Functionalized Protein Nanotubes Based on the Bacteriophage vB_KleM-RaK2 Tail Sheath Protein
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Gene Construction
2.3. Expression and Purification of 041 and Chimeras
2.4. Transmission Electron Microscopy (TEM)
2.5. Fluorescence Microscopy
2.6. Amidohydrolase Activity Assay
2.7. Bioinformatics Analysis
3. Results
3.1. Recombinant 041 Protein Analysis
3.2. Chimeric Protein Construction
3.3. Analysis of 041Δ200GFP and 041Δ200mCherry Nanotubes
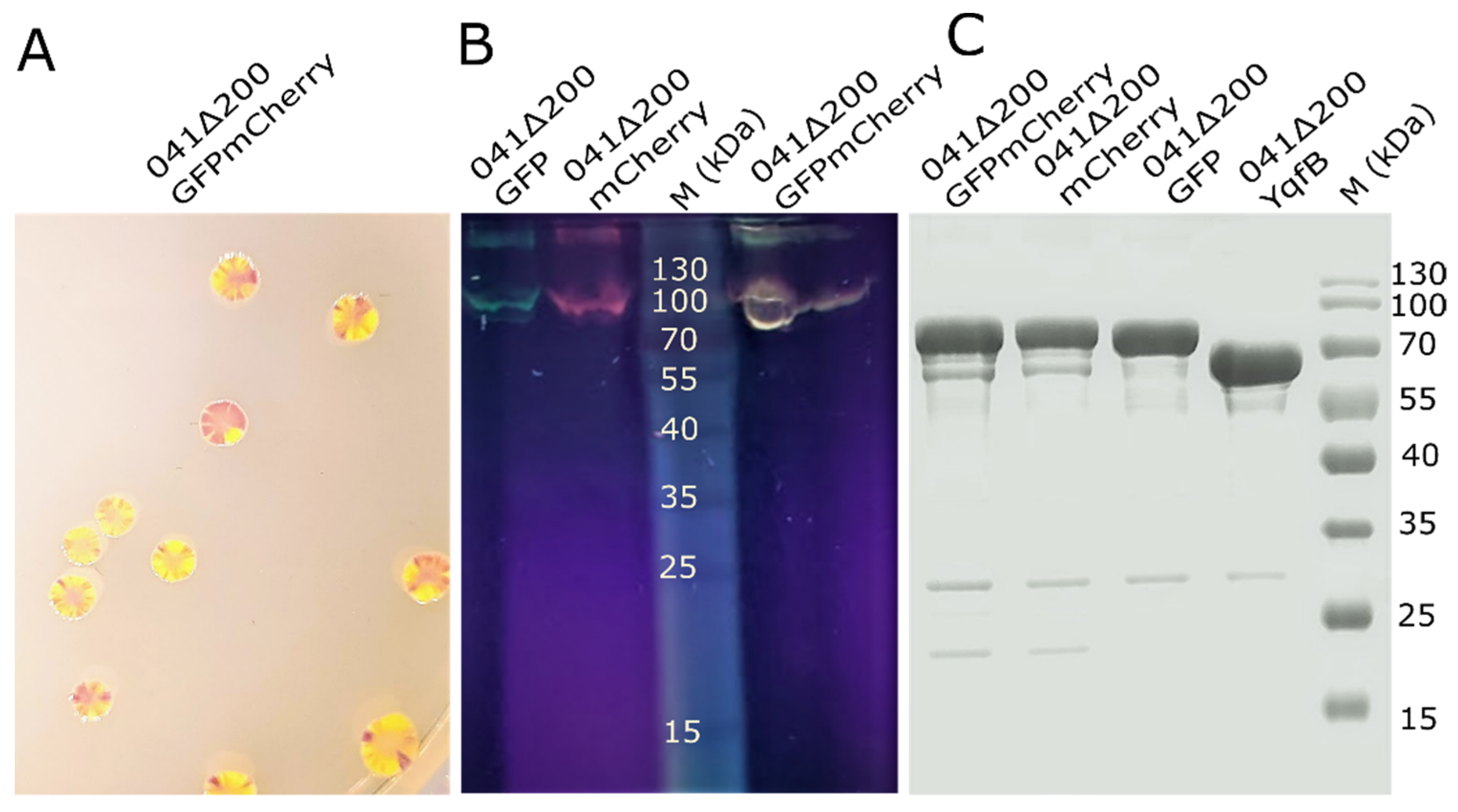
3.4. Analysis of 041Δ200GFPmCherry Hybrid Nanotubes
3.5. Analysis of 041Δ200YqfB Nanotubes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, M.; Fu, X.; Chen, Z.; Liu, J.; Zhong, W. Protein-Engineered Functional Materials for Bioelectronics. Adv. Funct. Mater. 2021, 31, 2006744. [Google Scholar] [CrossRef]
- Malisauskas, M.; Meskys, R.; Morozova-Roche, L.A. Ultrathin silver nanowires produced by amyloid biotemplating. Biotechnol. Prog. 2008, 24, 1166–1170. [Google Scholar] [CrossRef]
- Poviloniene, S.; Časaite, V.; Bukauskas, V.; Šetkus, A.; Staniulis, J.; Meškys, R. Functionalization of α-synuclein fibrils. Beilstein J. Nanotechnol. 2015, 6, 124–133. [Google Scholar] [CrossRef] [Green Version]
- Linares, R.; Arnaud, C.A.; Degroux, S.; Schoehn, G.; Breyton, C. Structure, function and assembly of the long, flexible tail of siphophages. Curr. Opin. Virol. 2020, 45, 34–42. [Google Scholar] [CrossRef]
- Paczesny, J.; Bielec, K. Application of Bacteriophages in Nanotechnology. Nanomaterials 2020, 10, 1944. [Google Scholar] [CrossRef] [PubMed]
- Cuenca, S.; Mansilla, C.; Aguado, M.; Yuste-Calvo, C.; Sánchez, F.; Sánchez-Montero, J.M.; Ponz, F. Nanonets derived from turnip mosaic virus as scaffolds for increased enzymatic activity of immobilized Candida antarctica lipase B. Front. Plant Sci. 2016, 7, 464. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koch, C.; Wabbel, K.; Eber, F.J.; Krolla-Sidenstein, P.; Azucena, C.; Gliemann, H.; Eiben, S.; Geiger, F.; Wege, C. Modified TMV particles as beneficial scaffolds to present sensor enzymes. Front. Plant Sci. 2015, 6, 1137. [Google Scholar] [CrossRef]
- Lim, A.A.L.; Tachibana, S.; Watanabe, Y.; Wong, S.M. Expression and purification of a neuropeptide nocistatin using two related plant viral vectors. Gene 2002, 289, 69–79. [Google Scholar] [CrossRef]
- Sugiyama, Y.; Hamamoto, H.; Takemoto, S.; Watanabe, Y.; Okada, Y. Systemic production of foreign peptides on the particle surface of tobacco mosaic virus. FEBS Lett. 1995, 359, 247–250. [Google Scholar] [CrossRef] [Green Version]
