Identification of Three Novel PmGRI1 Genomic Resistance Islands and One Multidrug Resistant Hybrid Structure of Tn7-like Transposon and PmGRI1 in Proteus mirabilis
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Antimicrobial Susceptibility Testing
2.3. DNA Extraction
2.4. Whole Genome Sequencing and Analysis
3. Results and Discussion
3.1. Antibiotic Susceptibility and Detection of Antimicrobial Resistance Genes
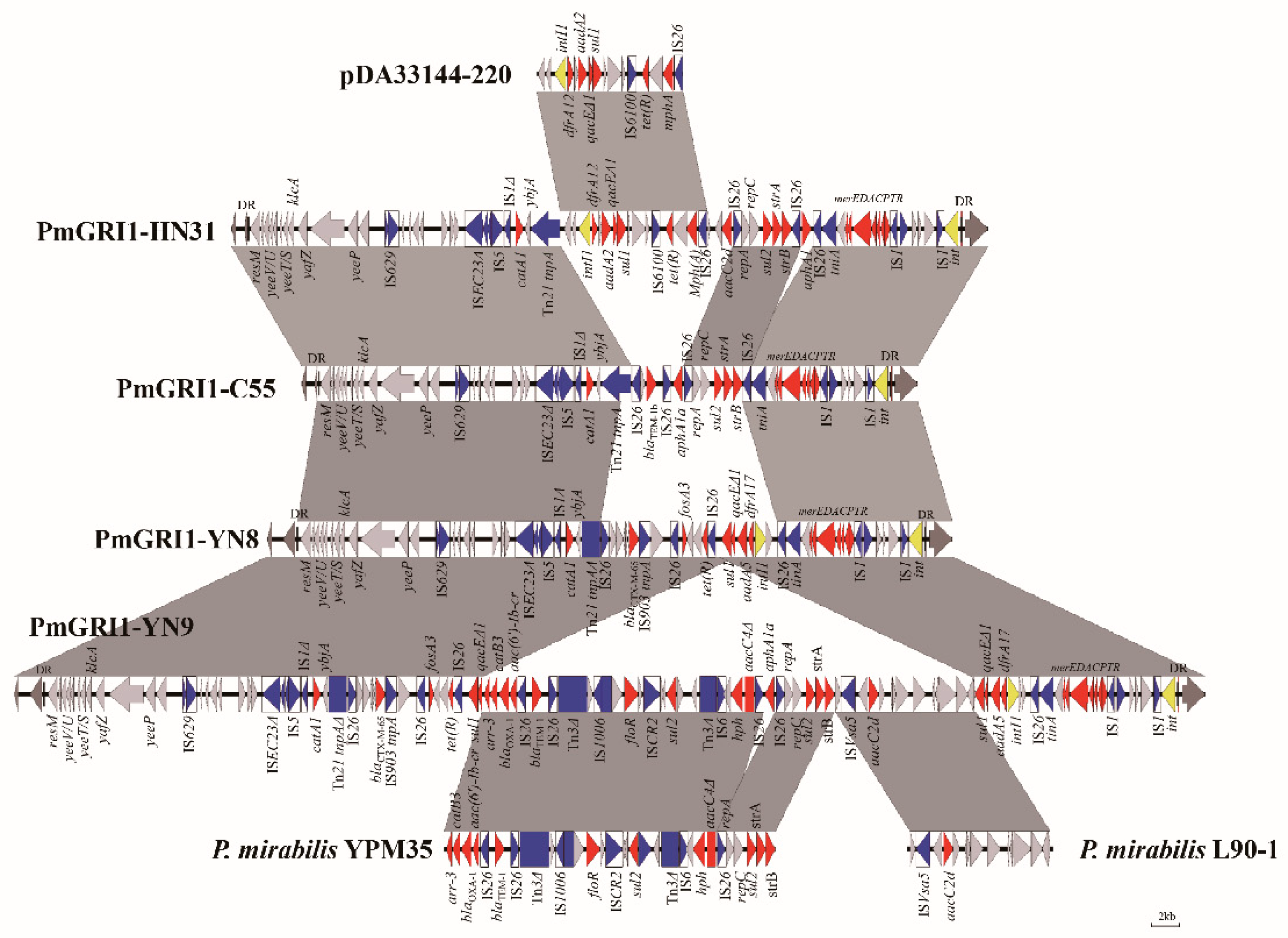
3.2. Analysis of Three Novel Variants of PmGRI1 in P. mirabilis
3.3. Characteristics of the Hybrid Structure in P. mirabilis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Schaffer, J.N.; Pearson, M.M. Proteus mirabilis and Urinary Tract Infections. Microbiol. Spectr. 2015, 3, 383–433. [Google Scholar] [CrossRef]
- Hu, Y.Y.; Cai, J.C.; Zhang, R.; Zhou, H.W.; Sun, Q.; Chen, G.X. Emergence of Proteus mirabilis harboring blaKPC-2 and qnrD in a Chinese Hospital. Antimicrob. Agents Chemother. 2012, 56, 2278–2282. [Google Scholar] [CrossRef]
- Ramos, A.C.; Cayô, R.; Carvalhaes, C.G.; Jové, T.; da Silva, G.P.; Sancho, F.M.P.; Chagas-Neto, T.; Medeiros, E.A.S.; Gales, A.C. Dissemination of Multidrug-Resistant Proteus mirabilis Clones Carrying a Novel Integron-Borne bla(IMP-1) in a Tertiary Hospital. Antimicrob. Agents Chemother. 2018, 62, e01321-17. [Google Scholar] [CrossRef] [PubMed]
- Sanches, M.S.; Baptista, A.A.S.; de Souza, M.; Menck-Costa, M.F.; Koga, V.L.; Kobayashi, R.K.T.; Rocha, S.P.D. Genotypic and phenotypic profiles of virulence factors and antimicrobial resistance of Proteus mirabilis isolated from chicken carcasses: Potential zoonotic risk. Braz. J. Microbiol. 2019, 50, 685–694. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Liu, D.; Zhang, X.; Tuo, H.; Lei, C.; Xie, X.; Gu, J.; Zhang, A. Characterization of Plasmid-Mediated Quinolone Resistance in Gram-Negative Bacterial Strains from Animals and Humans in China. Microb. Drug Resist. 2019, 25, 1050–1056. [Google Scholar] [CrossRef]
- Bushen, A.; Tekalign, E.; Abayneh, M. Drug- and Multidrug-Resistance Pattern of Enterobacteriaceae Isolated from Droppings of Healthy Chickens on a Poultry Farm in Southwest Ethiopia. Infect. Drug Resist. 2021, 14, 2051–2058. [Google Scholar] [CrossRef]
- Lei, C.W.; Chen, Y.P.; Kong, L.H.; Zeng, J.X.; Wang, Y.X.; Zhang, A.Y.; Wang, H.N. PGI2 Is a Novel SGI1-Relative Multidrug-Resistant Genomic Island Characterized in Proteus mirabilis. Antimicrob. Agents Chemother. 2018, 62, e00019-18. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.C.; Lei, C.W.; Kang, Z.Z.; Zhang, Y.; Wang, H.N. IS26-Mediated Genetic Rearrangements in Salmonella Genomic Island 1 of Proteus mirabilis. Front. Microbiol. 2019, 10, 2245. [Google Scholar] [CrossRef] [PubMed]
