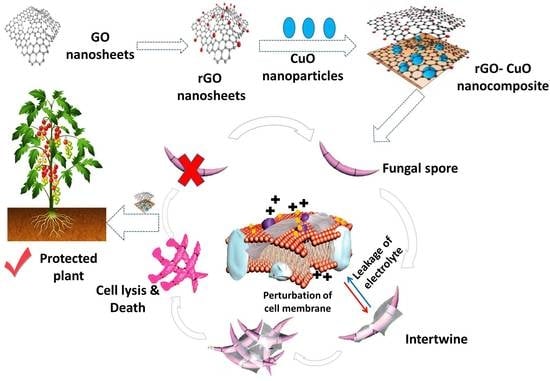
Reduced Graphene Oxide Nanosheet-Decorated Copper Oxide Nanoparticles: A Potent Antifungal Nanocomposite against Fusarium Root Rot and Wilt Diseases of Tomato and Pepper Plants
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemical Reagents
2.2. Fungal Strains and Culture Conditions
2.3. Molecular Identification of Fusarium Strains
2.4. Preparation of rGO Nanosheets
2.5. Synthesis and Characterization of rGO-CuO NPs
2.6. In Vitro Antifungal Activity of rGO-CuO NPs
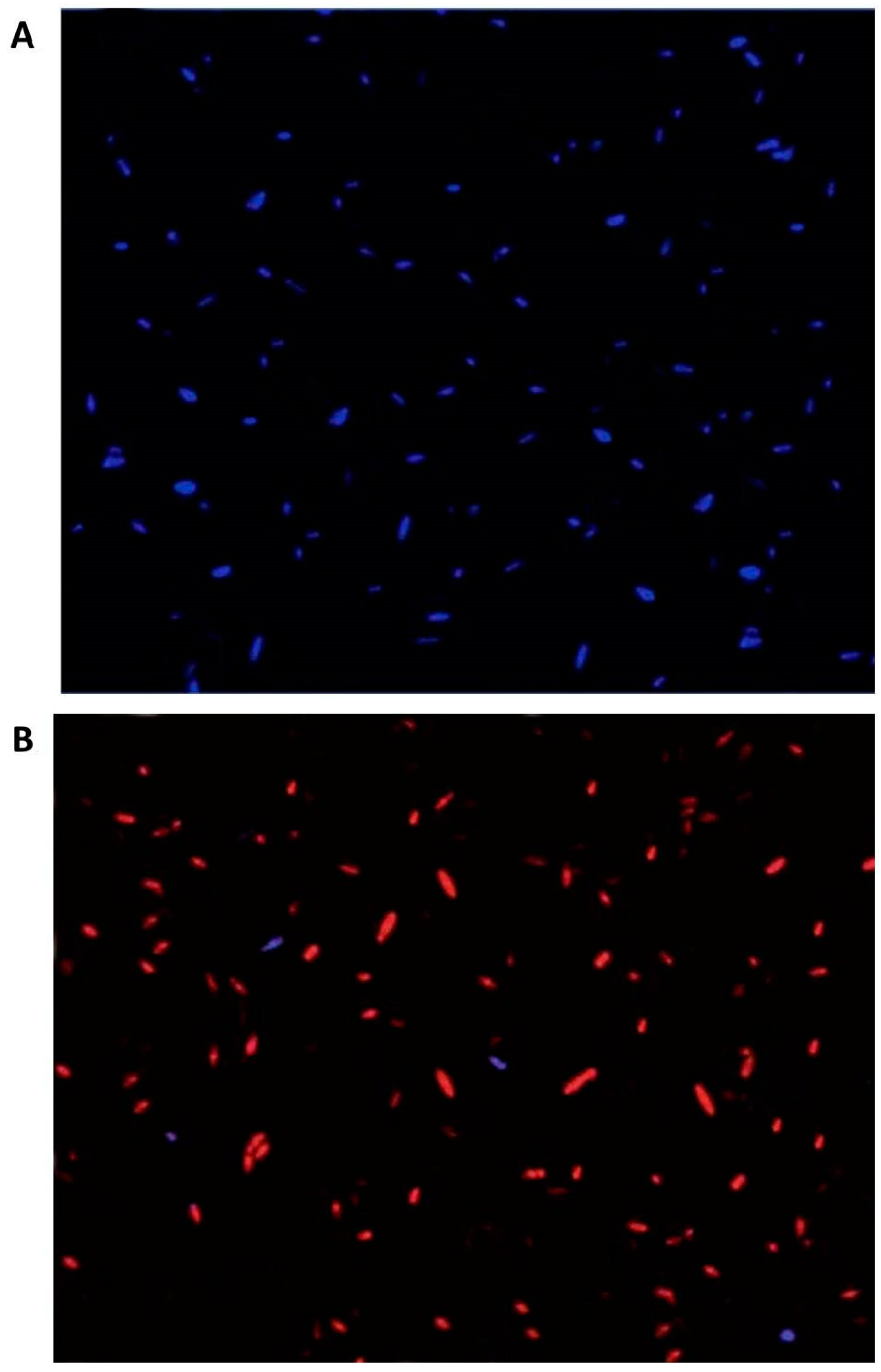
2.7. Fluorescence Microscopy Imaging
2.8. Fungal Cell Morphology Analysis
2.9. Inoculum Preparation for In Vivo Experiments
2.10. Antifungal Analysis In Vivo of rGO-CuO NPs
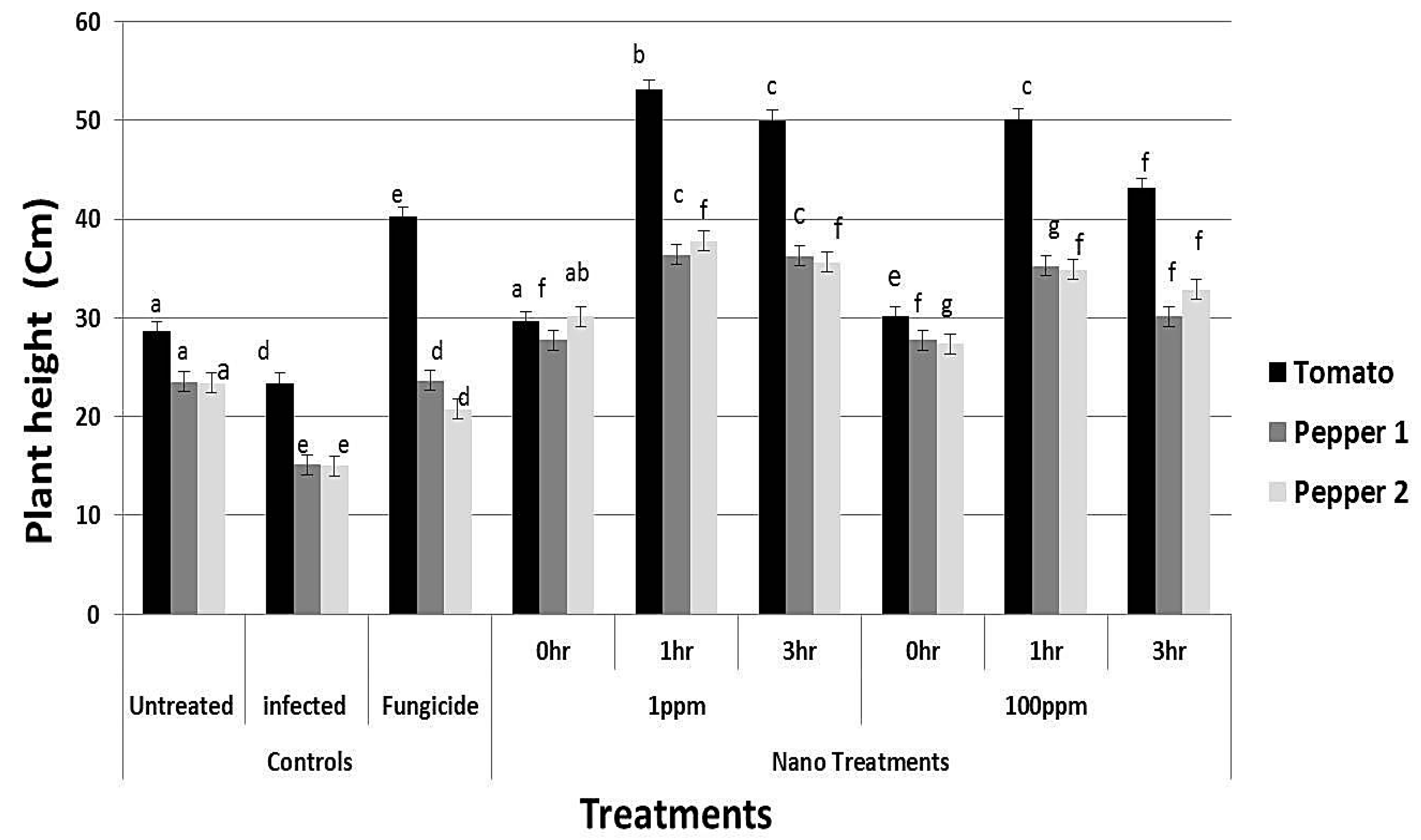
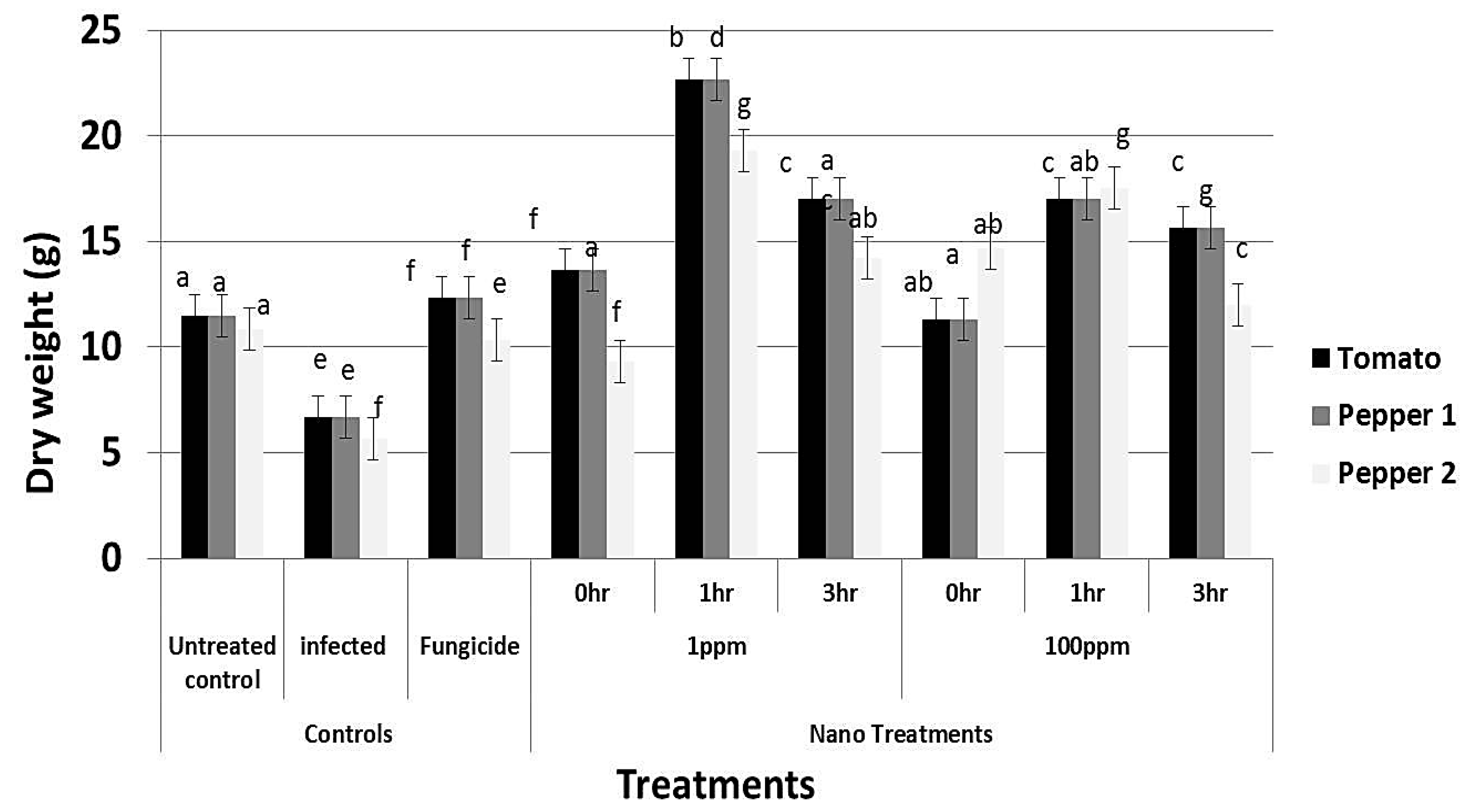
2.11. Analysis of Plant Height, Dry Weight and Photosynthetic Pigments
2.12. Translocation of rGO-CuO Nanoparticles in Tomato and Pepper Plants
2.13. Statistical Analysis
3. Results and Discussion
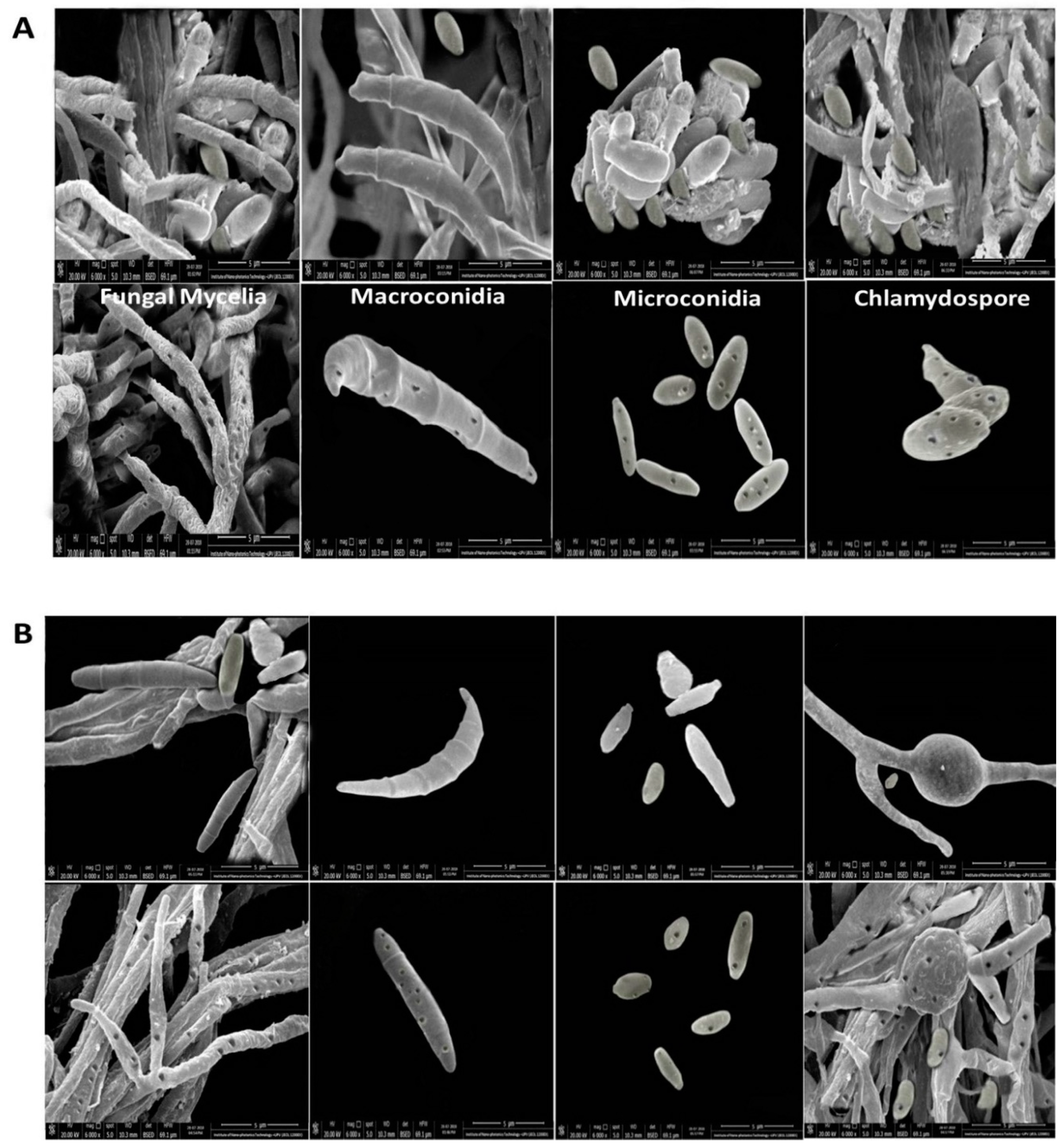
3.1. Characterization of Wild Fusarium Strains
3.2. Molecular Identification of Fusarium Strains
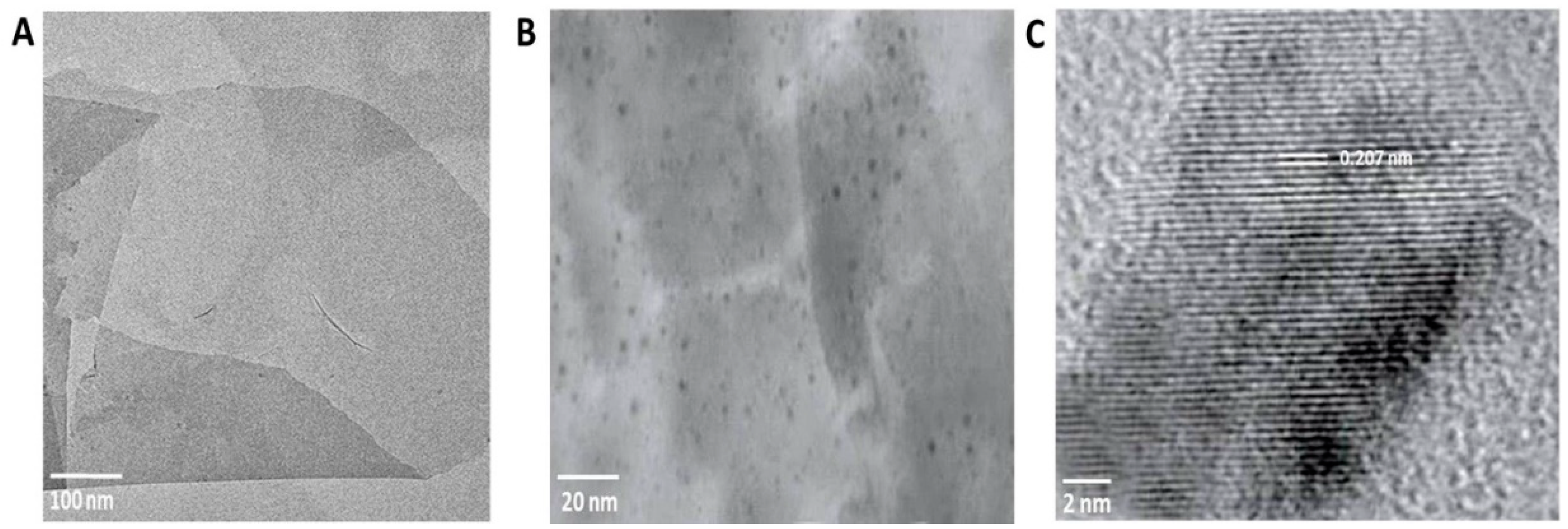
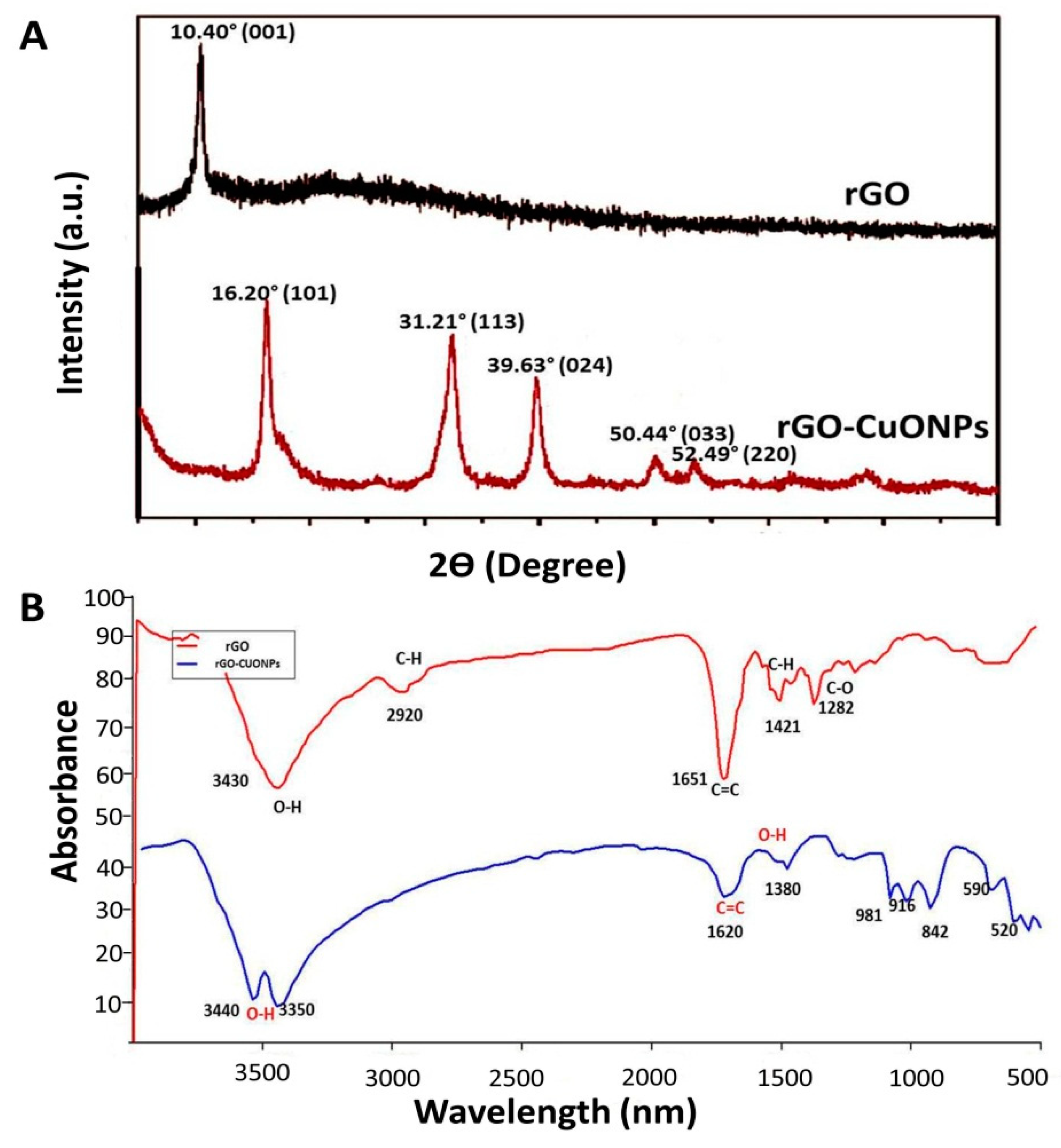
3.3. Characterization of rGO-CuO NPs
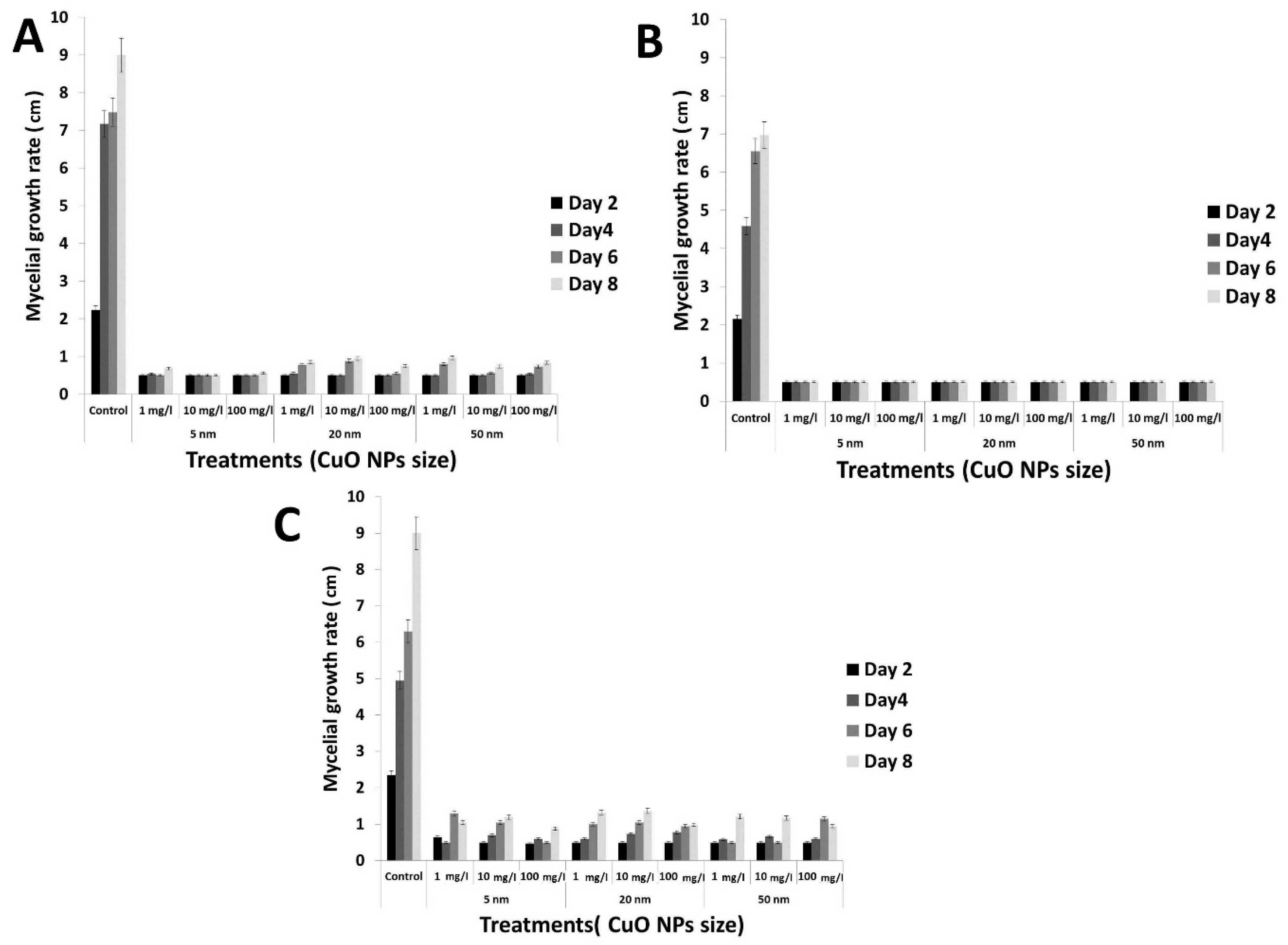
3.4. Antifungal Activity of the rGO-CuO NPs
3.5. Fungal Cell Viability after rGO-CuO NPs Treatment
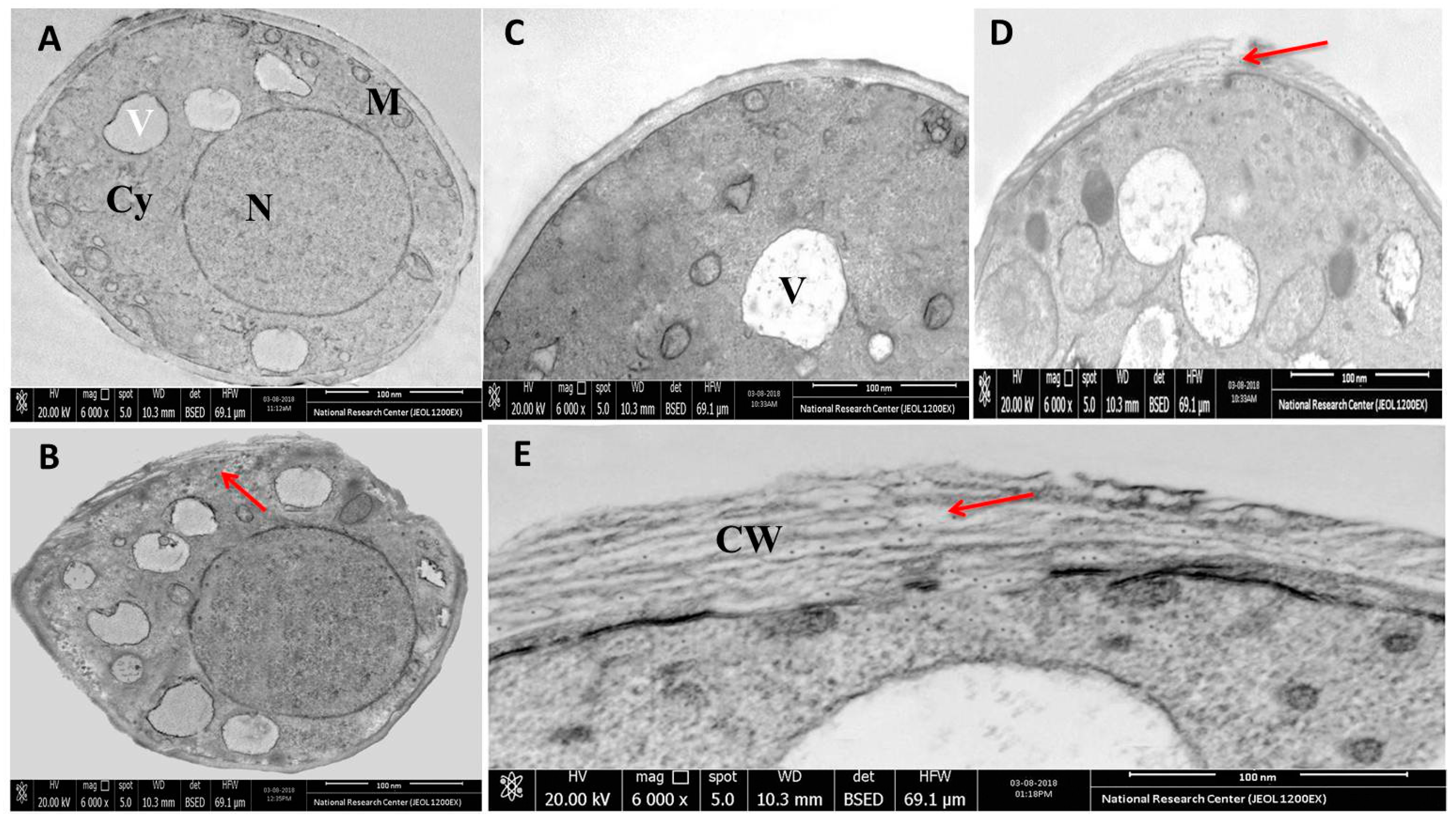
3.6. Microscopic Analysis of the Interaction between rGO-CuO NPs and Fungal Cells
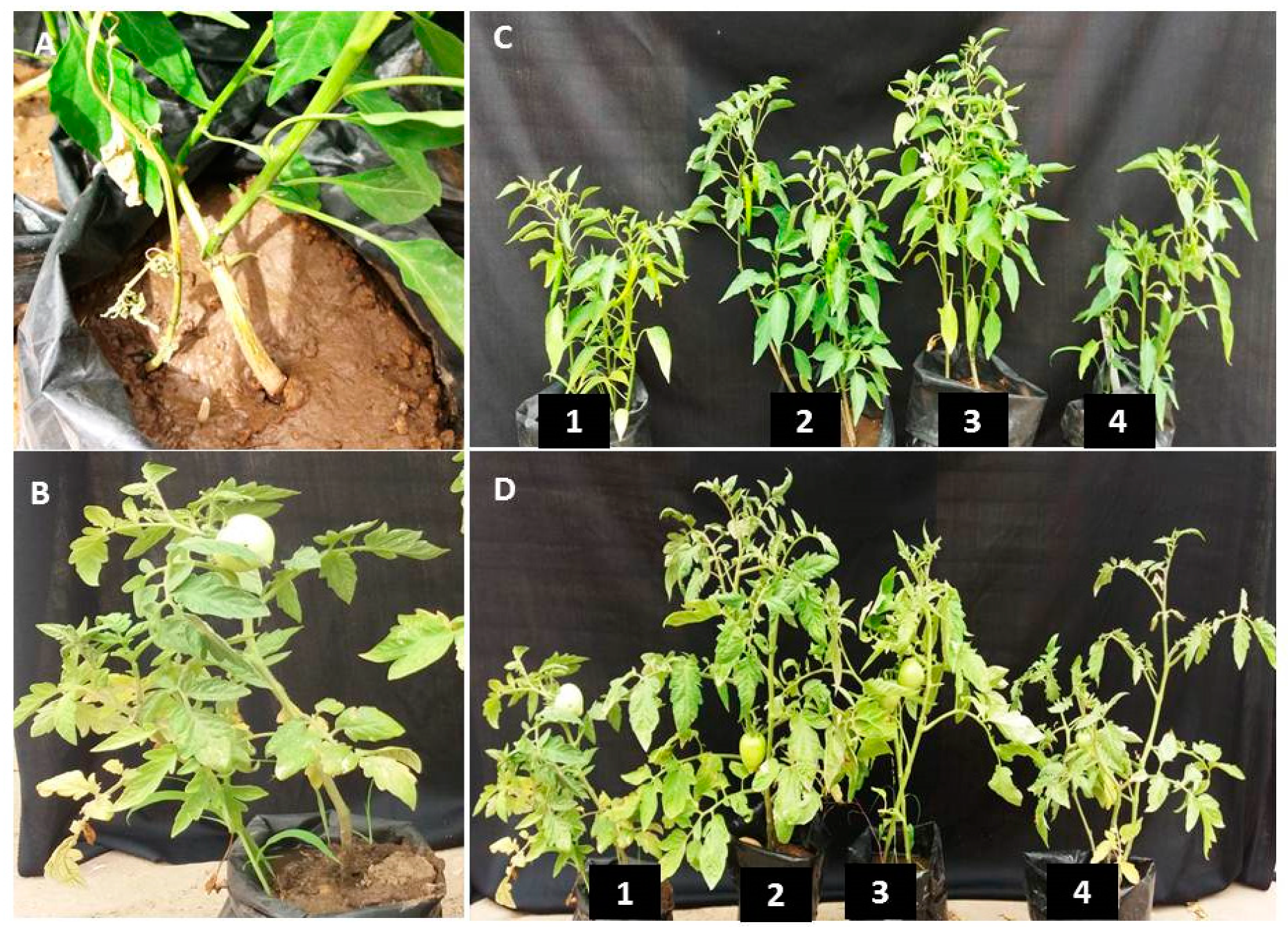
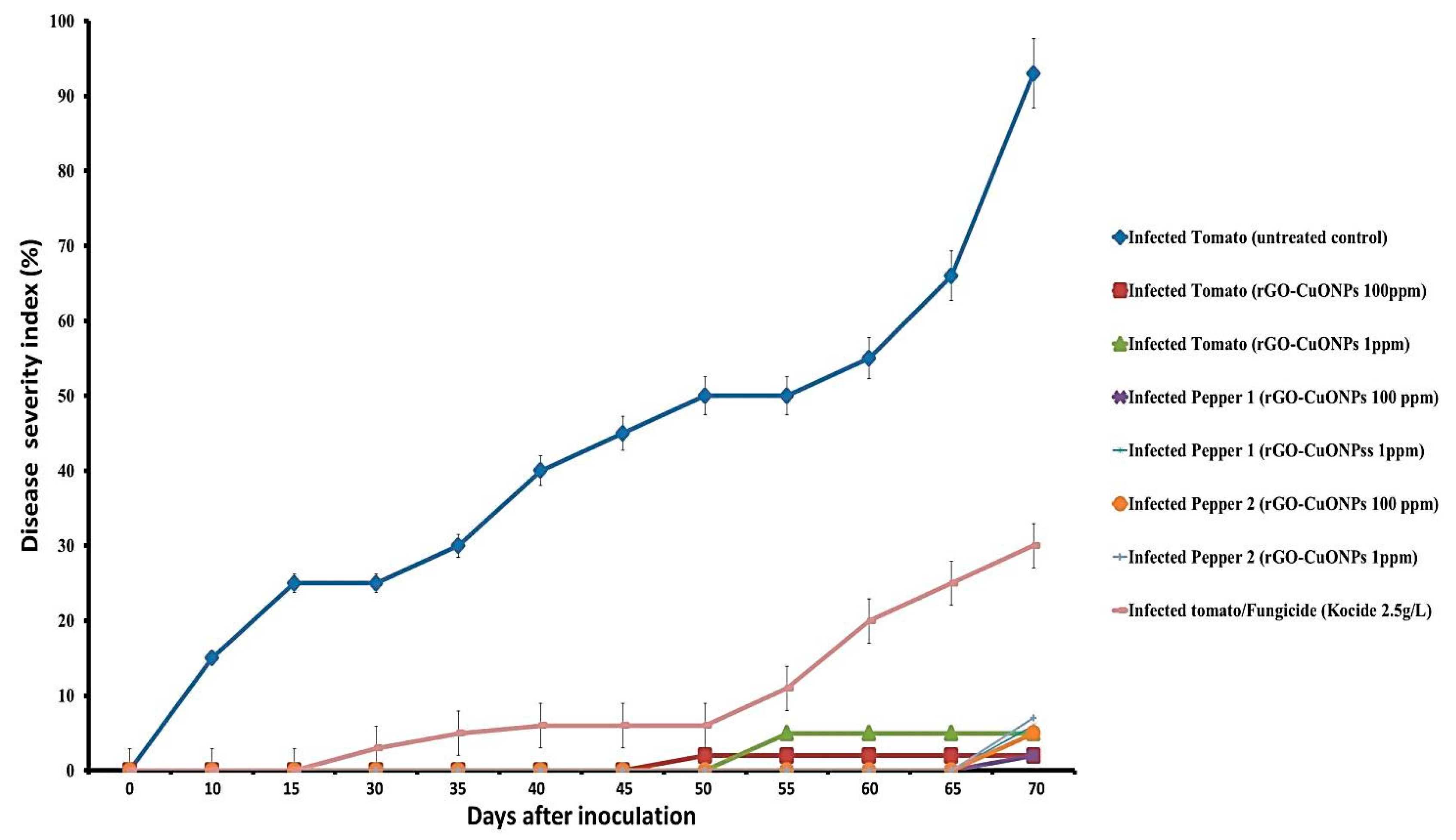
3.7. rGO-CuO NPs Effectiveness against Fusarium Diseases
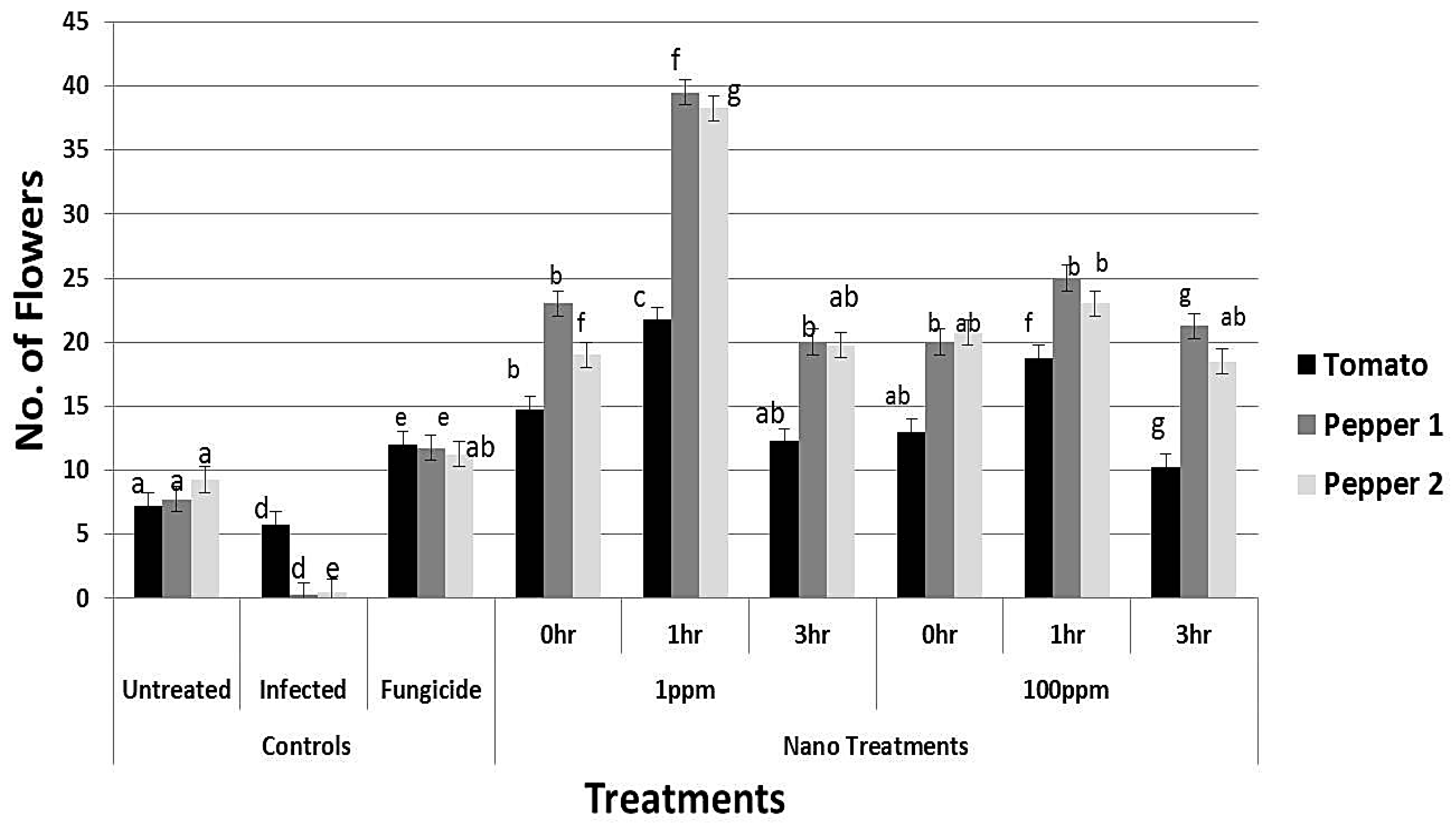
3.8. Effect of rGO-CuO on Flower Induction and Photosynthetic Pigments
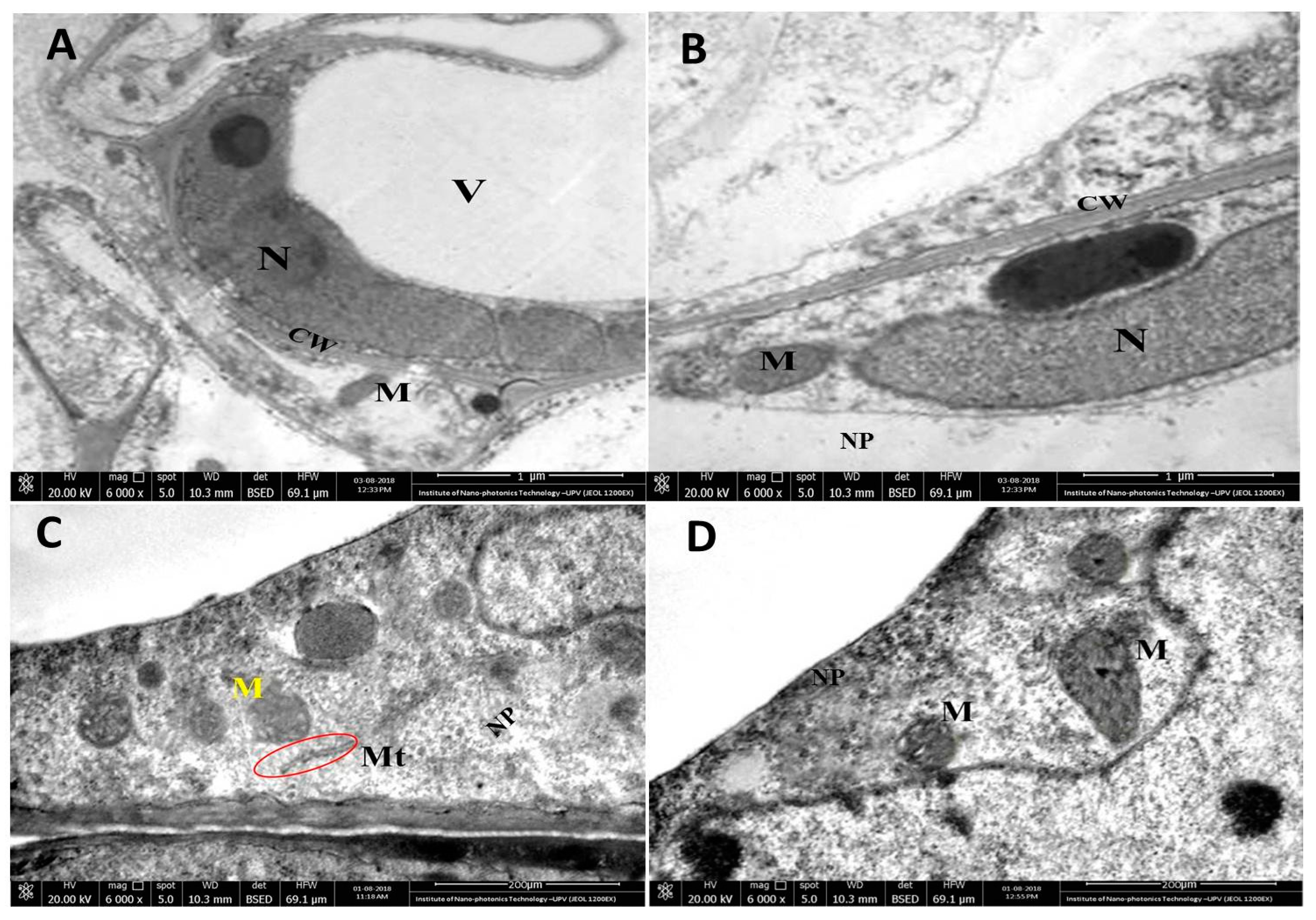
3.9. Translocation of CuO NPs in Plant Roots and Potential Phytotoxicity
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Dita, M.; Barquero, M.; Heck, D.; Mizubuti, E.S.; Staver, C.P. Fusarium wilt of banana: Current knowledge on epidemiology and research needs toward sustainable disease management. Front. Plant. Sci. 2018, 9, 1468. [Google Scholar] [CrossRef]
