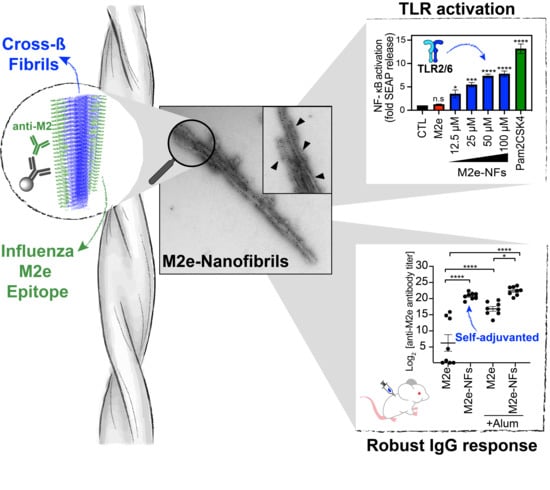
Harnessing the Activation of Toll-Like Receptor 2/6 by Self-Assembled Cross-β Fibrils to Design Adjuvanted Nanovaccines
Abstract
:1. Introduction
2. Materials and Methods
2.1. Peptide Synthesis, Purification, and Characterization
2.2. Peptide Self-Assembly
2.3. Critical Aggregation Concentration
2.4. Absorbancce and Fluorescence Spectroscopy
2.5. Circular Dichroism Spectroscopy
2.6. Powder X-ray Diffraction
2.7. Atomic Force Microscopy
2.8. Transmission Electron Microscopy
2.9. Enzyme-Linked Immunosorbent Assay for Epitope Accessibility
2.10. Measurement of Zeta potential
2.11. Cell Viability Assays
2.12. Evaluation of TLR2/6 Activation
2.13. Mice Immunization
2.14. Determination of Anti-M2e Antibody Titers
2.15. Data Analysis
3. Results and Discussion
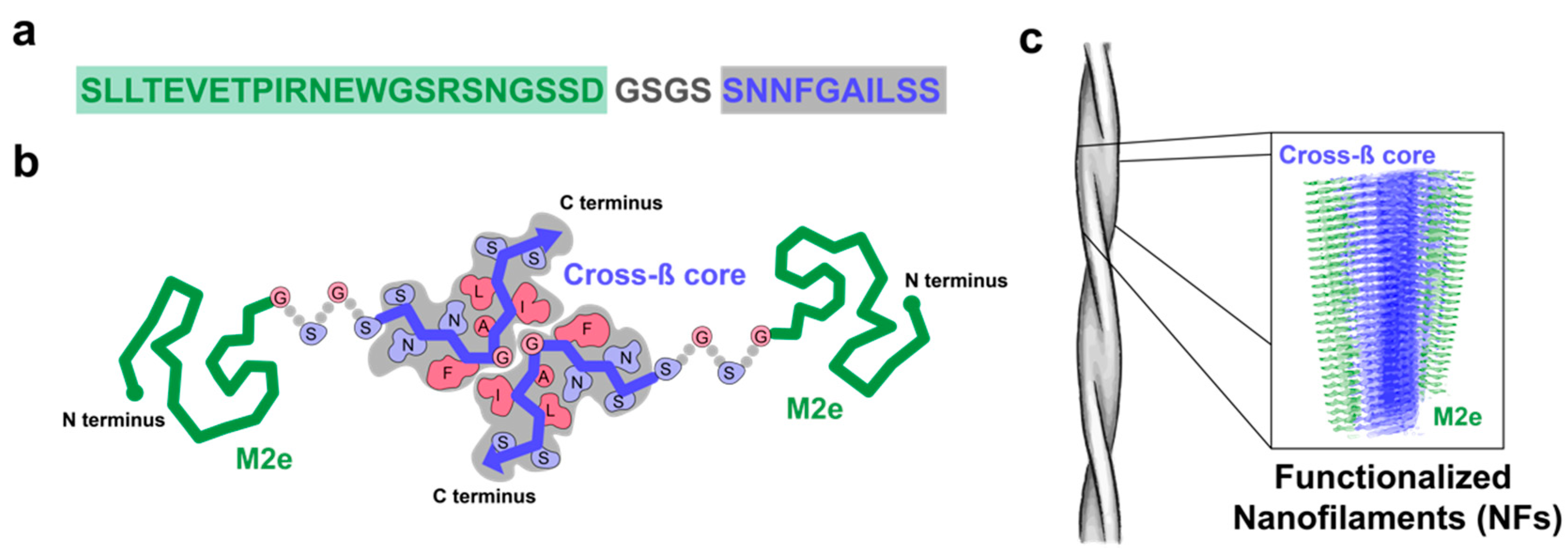
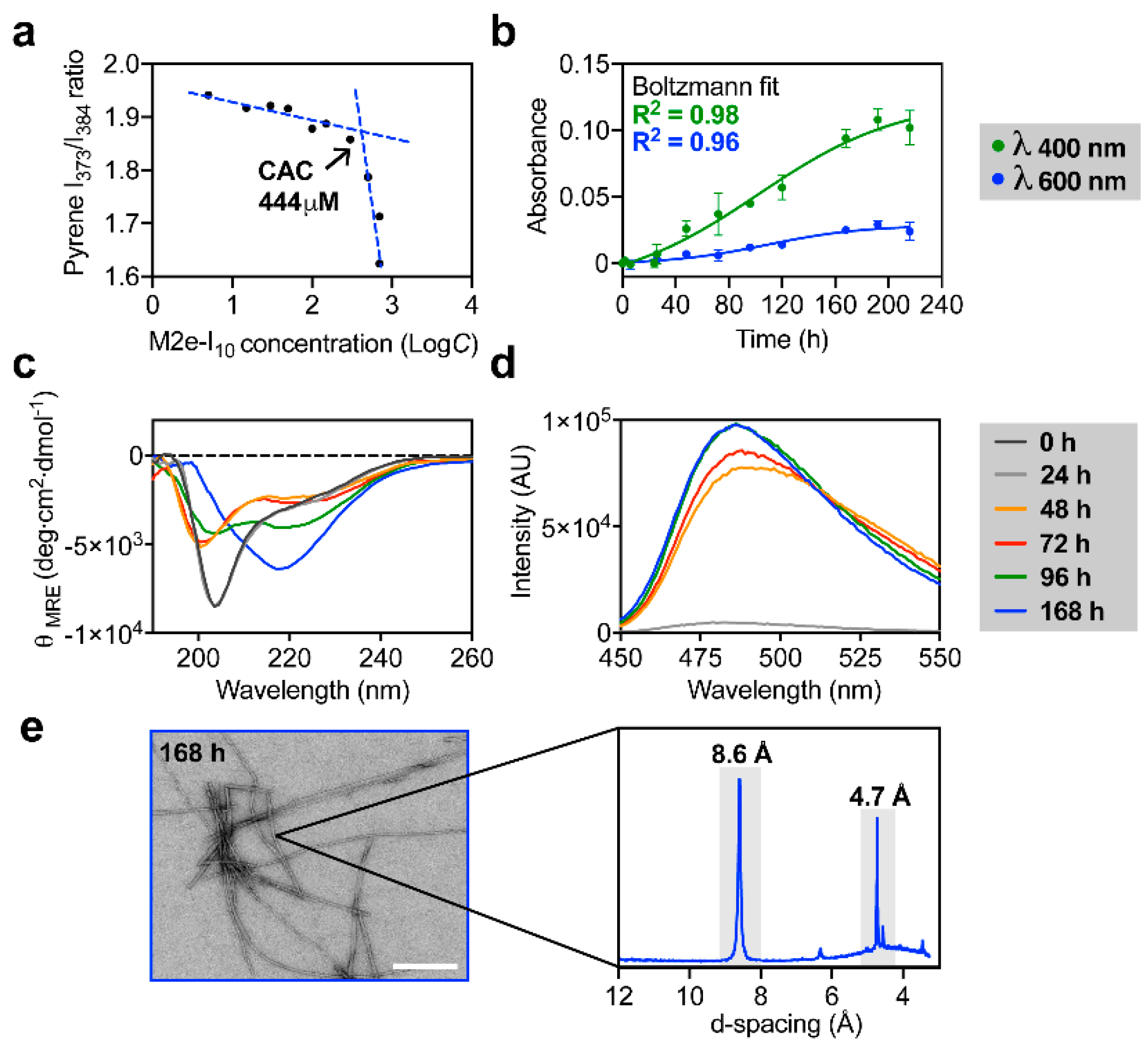
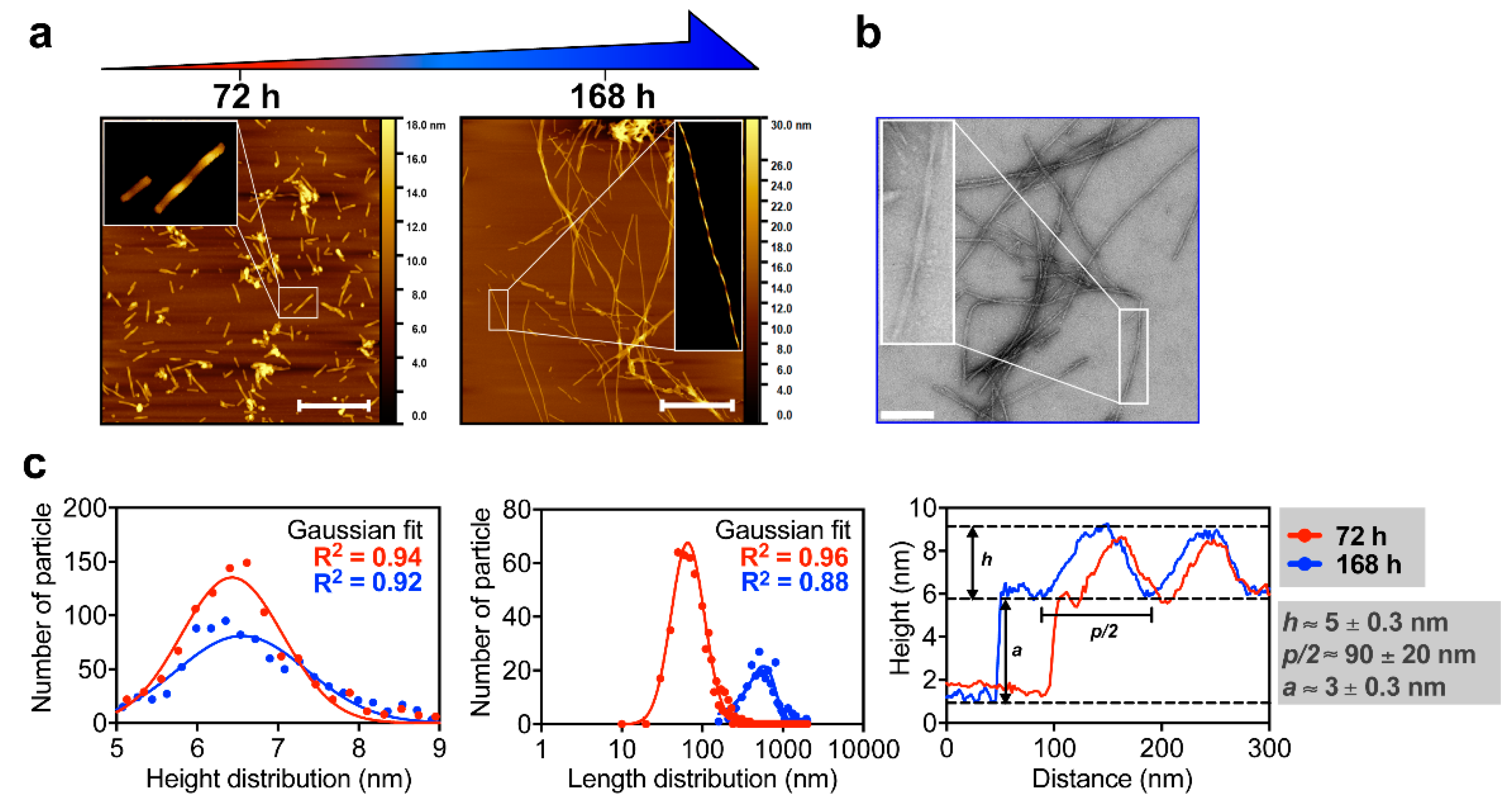
3.1. M2e-I10 Self-Assembles into Twisted Fibrils with a Cross-β-Sheet Quaternary Architecture
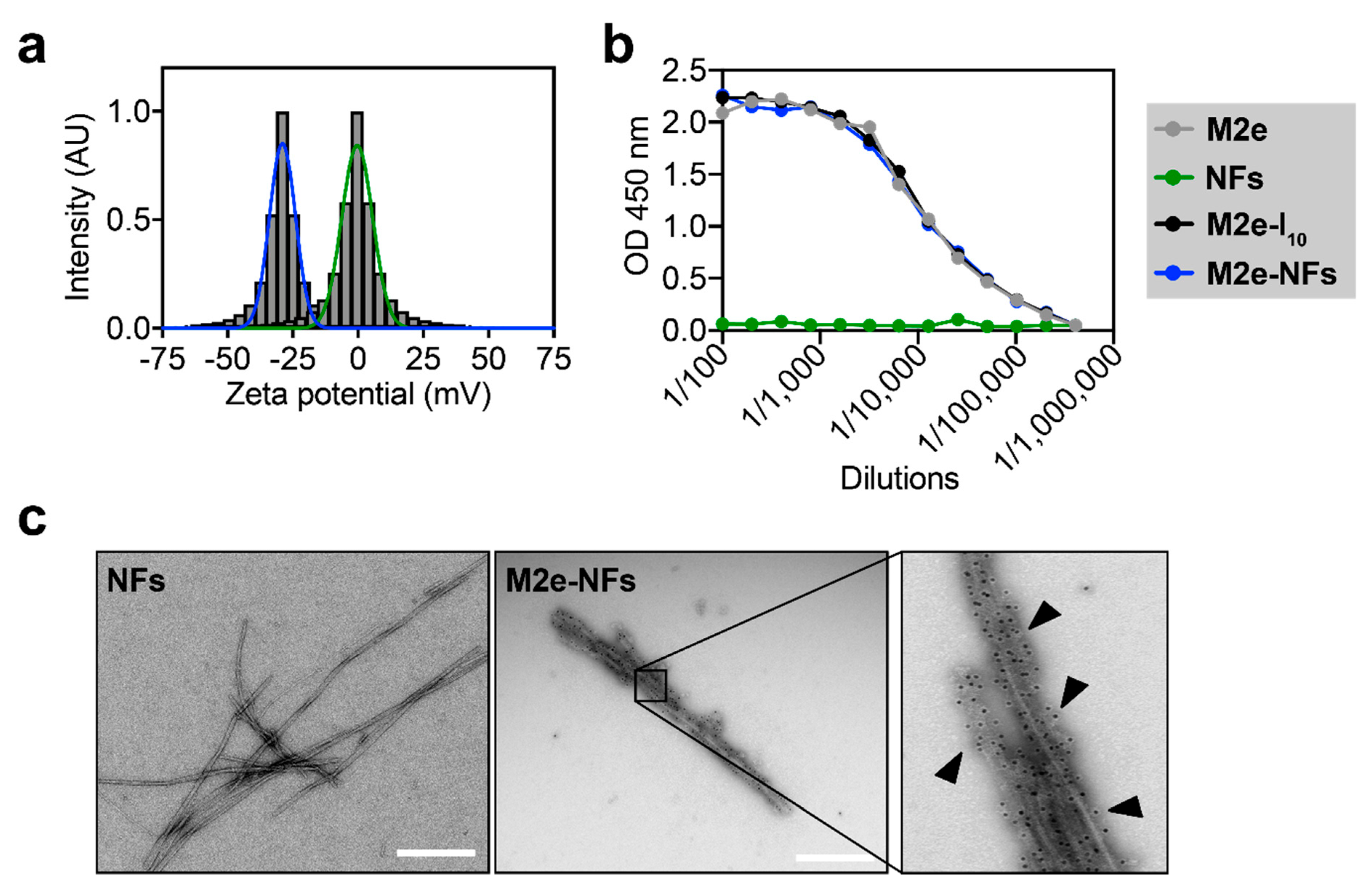
3.2. The M2e Epitope is Accesible on the Surface of the Fibrils
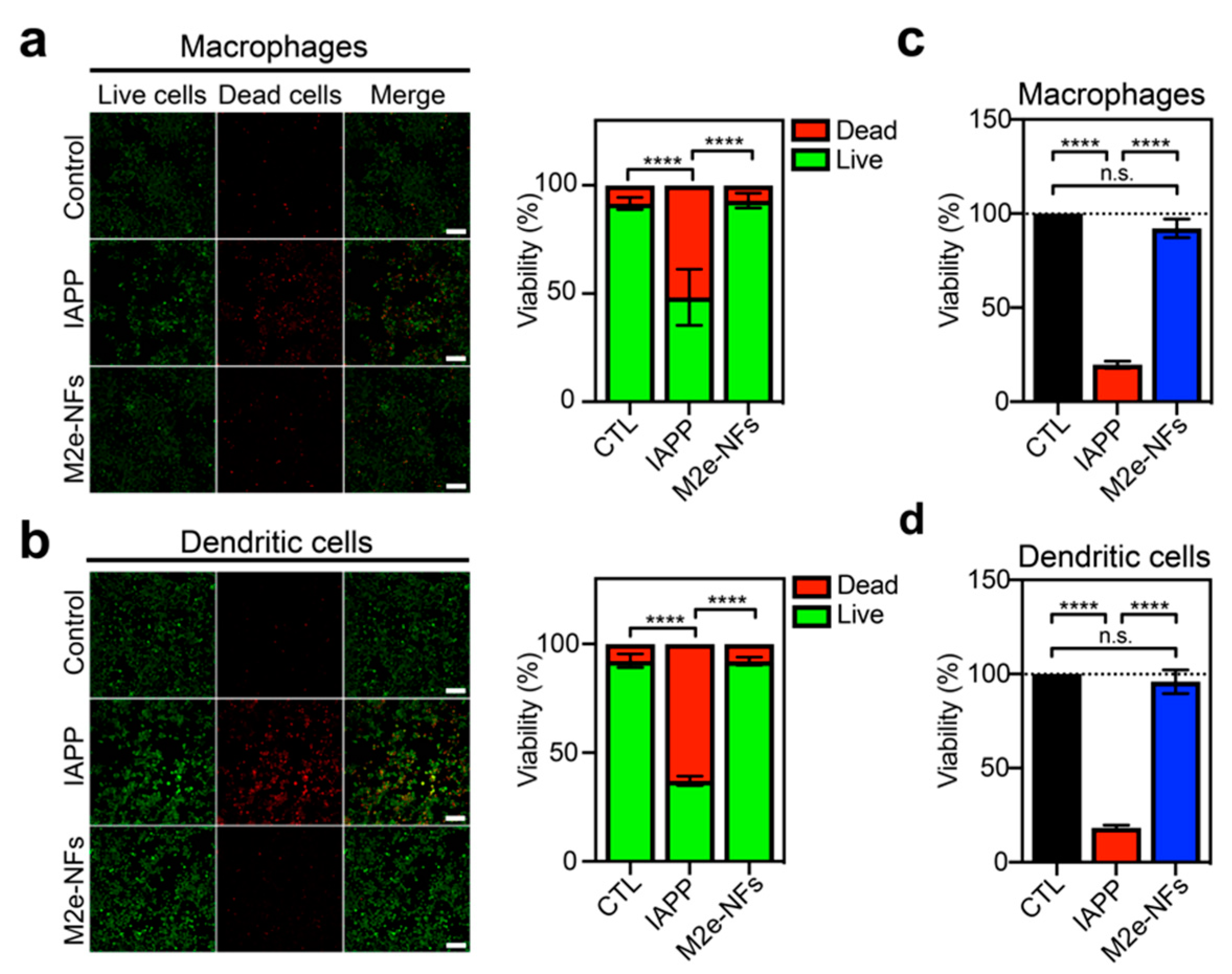
