Optical Biosensors for Therapeutic Drug Monitoring
Abstract
1. Introduction
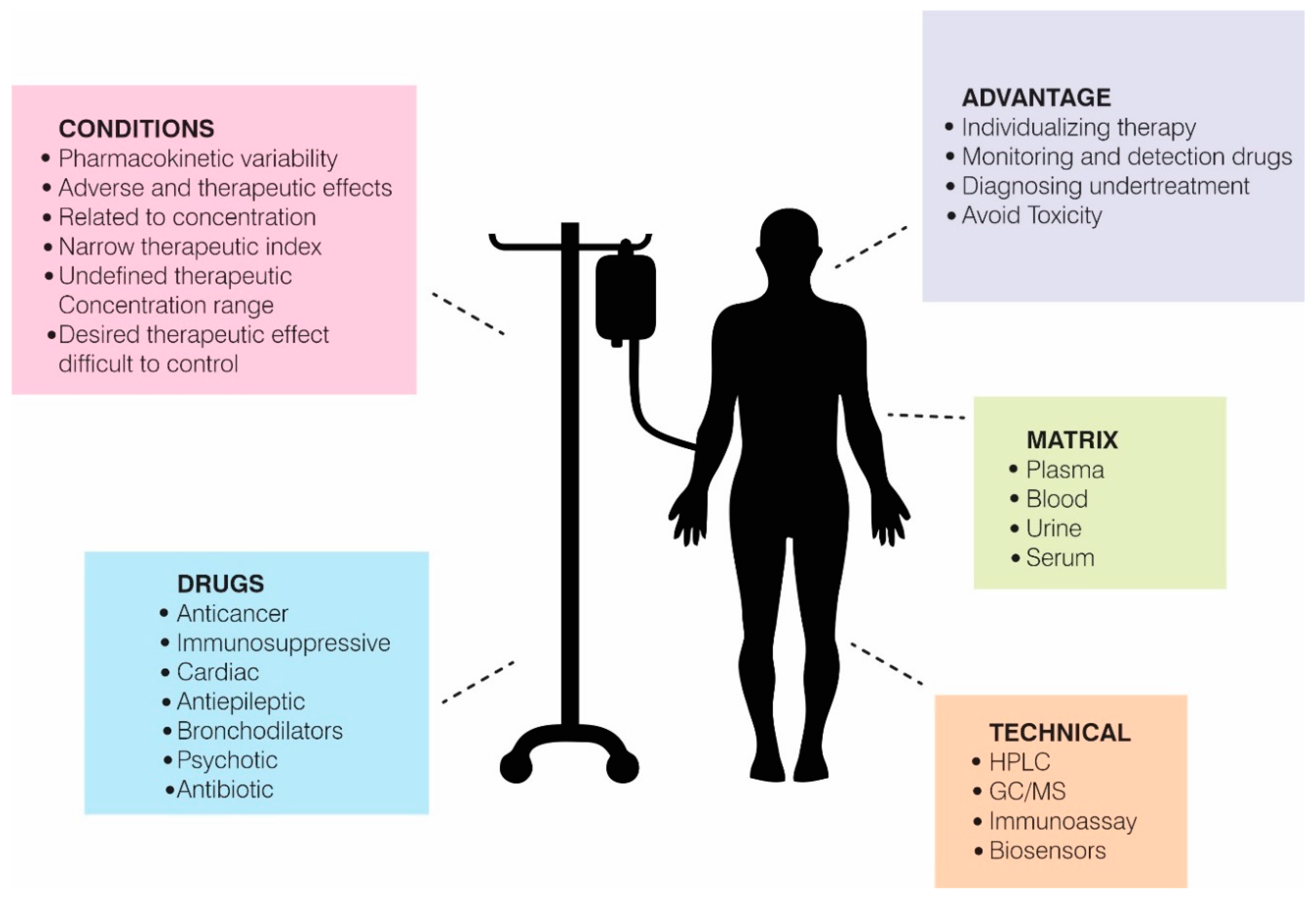
2. Therapeutic Drug Monitoring (TDM)
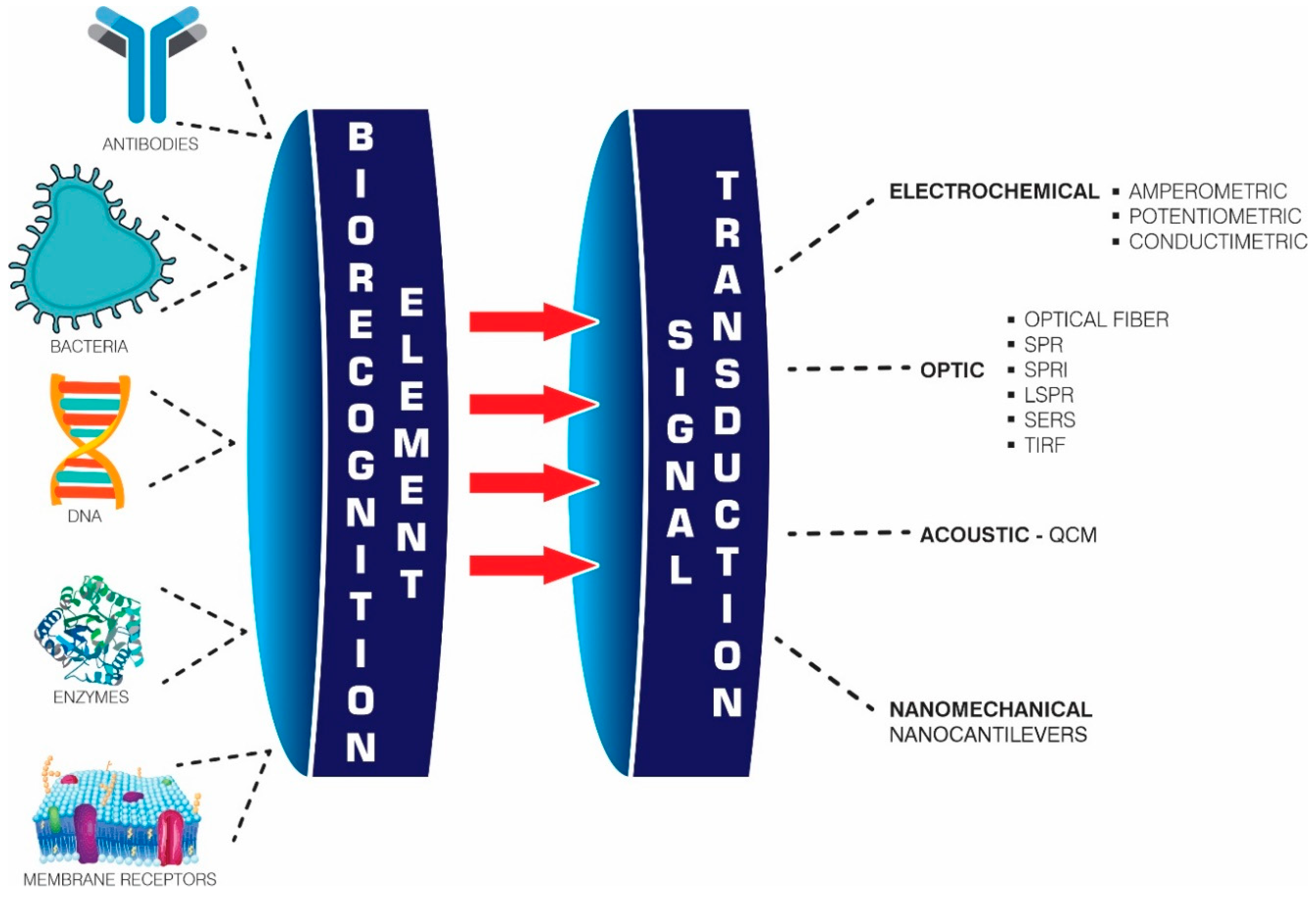
3. Biosensors
4. Optical Biosensors
4.1. Classifying Optical Biosensors
4.1.1. Fibre-Optic Biosensors
4.1.2. Surface Plasmon Resonance (SPR)-Based Optical Biosensors
4.1.3. Surface–Enhanced Raman Scattering (SERS)
4.1.4. Total Internal Reflection Fluorescence (TIRF) Biosensors
5. Surface Functionalization in TDM
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Figueras, A. Review of the Evidence to Include TDM in the Essential in Vitro Diagnostics List and Prioritization of Medicines to be Monitored; Fundació Institut Català de Farmacologia: Barcelona, Spain, 2019. [Google Scholar]
- Belmar-Liberato, R.; Gonzalez-Canga, A.; Tamame-Martin, P.; Escribano-Salazar, M. Amoxicillin and amoxicillin-clavulanic acid resistance in veterinary medicine—the situation in Europe: A review. Vet. Med. 2011, 56, 473–485. [Google Scholar] [CrossRef]
- Hauptman, P.J.; Blume, S.W.; Lewis, E.F.; Ward, S. Digoxin toxicity and use of digoxin immune fab: Insights from a national hospital database. JACC Heart Fail. 2016, 4, 357–364. [Google Scholar] [CrossRef] [PubMed]
- Alvarado-Miranda, A.; Calderillo-Ruiz, G.; Rodríguez-Ortiz, R.; Gallardo, L.; Aguilar-Flores, K.I.; Cabrera-Galeana, P. Risk factors for the development of hematological toxicity during the application of weekly paclitaxel in breast cancer. Gac. Mex. Oncol. 2019, 18, 8–12. [Google Scholar] [CrossRef]
- Gaus, K.; Hall, E.A. Surface plasmon resonance sensor for heparin measurements in blood plasma. Biosens. Bioelectron. 1998, 13, 1307–1315. [Google Scholar] [CrossRef]
- Tamargo, J.; Le Heuzey, J.-Y.; Mabo, P. Narrow therapeutic index drugs: A clinical pharmacological consideration to flecainide. Eur. J. Clin. Pharmacol. 2015, 71, 549–567. [Google Scholar] [CrossRef] [PubMed]
- Pacheco, T.; Bustos, R.-H.; González, D.; Garzón, V.; García, J.-C.; Ramírez, D. An approach to measuring colistin plasma levels regarding the treatment of multidrug-resistant bacterial infection. Antibiotics 2019, 8, 100. [Google Scholar] [CrossRef] [PubMed]
- Carroll, L.M.; Gaballa, A.; Guldimann, C.; Sullivan, G.; Henderson, L.O.; Wiedmann, M. Identification of novel mobilized colistin resistance gene mcr-9 in a multidrug-resistant, colistin-susceptible salmonella enterica serotype typhimurium isolate. mBio 2019, 10, e00853-19. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulos, A.; Ribera, A.; Mavrogenis, A.F.; Rodriguez-Pardo, D.; Bonnet, E.; Salles, M.J.; del Toro, M.D.; Nguyen, S.; Blanco-García, A.; Skaliczki, G. Multidrug-resistant and extensively drug-resistant Gram-negative prosthetic joint infections: Role of surgery and impact of colistin administration. Int. J. Antimicrob. Agents 2019, 53, 294–301. [Google Scholar] [CrossRef] [PubMed]
- Gervasini, G.; de Murillo, S.G.; Jiménez, M.; María, D.; Vagace, J.M. Effect of polymorphisms in transporter genes on dosing, efficacy and toxicity of maintenance therapy in children with acute lymphoblastic leukemia. Gene 2017, 628, 72–77. [Google Scholar] [CrossRef] [PubMed]
- McKeating, K.S.; Aubé, A.; Masson, J.-F. Biosensors and nanobiosensors for therapeutic drug and response monitoring. Analyst 2016, 141, 429–449. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, A.; Krasowski, M. Therapeutic Drug Monitoring Data: A Concise Guide; Academic Press: Cambridge, MA, USA, 2019. [Google Scholar]
- Reder-Christ, K.; Bendas, G. Biosensor applications in the field of antibiotic research--a review of recent developments. Sensors 2011, 11, 9450–9466. [Google Scholar] [CrossRef] [PubMed]
- Cennamo, N.; Massarotti, D.; Conte, L.; Zeni, L. Low cost sensors based on SPR in a plastic optical fiber for biosensor implementation. Sensors 2011, 11, 11752–11760. [Google Scholar] [CrossRef] [PubMed]
- Piliarik, M.; Vala, M.; Tichý, I.; Homola, J. Compact and low-cost biosensor based on novel approach to spectroscopy of surface plasmons. Biosens. Bioelectron. 2009, 24, 3430–3435. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.-S.; Lee, M.-H. Overview of therapeutic drug monitoring. Korean J. Intern. Med. 2009, 24, 1. [Google Scholar] [CrossRef] [PubMed]
- Leticia, J.; Betancourt, R.; Vigil, J.L.G.; Barnés, C.G.; Santillán, D.H.; Gutiérrez, L.J. Farmacovigilancia II. Las reacciones adversas y el Programa Internacional de Monitoreo de los Medicamentos. Rev. Med. IMSS 2004, 42, 419–423. [Google Scholar]
- Rowland, M.; Tozer, T.N. Clinical Pharmacokinetics/Pharmacodynamics; Lippincott Williams and Wilkins: Philadelphia, UK, 2005. [Google Scholar]
- Touw, D.J.; Neef, C.; Thomson, A.H.; Vinks, A.A. Cost-effectiveness of therapeutic drug monitoring: A systematic review. Ther. Drug Monit. 2005, 27, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Sime, F.B.; Roberts, M.S.; Peake, S.L.; Lipman, J.; Roberts, J.A. Does beta-lactam pharmacokinetic variability in critically ill patients justify therapeutic drug monitoring? A systematic review. Ann. Intensive Care 2012, 2, 35. [Google Scholar] [CrossRef] [PubMed]
- Ye, Z.-K.; Tang, H.-L.; Zhai, S.-D. Benefits of therapeutic drug monitoring of vancomycin: A systematic review and meta-analysis. PLoS ONE 2013, 8, e77169. [Google Scholar] [CrossRef] [PubMed]
- Knight, S.R.; Morris, P.J. Does the evidence support the use of mycophenolate mofetil therapeutic drug monitoring in clinical practice? A systematic review. Transplantation 2008, 85, 1675–1685. [Google Scholar] [CrossRef] [PubMed]
- Sparshatt, A.; Taylor, D.; Patel, M.X.; Kapur, S. A systematic review of aripiprazole—Dose, plasma concentration, receptor occupancy, and response: Implications for therapeutic drug monitoring. J. Clin. Psychiatry 2010, 71, 1447–1456. [Google Scholar] [CrossRef] [PubMed]
- McWhinney, B.C.; Wallis, S.C.; Hillister, T.; Roberts, J.A.; Lipman, J.; Ungerer, J.P. Analysis of 12 beta-lactam antibiotics in human plasma by HPLC with ultraviolet detection. J. Chromatogr. B 2010, 878, 2039–2043. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.-F.; Phillips, D.; Neue, U. Simple and rugged SPE method for the determination of tetracycline antibiotics in serum by HPLC using a volatile mobile phase. Chromatographia 1997, 44, 187–190. [Google Scholar] [CrossRef]