- Naguib, D.M. Control of Fusarium wilt in wheat seedlings by grain priming with defensin-like protein. Egypt. J. Biol. Pest. Control. 2018, 28, 68. [Google Scholar] [CrossRef]
- Shishido, M.; Miwa, C.; Usami, T.; Amemiya, Y.; Johnson, K.B. Biological control efficiency of Fusarium wilt of tomato by nonpathogenic Fusarium oxysporum Fo-B2 in different environments. Phytopathology 2005, 95, 1072–1080. [Google Scholar] [CrossRef] [PubMed]
- Gordon, T.R.; Martyn, R.D. The evolutionary biology of Fusarium oxysporum. Annu. Rev. Phytopathol. 1997, 35, 111–128. [Google Scholar] [CrossRef] [PubMed]
- Selim, M.E.; Khalifa, E.Z.; Amer, G.A.; Ely-kafrawy, A.A.; El-Gammal, N.A. Evaluation and characterization of some Egyptian Fusarium oxysporum isolates for their virulence on tomato and PCR detection of (SIX) effector genes. J. Bioprocess. Biotechnol. 2015, 5. [Google Scholar] [CrossRef]
- Heuvelink, E.; Dorais, M. Crop growth and yield. Crop. Prod. Sci. Hortic. 2005, 13, 85. [Google Scholar]
- Lukyanenko, A.N. Disease Resistance in Tomato; Springer: Berlin/Heidelberg, Germany, 1991; pp. 99–119. [Google Scholar] [CrossRef]
- Metwally, N.H.; Radwan, I.T.; El-Serwy, W.S.; Mohamed, M.A. Design, synthesis, DNA assessment and molecular docking study of novel 2-(pyridin-2-ylimino) thiazolidin-4-one derivatives as potent antifungal agents. Bioorg. Chem. 2019, 84, 456–467. [Google Scholar] [CrossRef]
- Buonaurio, R.; Moretti, C.; Bertona, A.; Scarponi, L. Investigations on the Systemic Acquired Resistance Induced by Acibenzolar-S-Methyl in Tomato Plants Against Pseudomonas syringae pv. tomato. In Pseudomonas Syringae and Related Pathogens; Springer: Dordrecht, The Netherlands, 2003; pp. 475–482. [Google Scholar] [CrossRef]
- Bower, C.K.; Daeschel, M.A. Resistance responses of microorganisms in food environments. Int. J. Food Microbiol. 1999, 50, 33–44. [Google Scholar] [CrossRef]
- Louws, F.J.; Wilson, M.; Campbell, H.L.; Cuppels, D.A.; Jones, J.B.; Shoemaker, P.B.; Miller, S.A. Field control of bacterial spot and bacterial speck of tomato using a plant activator. Plant. Dis. 2001, 85, 481–488. [Google Scholar] [CrossRef]