3.3. M2e-NFs are Cytocompatible
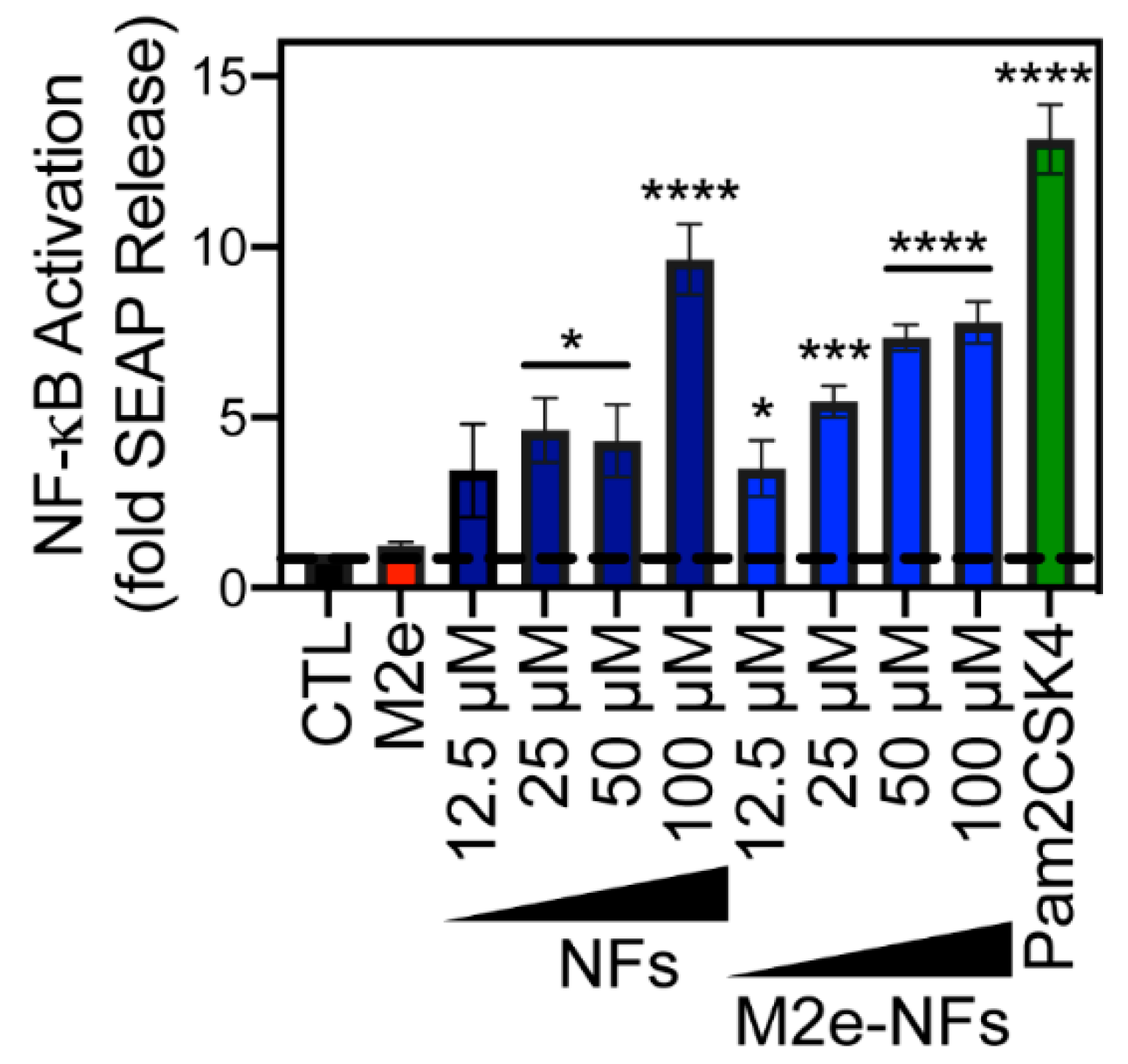
3.4. M2e-NFs Activate the Heterodimeric Toll-Like Receptor 2/6
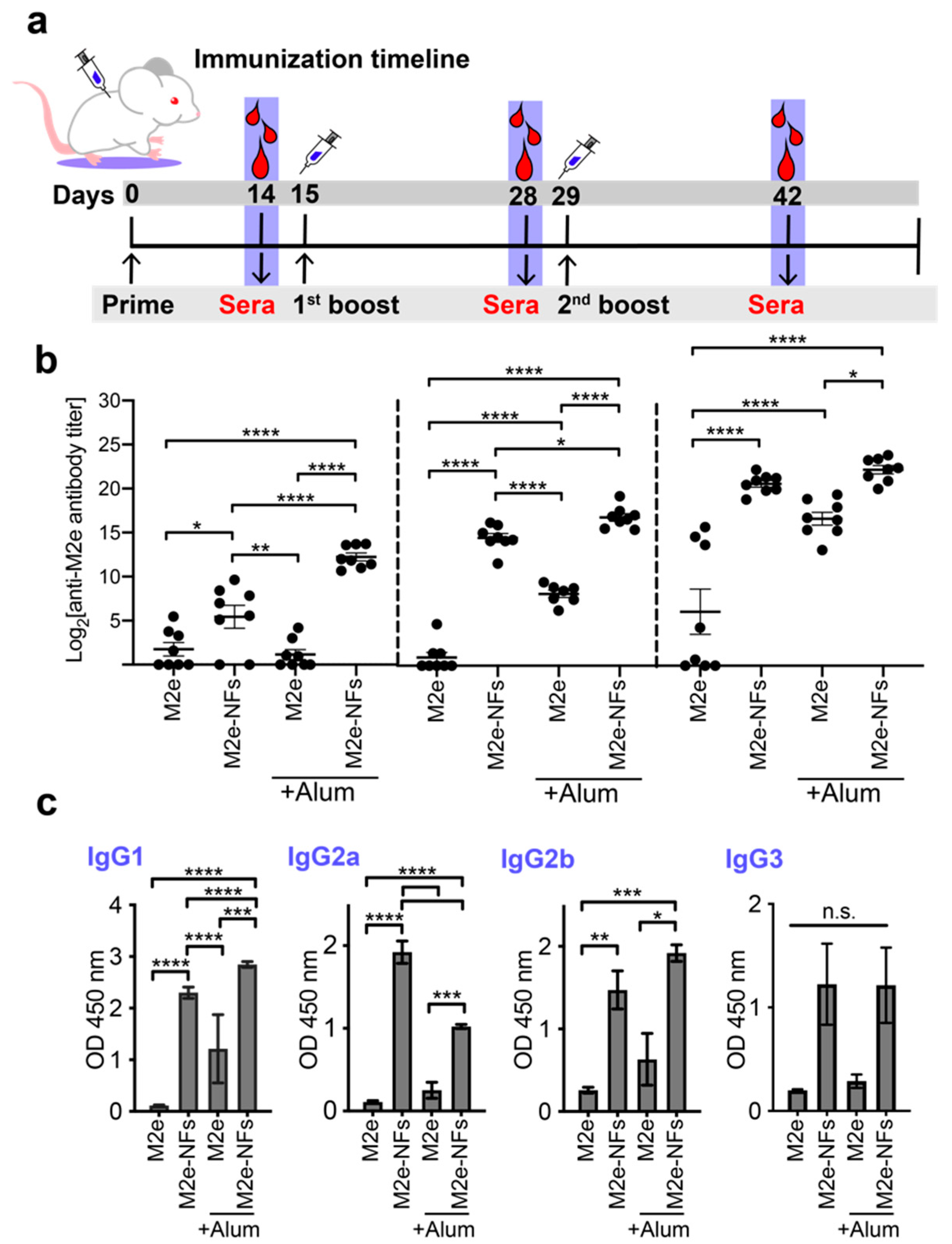
3.5. Cross-β Fibrils Potentiate the Anti-M2e Specific Immune Response
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Aguilar, J.; Rodríguez, E. Vaccine adjuvants revisited. Vaccine 2007, 25, 3752–3762. [Google Scholar] [CrossRef] [PubMed]
- Zaman, M.; Good, M.F.; Toth, I. Nanovaccines and their mode of action. Methods 2013, 60, 226–231. [Google Scholar] [CrossRef] [PubMed]
- Boraschi, D.; Italiani, P. From Antigen Delivery System to Adjuvanticy: The Board Application of Nanoparticles in Vaccinology. Vaccines 2015, 3, 930–939. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Al-Halifa, S.; Gauthier, L.; Arpin, D.; Bourgault, S.; Archambault, D. Nanoparticle-Based Vaccines Against Respiratory Viruses. Front. Immunol. 2019, 10, 22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laval, J.-M.; Thomas, D.; Mazeran, P.-E. Nanobiotechnology and its role in the development of new analytical devices. Analyst 2000, 125, 29–33. [Google Scholar] [CrossRef]