- Paal, M.; Zoller, M.; Schuster, C.; Vogeser, M.; Schütze, G. Simultaneous quantification of cefepime, meropenem, ciprofloxacin, moxifloxacin, linezolid and piperacillin in human serum using an isotope-dilution HPLC–MS/MS method. J. Pharm. Biomed. Anal. 2018, 152, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Begas, E.; Tsakalof, A.; Dardiotis, E.; Vatidis, G.E.; Kouvaras, E.; Asprodini, E.K. Development and validation of a reversed-phase HPLC method for licarbazepine monitoring in serum of patients under oxcarbazepine treatment. Biomed. Chromatogr. 2017, 31, e3950. [Google Scholar] [CrossRef] [PubMed]
- Esteve-Romero, J.; Albiol-Chiva, J.; Peris-Vicente, J. A review on development of analytical methods to determine monitorable drugs in serum and urine by micellar liquid chromatography using direct injection. Anal. Chim. Acta 2016, 926, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; You, J.; Sun, Z.; Ji, Z.; Hu, N.; Zhou, W.; Zhou, X. HPLC determination of γ-aminobutyric acid and its analogs in human serum using precolumn fluorescence labeling with 4-(carbazole-9-yl)-benzyl chloroformate. J. Sep. Sci. 2019, 42, 826–833. [Google Scholar] [CrossRef] [PubMed]
- Locatelli, M.; Tinari, N.; Grassadonia, A.; Tartaglia, A.; Macerola, D.; Piccolantonio, S.; Sperandio, E.; D’Ovidio, C.; Carradori, S.; Ulusoy, H.I. FPSE-HPLC-DAD method for the quantification of anticancer drugs in human whole blood, plasma, and urine. J. Chromatogr. B 2018, 1095, 204–213. [Google Scholar] [CrossRef] [PubMed]
- Crutchfield, C.A.; Marzinke, M.A.; Clarke, W.A. Quantification of docetaxel in serum using turbulent flow liquid chromatography electrospray tandem mass spectrometry (TFC-HPLC-ESI-MS/MS). In Clinical Applications of Mass Spectrometry in Drug Analysis; Humana Press: New York, NY, USA, 2016; pp. 121–124. [Google Scholar]
- Larson, R.R.; Khazaeli, M.; Dillon, H.K. Development of an HPLC method for simultaneous analysis of five antineoplastic agents. Appl. Occup. Environ. Hyg. 2003, 18, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S. Digoxin and other cardiac glycosides. In Principles and Practice of Critical Care Toxicology; Jaypee Brothers Medical Pub: New Delhi, India, 2019; pp. 194–199. [Google Scholar]
- Fridlund, J.; Woksepp, H.; Schön, T. A microbiological method for determining serum levels of broad spectrum β-lactam antibiotics in critically ill patients. J. Microbiol. Methods 2016, 129, 23–27. [Google Scholar] [CrossRef] [PubMed]
- Bustos, R.; Zapata, C.; Esteban, E.; García, J.-C.; Jáuregui, E.; Jaimes, D. Label-free quantification of anti-TNF-α in patients treated with adalimumab using an optical biosensor. Sensors 2018, 18, 691. [Google Scholar] [CrossRef] [PubMed]
- Vis, A.; van der Gaast, A.; van Rhijn, B.; Catsburg, T.; Schmidt, C.; Mickisch, G. A phase II trial of methotrexate-human serum albumin (MTX-HSA) in patients with metastatic renal cell carcinoma who progressed under immunotherapy. Cancer Chemother. Pharmacol. 2002, 49, 342–345. [Google Scholar] [PubMed]
- Bouquié, R.; Deslandes, G.; Bernáldez, B.N.; Renaud, C.; Dailly, E.; Jolliet, P. A fast LC-MS/MS assay for methotrexate monitoring in plasma: Validation, comparison to FPIA and application in the setting of carboxypeptidase therapy. Anal. Methods 2014, 6, 178–186. [Google Scholar] [CrossRef]
- Aherne, G.; Piall, E.; Marks, V. Development and application of a radioimmunoassay for methotrexate. Br. J. Cancer 1977, 36, 608. [Google Scholar] [CrossRef] [PubMed]
- Tartaggia, S.; Alvau, M.D.; Meneghello, A.; Casetta, B.; Polo, F.; Toffoli, G. Practical fluorimetric assay for the detection of anticancer drug SN-38 in human plasma. J. Pharm. Biomed. Anal. 2018, 159, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Wang, F.; Wang, H.; Kambam, S.; Chen, X. Colorimetric and fluorometric detection of neomycin based on conjugated polydiacetylene supramolecules. Macromol. Rapid Commun. 2013, 34, 944–948. [Google Scholar] [CrossRef] [PubMed]
- La Marca, G.; Villanelli, F.; Malvagia, S.; Ombrone, D.; Funghini, S.; De Gaudio, M.; Fallani, S.; Cassetta, M.I.; Novelli, A.; Chiappini, E. Rapid and sensitive LC–MS/MS method for the analysis of antibiotic linezolid on dried blood spot. J. Pharm. Biomed. Anal. 2012, 67, 86–91. [Google Scholar] [CrossRef] [PubMed]
- Edelbroek, P.M.; van der Heijden, J.; Stolk, L.M. Dried blood spot methods in therapeutic drug monitoring: Methods, assays, and pitfalls. Ther. Drug Monit. 2009, 31, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Goode, J.A.; Rushworth, J.V.; Millner, P.A. Biosensor regeneration: A review of common techniques and outcomes. Langmuir ACS J. Surf. Colloids 2015, 31, 6267–6276. [Google Scholar] [CrossRef] [PubMed]
- Vo-Dinh, T.; Cullum, B. Biosensors and biochips: Advances in biological and medical diagnostics. Fresenius J. Anal. Chem. 2000, 366, 540–551. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.J.; Bangsaruntip, S.; Drouvalakis, K.A.; Kam, N.W.; Shim, M.; Li, Y.; Kim, W.; Utz, P.J.; Dai, H. Noncovalent functionalization of carbon nanotubes for highly specific electronic biosensors. Proc. Natl. Acad. Sci. USA 2003, 100, 4984–4989. [Google Scholar] [CrossRef] [PubMed]
- Matsiuk Ia, R. Morphocytochemical reaction of the main exocrinocytes of the proper gastric glands to orchiectomy. Arkhiv Anat. Gistol. Embriol. 1989, 97, 79–84. [Google Scholar]
- Wang, X.; Niessner, R.; Tang, D.; Knopp, D. Nanoparticle-based immunosensors and immunoassays for aflatoxins. Anal. Chim. Acta 2016, 912, 10–23. [Google Scholar] [CrossRef] [PubMed]
- Reverte, L.; Prieto-Simon, B.; Campas, M. New advances in electrochemical biosensors for the detection of toxins: Nanomaterials, magnetic beads and microfluidics systems. A review. Anal. Chim. Acta 2016, 908, 8–21. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Lin, Y.; Guo, L.; Qiu, B.; Chen, G.; Yang, H.H.; Lin, Z. A universal multicolor immunosensor for semiquantitative visual detection of biomarkers with the naked eyes. Biosens. Bioelectron. 2016, 87, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Maldonado, J.; Gonzalez-Guerrero, A.B.; Dominguez, C.; Lechuga, L.M. Label-free bimodal waveguide immunosensor for rapid diagnosis of bacterial infections in cirrhotic patients. Biosens. Bioelectron. 2016, 85, 310–316. [Google Scholar] [CrossRef] [PubMed]