- Dimkpa, C.O.; McLean, J.E.; Britt, D.W.; Anderson, A.J. Antifungal activity of ZnO nanoparticles and their interactive effect with a biocontrol bacterium on growth antagonism of the plant pathogen Fusarium graminearum. Biometals 2013, 26, 913–924. [Google Scholar] [CrossRef]
- Liu, J.J.; Wei, Z.; Li, J.H. Effects of copper on leaf membrane structure and root activity of maize seedling. Bot. Stud. 2014, 55, 47. [Google Scholar] [CrossRef] [PubMed]
- Krishna, V.; Pumprueg, S.; Lee, S.H.; Zhao, J.; Sigmund, W.; Koopman, B.; Moudgil, B.M. Photocatalytic disinfection with titanium dioxide coated multi-wall carbon nanotubes. Process. Saf. Environ. 2005, 83, 393–397. [Google Scholar] [CrossRef]
- Ocsoy, I.; Paret, M.L.; Ocsoy, M.A.; Kunwar, S.; Chen, T.; You, M.; Tan, W. Nanotechnology in plant disease management: DNA-directed silver nanoparticles on graphene oxide as an antibacterial against Xanthomonas perforans. ACS Nano 2013, 7, 8972–8980. [Google Scholar] [CrossRef] [PubMed]
- Cordero, T.; Mohamed, M.A.; López-Moya, J.J.; Daròs, J.A. A recombinant potato virus y infectious clone tagged with the rosea1 visual marker (pvy-ros1) facilitates the analysis of viral infectivity and allows the production of large amounts of anthocyanins in plants. Front. Microbiol. 2017, 8, 611. [Google Scholar] [CrossRef]
- Salam, M.A.; Obaid, A.Y.; El-Shishtawy, R.M.; Mohamed, S.A. Synthesis of nanocomposites of polypyrrole/carbon nanotubes/silver nano particles and their application in water disinfection. RSC Adv. 2017, 7, 16878–16884. [Google Scholar] [CrossRef]
- Cui, J.; Liu, Y. Preparation of graphene oxide with silver nanowires to enhance antibacterial properties and cell compatibility. RSC Adv. 2015, 5, 85748–85755. [Google Scholar] [CrossRef]
- Pradhan, S.; Patra, P.; Mitra, S.; Dey, K.K.; Jain, S.; Sarkar, S.; Goswami, A. Manganese nanoparticles: Impact on non-nodulated plant as a potent enhancer in nitrogen metabolism and toxicity study both in vivo and in vitro. J. Agric. Food Chem. 2014, 62, 8777–8785. [Google Scholar] [CrossRef]
- Youssef, K.; de Oliveira, A.G.; Tischer, C.A.; Hussain, I.; Roberto, S.R. Synergistic effect of a novel chitosan/silica nanocomposites-based formulation against gray mold of table grapes and its possible mode of action. Int. J. Biol. Macromol. 2019, 141, 247–258. [Google Scholar] [CrossRef]
