] [CrossRef] [PubMed]
- Charnot-Katsikas, A.; Tesic, V.; Love, N.; Hill, B.; Bethel, C.; Boonlayangoor, S.; Beavis, K.G. Use of the Accelerate Pheno system for identification and antimicrobial susceptibility testing of pathogens in positive blood cultures and impact on time to results and workflow. J. Clin. Microbiol. 2017, 56, e01166-17. [Google Scholar] [CrossRef] [PubMed]
- Nölling, J.; Rapireddy, S.; Amburg, J.I.; Crawford, E.M.; Prakash, R.A.; Rabson, A.R.; Tang, Y.W.; Singer, A. Duplex DNA-invading γ-modified peptide nucleic acids enable rapid identification of bloodstream infections in whole blood. mBio 2016, 7, e00345-16. [Google Scholar] [CrossRef] [PubMed]
- Duan, H.; Li, X.; Mei, A.; Li, P.; Liu, Y.; Li, X.; Li, W.; Wang, C.; Xie, S. The diagnostic value of metagenomic next-generation sequencing in infectious diseases. BMC Infect. Dis. 2021, 21, 62. [Google Scholar] [CrossRef] [PubMed]
- Blauwkamp, T.A.; Thair, S.; Rosen, M.J.; Blair, L.; Lindner, M.S.; Vilfan, I.D.; Kawli, T.; Christians, F.C.; Venkatasubrahmanyam, S.; Wall, G.D.; et al. Analytical and clinical validation of a microbial cell-free DNA sequencing test for infectious disease. Nat. Microbiol. 2019, 4, 663–674. [Google Scholar] [CrossRef] [PubMed]
- Horiba, K.; Kawada, J.; Okuno, Y.; Tetsuka, N.; Suzuki, T.; Ando, S.; Kamiya, Y.; Torii, Y.; Yagi, T.; Takahashi, Y.; et al. Comprehensive detection of pathogens in immunocompromised children with bloodstream infections by next-generation sequencing. Sci. Rep. 2018, 8, 3784. [Google Scholar] [CrossRef]
- Kjaergaard Nielsen, T.; Denis Browne, P.; Hansen, L.H. Mobilization of antibiotic resistance genes differ by resistance mechanism. bioRxiv 2021. [Google Scholar] [CrossRef]
- Eyre, D.W.; De Silva, D.; Cole, K.; Peters, J.; Cole, M.J.; Grad, Y.H.; Demczuk, W.; Martin, I.; Mulvey, M.R.; Crook, D.W.; et al. WGS to predict antibiotic MICs for Neisseria gonorrhoeae. J. Antimicrob. Chemother. 2017, 72, 1937–1947. [Google Scholar] [CrossRef]
- Mensah, N.; Tang, Y.; Cawthraw, S.; AbuOun, M.; Fenner, J.; Thomson, N.R.; Mather, A.E.; Petrovska-Holmes, L. Determining antimicrobial susceptibility in Salmonella enterica serovar Typhimurium through whole genome sequencing: A comparison against multiple phenotypic susceptibility testing methods. BMC Microbiol. 2019, 19, 148. [Google Scholar] [CrossRef]
- Ma, K.C.; Mortimer, T.D.; Hicks, A.L.; Wheeler, N.E.; Sánchez-Busó, L.; Golparian, D.; Taiaroa, G.; Rubin, D.H.F.; Wang, Y.; Williamson, D.A.; et al. Adaptation to the cervical environment is associated with increased antibiotic susceptibility in Neisseria gonorrhoeae. Nat. Commun. 2020, 11, 4126. [Google Scholar] [CrossRef]
- Demczuk, W.; Martin, I.; Sawatzky, P.; Allen, V.; Lefebvre, B.; Hoang, L.; Naidu, P.; Minion, J.; VanCaeseele, P.; Haldane, D.; et al. Equations to predict antimicrobial MICs in Neisseria gonorrhoeae using molecular antimicrobial resistance determinants. Antimicrob. Agents Chemother. 2019, 64, e02005-19. [Google Scholar] [CrossRef] [PubMed]
- Ram-Mohan, N.; Thair, S.A.; Litzenburger, U.M.; Cogill, S.; Andini, N.; Yang, X.; Chang, H.Y.; Yang, S. Profiling chromatin accessibility responses in human neutrophils with sensitive pathogen detection. Life Sci. Alliance 2021, 4, e202000976. [Google Scholar] [CrossRef] [PubMed]