- Lei, C.W.; Zhang, A.Y.; Liu, B.H.; Wang, H.N.; Yang, L.Q.; Guan, Z.B.; Xu, C.W.; Zhang, D.D.; Yang, Y.Q. Two novel Salmonella genomic island 1 variants in Proteus mirabilis isolates from swine farms in China. Antimicrob. Agents Chemother. 2015, 59, 4336–4338. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.P.; Lei, C.W.; Kong, L.H.; Zeng, J.X.; Zhang, X.Z.; Liu, B.H.; Li, Y.; Xiang, R.; Wang, Y.X.; Chen, D.Y.; et al. Tn6450, a Novel Multidrug Resistance Transposon Characterized in a Proteus mirabilis Isolate from Chicken in China. Antimicrob. Agents Chemother. 2018, 62, e02192-17. [Google Scholar] [CrossRef] [PubMed]
- Delavat, F.; Miyazaki, R.; Carraro, N.; Pradervand, N.; van der Meer, J.R. The hidden life of integrative and conjugative elements. FEMS Microbiol. Rev. 2017, 41, 512–537. [Google Scholar] [CrossRef]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef] [PubMed]
- Bellanger, X.; Payot, S.; Leblond-Bourget, N.; Guédon, G. Conjugative and mobilizable genomic islands in bacteria: Evolution and diversity. FEMS Microbiol. Rev. 2014, 38, 720–760. [Google Scholar] [CrossRef]
- Liu, D.; Liu, W.; Lv, Z.; Xia, J.; Li, X.; Hao, Y.; Zhou, Y.; Yao, H.; Liu, Z.; Wang, Y.; et al. Emerging erm(B)-Mediated Macrolide Resistance Associated with Novel Multidrug Resistance Genomic Islands in Campylobacter. Antimicrob. Agents Chemother. 2019, 63, e00153-19. [Google Scholar] [CrossRef]
- Yao, H.; Jiao, D.; Zhao, W.; Li, A.; Li, R.; Du, X.D. Emergence of a Novel tet(L) Variant in Campylobacter spp. of Chicken Origin in China. Antimicrob. Agents Chemother. 2020, 65, e01622-20. [Google Scholar] [CrossRef]
- Zhu, D.; Yang, Z.; Xu, J.; Wang, M.; Jia, R.; Chen, S.; Liu, M.; Zhao, X.; Yang, Q.; Wu, Y.; et al. Pan-genome analysis of Riemerella anatipestifer reveals its genomic diversity and acquired antibiotic resistance associated with genomic islands. Funct. Integr. Genomics 2020, 20, 307–320. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.L.; Xu, Q.J.; Atiah, L.A.; Odah, K.A.; Gao, Y.H.; Kong, L.C.; Ma, H.X. Genomic island type IV secretion system and transposons in genomic islands involved in antimicrobial resistance in Trueperella pyogenes. Vet. Microbiol. 2020, 242, 108602. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, A.M.; Hussein, A.I.; Shimamoto, T. Proteus mirabilis clinical isolate harbouring a new variant of Salmonella genomic island 1 containing the multiple antibiotic resistance region. J. Antimicrob. Chemother. 2007, 59, 184–190. [Google Scholar] [CrossRef] [PubMed]
- Doublet, B.; Poirel, L.; Praud, K.; Nordmann, P.; Cloeckaert, A. European clinical isolate of Proteus mirabilis harbouring the Salmonella genomic island 1 variant SGI1-O. J. Antimicrob. Chemother. 2010, 65, 2260–2262. [Google Scholar] [CrossRef]