- Choudhary, R.C.; Kumaraswamy, R.V.; Kumari, S.; Sharma, S.S.; Pal, A.; Raliya, R.; Saharan, V. Cu-chitosan nanoparticle boost defense responses and plant growth in maize (Zea mays L.). Sci. Rep. 2017, 7, 9754. [Google Scholar] [CrossRef]
- Karlsson, K.R.; Cowley, S.; Martinez, F.O.; Shaw, M.; Minger, S.L.; James, W. Homogeneous monocytes and macrophages from human embryonic stem cells following co culture-free differentiation in M-CSF and IL-3. Exp. Hematol. 2008, 36, 1167–1175. [Google Scholar] [CrossRef]
- Karthik, R.; Govindasamy, M.; Chen, S.M.; Chen, T.W.; Elangovan, A.; Muthuraj, V.; Yu, M.C. A facile graphene oxide based sensor for electrochemical detection of prostate anti-cancer (anti-testosterone) drug flutamide in biological samples. RSC Adv. 2017, 7, 25702–25709. [Google Scholar] [CrossRef]
- Li, Y.; Yang, D.; Cui, J. Graphene oxide loaded with copper oxide nanoparticles as an antibacterial agent against Pseudomonas syringae pv. tomato. RSC Adv. 2017, 7, 38853–38860. [Google Scholar] [CrossRef]
- Maiti, M.; Sarkar, M.; Maiti, S.; Liu, D. Efficacy of shape-monitored reduced graphene oxide–copper nanohybrids: Anti-bacterial attributes for food safety and dye degradation studies. New J. Chem. 2019, 43, 662–674. [Google Scholar] [CrossRef]
- Alayande, A.B.; Obaid, M.; Kim, I.S. Antimicrobial mechanism of reduced graphene oxide-copper oxide (rGO-CuO) nanocomposite films: The case of Pseudomonas aeruginosa PAO1. Mater. Sci. Eng. C 2020, 109, 110596. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Cao, X.; Ma, C.; Zhang, Z.; Zhao, N.; Ali, A.; Xing, B. Potential applications and antifungal activities of engineered nanomaterials against gray mold disease agent Botrytis cinerea on rose petals. Front. Plant. Sci. 2017, 8, 1332. [Google Scholar] [CrossRef]
- Yang, H.; Feng, S.; Ma, Q.; Ming, Z.; Bai, Y.; Chen, L.; Yang, S.T. Influence of reduced graphene oxide on the growth, structure and decomposition activity of white-rot fungus Phanerochaete chrysosporium. RSC Adv. 2018, 9, 5026–5033. [Google Scholar] [CrossRef]