- Fitchen, J.; Beachy, R.N.; Hein, M.B. Plant virus expressing hybrid coat protein with added murine epitope elicits autoantibody response. Vaccine 1995, 13, 1051–1057. [Google Scholar] [CrossRef]
- Gamper, C.; Spenlé, C.; Boscá, S.; Van Der Heyden, M.; Erhardt, M.; Orend, G.; Bagnard, D.; Heinlein, M. Functionalized Tobacco Mosaic Virus Coat Protein Monomers and Oligomers as Nanocarriers for Anti-Cancer Peptides. Cancers 2019, 11, 1609. [Google Scholar] [CrossRef] [Green Version]
- Smith, M.L.; Lindbo, J.A.; Dillard-Telm, S.; Brosio, P.M.; Lasnik, A.B.; McCormick, A.A.; Nguyen, L.V.; Palmer, K.E. Modified Tobacco mosaic virus particles as scaffolds for display of protein antigens for vaccine applications. Virology 2006, 348, 475–488. [Google Scholar] [CrossRef] [Green Version]
- Carette, N.; Engelkamp, H.; Akpa, E.; Pierre, S.J.; Cameron, N.R.; Christianen, P.C.M.; Maan, J.C.; Thies, J.C.; Weberskirch, R.; Rowan, A.E.; et al. A virus-based biocatalyst. Nat. Nanotechnol. 2007, 2, 226–229. [Google Scholar] [CrossRef] [PubMed]
- Wi, S.; Hwang, I.S.; Jo, B.H. Engineering a Plant Viral Coat Protein for in Vitro Hybrid Self-Assembly of CO2-Capturing Catalytic Nanofilaments. Biomacromolecules 2020, 21, 3847–3856. [Google Scholar] [CrossRef]
- Bustos-Jaimes, I.; Soto-Román, R.A.; Gutiérrez-Landa, I.A.; Valadez-García, J.; Segovia-Trinidad, C.L. Construction of protein-functionalized virus-like particles of parvovirus B19. J. Biotechnol. 2017, 263, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Špakova, A.; Dalgėdienė, I.; Insodaitė, R.; Sasnauskienė, A.; Žvirblienė, A.; Petraitytė-Burneikienė, R. vB_EcoS_NBD2 bacteriophage-originated polytubes as a carrier for the presentation of foreign sequences. Virus Res. 2020, 290, 198194. [Google Scholar] [CrossRef] [PubMed]
- Špakova, A.; Šimoliunas, E.; Raminta, B.; Pajeda, S.; Meškys, R.; Petraitytė-Burneikienė, R. vB _ EcoS _ NBD2 into Extremely Long Polytubes in. Viruses 2019, 11, 208. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhardwaj, A.; Walker-Kopp, N.; Wilkens, S.; Cingolani, G. Foldon-guided self-assembly of ultra-stable protein fibers. Protein Sci. 2008, 17, 1475–1485. [Google Scholar] [CrossRef] [Green Version]
- Hyman, P.; Valluzzi, R.; Goldberg, E. Design of protein struts for self-assembling nanoconstructs. Proc. Natl. Acad. Sci. USA 2002, 99, 8488–8493. [Google Scholar] [CrossRef] [Green Version]