- Siebor, E.; Neuwirth, C. Emergence of Salmonella genomic island 1 (SGI1) among Proteus mirabilis clinical isolates in Dijon, France. J. Antimicrob. Chemother. 2013, 68, 1750–1756. [Google Scholar] [CrossRef]
- Siebor, E.; Neuwirth, C. Proteus genomic island 1 (PGI1), a new resistance genomic island from two Proteus mirabilis French clinical isolates. J. Antimicrob. Chemother. 2014, 69, 3216–3220. [Google Scholar] [CrossRef] [PubMed]
- Girlich, D.; Dortet, L.; Poirel, L.; Nordmann, P. Integration of the blaNDM-1 carbapenemase gene into Proteus genomic island 1 (PGI1-PmPEL) in a Proteus mirabilis clinical isolate. J. Antimicrob. Chemother. 2015, 70, 98–102. [Google Scholar] [CrossRef]
- Cummins, M.L.; Roy Chowdhury, P.; Marenda, M.S.; Browning, G.F.; Djordjevic, S.P. Salmonella Genomic Island 1B Variant Found in a Sequence Type 117 Avian Pathogenic Escherichia coli Isolate. mSphere 2019, 4, e00169-19. [Google Scholar] [CrossRef]
- Schultz, E.; Barraud, O.; Madec, J.Y.; Haenni, M.; Cloeckaert, A.; Ploy, M.C.; Doublet, B. Multidrug Resistance Salmonella Genomic Island 1 in a Morganella morganii subsp. morganii Human Clinical Isolate from France. mSphere 2017, 2, e00118-17. [Google Scholar] [CrossRef]
- Soliman, A.M.; Shimamoto, T.; Nariya, H.; Shimamoto, T. Emergence of Salmonella Genomic Island 1 Variant SGI1-W in a Clinical Isolate of Providencia stuartii from Egypt. Antimicrob. Agents Chemother. 2019, 63, e01793-18. [Google Scholar] [CrossRef] [PubMed]
- Siebor, E.; de Curraize, C.; Neuwirth, C. Genomic context of resistance genes within a French clinical MDR Proteus mirabilis: Identification of the novel genomic resistance island GIPmi1. J. Antimicrob. Chemother. 2018, 73, 1808–1811. [Google Scholar] [CrossRef] [PubMed]
- Lei, C.W.; Yao, T.G.; Yan, J.; Li, B.Y.; Wang, X.C.; Zhang, Y.; Gao, Y.F.; Wang, H.N. Identification of Proteus genomic island 2 variants in two clonal Proteus mirabilis isolates with coexistence of a novel genomic resistance island PmGRI1. J. Antimicrob. Chemother. 2020, 75, 2503–2507. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Lei, C.; Zuo, L.; Kong, L.; Kang, Z.; Zeng, J.; Zhang, X.; Wang, H. A novel cfr-carrying Tn7 transposon derivative characterized in Morganella morganii of swine origin in China. J. Antimicrob. Chemother. 2019, 74, 603–606. [Google Scholar] [CrossRef]
- Peters, J.E.; Craig, N.L. Tn7: Smarter than we thought. Nat. Rev. Mol. Cell Biol. 2001, 2, 806–814. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed]
- Cingolani, P.; Platts, A.; Wang le, L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Abyzov, A.; Urban, A.E.; Snyder, M.; Gerstein, M. CNVnator: An approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome Res. 2011, 21, 974–984. [Google Scholar] [CrossRef]