- Farina, D.; Zinellu, M.; Fanari, M.; Porcu, M.C.; Scognamillo, S.; Puggioni, G.M.; Rocchitta, G.; Serra, P.A.; Pretti, L. Development of a biosensor telemetry system for monitoring fermentation in craft breweries. Food Chem. 2017, 218, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Pashazadeh, P.; Mokhtarzadeh, A.; Hasanzadeh, M.; Hejazi, M.; Hashemi, M.; de la Guardia, M. Nano-materials for use in sensing of salmonella infections: Recent advances. Biosens. Bioelectron. 2016, 87, 1050–1064. [Google Scholar] [CrossRef] [PubMed]
- Doldan, X.; Fagundez, P.; Cayota, A.; Laiz, J.; Tosar, J.P. Electrochemical sandwich immunosensor for determination of exosomes based on surface marker-mediated signal amplification. Anal. Chem. 2016, 88, 10466–10473. [Google Scholar] [CrossRef] [PubMed]
- Dai, H.; Zhang, S.; Hong, Z.; Lin, Y. A potentiometric addressable photoelectrochemical biosensor for sensitive detection of two biomarkers. Anal. Chem. 2016, 88, 9532–9538. [Google Scholar] [CrossRef] [PubMed]
- Ditto, N.T.; Brooks, B.D. The emerging role of biosensor-based epitope binning and mapping in antibody-based drug discovery. Expert Opin. Drug Discov. 2016, 11, 925–937. [Google Scholar] [CrossRef] [PubMed]
- Zanganeh, S.; Khosravi, S.; Namdar, N.; Amiri, M.H.; Gharooni, M.; Abdolahad, M. Electrochemical approach for monitoring the effect of anti tubulin drugs on breast cancer cells based on silicon nanograss electrodes. Anal. Chim. Acta 2016, 938, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Huang, Z.; Rogers, M.; Boutelle, M.; Cass, A.E. Evaluation of a minimally invasive glucose biosensor for continuous tissue monitoring. Anal. Bioanal. Chem. 2016, 408, 8427–8435. [Google Scholar] [CrossRef] [PubMed]
- Dubiak-Szepietowska, M.; Karczmarczyk, A.; Winckler, T.; Feller, K.H. A cell-based biosensor for nanomaterials cytotoxicity assessment in three dimensional cell culture. Toxicology 2016, 370, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Bustos, R.H.; Suesca, E.; Millan, D.; Gonzalez, J.M.; Fontanilla, M.R. Real-time quantification of proteins secreted by artificial connective tissue made from uni- or multidirectional collagen I scaffolds and oral mucosa fibroblasts. Anal. Chem. 2014, 86, 2421–2428. [Google Scholar] [CrossRef] [PubMed]
- Thevenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Biosens. Bioelectron. 2001, 16, 121–131. [Google Scholar] [CrossRef] [PubMed]
- Mehrotra, P. Biosensors and their applications—A review. J. Oral Biol. Craniofacial Res. 2016, 6, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Bueno, J. Biosensors in antimicrobial drug discovery: Since biology until screening platforms. J. Microb. Biochem. Technol. 2014, S10, 1–10. [Google Scholar]
- Grieshaber, D.; MacKenzie, R.; Voros, J.; Reimhult, E. Electrochemical biosensors—Sensor principles and architectures. Sensors 2008, 8, 1400–1458. [Google Scholar] [CrossRef] [PubMed]
- Wang, J. Analytical Electrochemistry; Wyley-VCH: Weinheim, Germany, 2006; p. 272. [Google Scholar]
- Hammond, J.L.; Formisano, N.; Estrela, P.; Carrara, S.; Tkac, J. Electrochemical biosensors and nanobiosensors. Essays Biochem. 2016, 60, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Rowe, A.A.; Miller, E.A.; Plaxco, K.W. Reagentless measurement of aminoglycoside antibiotics in blood serum via an electrochemical, ribonucleic acid aptamer-based biosensor. Anal. Chem. 2010, 82, 7090–7095. [Google Scholar] [CrossRef] [PubMed]
- Ferapontova, E.E.; Olsen, E.M.; Gothelf, K.V. An RNA aptamer-based electrochemical biosensor for detection of theophylline in serum. J. Am. Chem. Soc. 2008, 130, 4256–4258. [Google Scholar] [CrossRef] [PubMed]
- Bagheri, H.; Talemi, R.; Afkhami, A. Gold nanoparticles deposited on fluorine-doped tin oxide surface as an effective platform for fabricating a highly sensitive and specific digoxin aptasensor. RSC Adv. 2015, 5, 58491–58498. [Google Scholar] [CrossRef]
- Baj-Rossi, C.; Micheli, G.D.; Carrara, S. Electrochemical detection of anti-breast-cancer agents in human serum by cytochrome P450-coated carbon nanotubes. Sensors 2012, 12, 6520–6537. [Google Scholar] [CrossRef] [PubMed]
- Bonazza, G.; Tartaggia, S.; Toffoli, G.; Polo, F.; Daniele, S. Voltammetric behaviour of the anticancer drug irinotecan and its metabolites in acetonitrile. Implications for electrochemical therapeutic drug monitoring. Electrochim. Acta 2018, 289, 483–493. [Google Scholar] [CrossRef]
- Ensafi, A.A.; Amini, M.; Rezaei, B. Impedimetric DNA-biosensor for the study of anti-cancer action of mitomycin C: Comparison between acid and electroreductive activation. Biosens. Bioelectron. 2014, 59, 282–288. [Google Scholar] [CrossRef] [PubMed]
- Erdem, A.; Congur, G. Impedimetric detection of in situ interaction between anti-cancer drug bleomycin and DNA. Int. J. Biol. Macromol. 2013, 61, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Karimi-Maleh, H.; Bananezhad, A.; Ganjali, M.R.; Norouzi, P.; Sadrnia, A. Surface amplification of pencil graphite electrode with polypyrrole and reduced graphene oxide for fabrication of a guanine/adenine DNA based electrochemical biosensors for determination of didanosine anticancer drug. Appl. Surf. Sci. 2018, 441, 55–60. [Google Scholar] [CrossRef]
- Meneghello, A.; Tartaggia, S.; Alvau, M.D.; Polo, F.; Toffoli, G. Biosensing technologies for therapeutic drug monitoring. Curr. Med. Chem. 2018, 25, 4354–4377. [Google Scholar] [CrossRef] [PubMed]
- Radhapyari, K.; Khan, R. Biosensor for detection of selective anticancer drug gemcitabine based on polyaniline-gold nanocomposite. Adv. Mater. Lett. 2015, 6, 13–18. [Google Scholar] [CrossRef]
- Radhapyari, K.; Kotoky, P.; Khan, R. Detection of anticancer drug tamoxifen using biosensor based on polyaniline probe modified with horseradish peroxidase. Mater. Sci. Eng. C 2013, 33, 583–587. [Google Scholar] [CrossRef] [PubMed]