- Tadesse, L.F.; Safir, F.; Ho, C.-S.; Hasbach, X.; Khuri-Yakub, B.P.; Jeffrey, S.S.; Saleh, A.A.E.; Dionne, J. Toward rapid infectious disease diagnosis with advances in surface-enhanced Raman spectroscopy. J. Chem. Phys. 2020, 152, 240902. [Google Scholar] [CrossRef] [PubMed]
- Stöckel, S.; Kirchhoff, J.; Neugebauer, U.; Rösch, P.; Popp, J. The application of Raman spectroscopy for the detection and identification of microorganisms. J. Raman Spectrosc. 2016, 47, 89–109. [Google Scholar] [CrossRef]
- Han, Y.-Y.; Lin, Y.-C.; Cheng, W.-C.; Lin, Y.-T.; Teng, L.-J.; Wang, J.-K.; Wang, Y.-L. Rapid antibiotic susceptibility testing of bacteria from patients’ blood via assaying bacterial metabolic response with surface-enhanced Raman spectroscopy. Sci. Rep. 2020, 10, 12538. [Google Scholar] [CrossRef]
- Cheng, I.F.; Chang, H.-C.; Chen, T.-Y.; Hu, C.; Yang, F.-L. Rapid (<5 min) identification of pathogen in human blood by electrokinetic concentration and surface-enhanced Raman spectroscopy. Sci. Rep. 2013, 3, 2365. [Google Scholar] [CrossRef]
- Ho, C.S.; Jean, N.; Hogan, C.A.; Blackmon, L.; Jeffrey, S.S.; Holodniy, M.; Banaei, N.; Saleh, A.A.E.; Ermon, S.; Dionne, J. Rapid identification of pathogenic bacteria using Raman spectroscopy and deep learning. Nat. Commun. 2019, 10, 4927. [Google Scholar] [CrossRef]
- Novelli-Rousseau, A.; Espagnon, I.; Filiputti, D.; Gal, O.; Douet, A.; Mallard, F.; Josso, Q. Culture-free Antibiotic-susceptibility Determination From Single-bacterium Raman Spectra. Sci. Rep. 2018, 8, 3957. [Google Scholar] [CrossRef]
- Ambriz-Aviña, V.; Contreras-Garduño, J.A.; Pedraza-Reyes, M. Applications of flow cytometry to characterize bacterial physiological responses. Biomed. Res. Int. 2014, 2014, 461941. [Google Scholar] [CrossRef]
- Fonseca e Silva, D.; Andrade, F.F.; Gomes, R.; Silva-Dias, A.; Martins-Oliveira, I.; Pérez-Viso, B.; Ramos, M.H.; Rodrigues, A.G.; Cantón, R.; Pina-Vaz, C. Ultra-rapid flow cytometry assay for colistin MIC determination in Enterobacterales, Pseudomonas aeruginosa and Acinetobacter baumannii. Clin. Microbiol. Infect. 2020, 26, 1559.e1–1559.e4. [Google Scholar] [CrossRef]
- Costa-de-Oliveira, S.; Teixeira-Santos, R.; Silva, A.P.; Pinho, E.; Mergulhão, P.; Silva-Dias, A.; Marques, N.; Martins-Oliveira, I.; Rodrigues, A.G.; Paiva, J.A.; et al. Potential impact of flow cytometry antimicrobial susceptibility testing on the clinical management of gram-negative bacteremia using the FASTinov® kit. Front. Microbiol. 2017, 8, 2455. [Google Scholar] [CrossRef] [PubMed]
- Wiegand, I.; Hilpert, K.; Hancock, R.E.W. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef] [PubMed]
- El-Halfawy, O.M.; Valvano, M.A. Antimicrobial heteroresistance: An emerging field in need of clarity. Clin. Microbiol. Rev. 2015, 28, 191–207. [Google Scholar] [CrossRef] [PubMed]
- Nicoloff, H.; Hjort, K.; Levin, B.R.; Andersson, D.I. The high prevalence of antibiotic heteroresistance in pathogenic bacteria is mainly caused by gene amplification. Nat. Microbiol. 2019, 4, 504–514. [Google Scholar] [CrossRef]
- Li, X.Z.; Plésiat, P.; Nikaido, H. The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin. Microbiol. Rev. 2015, 28, 337–418. [Google Scholar] [CrossRef]