- Wang, J.; Mullighan, C.G.; Easton, J.; Roberts, S.; Heatley, S.L.; Ma, J.; Rusch, M.C.; Chen, K.; Harris, C.C.; Ding, L.; et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nat. Methods 2011, 8, 652–654. [Google Scholar] [CrossRef] [PubMed]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef]
- Zankari, E.; Hasman, H.; Cosentino, S.; Vestergaard, M.; Rasmussen, S.; Lund, O.; Aarestrup, F.M.; Larsen, M.V. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 2012, 67, 2640–2644. [Google Scholar] [CrossRef]
- Siguier, P.; Perochon, J.; Lestrade, L.; Mahillon, J.; Chandler, M. ISfinder: The reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006, 34, D32–D36. [Google Scholar] [CrossRef]
- Carattoli, A.; Zankari, E.; García-Fernández, A.; Voldby Larsen, M.; Lund, O.; Villa, L.; Møller Aarestrup, F.; Hasman, H. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef]
- Clausen, P.; Aarestrup, F.M.; Lund, O. Rapid and precise alignment of raw reads against redundant databases with KMA. BMC Bioinform. 2018, 19, 307. [Google Scholar] [CrossRef] [PubMed]
- Blackwell, G.A.; Nigro, S.J.; Hall, R.M. Evolution of AbGRI2-0, the Progenitor of the AbGRI2 Resistance Island in Global Clone 2 of Acinetobacter baumannii. Antimicrob. Agents Chemother. 2015, 60, 1421–1429. [Google Scholar] [CrossRef]
- He, S.; Hickman, A.B.; Varani, A.M.; Siguier, P.; Chandler, M.; Dekker, J.P.; Dyda, F. Insertion Sequence IS26 Reorganizes Plasmids in Clinically Isolated Multidrug-Resistant Bacteria by Replicative Transposition. mBio 2015, 6, e00762. [Google Scholar] [CrossRef]
- Proença, J.T.; Barral, D.C.; Gordo, I. Commensal-to-pathogen transition: One-single transposon insertion results in two pathoadaptive traits in Escherichia coli-macrophage interaction. Sci. Rep. 2017, 7, 4504. [Google Scholar] [CrossRef]
- Harmer, C.J.; Hall, R.M. IS26-Mediated Formation of Transposons Carrying Antibiotic Resistance Genes. mSphere 2016, 1, e00038-16. [Google Scholar] [CrossRef]
- Harmer, C.J.; Hall, R.M. IS26-Mediated Precise Excision of the IS26-aphA1a Translocatable Unit. mBio 2015, 6, e01866-15. [Google Scholar] [CrossRef] [PubMed]
- Vinué, L.; Sater, M.R.A.; Herriott, I.; Huntley, M.H.; Jacoby, G.A.; Hooper, D.C. Multiple Copies of qnrA1 on an IncA/C(2) Plasmid Explain Enhanced Quinolone Resistance in an Escherichia coli Mutant. Antimicrob. Agents Chemother. 2019, 63, e00718-19. [Google Scholar] [CrossRef]
- Fritsche, T.R.; Castanheira, M.; Miller, G.H.; Jones, R.N.; Armstrong, E.S. Detection of methyltransferases conferring high-level resistance to aminoglycosides in enterobacteriaceae from Europe, North America, and Latin America. Antimicrob. Agents Chemother. 2008, 52, 1843–1845. [Google Scholar] [CrossRef] [PubMed]