- Rafique, B.; Khalid, A.M.; Akhtar, K.; Jabbar, A. Interaction of anticancer drug methotrexate with DNA analyzed by electrochemical and spectroscopic methods. Biosens. Bioelectron. 2013, 44, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Rauf, S.; Gooding, J.; Akhtar, K.; Ghauri, M.; Rahman, M.; Anwar, M.; Khalid, A. Electrochemical approach of anticancer drugs–DNA interaction. J. Pharm. Biomed. Anal. 2005, 37, 205–217. [Google Scholar] [CrossRef] [PubMed]
- Santiago-Lopez, A.J.; Vera, J.L.; Meléndez, E. DNA electrochemical biosensor for metallic drugs at physiological conditions. J. Electroanal. Chem. 2014, 731, 139–144. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sengiz, C.; Congur, G.; Eksin, E.; Erdem, A. Multiwalled carbon nanotubes-chitosan modified single-use biosensors for electrochemical monitoring of drug-DNA interactions. Electroanalysis 2015, 27, 1855–1863. [Google Scholar] [CrossRef]
- Tajik, S.; Taher, M.A.; Beitollahi, H.; Torkzadeh-Mahani, M. Electrochemical determination of the anticancer drug taxol at a ds-DNA modified pencil-graphite electrode and its application as a label-free electrochemical biosensor. Talanta 2015, 134, 60–64. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.U.; Chen, C.; Lin, K.-h.; Fang, Y.; Lieber, C.M. Label-free detection of small-molecule–protein interactions by using nanowire nanosensors. Proc. Natl. Acad. Sci. USA 2005, 102, 3208–3212. [Google Scholar] [CrossRef] [PubMed]
- Alvau, M.D.; Tartaggia, S.; Meneghello, A.; Casetta, B.; Calia, G.; Serra, P.A.; Polo, F.; Toffoli, G. Enzyme-based electrochemical biosensor for therapeutic drug monitoring of anticancer drug irinotecan. Anal. Chem. 2018, 90, 6012–6019. [Google Scholar] [CrossRef] [PubMed]
- Babacan, S.; Pivarnik, P.; Letcher, S.; Rand, A.G. Evaluation of antibody immobilization methods for piezoelectric biosensor application. Biosens. Bioelectron. 2000, 15, 615–621. [Google Scholar] [CrossRef]
- Lec, R.M. Piezoelectric biosensor: Recent advances and applications. In Proceedings of the IEEE International Frequency Control Symposium and PDA Exhibition, Seattle, WA, USA, 8 June 2001; pp. 419–429. [Google Scholar]
- Suri, C.R.; Jain, P.K.; Mishra, G.C. Development of piezoelectric crystal based microgravimetric immunoassay for determination of insulin concentration. J. Biotechnol. 1995, 39, 27–34. [Google Scholar] [CrossRef]
- Miura, N.; Higobashi, H.; Sakai, G.; Takeyasu, A.; Uda, T.; Yamazoe, N. Piezoelectric crystal immunosensor for sensitive detection of methamphetamine (stimulant drug) in human urine. Sens. Actuators B Chem. 1993, 13, 188–191. [Google Scholar] [CrossRef]
- Halámek, J.; Makower, A.; Skládal, P.; Scheller, F.W. Highly sensitive detection of cocaine using a piezoelectric immunosensor. Biosens. Bioelectron. 2002, 17, 1045–1050. [Google Scholar] [CrossRef]
- Carrascosa, L.; Moreno, M.; Alvarez, M.; Lechuga, L. Nanomechanical biosensors: A new sensing tool. Trends Anal. Chem. 2006, 25, 196–206. [Google Scholar] [CrossRef]
- Dutta, P.; Hill, K.; Datskos, P.G.; Sepaniak, M.J. Development of a nanomechanical biosensor for analysis of endocrine disrupting chemicals. Lab A Chip 2007, 7, 1184–1191. [Google Scholar] [CrossRef] [PubMed]
- Fritz, J.; Baller, M.K.; Lang, H.P.; Rothuizen, H.; Vettiger, P.; Meyer, E.; Guntherodt, H.; Gerber, C.; Gimzewski, J.K. Translating biomolecular recognition into nanomechanics. Science 2000, 288, 316–318. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Datar, R.H.; Hansen, K.M.; Thundat, T.; Cote, R.J.; Majumdar, A. Bioassay of prostate-specific antigen (PSA) using microcantilevers. Nat. Biotechnol. 2001, 19, 856–860. [Google Scholar] [CrossRef] [PubMed]
- Gfeller, K.Y.; Nugaeva, N.; Hegner, M. Rapid biosensor for detection of antibiotic-selective growth of Escherichia coli. Appl. Environ. Microbiol. 2005, 71, 2626–2631. [Google Scholar] [CrossRef] [PubMed]
- McAleer, J.F.; Law, J.T.; Morris, R.A.; Scott, L.; Mellor, J.M.; Dennison, M. Enhanced Amperometric Sensor. U.S. Patent 5,264,106, 23 November 1993. [Google Scholar]
- Wang, J. Amperometric biosensors for clinical and therapeutic drug monitoring: A review. J. Pharm. Biomed. Anal. 1999, 19, 47–53. [Google Scholar] [CrossRef]
- Yeh, W.-M.; Ho, K.-C. Amperometric morphine sensing using a molecularly imprinted polymer-modified electrode. Anal. Chim. Acta 2005, 542, 76–82. [Google Scholar] [CrossRef]
- Khaldeeva, E.; Medyantseva, E.; Imanaeva, N.; Budnikov, G. Determination of gentamicin with an amperometric enzyme immunosensor. J. Anal. Chem. 2002, 57, 1097–1102. [Google Scholar] [CrossRef]
- Gupta, V.K.; Nayak, A.; Agarwal, S.; Singhal, B. Recent advances on potentiometric membrane sensors for pharmaceutical analysis. Comb. Chem. High Throughput Screen. 2011, 14, 284–302. [Google Scholar] [CrossRef] [PubMed]
- Shamsipur, M.; Jalali, F.; Ershad, S. Preparation of a diclofenac potentiometric sensor and its application to pharmaceutical analysis and to drug recovery from biological fluids. J. Pharm. Biomed. Anal. 2005, 37, 943–947. [Google Scholar] [CrossRef] [PubMed]
- Kulapina, E.; Snesarev, S.; Makarova, N.; Pogorelova, E. Potentiometric sensor arrays for the individual determination of penicillin class antibiotics using artificial neural networks. J. Anal. Chem. 2011, 66, 78–83. [Google Scholar] [CrossRef]
- Simpson, D.; Kobos, R. Microbiological assay of tetracycline with a potentiometric CO2 gas sensor. Anal. Lett. 1982, 15, 1345–1359. [Google Scholar] [CrossRef]
- Saber, A.L.; Elmosallamy, M.A.; Killa, H.M.; Ghoneim, M.M. Selective potentiometric method for determination of flucloxacillin antibiotic. J. Taibah Univ. Sci. 2013, 7, 195–201. [Google Scholar] [CrossRef][Green Version]
- Kamel, A.H.; Moreira, F.T.; Sales, F.; Goreti, M. Biomimetic sensor potentiometric system for doxycycline antibiotic using a molecularly imprinted polymer as an artificial recognition element. Sens. Lett. 2011, 9, 1654–1660. [Google Scholar] [CrossRef]
- Abouzar, M.H.; Poghossian, A.; Razavi, A.; Besmehn, A.; Bijnens, N.; Williams, O.A.; Haenen, K.; Wagner, P.; Schöning, M.J. Penicillin detection with nanocrystalline-diamond field-effect sensor. Phys. Status Solidi (A) 2008, 205, 2141–2145. [Google Scholar] [CrossRef]