- Sandegren, L.; Andersson, D.I. Bacterial gene amplification: Implications for the evolution of antibiotic resistance. Nat. Rev. Microbiol. 2009, 7, 578–588. [Google Scholar] [CrossRef]
- Abe, R.; Akeda, Y.; Sugawara, Y.; Matsumoto, Y.; Motooka, D.; Kawahara, R.; Yamamoto, N.; Tomono, K.; Iida, T.; Hamada, S. Enhanced Carbapenem Resistance through Multimerization of Plasmids Carrying Carbapenemase Genes. mBio 2021, 12, e0018621. [Google Scholar] [CrossRef]

| No. | Company | System | Approach | Status * | Sample Prep ‡ | Detection/ ID | AST | TAT ^ |
|---|---|---|---|---|---|---|---|---|
| 1 | Abacus Diagnostica | Genomera CDX | Rapid/ Real-Time PCR | Dev. | BC (+) | ✓ | 50 min | |
| 2 | Affinity Biosensors | LifeScale AST | Microorganism mass measurement | CE-IVD | BC (+) | ✓ | 4 h | |
| 3 | Amplex Diagnostics, GmbH, Germany | Eazyplex MRSA | LAMP ultra-rapid MRSA detection | CE-IVD | BC (+) | ✓ | 30 min | |
| 4 | Arc Bio | Galileo pathogen solution | Shotgun Sequencing | Dev. | WB | ✓ | 48 h | |
| 5 | BD | GeneOhm MRSA | Real-Time PCR | FDA, CE-IVD | BC (+) | ✓ | 2 h | |
| 6 | Becton Dickinson | BD Max StaphSR | Real-Time PCR | FDA | BC (+) | ✓ | ~1.5 h | |
| 7 | BioFire/bioMerieux Diagnostics | FilmArray DIRECT (new) | Nested PCR | FDA, CE-IVD | WB | ✓ | 1 h | |
| 8 | BioRad | Droplet dPCR | dPCR; absolute quantification using Poisson’s statistics without requiring a standard curve | CE-IVD | BC (+) | ✓ | No report | |
| 9 | BioSense Solutions (Denmark) | oCelloScope | 3D optical scanning microscopy imaging | Dev. | BC (+) | ✓ | 1 to 4 h | |
| 10 | Bruker Daltonics | MALDI Biotyper + DxM MicroScan WalkAway System | Mass spectrometry | FDA, CE-IVD | BC (+) | ✓ | 12 to 24 h | |
| 11 | DNAe (electronic) | LiDia Bloodstream Infection Test | WGS/NGS/miniaturised sequencing | Dev. | WB | ✓ | 3 to 4 h | |
| 12 | FASTinov | Flow cytometry | Cell sorting fluorescence-based AST | Dev. | BC (+) | ✓ | <2 h | |
| 14 | Roche | Smarticles | Bacteriophage-based | Dev. | BC (+) | ✓ | No report | |
| 15 | GenMarkDx USA | ePlex BCID | Multiplex PCR | CE-IVD | BC (+) | ✓ | 1.5 h | |
| 16 | Gradientech AB | Rapid IVD; QuickMIC and CellDirector | Microfluidics Phenotypic multiplex chip | Dev. | BC (+) | ✓ | 2 h | |
| 17 | Great Basin Corporation (Bringham Young Univ.) | OptoFluidic Platform | Single molecule fluorescence hybridization | Dev. | WB | ✓ | 1 h | |
| 18 | Hologic | AccuProbe | In situ hybridization | CE-IVD | BC (+) | ✓ | 1 h | |
| 19 | iCubate | iC GPC | Multiplex amplification assay | FDA, CE-IVD | BC (+) | ✓ | 4 to 5 h | |
| 20 | IRIDICA | BAC BSI Assay | PCR/ESI-MS | withdrawn | WB | ✓ | 8 h | |
| 21 | Karius, Inc. | Karius Test | NextGen Seq cfDNA; Genomic; Bioinformatics | Dev. | WB | ✓ | 48 h | |
| 22 | Luminex | Verigene Gram+ BC | Microarray | FDA. | BC (+) | ✓ | 2.5 h | |
| 23 | Luminex | Verigene Gram− BC | Microarray | FDA | BC (+) | ✓ | 2.5 h | |
| 24 | Master Diagnostica, Spain | Sepsis Flow Chip | Microarray | CE-IVD | BC (+) | ✓ | 3 to 4 h | |
| 25 | Molzym, Germany | SeptiTest; UMD SelectNA | Real Time PCR | CE-IVD | WB | ✓ | 8 to 12 h | |
| 26 | Momentum Biosciences (Cardiff, UK) | TBD Cognitor Minus | Enzymatic template generation and amplification | awaiting clearance | BC (+) | ✓ | No report | |
| 27 | OpGen USA | PNA FISH | In situ hybridization | CE-IVD | BC (+) | ✓ | 2.5 h | |