- Cassu-Corsi, D.; Martins, W.M.; Nicoletti, A.G.; Almeida, L.G.; Vasconcelos, A.T.; Gales, A.C. Characterisation of plasmid-mediated rmtB-1 in Enterobacteriaceae clinical isolates from São Paulo, Brazil. Memórias do Instituto Oswaldo Cruz 2018, 113, e180392. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, K.; Doi, Y.; Yamane, K.; Kurokawa, H.; Shibata, N.; Shibayama, K.; Yagi, T.; Kato, H.; Arakawa, Y. Acquisition of 16S rRNA methylase gene in Pseudomonas aeruginosa. Lancet 2003, 362, 1888–1893. [Google Scholar] [CrossRef]
- Yamane, K.; Wachino, J.; Doi, Y.; Kurokawa, H.; Arakawa, Y. Global spread of multiple aminoglycoside resistance genes. Emerg. Infect. Dis. 2005, 11, 951–953. [Google Scholar] [CrossRef]
- Doi, Y.; Yokoyama, K.; Yamane, K.; Wachino, J.; Shibata, N.; Yagi, T.; Shibayama, K.; Kato, H.; Arakawa, Y. Plasmid-mediated 16S rRNA methylase in Serratia marcescens conferring high-level resistance to aminoglycosides. Antimicrob. Agents Chemother. 2004, 48, 491–496. [Google Scholar] [CrossRef] [PubMed]
- Wachino, J.; Yamane, K.; Shibayama, K.; Kurokawa, H.; Shibata, N.; Suzuki, S.; Doi, Y.; Kimura, K.; Ike, Y.; Arakawa, Y. Novel plasmid-mediated 16S rRNA methylase, RmtC, found in a Proteus mirabilis isolate demonstrating extraordinary high-level resistance against various aminoglycosides. Antimicrob. Agents Chemother. 2006, 50, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Du, X.D.; Wu, C.M.; Liu, H.B.; Li, X.S.; Beier, R.C.; Xiao, F.; Qin, S.S.; Huang, S.Y.; Shen, J.Z. Plasmid-mediated ArmA and RmtB 16S rRNA methylases in Escherichia coli isolated from chickens. J. Antimicrob. Chemother. 2009, 64, 1328–1330. [Google Scholar] [CrossRef][Green Version]
- Doi, Y.; Adams-Haduch, J.M.; Paterson, D.L. Escherichia coli isolate coproducing 16S rRNA Methylase and CTX-M-type extended-spectrum beta-lactamase isolated from an outpatient in the United States. Antimicrob. Agents Chemother. 2008, 52, 1204–1205. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Siebor, E.; Neuwirth, C. The new variant of Salmonella genomic island 1 (SGI1-V) from a Proteus mirabilis French clinical isolate harbours blaVEB-6 and qnrA1 in the multiple antibiotic resistance region. J. Antimicrob. Chemother. 2011, 66, 2513–2520. [Google Scholar] [CrossRef]
- Dong, W.L.; Odah, K.A.; Liu, L.; Xu, Q.J.; Gao, Y.H.; Kong, L.C.; Ma, H.X. Multidrug resistance genes are associated with a 42-kb island TGI1 carrying a complex class 1 integron in Trueperella pyogenes. J. Glob. Antimicrob. Resist. 2020, 22, 1–4. [Google Scholar] [CrossRef] [PubMed]

| Strain | Source of Samples | SGI1/SGI1-like Gene Island Type | Antibiotic Resistance Genes a | Strain Resistance Profile | Size (bp) |
|---|---|---|---|---|---|
| HN31 | Swine | PmGRI1-HN31 | catA1, dfrA12, aadA2, sul1, mph(A), aacC2d, sul2, strA, strB, aphA1, floR, tet(A) | AMP-CHL-NAL-STR-SPT-GEN-TMP-SUL-SXT | 62,862 |