- Luo, M.; Samandi, L.; Wang, Y.; Chen, Z.J.; Gao, J. Synthetic nanovaccines for immunotherapy. J. Control. Release 2017, 263, 200–210. [Google Scholar] [CrossRef]
- Mamo, T.; Poland, G.A. Nanovaccinology: The next generation of vaccines meets 21st century materials science and engineering. Vaccine 2012, 30, 6609–6611. [Google Scholar] [CrossRef] [PubMed]
- Kubackova, J.; Zbytovska, J.; Holas, O. Nanomaterials for direct and indirect immunomodulation: A review of applications. Eur. J. Pharm. Sci. 2020, 142, 105139. [Google Scholar] [CrossRef]
- Yan, X.; Zhou, M.; Yu, S.; Jin, Z.; Zhao, K. An overview of biodegradable nanomaterials and applications in vaccines. Vaccine 2020, 38, 1096–1104. [Google Scholar] [CrossRef]
- Zottig, X.; Côté-Cyr, M.; Arpin, D.; Archambault, D.; Bourgault, S. Protein Supramolecular Structures: From Self-Assembly to Nanovaccine Design. Nanomaterials 2020, 10, 1008. [Google Scholar] [CrossRef]
- Noad, R.J.; Roy, P. Virus-like particles as immunogens. Trends Microbiol. 2003, 11, 438–444. [Google Scholar] [CrossRef]
- Malik, H.; Khan, F.H.; Ahsan, H. Human papillomavirus: Current status and issues of vaccination. Arch. Virol. 2013, 159, 199–205. [Google Scholar] [CrossRef] [PubMed]