- Moody, M.F. Structure of the sheath of bacteriophage T4. II. Rearrangement of the sheath subunits during contraction. J. Mol. Biol. 1967, 25, 201–208. [Google Scholar] [CrossRef]
- Simoliunas, E.; Kaliniene, L.; Truncaite, L.; Klausa, V.; Zajanckauskaite, A.; Meskys, R. Genome of Klebsiella sp.-Infecting Bacteriophage vB_KleM_RaK2. J. Virol. 2012, 86, 5406. [Google Scholar] [CrossRef] [Green Version]
- Šimoliunas, E.; Kaliniene, L.; Truncaite, L.; Zajančkauskaite, A.; Staniulis, J.; Kaupinis, A.; Ger, M.; Valius, M.; Meškys, R. Klebsiella Phage vB_KleM-RaK2—A Giant Singleton Virus of the Family Myoviridae. PLoS ONE 2013, 8, e60717. [Google Scholar] [CrossRef] [Green Version]
- Chirico, G.; Cannone, F.; Beretta, S.; Diaspro, A.; Campanini, B.; Bettati, S.; Ruotolo, R.; Mozzarelli, A. Dynamics of green fluorescent protein mutant2 in solution, on spin-coated glasses, and encapsulated in wet silica gels. Protein Sci. 2002, 11, 1152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shaner, N.C.; Campbell, R.E.; Steinbach, P.A.; Giepmans, B.N.G.; Palmer, A.E.; Tsien, R.Y. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 2004, 22, 1567–1572. [Google Scholar] [CrossRef]
- LAEMMLI, U.K. Cleavage of Structural Proteins during the Assembly of the Head of Bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Šimoliūnas, E.; Truncaitė, L.; Rutkienė, R.; Povilonienė, S.; Goda, K.; Kaupinis, A.; Valius, M.; Meškys, R. The robust self-assembling tubular nanostructures formed by gp053 from phage vb_ecom_fv3. Viruses 2019, 11, 50. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gabler, F.; Nam, S.Z.; Till, S.; Mirdita, M.; Steinegger, M.; Söding, J.; Lupas, A.N.; Alva, V. Protein Sequence Analysis Using the MPI Bioinformatics Toolkit. Curr. Protoc. Bioinform. 2020, 72, e108. [Google Scholar] [CrossRef]
- Huang, T.-T.; Hwang, J.-K.; Chen, C.-H.; Chu, C.-S.; Lee, C.-W.; Chen, C.-C. (PS)2: Protein structure prediction server version 3.0. Nucleic Acids Res. 2015, 43, W338–W342. [Google Scholar] [CrossRef] [Green Version]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [Green Version]
- Pan, Y.J.; Lin, T.L.; Lin, Y.T.; Su, P.A.; Chen, C.T.; Hsieh, P.F.; Hsu, C.R.; Chen, C.C.; Hsieh, Y.C.; Wang, J.T. Identification of capsular types in carbapenem-resistant klebsiella pneumoniae strains by wzc sequencing and implications for capsule depolymerase treatment. Antimicrob. Agents Chemother. 2015, 59, 1038–1047. [Google Scholar] [CrossRef] [Green Version]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Nei, M.; Kumar, S. Molecular Evolutionand Phylogenetics; Oxford University Press: Oxford, UK, 2000; ISBN 0195135849. [Google Scholar]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aksyuk, A.A.; Kurochkina, L.P.; Fokine, A.; Forouhar, F.; Mesyanzhinov, V.V.; Tong, L.; Rossmann, M.G. Structural conservation of the myoviridae phage tail sheath protein fold. Structure 2011, 19, 1885–1894. [Google Scholar] [CrossRef] [Green Version]