| YN8 | Chicken | PmGRI1-YN8 | catA1, blaCTX-M-65, fosA3, sul1, aadA5, dfrA17, aadA1, dfrA1, sul2 | NAL-SPT-TMP-SUL-SXT | 62,784 |
| YN9 | Chicken | PmGRI1-YN9 | catA1, blaCTX-M-65, fosA3, sul1, arr-3, catB3, blaOXA-1, aac(6′)-Ib-cr, blaTEM-1, floR, sul2, hph, aphA1a, strA, strB, aacC2d, aadA5, dfrA17, aadA1, dfrA1, aacC4 | AMP-CHL-FFC-NAL-CIP-STR-SPT-GEN-TMP-SUL-SXT | 55,239 |
| HN2p | Swine | Hybrid structure of Tn7-like and PmGRI1 | rmtB, hph, aacC4, aphA1a, strA, strB, aadA1, aadA2, blaTEM-1b, qnrA1, lun(F), erm(42), catA1, floR, sul1, sul2, tet(G), dfrA12, dfrA1, blaCMY-2 | AMP-AMC-AMK-SXT-LEV-CIP-NAL-FFC-CHL-GEN | 123,622 |
| HN31 | Swine | PmGRI1-HN31 | catA1, dfrA12, aadA2, sul1, mph(A), aacC2d, sul2, strA, strB, aphA1, floR, tet(A) | AMP-CHL-NAL-STR-SPT-GEN-TMP-SUL-SXT | 62,862 |
| YN8 | Chicken | PmGRI1-YN8 | catA1, blaCTX-M-65, fosA3, sul1, aadA5, dfrA17, aadA1, dfrA1, sul2 | NAL-SPT-TMP-SUL-SXT | 62,784 |
| YN9 | Chicken | PmGRI1-YN9 | catA1, blaCTX-M-65, fosA3, sul1, arr-3, catB3, blaOXA-1, aac(6′)-Ib-cr, blaTEM-1, floR, sul2, hph, aphA1a, strA, strB, aacC2d, aadA5, dfrA17, aadA1, dfrA1, aacC4 | AMP-CHL-FFC-NAL-CIP-STR-SPT-GEN-TMP-SUL-SXT | 55,239 |
| HN2p | Swine | Hybrid structure of Tn7-like and PmGRI1 | rmtB, hph, aacC4, aphA1a, strA, strB, aadA1, aadA2, blaTEM-1b, qnrA1, lun(F), erm(42), catA1, floR, sul1, sul2, tet(G), dfrA12, dfrA1, blaCMY-2 | AMP-AMC-AMK-SXT-LEV-CIP-NAL-FFC-CHL-GEN | 123,622 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, B.; Wang, X.; Lei, C.; Tang, Y.; He, J.; Gao, Y.; Zhang, Y.; Wang, H. Identification of Three Novel PmGRI1 Genomic Resistance Islands and One Multidrug Resistant Hybrid Structure of Tn7-like Transposon and PmGRI1 in Proteus mirabilis. Antibiotics 2021, 10, 1268. https://doi.org/10.3390/antibiotics10101268
Ma B, Wang X, Lei C, Tang Y, He J, Gao Y, Zhang Y, Wang H. Identification of Three Novel PmGRI1 Genomic Resistance Islands and One Multidrug Resistant Hybrid Structure of Tn7-like Transposon and PmGRI1 in Proteus mirabilis. Antibiotics. 2021; 10(10):1268. https://doi.org/10.3390/antibiotics10101268
Chicago/Turabian StyleMa, Boheng, Xuechun Wang, Changwei Lei, Yizhi Tang, Juan He, Yufeng Gao, Yu Zhang, and Hongning Wang. 2021. "Identification of Three Novel PmGRI1 Genomic Resistance Islands and One Multidrug Resistant Hybrid Structure of Tn7-like Transposon and PmGRI1 in Proteus mirabilis" Antibiotics 10, no. 10: 1268. https://doi.org/10.3390/antibiotics10101268
APA StyleMa, B., Wang, X., Lei, C., Tang, Y., He, J., Gao, Y., Zhang, Y., & Wang, H. (2021). Identification of Three Novel PmGRI1 Genomic Resistance Islands and One Multidrug Resistant Hybrid Structure of Tn7-like Transposon and PmGRI1 in Proteus mirabilis. Antibiotics, 10(10), 1268. https://doi.org/10.3390/antibiotics10101268