- Aliakbarinodehi, N.; Jolly, P.; Bhalla, N.; Miodek, A.; De Micheli, G.; Estrela, P.; Carrara, S. Aptamer-based field-effect biosensor for tenofovir detection. Sci. Rep. 2017, 7, 44409. [Google Scholar] [CrossRef] [PubMed]
- Chou, J.-C.; Chiang, J.-L.; Wu, C.-L. pH and procaine sensing characteristics of extended-gate field-effect transistor based on indium tin oxide glass. Jpn. J. Appl. Phys. 2005, 44, 4838. [Google Scholar] [CrossRef]
- Milović, N.M.; Behr, J.R.; Godin, M.; Hou, C.-S.J.; Payer, K.R.; Chandrasekaran, A.; Russo, P.R.; Sasisekharan, R.; Manalis, S.R. Monitoring of heparin and its low-molecular-weight analogs by silicon field effect. Proc. Natl. Acad. Sci. USA 2006, 103, 13374–13379. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.-J.; Luo, X.-L.; Chen, H.-Y. Analytical aspects of FET-based biosensors. Front. Biosci. 2005, 10, 420–430. [Google Scholar] [CrossRef] [PubMed]
- de-los-Santos-Álvarez, N.; Lobo-Castañón, M.J.; Miranda-Ordieres, A.J.; Tuñón-Blanco, P. Modified-RNA aptamer-based sensor for competitive impedimetric assay of neomycin B. J. Am. Chem. Soc. 2007, 129, 3808–3809. [Google Scholar] [CrossRef] [PubMed]
- Paniel, N.; Istamboulié, G.; Triki, A.; Lozano, C.; Barthelmebs, L.; Noguer, T. Selection of DNA aptamers against penicillin G using Capture-SELEX for the development of an impedimetric sensor. Talanta 2017, 162, 232–240. [Google Scholar] [CrossRef] [PubMed]
- Ionescu, R.E.; Jaffrezic-Renault, N.; Bouffier, L.; Gondran, C.; Cosnier, S.; Pinacho, D.G.; Marco, M.-P.; Sánchez-Baeza, F.J.; Healy, T.; Martelet, C. Impedimetric immunosensor for the specific label free detection of ciprofloxacin antibiotic. Biosens. Bioelectron. 2007, 23, 549–555. [Google Scholar] [CrossRef] [PubMed]
- Karaseva, N.; Ermolaeva, T.; Mizaikoff, B. Piezoelectric sensors using molecularly imprinted nanospheres for the detection of antibiotics. Sens. Actuators B Chem. 2016, 225, 199–208. [Google Scholar] [CrossRef]
- Long, Y.; Nie, L.; Chen, J.; Yao, S. Piezoelectric quartz crystal impedance and electrochemical impedance study of HSA–diazepam interaction by nanogold-structured sensor. J. Colloid Interface Sci. 2003, 263, 106–112. [Google Scholar] [CrossRef]
- Skládal, P. Piezoelectric biosensors. TrAC Trends Anal. Chem. 2016, 79, 127–133. [Google Scholar] [CrossRef]
- Wu, S.; Liu, X.; Zhou, X.; Liang, X.M.; Gao, D.; Liu, H.; Zhao, G.; Zhang, Q.; Wu, X. Quantification of cell viability and rapid screening anti-cancer drug utilizing nanomechanical fluctuation. Biosens. Bioelectron. 2016, 77, 164–173. [Google Scholar] [CrossRef] [PubMed]
- Ndieyira, J.W.; Watari, M.; Barrera, A.D.; Zhou, D.; Vögtli, M.; Batchelor, M.; Cooper, M.A.; Strunz, T.; Horton, M.A.; Abell, C. Nanomechanical detection of antibiotic–mucopeptide binding in a model for superbug drug resistance. Nat. Nanotechnol. 2008, 3, 691. [Google Scholar] [CrossRef] [PubMed]
- Regatos, D. Biosensores ópticos de alta sensibilidad basados en técnicas de modulación plasmónica; Universidad de Santiago de Compostela: Barcelona, Spain, 2012. [Google Scholar]
- Damborsky, P.; Svitel, J.; Katrlik, J. Optical biosensors. Essays Biochem. 2016, 60, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Dey, D.; Goswami, T. Optical biosensors: A revolution towards quantum nanoscale electronics device fabrication. J. Biomed. Biotechnol. 2011, 2011, 348218. [Google Scholar] [CrossRef] [PubMed]
- Losoya-Leal, A.; Estevez, M.-C.; Martínez-Chapa, S.O.; Lechuga, L.M. Design of a surface plasmon resonance immunoassay for therapeutic drug monitoring of amikacin. Talanta 2015, 141, 253–258. [Google Scholar] [CrossRef] [PubMed]
- Yola, M.L.; Atar, N.; Eren, T. Determination of amikacin in human plasma by molecular imprinted SPR nanosensor. Sens. Actuators B Chem. 2014, 198, 70–76. [Google Scholar] [CrossRef]
- Salvati, E.; Stellacci, F.; Krol, S. Nanosensors for early cancer detection and for therapeutic drug monitoring. Nanomedicine 2015, 10, 3495–3512. [Google Scholar] [CrossRef] [PubMed]
- Hon, Y.Y.; Evans, W.E. Making TDM work to optimize cancer chemotherapy: A multidisciplinary team approach. Clin. Chem. 1998, 44, 388–400. [Google Scholar] [PubMed]
- Gao, B.; Yeap, S.; Clements, A.; Balakrishnar, B.; Wong, M.; Gurney, H. Evidence for therapeutic drug monitoring of targeted anticancer therapies. J. Clin. Oncol. 2012, 30, 4017–4025. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.S.; Bukar, N.; Toulouse, J.L.; Pelechacz, D.; Robitaille, R.; Pelletier, J.N.; Masson, J.-F. Miniature multi-channel SPR instrument for methotrexate monitoring in clinical samples. Biosens. Bioelectron. 2015, 64, 664–670. [Google Scholar] [CrossRef] [PubMed]
- Masson, J.-F.; Pelletier, J.N. Will nanobiosensors change therapeutic drug monitoring? The case of methotrexate. Nanomedicine 2015, 10, 521–524. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.S.; Bichelberger, M.A.; Colin, D.Y.; Robitaille, R.; Pelletier, J.N.; Masson, J.-F. Monitoring methotrexate in clinical samples from cancer patients during chemotherapy with a LSPR-based competitive sensor. Analyst 2012, 137, 4742–4750. [Google Scholar] [CrossRef] [PubMed]
- Kivirand, K.; Floren, A.; Kagan, M.; Avarmaa, T.; Rinken, T.; Jaaniso, R. Analyzing the biosensor signal in flows: Studies with glucose optrodes. Talanta 2015, 131, 74–80. [Google Scholar] [CrossRef] [PubMed]
- Hutchinson, A.M. Evanescent wave biosensors. Real-time analysis of biomolecular interactions. Mol. Biotechnol. 1995, 3, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Leung, A.; Shankar, P.; Mutharasan, R. A review of fiber-optic biosensors. Sens. Actuators B Chem. 2007, 125, 688–703. [Google Scholar] [CrossRef]
- Ogert, R.A.; Brown, J.E.; Singh, B.R.; Shriver-Lake, L.C.; Ligler, F.S. Detection of Clostridium botulinum toxin A using a fiber optic-based biosensor. Anal. Biochem. 1992, 205, 306–312. [Google Scholar] [CrossRef]
- Eivazzadeh-Keihan, R.; Pashazadeh-Panahi, P.; Baradaran, B.; de la Guardia, M.; Hejazi, M.; Sohrabi, H.; Mokhtarzadeh, A.; Maleki, A. Recent progress in optical and electrochemical biosensors for sensing of Clostridium botulinum neurotoxin. TRAC Trends Anal. Chem. 2018, 103, 184–197. [Google Scholar] [CrossRef]