- Aksyuk, A.A.; Leiman, P.G.; Kurochkina, L.P.; Shneider, M.M.; Kostyuchenko, V.A.; Mesyanzhinov, V.V.; Rossmann, M.G. The tail sheath structure of bacteriophage T4: A molecular machine for infecting bacteria. EMBO J. 2009, 28, 821–829. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsova, T.; Efimov, A.V.; Aijrich, L.G.; Kireeva, I.; Marusich, E.; Cappuccinelli, P.; Fiori, P.; Rappelli, P.; Kurochkina, L.; Poglazov, B.; et al. Properties of recombinant bacteriophage T4 tail sheath protein and its deletion fragments. Biochemistry 1998, 63, 702–709. [Google Scholar] [PubMed]
- Poglazov, B.F.; Efimov, A.V.; Marco, S.; Carrascosa, J.; Kuznetsova, T.A.; Aijrich, L.G.; Kurochkina, L.P.; Mesyanzhinov, V.V. Polymerization of bacteriophage T4 tail sheath protein mutants truncated at the C-termini. J. Struct. Biol. 1999, 127, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Stanislauskienė, R.; Laurynėnas, A.; Rutkienė, R.; Aučynaitė, A.; Tauraitė, D.; Meškienė, R.; Urbelienė, N.; Kaupinis, A.; Valius, M.; Kaliniene, L.; et al. YqfB protein from Escherichia coli: An atypical amidohydrolase active towards N 4-acylcytosine derivatives. Sci. Rep. 2020, 10, 788. [Google Scholar] [CrossRef]
- Fiedler, J.D.; Brown, S.D.; Lau, J.L.; Finn, M.G. RNA-directed packaging of enzymes within virus-like particles. Angew. Chemie Int. Ed. 2010, 49, 9648–9651. [Google Scholar] [CrossRef] [Green Version]
- Inoue, T.; Kawano, M.-A.; Takahashi, R.-U.; Tsukamoto, H.; Enomoto, T.; Imai, T.; Kataoka, K.; Handa, H. Engineering of SV40-based nano-capsules for delivery of heterologous proteins as fusions with the minor capsid proteins VP2/3. J. Biotechnol. 2008, 134, 181–192. [Google Scholar] [CrossRef]
- Minten, I.J.; Claessen, V.I.; Blank, K.; Rowan, A.E.; Nolte, R.J.M.; Cornelissen, J.J.L.M. Catalytic capsids: The art of confinement. Chem. Sci. 2011, 2, 358–362. [Google Scholar] [CrossRef] [Green Version]
- Patterson, D.P.; Prevelige, P.E.; Douglas, T. Nanoreactors by programmed enzyme encapsulation inside the capsid of the bacteriophage P22. ACS Nano 2012, 6, 5000–5009. [Google Scholar] [CrossRef] [PubMed]








| Primer | Sequence 5′-3′ | Resulting Plasmid |
|---|---|---|