- Branco, M.C.; Sigano, D.M.; Schneider, J.P. Materials from peptide assembly: Towards the treatment of cancer and transmittable disease. Curr. Opin. Chem. Biol. 2011, 15, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Eskandari, S.; Guerin, T.; Toth, I.; Stephenson, R.J. Recent advances in self-assembled peptides: Implications for targeted drug delivery and vaccine engineering. Adv. Drug Deliv. Rev. 2017, 110, 169–187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joo, J.; Poon, C.; Yoo, S.P.; Chung, E.J. Shape Effects of Peptide Amphiphile Micelles for Targeting Monocytes. Molecules 2018, 23, 2786. [Google Scholar] [CrossRef] [Green Version]
- Zaman, M.; Abdel-Aal, A.-B.M.; Fujita, Y.; Phillipps, K.S.M.; Batzloff, M.R.; Good, M.F.; Toth, I. Immunological Evaluation of Lipopeptide Group A Streptococcus (GAS) Vaccine: Structure-Activity Relationship. PLoS ONE 2012, 7, e30146. [Google Scholar] [CrossRef] [Green Version]
- Zaman, M.; Abdel-Aal, A.-B.M.; Fujita, Y.; Ziora, Z.M.; Batzloff, M.R.; Good, M.F.; Toth, I. Structure–Activity Relationship for the Development of a Self-Adjuvanting Mucosally Active Lipopeptide Vaccine against Streptococcus pyogenes. J. Med. Chem. 2012, 55, 8515–8523. [Google Scholar] [CrossRef]