- Nelson, P.E.; Toussoun, T.A.; Marasas, W.F.O. Fusarium Species: An Illustrated Manual for Identification; Pennsylvania State University Press: University Park, TX, USA, 1983. [Google Scholar]
- Hirano, Y.; Arie, T. PCR-based differentiation of Fusarium oxysporum ff. sp. lycopersici and radicis-lycopersici and races of F. oxysporum f. sp. lycopersici. J. Gen. Plant. Pathol. 2006, 72, 273–283. [Google Scholar] [CrossRef]
- Çolak, A.; Biçici, M. PCR detection of Fusarium oxysporum f. sp. radicis-lycopersici and races of F. oxysporum f. sp. lycopersici of tomato in protected tomato-growing areas of the eastern Mediterranean region of Turkey. Turk. J. Agric. For. 2013, 37, 457–467. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.J.W.T.; Taylor, J.; Innis, M.A.; Gelfand, D.H.; Sninsky, J.J. PCR Protocols: A Guide to Methods and Applications; The University of Michigan: Arbor, MI, USA; Academic Press: Cambridge, MA, USA, 1990. [Google Scholar] [CrossRef]
- Pandey, V.; Mishra, G.; Verma, S.; Wan, M.; Yadav, R. Synthesis and Ultrasonic Investigations of CuOPVA Nanofluid. J. Mater. Sci. Appl. 2012, 3, 653–664. [Google Scholar] [CrossRef]
- Chen, J.; Peng, H.; Wang, X.; Shao, F.; Yuan, Z.; Han, H. Graphene oxide exhibits broad-spectrum antimicrobial activity against bacterial phytopathogens and fungal conidia by intertwining and membrane perturbation. Nanoscale 2014, 6, 1879–1889. [Google Scholar] [CrossRef]
- Lichtenthaler, H.K. Chlorophylls and carotenoids: Pigments of photosynthetic biomembranes. In Methods in Enzymology; Academic Press: New York, NY, USA, 1987; Volume 148, pp. 350–382. [Google Scholar] [CrossRef]
- Kim, S.W.; Jung, J.H.; Lamsal, K.; Kim, Y.S.; Min, J.S.; Lee, Y.S. Antifungal effects of silver nanoparticles (AgNPs) against various plant pathogenic fungi. Mycobiology 2012, 40, 53–58. [Google Scholar] [CrossRef] [PubMed]
- Goyal, P.; Chakraborty, S.; Misra, S.K. Multifunctional Fe3O4-ZnO nanocomposites for environmental remediation applications. Environ. Nanotechnol. Monit. Manag. 2018, 10, 28–35. [Google Scholar] [CrossRef]