| 28 | OpGen USA | Quick FISH | In situ hybridization | CE-IVD | BC (+) | ✓ | 30 min | |
| 29 | QLinea (Uppsala, Sweden) | AsTAR | High-speed time-lapse microscopy imaging of bacteria in broth | Dev. | BC (+) | ✓ | 6 h | |
| 30 | Resistell (Switzerland) | Rapid AST antibiogram | AFM, Cantilever, Nanomotion detection-based AST | unknown | BC (+) | ✓ | No report | |
| 31 | Roche Molecular System, Switzerland | LightCycler SeptiFast | Real-Time PCR | CE-IVD | WB | ✓ | 6 h | |
| 32 | SeeGene, Korea | Magicplex Sepsis RT test | Real-Time PCR | CE-IVD | WB | ✓ | 3 to 6 h | |
| 33 | Specific Diagnostics Inc | Reveal phenotypic AST | Detection of volatile organic compounds | Dev. | BC (+) | ✓ | ~5 h (with MIC) | |
| 34 | T2Biosystem | T2 Candida Panel T2MR | Nuclear Magnetic Resonance | FDA, CE-IVD | WB | ✓ | 3 to 5 h | |
| 35 | QuantaMatrix | QMAC-dRAST | Optical Microscopy | Dev. | BC (+) | ✓ | 4 to 6 h |
| Isolation Method | Recovery Rate # | Throughput ## | Pathogens | Sample Type | Conc. (cell/mL) | Ref |
|---|---|---|---|---|---|---|
| Affinity capture (magnetic; antibody) | 78% | 0.025 mL/h | E. coli | Red blood cells | 5 × 106 | [26] |
| Affinity capture (magnetic; antibody) | 80% | 20 mL/h | C. albicans | Whole blood | 1 × 106 | [27] |
| Affinity capture (magnetic; lectin) | 60–90% | 10 mL/h | S. aureus, C. albicans, E. coli | Whole blood | 1 × 104 | [28] |
| Filtration Affinity capture (magnetic; MBL) | 68–76% | 1.2 mL/h | E. coli, P. aeruginosa, K. pneumoniae, S. saprophyticus, S. epidermidis | Whole blood | 101–102 | [29] |
| 56–77% | 3 mL/h | Filtered blood | ||||
| Affinity capture (magnetic; Zn-DPA) | >88% | 60 mL/h | E. coli | Whole blood | 5 × 106 | [30] |
| Erythrocyte depletion (detergent + water) | ~100% | 2.88 mL/h | E. coli, M. luteus | Whole blood | 1 × 107 | [31] |
| Erythrocyte depletion (lysis) | >90% | 20 mL/h | E. coli | Whole blood | 1 × 103 | [32] |
| Erythrocyte depletion (dextran sedimentation) | 50–60% | 20 mL/h | E. coli, E. faecalis, K. pneumoniae | Whole blood | 101–102 | [33] |
| Acoustophoresis | 95.65% | 0.03 mL/h | E. coli | PBMC | 3 × 106 | [34] |
| Acoustophoresis | 91% | 3 mL/h | P. putida, E. coli | Diluted blood | 5 × 105 | [35] |
| Acoustophoresis (GAF) | 79.77% | 0.72 mL/h | S. aureus, S. pneumoniae, E. coli | Blood lysates | 1 × 105 | [36] |
| Electrokinetics (DEP) | 30% | 0.035 mL/h | E. coli | Red blood cells | 1 × 106 | [37] |
| Electrokinetics (DEP) | 97% | 0.0009 mL/h | E. coli, S. epidermidis, and C. albicans | Diluted blood | 1 × 104 | [38] |
| Electrokinetics (DEP and ACEF) | 30–80% | 0.006–0.06 mL/h | E. coli, A. baumannii, B. globigii | Buffy coat | 1 × 105 | [39] |
| Inertial focusing | >60% | 12 mL/h | E. coli | Whole blood | 1 × 108 | [40] |
| Inertial focusing (Dean flow) | >65% | 0.6 mL/h | E. coli, S. aureus, P. aeruginosa, E. faecalis | Diluted blood | 1 × 101 | [41] |
| Elasto-inertial | 76% | 0.03 mL/h | E. coli | Whole blood | 1 × 106 | [42] |
| Elasto-inertial | >80% | 0.3–1.5 mL/h | E. coli, S. capitis | Diluted blood | 1 × 103 | [43] |
| Elasto-inertial | 60–80% | 0.3 mL/h | K. pneumoniae, S. pneumoniae | Diluted blood | 1 × 102 | [44] |
| Margination | 80–90% | 1 mL/h | E. coli and S. cerevisiae | Whole blood | 1 × 106 | [45] |
| Technologies | Sample | Company | ID | AST | ||||
|---|---|---|---|---|---|---|---|---|