- Riedel, T.; Ghasparian, A.; Moehle, K.; Rusert, P.; Trkola, A.; Robinson, J.A. Synthetic Virus-Like Particles and Conformationally Constrained Peptidomimetics in Vaccine Design. ChemBioChem 2011, 12, 2829–2836. [Google Scholar] [CrossRef] [Green Version]
- Ghasparian, A.; Riedel, T.; Koomullil, J.; Moehle, K.; Gorba, C.; Svergun, D.I.; Perriman, A.W.; Mann, S.; Tamborrini, M.; Pluschke, G.; et al. Engineered Synthetic Virus-Like Particles and Their Use in Vaccine Delivery. ChemBioChem 2011, 12, 100–109. [Google Scholar] [CrossRef]
- Tamborrini, M.; Geib, N.; Marrero-Nodarse, A.; Jud, M.; Hauser, J.; Aho, C.; Lamelas, A.; Zuniga, A.; Pluschke, G.; Ghasparian, A.; et al. A Synthetic Virus-Like Particle Streptococcal Vaccine Candidate Using B-Cell Epitopes from the Proline-Rich Region of Pneumococcal Surface Protein A. Vaccines 2015, 3, 850–874. [Google Scholar] [CrossRef]
- Rudra, J.S.; Tian, Y.F.; Jung, J.P.; Collier, J.H. A self-assembling peptide acting as an immune adjuvant. Proc. Natl. Acad. Sci. USA 2010, 107, 622–627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wen, Y.; Collier, J.H. Supramolecular peptide vaccines: Tuning adaptive immunity. Curr. Opin. Immunol. 2015, 35, 73–79. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Azmi, F.; Fuaad, A.A.; Giddam, A.K.; Batzloff, M.R.; Good, M.F.; Skwarczynski, M.; Toth, I. Self-adjuvanting vaccine against group A streptococcus: Application of fibrillized peptide and immunostimulatory lipid as adjuvant. Bioorganic Med. Chem. 2014, 22, 6401–6408. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rudra, J.S.; Mishra, J.; Chong, A.S.; Mitchell, R.A.; Nardin, E.H.; Nussenzweig, V.; Collier, J.H. Self-assembled peptide nanofibers raising durable antibody responses against a malaria epitope. Biomaterials 2012, 33, 6476–6484. [Google Scholar] [CrossRef] [Green Version]
- Babych, M.; Bertheau-Mailhot, G.; Zottig, X.; Dion, J.; Gauthier, L.; Archambault, D.; Bourgault, S.; Gauhier, L. Engineering and evaluation of amyloid assemblies as a nanovaccine against the Chikungunya virus. Nanoscale 2018, 10, 19547–19556. [Google Scholar] [CrossRef]