| 041XhoR | gtgctcgagagtattttcaatac | pET21_041 |
| 041NcoF | tataccatggcagatttaatcag | |
| 041Δ200HindR | gccaagctttctagtactcaggc | pET28_041Δ200 |
| mcheryHindF | cgtgtaaagcttgttagtaaaggagaaga | pET21_041Δ200mCherry |
| mcheryXhoR | tagttctcgagttatgcggtaccaga | |
| gfpHindF | agaaggaaagcttcatatgagtaaagga | pET28_041Δ200GFP |
| gfpXhoR | gtgctcgagtgatttgtatag | |
| YqfbbukF | atgcagccaaacgacatcac | pET21_041Δ200YqfB |
| YqfbbukR | ttaaagacatttaaattcaatcacata | |
| GS_F | agcttggaggaggaggaagtggaggaggaggaagtcctagggtcgactacgtac | pET21_041Δ200GS |
| GS_R | tcgagtacgtagtcgaccctaggacttcctcctcctccacttcctcctcca |
| Protein | Excititation (nm) | Emission (nm) |
|---|---|---|
| 041Δ200GFP | 488 | 493–582 |
| 041Δ200mCherry | 587 | 600–700 |
| Construct | Length, nm | Diameter, nm |
|---|---|---|
| 041 | 319.7 ± 76.1 | 41.1 ± 4.9 |
| 041Δ200 | 311.7 ± 87.7 | 43.4 ± 4.2 |
| 041Δ200GFP | 276.0 ± 68.2 | 43.9 ± 4.1 |
| 041Δ200mCherry | 237.3 ± 76.4 | 49.2 ± 3.4 |
| 041Δ200GFPmCherry | 283.8 ± 86.2 | 46.3 ± 3.4 |
| 041Δ200YqfB | 231.0 ± 106 | 51.5 ± 9.4 (internal diameter 13 nm) |
| Protein | Activity, U/mg | Km, M | kcat, s−1 | kcat/Km, M−1s−1 |
|---|---|---|---|---|
| 041Δ200YqfB | 33 ± 4.3 | (7.6 ± 0.05) × 10−4 | 241 ± 2 | 3.2 × 105 |
| YqfB | 101 ± 9.2 | (4.9 ± 0.04) × 10−4 | 113 ± 1 | 2.3 × 105 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Labutytė, G.; Povilonienė, S.; Šimoliūnas, E.; Gabrielaitis, D.; Skapas, M.; Noreika, A.; Meškys, R.; Časaitė, V. Functionalized Protein Nanotubes Based on the Bacteriophage vB_KleM-RaK2 Tail Sheath Protein. Nanomaterials 2021, 11, 3031. https://doi.org/10.3390/nano11113031
Labutytė G, Povilonienė S, Šimoliūnas E, Gabrielaitis D, Skapas M, Noreika A, Meškys R, Časaitė V. Functionalized Protein Nanotubes Based on the Bacteriophage vB_KleM-RaK2 Tail Sheath Protein. Nanomaterials. 2021; 11(11):3031. https://doi.org/10.3390/nano11113031
Chicago/Turabian StyleLabutytė, Greta, Simona Povilonienė, Eugenijus Šimoliūnas, Dovydas Gabrielaitis, Martynas Skapas, Algirdas Noreika, Rolandas Meškys, and Vida Časaitė. 2021. "Functionalized Protein Nanotubes Based on the Bacteriophage vB_KleM-RaK2 Tail Sheath Protein" Nanomaterials 11, no. 11: 3031. https://doi.org/10.3390/nano11113031
APA StyleLabutytė, G., Povilonienė, S., Šimoliūnas, E., Gabrielaitis, D., Skapas, M., Noreika, A., Meškys, R., & Časaitė, V. (2021). Functionalized Protein Nanotubes Based on the Bacteriophage vB_KleM-RaK2 Tail Sheath Protein. Nanomaterials, 11(11), 3031. https://doi.org/10.3390/nano11113031