- Preejith, P.V.; Lim, C.S.; Kishen, A.; John, M.S.; Asundi, A. Total protein measurement using a fiber-optic evanescent wave-based biosensor. Biotechnol. Lett. 2003, 25, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Astles, J.R.; Miller, W.G. Measurement of free phenytoin in blood with a self-contained fiber-optic immunosensor. Anal. Chem. 1994, 66, 1675–1682. [Google Scholar] [CrossRef] [PubMed]
- Hanbury, C.M.; Miller, W.G.; Harris, R.B. Antibody characteristics for a continuous response fiber optic immunosensor for theophylline. Biosens. Bioelectron. 1996, 11, 1129–1138. [Google Scholar] [CrossRef]
- Arjmand, M.; Saghafifar, H.; Alijanianzadeh, M.; Soltanolkotabi, M. A sensitive tapered-fiber optic biosensor for the label-free detection of organophosphate pesticides. Sens. Actuators B Chem. 2017, 249, 523–532. [Google Scholar] [CrossRef]
- Biran, I.; Rissin, D.M.; Ron, E.Z.; Walt, D.R. Optical imaging fiber-based live bacterial cell array biosensor. Anal. Biochem. 2003, 315, 106–113. [Google Scholar] [CrossRef]
- Sun, W.; Yuan, S.; Huang, H.; Liu, N.; Tan, Y. A label-free biosensor based on localized surface plasmon resonance for diagnosis of tuberculosis. J. Microbiol. Methods 2017, 142, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Bustos, R.H.; Magdy Sanchez, M.; Dominguez, M.A.; Barreto, G.E.; Lancheros, D.; Reynolds, J. Nanotechnology in Neurosciences: An Approach. Curr. Pharm. Des. 2017, 23, 4154–4169. [Google Scholar] [CrossRef] [PubMed]
- Nie, W.; Wang, Q.; Yang, X.; Zhang, H.; Li, Z.; Gao, L.; Zheng, Y.; Liu, X.; Wang, K. High sensitivity surface plasmon resonance biosensor for detection of microRNA based on gold nanoparticles-decorated molybdenum sulfide. Anal. Chim. Acta 2017, 993, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Guo, X. Surface plasmon resonance based biosensor technique: A review. J. Biophotonics 2012, 5, 483–501. [Google Scholar] [CrossRef] [PubMed]
- Cooper, M.A.; Fiorini, M.T.; Abell, C.; Williams, D.H. Binding of vancomycin group antibiotics to D-alanine and D-lactate presenting self-assembled monolayers. Bioorg. Med. Chem. 2000, 8, 2609–2616. [Google Scholar] [CrossRef]
- Rao, J.; Yan, L.; Xu, B.; Whitesides, G.M. Using surface plasmon resonance to study the binding of vancomycin and its dimer to self-assembled monolayers presenting D-Ala-D-Ala. J. Am. Chem. Soc. 1999, 121, 2629–2630. [Google Scholar] [CrossRef]
- Luo, Q.; Yu, N.; Shi, C.; Wang, X.; Wu, J. Surface plasmon resonance sensor for antibiotics detection based on photo-initiated polymerization molecularly imprinted array. Talanta 2016, 161, 797–803. [Google Scholar] [CrossRef] [PubMed]
- Tomassetti, M.; Conta, G.; Campanella, L.; Favero, G.; Sanzò, G.; Mazzei, F.; Antiochia, R. A flow SPR immunosensor based on a sandwich direct method. Biosensors 2016, 6, 22. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Qu, C.; Kuang, H.; Xu, L.; Liu, L.; Hua, Y.; Wang, L.; Xu, C. Simple, rapid and sensitive detection of antibiotics based on the side-by-side assembly of gold nanorod probes. Biosens. Bioelectron. 2011, 26, 4387–4392. [Google Scholar] [CrossRef] [PubMed]
- de-los-Santos-Álvarez, N.; Lobo-Castañón, M.J.; Miranda-Ordieres, A.J.; Tuñón-Blanco, P. SPR sensing of small molecules with modified RNA aptamers: Detection of neomycin B. Biosens. Bioelectron. 2009, 24, 2547–2553. [Google Scholar] [CrossRef] [PubMed]
- Nieciecka, D.; Krysinski, P. Interactions of doxorubicin with self-assembled monolayer-modified electrodes: Electrochemical, surface plasmon resonance (SPR), and gravimetric studies. Langmuir Acs J. Surf. Colloids 2011, 27, 1100–1107. [Google Scholar] [CrossRef] [PubMed]
- Rusnati, M.; Bugatti, A. Surface plasmon resonance analysis of heparin-binding angiogenic growth factors. In Tumor Angiogenesis Assays; Springer: Berlin, Germany, 2016; pp. 73–84. [Google Scholar]
- Dillon, P.P.; Daly, S.J.; Manning, B.M.; O’Kennedy, R. Immunoassay for the determination of morphine-3-glucuronide using a surface plasmon resonance-based biosensor. Biosens. Bioelectron. 2003, 18, 217–227. [Google Scholar] [CrossRef]
- Sankiewicz, A.; Puzan, B.; Gorodkiewicz, E. Biosensors SPRI as a diagnostic tool in the future. Chemik 2014, 68, 528–535. [Google Scholar]
- Safina, G. Application of surface plasmon resonance for the detection of carbohydrates, glycoconjugates, and measurement of the carbohydrate-specific interactions: A comparison with conventional analytical techniques. A critical review. Anal. Chim. Acta 2012, 712, 9–29. [Google Scholar] [CrossRef] [PubMed]
- Guner, H.; Ozgur, E.; Kokturk, G.; Celik, M.; Esen, E.; Topal, A.E.; Ayas, S.; Uludag, Y.; Elbuken, C.; Dana, A. A smartphone based surface plasmon resonance imaging (SPRi) platform for on-site biodetection. Sens. Actuators B Chem. 2017, 239, 571–577. [Google Scholar] [CrossRef]
- Cottat, M.; Thioune, N.; Gabudean, A.; Lidgi-Guigui, N.; Focsan, M.; Astilean, S.; De la Chapelle, M. Localized surface plasmon resonance (LSPR) biosensor for the protein detection. Plasmonics 2013, 8, 699–704. [Google Scholar] [CrossRef]
- Cappi, G.; Spiga, F.M.; Moncada, Y.; Ferretti, A.; Beyeler, M.; Bianchessi, M.; Decosterd, L.; Buclin, T.; Guiducci, C. Label-free detection of tobramycin in serum by transmission-localized surface plasmon resonance. Anal. Chem. 2015, 87, 5278–5285. [Google Scholar] [CrossRef] [PubMed]
- Caglayan, M.G.; Onur, F. Silver nanoparticle based analysis of aminoglycosides. Spectrosc. Lett. 2014, 47, 771–780. [Google Scholar] [CrossRef]
- McKeating, K.S.; Couture, M.; Dinel, M.-P.; Garneau-Tsodikova, S.; Masson, J.-F. High throughput LSPR and SERS analysis of aminoglycoside antibiotics. Analyst 2016, 141, 5120–5126. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Yin, Y.; Huang, R.; Qiu, B.; Lin, Z.; Yang, H.-H.; Li, J.; Chen, G. Enantioselective analysis of melagatran via an LSPR biosensor integrated with a microfluidic chip. Lab A Chip 2012, 12, 3901–3906. [Google Scholar] [CrossRef] [PubMed]
- Peláez-Gutierrez, E.C.; Estévez, M.C.; Salvador, J.; Marco, M.; Lechuga, L.M. Localised Surface Plasmon Resonance Biosensor for the Monitoring of sintrom® Therapeutic Drug in Plasma. Available online: https://digital.csic.es/handle/10261/161311 (accessed on 1 November 2019).