- Kolpe, A.; Schepens, B.; Fiers, W.; Saelens, X. M2-based influenza vaccines: Recent advances and clinical potential. Expert Rev. Vaccines 2017, 16, 123–136. [Google Scholar] [CrossRef]
- Watkins, H.C.; Rappazzo, C.G.; Higgins, J.S.; Sun, X.; Brock, N.; Chau, A.; Misra, A.; Cannizzo, J.P.; King, M.R.; Maines, T.R.; et al. Safe Recombinant Outer Membrane Vesicles that Display M2e Elicit Heterologous Influenza Protection. Mol. Ther. 2017, 25, 989–1002. [Google Scholar] [CrossRef] [Green Version]
- Schepens, B.; De Vlieger, D.; Saelens, X. Vaccine options for influenza: Thinking small. Curr. Opin. Immunol. 2018, 53, 22–29. [Google Scholar] [CrossRef]
- De Filette, M.; Jou, W.M.; Birkett, A.; Lyons, K.; Schultz, B.; Tonkyro, A.; Resch, S.; Fiers, W. Universal influenza A vaccine: Optimization of M2-based constructs. Virolology 2005, 337, 149–161. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, P.T.; Zottig, X.; Sebastiao, M.; Bourgault, S. Role of Site-Specific Asparagine Deamidation in Islet Amyloid Polypeptide Amyloidogenesis: Key Contributions of Residues 14 and 21. Biochemistry 2017, 56, 3808–3817. [Google Scholar] [CrossRef]