- Bondarenko, O.; Juganson, K.; Ivask, A.; Kasemets, K.; Mortimer, M.; Kahru, A. Toxicity of Ag, CuO and ZnO nanoparticles to selected environmentally relevant test organisms and mammalian cells in vitro: A critical review. Arch. Toxicol. 2013, 87, 1181–1200. [Google Scholar] [CrossRef] [PubMed]
- Dreyer, D.R.; Park, S.; Bielawski, C.W.; Ruoff, R.S. The chemistry of graphene oxide. Chem. Soc. Rev. 2010, 39, 228–240. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Pinault, M.; Pfefferle, L.D.; Elimelech, M. Single-walled carbon nanotubes exhibit strong antimicrobial activity. Langmuir 2007, 23, 8670–8673. [Google Scholar] [CrossRef] [PubMed]
- Raghunath, A.; Perumal, E. Metal oxide nanoparticles as antimicrobial agents: A promise for the future. Int. J. Antimicrob. Agents 2017, 49, 137–152. [Google Scholar] [CrossRef]
- Raffi, M.; Mehrwan, S.; Bhatti, T.M.; Akhter, J.I.; Hameed, A.; Yawar, W.; ul Hasan, M.M. Investigations into the antibacterial behavior of copper nanoparticles against Escherichia coli. Ann. Microbiol. 2010, 60, 75–80. [Google Scholar] [CrossRef]
- Kanmani, P.; Lim, S.T. Synthesis and characterization of pullulan-mediated silver nanoparticles and its antimicrobial activities. Carbohydr. Polym. 2013, 97, 421–428. [Google Scholar] [CrossRef]
- Jasuja, N.D.; Gupta, D.K.; Reza, M.; Joshi, S.C. Green Synthesis of AgNPs Stabilized with biowaste and their antimicrobial activities. Braz. J. Microbiol. 2014, 45, 1325–1332. [Google Scholar] [CrossRef]
- Tripathi, D.K.; Tripathi, A.; Singh, S.; Singh, Y.; Vishwakarma, K.; Yadav, G.; Dubey, N.K. Uptake, accumulation and toxicity of silver nanoparticle in autotrophic plants, and heterotrophic microbes: A concentric review. Front. Microbiol. 2017, 8, 7. [Google Scholar] [CrossRef]
- Abdel-Hafez, S.I.; Nafady, N.A.; Abdel-Rahim, I.R.; Shaltout, A.M.; Daròs, J.A.; Mohamed, M.A. Assessment of protein silver nanoparticles toxicity against pathogenic Alternaria solani. 3 Biotech. 2016, 6, 199. [Google Scholar] [CrossRef] [PubMed]
- Moore, C.W.; McKoy, J.; Del Valle, R.; Armstrong, D.; Bernard, E.M.; Katz, N.; Gordon, R.E. Fungal cell wall septation and cytokinesis are inhibited by bleomycins. Antimicrob. Agents Chemother. 2003, 47, 3281–3289. [Google Scholar] [CrossRef] [PubMed]