| Sens. (CFU/mL) | Spec. | Breadth | TAT | Output | TAT | |||
| EMERGING | ||||||||
| qPCR-HRM | WB * | Non-commercial | 1 | 100% | 37 bacteria (expandable) | 8 h (with AST) | MIC | 8 h (with ID) |
| SERS | WB | Spectral Platforms | 1 | 94% | >30 pathogens | 20 min | S/R (enzyme-based) | unspecified |
| ddPCR/ IC3D | WB | Velox Bio | 10 | 100% | unspecified | 1–4 h (with AMR) | resistance marker | 1–4 h (with ID) |
| Flow Cytometry | BC (+) | FASTinov | N/A | N/A | N/A | N/A | MIC | <26 h |
| PNA-FISH | WB | HelixBind | <10 | 95% | 21 pathogens | 2.5 h (with AMR) | resistance marker | 2.5 h (with ID) |
| EXISTING | ||||||||
| PCR+T2MR | WB | T2 Biosystems | 1–10 | 91% | 5 candida species, ESKAPE organisms | 27–29 h (with AMR) | resistance marker | 27–29 h (with ID) |
| Multiplex PCR | WB | MagicPlex (SeeGene) | 30 | 66–92% | >90 pathogens with 27 pathogens at species level | 27–30 h (with AMR) | resistance marker | 27–30 h (with ID) |
| Real-time PCR+Sequencing | WB | SepsiTest (Molzym) | 10–40 | 86–100% | >1350 pathogens | 30–31 h | N/A | N/A |
| Multiplex PCR | BC (+) | BioFire (FilmArray) | 106 to 108 | 82–92% | 8 Gram+/11 Gram−/5 fungi | 25 h (with AMR) | resistance marker | 25 h (with ID) |
| DNA Microarray | BC (+) | Luminex (Verigene) | 10–100 | 84–99% | 8 Gram+/5 Gram− | 26.5 h (with AMR) | resistance marker | 26.5 h (with ID) |
| MALDI-TOF + AST cards | BC (+) | Biomerieux (VITEK 2) | 106 | 61–98 | 1316 pathogens | 30–36 h (with AST) | MIC | 30–36 h (with ID) |
| PNA FISH + morphokinetic cellular analysis | BC (+) | Accelerate Diagnostics (Accelerate Pheno) | 0.8 to 1.7 | 86–100 | 7 Gram+/8 Gram−/2 fungi | 32 h (with AST) | MIC | 32 h (with ID) |
| Traditional Blood Culture | WB | BD (BACTEC) | 1 | 100% | Broad | 30 h | MIC | 54 h (with ID) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tjandra, K.C.; Ram-Mohan, N.; Abe, R.; Hashemi, M.M.; Lee, J.-H.; Chin, S.M.; Roshardt, M.A.; Liao, J.C.; Wong, P.K.; Yang, S. Diagnosis of Bloodstream Infections: An Evolution of Technologies towards Accurate and Rapid Identification and Antibiotic Susceptibility Testing. Antibiotics 2022, 11, 511. https://doi.org/10.3390/antibiotics11040511
Tjandra KC, Ram-Mohan N, Abe R, Hashemi MM, Lee J-H, Chin SM, Roshardt MA, Liao JC, Wong PK, Yang S. Diagnosis of Bloodstream Infections: An Evolution of Technologies towards Accurate and Rapid Identification and Antibiotic Susceptibility Testing. Antibiotics. 2022; 11(4):511. https://doi.org/10.3390/antibiotics11040511
Chicago/Turabian StyleTjandra, Kristel C., Nikhil Ram-Mohan, Ryuichiro Abe, Marjan M. Hashemi, Jyong-Huei Lee, Siew Mei Chin, Manuel A. Roshardt, Joseph C. Liao, Pak Kin Wong, and Samuel Yang. 2022. "Diagnosis of Bloodstream Infections: An Evolution of Technologies towards Accurate and Rapid Identification and Antibiotic Susceptibility Testing" Antibiotics 11, no. 4: 511. https://doi.org/10.3390/antibiotics11040511
APA StyleTjandra, K. C., Ram-Mohan, N., Abe, R., Hashemi, M. M., Lee, J.-H., Chin, S. M., Roshardt, M. A., Liao, J. C., Wong, P. K., & Yang, S. (2022). Diagnosis of Bloodstream Infections: An Evolution of Technologies towards Accurate and Rapid Identification and Antibiotic Susceptibility Testing. Antibiotics, 11(4), 511. https://doi.org/10.3390/antibiotics11040511