- Nikfarjam, A.; Rezayan, A.H.; Mohammadkhani, G.; Mohammadnejad, J. Label-free detection of digoxin using localized surface plasmon resonance-based nanobiosensor. Plasmonics 2017, 12, 157–164. [Google Scholar] [CrossRef]
- Yang, J.; Tan, X.; Shih, W.-C.; Cheng, M.M.-C. A sandwich substrate for ultrasensitive and label-free SERS spectroscopic detection of folic acid/methotrexate. Biomed. Microdevices 2014, 16, 673–679. [Google Scholar] [CrossRef] [PubMed]
- Farquharson, S.; Shende, C.; Inscore, F.E.; Maksymiuk, P.; Gift, A. Analysis of 5-fluorouracil in saliva using surface-enhanced Raman spectroscopy. J. Raman Spectrosc. Int. J. Orig. Work All Asp. Raman Spectrosc. Incl. High. Order Process. Brillouin Rayleigh Scatt. 2005, 36, 208–212. [Google Scholar] [CrossRef]
- El-Zahry, M.R.; Refaat, I.H.; Mohamed, H.A.; Rosenberg, E.; Lendl, B. Utility of surface enhanced Raman spectroscopy (SERS) for elucidation and simultaneous determination of some penicillins and penicilloic acid using hydroxylamine silver nanoparticles. Talanta 2015, 144, 710–716. [Google Scholar] [CrossRef] [PubMed]
- Tschmelak, J.; Kumpf, M.; Kappel, N.; Proll, G.; Gauglitz, G. Total internal reflectance fluorescence (TIRF) biosensor for environmental monitoring of testosterone with commercially available immunochemistry: Antibody characterization, assay development and real sample measurements. Talanta 2006, 69, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Kappel, N.D.; Proll, F.; Gauglitz, G. Development of a TIRF-based biosensor for sensitive detection of progesterone in bovine milk. Biosens. Bioelectron. 2007, 22, 2295–2300. [Google Scholar] [CrossRef] [PubMed]
- Tschmelak, J.; Proll, G.; Gauglitz, G. Verification of performance with the automated direct optical TIRF immunosensor (River Analyser) in single and multi-analyte assays with real water samples. Biosens. Bioelectron. 2004, 20, 743–752. [Google Scholar] [CrossRef] [PubMed]
- Ehrentreich-Forster, E.; Orgel, D.; Krause-Griep, A.; Cech, B.; Erdmann, V.A.; Bier, F.; Scheller, F.W.; Rimmele, M. Biosensor-based on-site explosives detection using aptamers as recognition elements. Anal. Bioanal. Chem. 2008, 391, 1793–1800. [Google Scholar] [CrossRef] [PubMed]
- Adinolfi, B.; Baldini, F.; Berrettoni, C.; Berneschi, S.; Giannetti, A.; Tombelli, S.; Trono, C.; Bernini, R.; Grimaldi, I.A.; Persichetti, G. Total internal reflection fluorescence-based optical biochip for the detection of immunosuppressants in transplanted patients. In Proceedings of the 2015 1st Workshop on Nanotechnology in Instrumentation and Measurement (NANOFIM), Lecce, Italy, 24–25 July 2015; pp. 39–42. [Google Scholar]
- Klinth, J.; Larsson, R.; Andersson, P.; Ekdahl, K.N. A novel application of multi-wavelength TIRF spectroscopy for real time monitoring of antithrombin interactions with immobilized heparin. Biosens. Bioelectron. 2006, 21, 1973–1980. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, K.; Schirmer, B.; Hoffmann, C.; Brandenburg, A.; Meyrueis, P. Interferometric biosensor based on planar optical waveguide sensor chips for label-free detection of surface bound bioreactions. Biosens. Bioelectron. 2007, 22, 2591–2597. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Suarez, D.; Gottfried, D.S. Detection of avian influenza virus using an interferometric biosensor. Anal. Bioanal. Chem. 2007, 389, 1193–1199. [Google Scholar] [CrossRef] [PubMed]
- Fei, Y.; Sun, Y.S.; Li, Y.; Yu, H.; Lau, K.; Landry, J.P.; Luo, Z.; Baumgarth, N.; Chen, X.; Zhu, X. Characterization of receptor binding profiles of influenza a viruses using an ellipsometry-based label-free glycan microarray assay platform. Biomolecules 2015, 5, 1480–1498. [Google Scholar] [CrossRef] [PubMed]
- Kumeria, T.; Kurkuri, M.D.; Diener, K.R.; Parkinson, L.; Losic, D. Label-free reflectometric interference microchip biosensor based on nanoporous alumina for detection of circulating tumour cells. Biosens. Bioelectron. 2012, 35, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Proll, G.; Markovic, G.; Fechner, P.; Proell, F.; Gauglitz, G. Reflectometric interference spectroscopy. Methods Mol. Biol. 2017, 1571, 207–220. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Campbell, A.S.; de Ávila, B.E.-F.; Wang, J. Wearable biosensors for healthcare monitoring. Nat. Biotechnol. 2019, 37, 389–406. [Google Scholar] [CrossRef] [PubMed]
- Elsherif, M.; Hassan, M.U.; Yetisen, A.K.; Butt, H. Wearable contact lens biosensors for continuous glucose monitoring using smartphones. ACS Nano 2018, 12, 5452–5462. [Google Scholar] [CrossRef] [PubMed]
- Rachim, V.P.; Chung, W.-Y. Wearable-band type visible-near infrared optical biosensor for non-invasive blood glucose monitoring. Sens. Actuators B Chem. 2019, 286, 173–180. [Google Scholar] [CrossRef]
- Fu, E.; Chinowsky, T.; Nelson, K.; Johnston, K.; Edwards, T.; Helton, K.; Grow, M.; Miller, J.W.; Yager, P. SPR imaging-based salivary diagnostics system for the detection of small molecule analytes. Ann. New York Acad. Sci. 2007, 1098, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Shriver-Lake, L.C.; Donner, B.L.; Ligler, F.S. On-site detection of TNT with a portable fiber optic biosensor. Environ. Sci. Technol. 1997, 31, 837–841. [Google Scholar] [CrossRef]
- Mauriz, E.; Calle, A.; Montoya, A.; Lechuga, L.M. Determination of environmental organic pollutants with a portable optical immunosensor. Talanta 2006, 69, 359–364. [Google Scholar] [CrossRef] [PubMed]
- Hildebrandt, A.; Bragos, R.; Lacorte, S.; Marty, J. Performance of a portable biosensor for the analysis of organophosphorus and carbamate insecticides in water and food. Sens. Actuators B Chem. 2008, 133, 195–201. [Google Scholar] [CrossRef]
- Fernández, F.; Pinacho, D.G.; Sánchez-Baeza, F.; Marco, M.P. Portable surface plasmon resonance immunosensor for the detection of fluoroquinolone antibiotic residues in milk. J. Agric. Food Chem. 2011, 59, 5036–5043. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, S.; Ibrahim, F.; Djordjevic, I.; Koole, L.H. Recent advances in surface functionalization techniques on polymethacrylate materials for optical biosensor applications. Analyst 2014, 139, 2933–2943. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, H.; Huang, J.; Yang, J.; Liu, B.; Yang, P. Microchip-based ELISA strategy for the detection of low-level disease biomarker in serum. Anal. Chim. Acta 2009, 650, 77–82. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Shamsi, M.H. Biosensors-on-chip: A topical review. J. Micromech. Microengin. 2017, 27, 083001. [Google Scholar] [CrossRef]
- Noh, J.; Kim, H.C.; Chung, T.D. Biosensors in microfluidic chips. Top. Curr. Chem. 2011, 304, 117–152. [Google Scholar] [CrossRef] [PubMed]
- Ranamukhaarachchi, S.A.; Padeste, C.; Dübner, M.; Häfeli, U.O.; Stoeber, B.; Cadarso, V.J. Integrated hollow microneedle-optofluidic biosensor for therapeutic drug monitoring in sub-nanoliter volumes. Sci. Rep. 2016, 6, 29075. [Google Scholar] [CrossRef] [PubMed]