- Priester, J.H.; Ge, Y.; Mielke, R.E.; Horst, A.M.; Moritz, S.C.; Espinosa, K.; Schimel, J.P. Soybean susceptibility to manufactured nanomaterials with evidence for food quality and soil fertility interruption. Proc. Natl. Acad. Sci. USA 2012, 109, E2451–E2456. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.F.; Gurunathan, S. Biofabrication of a novel biomolecule-assisted reduced graphene oxide: An excellent biocompatible nanomaterial. Int. J. Nanomed. 2016, 11, 6635. [Google Scholar] [CrossRef] [PubMed]
- Szmidt, M.; Stankiewicz, A.; Urbańska, K.; Jaworski, S.; Kutwin, M.; Wierzbicki, M.; Sawosz, E. Graphene oxide down-regulates genes of the oxidative phosphorylation complexes in a glioblastoma. BMC Mol. Biol. 2019, 20, 1–9. [Google Scholar] [CrossRef]
- Elmer, W.H.; White, J.C. The use of metallic oxide nanoparticles to enhance growth of tomatoes and eggplants in disease infested soil or soilless medium. Environ. Sci. Nano 2016, 3, 1072–1079. [Google Scholar] [CrossRef]
- Arias, M.M.D.; Leandro, L.F.; Munkvold, G.P. Aggressiveness of Fusarium species and impact of root infection on growth and yield of soybeans. Phytopathology 2013, 103, 822–832. [Google Scholar] [CrossRef]
- Fernández, V.; Brown, P.H. From plant surface to plant metabolism: The uncertain fate of foliar-applied nutrients. Front. Plant. Sci. 2013, 4, 289. [Google Scholar] [CrossRef]
- Sathiyabama, M.; Charles, R.E. Fungal cell wall polymer based nanoparticles in protection of tomato plants from wilt disease caused by Fusarium oxysporum f. sp. lycopersici. Carbohydr. Polym. 2015, 133, 400–407. [Google Scholar] [CrossRef]




| Pathogen | Treatment | Concentration | Inhibition Rate (%) | |||
|---|---|---|---|---|---|---|
| 4 Days | 6 Days | 8 Days | ||||
| F. oxysporum (FORL) | rGO-CuO nanocomposite | 1 mg/L | 92.35 ± 2.047 | 93.15 ± 1.09 | 92.37 ± 1.35 | |
| 10 mg/L | 92.90 ± 1.1 | 93.153 ± 1.09 | 93.7 ± 1.28 | |||
| 100 mg/L | 92.89 ± 1.13 | 93.17 ± 1.08 | 94.44 ± 0.0 | |||
| Controls | rGO | 1 g/L | 5.55 ± 0.0 | 8.33 ± 1.1 | 11.11 ± 0.0 | |
| CuO NPs | 1 g/L | 19.22 ± 0.0 | 22.22 ± 0.0 | 33.33 ± 1.1 | ||
| Kocide 2000 | 2.5 g/L | 65.3 ± 0.2 | 67.2 ± 0.4 | 68.3 ± 1.1 | ||
| F. oxysporum (FOC1) | rGO-CuO nanocomposite | 1 mg/L | 89.06 ± 0.31 | 92.35 ± 0.30 | 92.80 ± 0.3 | |
| 10 mg/L | 89.06 ± 0.31 | 92.36 ± 0.35 | 92.80 ± 0.33 | |||
| 100 mg/L | 89.08 ± 0.27 | 92.35 ± 0.35 | 92.80 ± 0.33 | |||
| Controls | rGO | 1 g/L | 2.27 ± 0.0 | 5.55 ± 1.1 | 8.33 ± 0.20 | |
| CuO NPs | 1 g/L | 25.00 ± 1.0 | 33.33 ± 1.1 | 38.88 ± 1.1 | ||
| Kocide 2000 | 2.5 g/L | 64.5 ± 0.5 | 69.2 ± 0.2 | 68.9 ± 0.4 | ||
| F. oxysporum (FOC2) | rGO-CuO nanocomposite | 1 mg/L | 85.95 ± 6.30 | 81.24 ± 7.1 | 83.34 ± 3.4 | |
| 10 mg/L | 87.13 ± 2.49 | 82.07 ± 1.8 | 82.13 ± 4.5 | |||
| 100 mg/L | 87.72 ± 2.8 | 88.77 ± 2.8 | 87.8 ± 2.5 | |||
| Controls | rGO | 1 g/L | 5.55 ± 0.0 | 8.33 ± 1.1 | 11.11 ± 0.0 | |
| CuO NPs | 1 g/L | 19.22 ± 0.0 | 22.22 ± 0.0 | 30.55 ± 0.0 | ||
| Kocide 2000 | 2.5 g/L | 63.3 ± 0.5 | 65.2 ± 0.4 | 70.5 ± 1.0 | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
El-Abeid, S.E.; Ahmed, Y.; Daròs, J.-A.; Mohamed, M.A. Reduced Graphene Oxide Nanosheet-Decorated Copper Oxide Nanoparticles: A Potent Antifungal Nanocomposite against Fusarium Root Rot and Wilt Diseases of Tomato and Pepper Plants. Nanomaterials 2020, 10, 1001. https://doi.org/10.3390/nano10051001
El-Abeid SE, Ahmed Y, Daròs J-A, Mohamed MA. Reduced Graphene Oxide Nanosheet-Decorated Copper Oxide Nanoparticles: A Potent Antifungal Nanocomposite against Fusarium Root Rot and Wilt Diseases of Tomato and Pepper Plants. Nanomaterials. 2020; 10(5):1001. https://doi.org/10.3390/nano10051001
Chicago/Turabian StyleEl-Abeid, Sozan E., Yosra Ahmed, José-Antonio Daròs, and Mohamed A. Mohamed. 2020. "Reduced Graphene Oxide Nanosheet-Decorated Copper Oxide Nanoparticles: A Potent Antifungal Nanocomposite against Fusarium Root Rot and Wilt Diseases of Tomato and Pepper Plants" Nanomaterials 10, no. 5: 1001. https://doi.org/10.3390/nano10051001
APA StyleEl-Abeid, S. E., Ahmed, Y., Daròs, J.-A., & Mohamed, M. A. (2020). Reduced Graphene Oxide Nanosheet-Decorated Copper Oxide Nanoparticles: A Potent Antifungal Nanocomposite against Fusarium Root Rot and Wilt Diseases of Tomato and Pepper Plants. Nanomaterials, 10(5), 1001. https://doi.org/10.3390/nano10051001