- Masson, J.-F.; Zhao, S.S.; Bukar, N.; Pelletier, J.N.; Labrecque-Carbonneau, J.; McKeating, K.; Yockell-Lelièvre, H. Surface plasmon resonance (SPR) sensing for small molecules in biofluids. In Proceedings of the Advanced Photonics 2015, Boston, MA, USA, 27 June–1 July 2015; p. SeW1B.2. [Google Scholar]
- Beeg, M.; Nobili, A.; Orsini, B.; Rogai, F.; Gilardi, D.; Fiorino, G.; Danese, S.; Salmona, M.; Garattini, S.; Gobbi, M. A surface plasmon resonance-based assay to measure serum concentrations of therapeutic antibodies and anti-drug antibodies. Sci. Rep. 2019, 9, 2064. [Google Scholar] [CrossRef] [PubMed]
- Bhalla, N.; Jamshaid, A.; Leung, M.H.M.; Ishizu, N.; Shen, A.Q. Electrical contact of metals at the nanoscale overcomes the oxidative susceptibility of silver-based nanobiosensors. ACS Appl. Nano Mater. 2019, 2, 2064–2075. [Google Scholar] [CrossRef]
| Biosensor | Measurement | Graph | TDM Applications | Ref |
|---|---|---|---|---|
| Electrochemical | ||||
| Amperometric | Measuring current flows produced by an electrochemical reaction | Amperogram | Morphine, Metotrexate, Gentamycin, Tamoxifen, Gemcitabine, Didanosine, Irinotecan, Cyclophosphamide, Ifosfamide, Ftorafur, Etoposide | [69,70,73,75,76,94,95,96,97] |
| Potentiometric | Measuring variations potential (VP) on the electrode’s surface | Potentiogram | Diclofenac, Penicillin, Tetracycline, Flucloxacillin, Doxycycline, Methotrexate, Cisplatin, Titanocene dichloride | [79,98,99,100,101,102,103] |
| Field-effect transistor-based biosensor (FET) | Measuring variations in the current on the sensor’s surface | Time vs. current | Penicillins, Tenofovir, Procaine, Heparin, Imatinib | [82,104,105,106,107,108] |
| Impedimetric | Measuring changes via the impedance between electrodes or the perturbation caused by electrolytes/electrodes | Impedance graph | Neomycin, Penicillin, Ciprofloxacin, Bleomycin, Mitomycin C | [65,71,72,109,110,111] |
| Piezoelectric | ||||
| Quartz crystal microbalance (QCM) | Measuring the vibration frequency and displacement producing changes in electric current | Frequency variation | Penicillins, Sulfamides, Diazepam | [87,112,113,114] |
| Nanomechanical | ||||
| Nanocantilevers | Measuring the cantilever flexion when a molecular interaction occurs on a surface that becomes a nanomechanical movement | Time vs. deflection | Paclitaxel, Vancomycin | [89,115,116] |
| Type of Biosensor | Type of Drug | Drug | Biosensor Characteristics | Matrix | Limit/Detection Range | Results | Ref. |
|---|---|---|---|---|---|---|---|
| Fibre optic | Anticonvulsant | Phenytoin | Autonomous reversible immunosensor: mouse monoclonal IgG | Blood and plasma | 4.45 µM | Viability for quantifying phenytoin in blood, applicable for other haptens in blood | [134] |
| Bronchodilator | Theophylline | Fluorescence-based autonomous reversible immunosensor: mouse monoclonal IgG | Serum | 55 µM | Analyte concentrations give rise to a change in the antibody binding equilibrium with changes in fluorescence | [135] | |
| SPR | Antibiotic | Vancomycin and Chloroeremomycin | Covalent bacterial wall peptide coupling to a self-assembled monolayer (SAM) on a gold film | Solution buffer | 20 mM 2.5 mM | Chloroeremomycin is related to bacterial wall peptides, thereby facilitating quantification | [142,143] |
| Ciprofloxacin | SPR with a molecularly imprinted polymer (MIP) | Solution buffer | 0.08 µg/L | Sensitive technique for quantifying this type of molecule | [144] | ||
| Ampicillin | SPR operated in flow conditions | Solution buffer | 10−3 M to 10−1 M | Technique requiring less time (20 min) without losing sensitivity | [145] | ||
| Gentamycin | SPR with a Doppler laser using UV-Vis spectroscopy | Solution buffer | 0.05 ng/mL | Lower detection limit compared to ELISA | [146] | ||
| Anticancer | MTX | LSPR with functionalized gold nanoparticles with folic acid (FA-AuNPs) in completion with MTX | Serum | 28 nM | Lower detection limit than that reported for LSPR biosensors (155 Nm) | [125] | |
| Anticoagulant | Heparin | Using prolamine and polyethyleneimine as affinity surface | Plasma | 0.2 U/mL | Lower detection limit than that found for previously cited techniques | [149] | |
| Opioid | Morphine | Immunoassays using polyclonal antibodies from New Zealand rabbits | Urine | 762–24,4000 pg/mL | This technique enables the sensitive and specific quantification of different opioids such as heroin and morphine | [150] | |
| LSPR | Anticancer | MTX | LSPR with functionalized gold nanoparticles with folic acid (FA-AuNPs) | Plasma | 155 nM | This technique provides a new index for quantifying this drug by this type of biosensor | [127] |
| Antibiotic | Tobramycin | Transmission localized surface plasmon resonance (T-LSPR), using DNA aptamers | Serum | 0.34 µM | This modification enables the direct detection (without using labels) of a small molecule in a complex matrix | [155] | |
| Anticoagulant | Megalatran | LSPR integrated with a microfluidic chip | Solution buffer | 0.9 nM | A pioneering study regarding the use of enantioselective biosensors | [141] | |
| Anti-arrhythmic | Digoxin | LSPR with gold nanoparticles | Solution buffer | 2 ng/mL | This device enables the direct low-cost detection of digoxin, as well as being a device that is easy to make and use | [160] | |
| SERS | Anticancer | 5-fluorouracil | SERS with silver nanoparticles | Saliva | 150 ng/mL | This study provides a great opportunity since it enables one to quantify a highly toxic drug with genetic variations in its metabolism | [162] |
| Antibiotic | Ampicillin | SERS with silver on nanoparticles using hydroxylamine—HCl | Solution buffer | 27 ng/mL | This technique has been compared to LC/MS, with greater sensitivity. It provides an index for quantifying drugs with this type of device | [163] | |
| Penicillin G | 29 ng/mL | ||||||
| Carbenicillin | 30 ng/mL | ||||||
| Penicilloic acid | 28 ng/mL |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garzón, V.; Pinacho, D.G.; Bustos, R.-H.; Garzón, G.; Bustamante, S. Optical Biosensors for Therapeutic Drug Monitoring. Biosensors 2019, 9, 132. https://doi.org/10.3390/bios9040132
Garzón V, Pinacho DG, Bustos R-H, Garzón G, Bustamante S. Optical Biosensors for Therapeutic Drug Monitoring. Biosensors. 2019; 9(4):132. https://doi.org/10.3390/bios9040132
Chicago/Turabian StyleGarzón, Vivian, Daniel G. Pinacho, Rosa-Helena Bustos, Gustavo Garzón, and Sandra Bustamante. 2019. "Optical Biosensors for Therapeutic Drug Monitoring" Biosensors 9, no. 4: 132. https://doi.org/10.3390/bios9040132
APA StyleGarzón, V., Pinacho, D. G., Bustos, R.-H., Garzón, G., & Bustamante, S. (2019). Optical Biosensors for Therapeutic Drug Monitoring. Biosensors, 9(4), 132. https://doi.org/10.3390/bios9040132