- De Carufel, C.A.; Quittot, N.; Nguyen, P.T.; Bourgault, S. Delineating the Role of Helical Intermediates in Natively Unfolded Polypeptide Amyloid Assembly and Cytotoxicity. Angew. Chem. 2015, 127, 14591–14595. [Google Scholar] [CrossRef]
- Zottig, X.; Al-Halifa, S.; Babych, M.; Quittot, N.; Archambault, D.; Bourgault, S. Guiding the Morphology of Amyloid Assemblies by Electrostatic Capping: From Polymorphic Twisted Fibrils to Uniform Nanorods. Small 2019, 15, e1901806. [Google Scholar] [CrossRef] [PubMed]
- Hervé, P.-L.; Raliou, M.; Bourdieu, C.; Dubuquoy, C.; Petit-Camurdan, A.; Bertho, N.; Eleouet, J.-F.; Chevalier, C.; Riffault, S. A Novel Subnucleocapsid Nanoplatform for Mucosal Vaccination against Influenza Virus That Targets the Ectodomain of Matrix Protein 2. J. Virol. 2014, 88, 325–338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tovar, J.D.; Claussen, R.C.; Stupp, S.I. Probing the Interior of Peptide Amphiphile Supramolecular Aggregates. J. Am. Chem. Soc. 2005, 127, 7337–7345. [Google Scholar] [CrossRef] [PubMed]
- Bourgault, S.; Solomon, J.P.; Reixach, N.; Kelly, J.W. Sulfated Glycosaminoglycans Accelerate Transthyretin Amyloidogenesis by Quaternary Structural Conversion. Biochemistry 2011, 50, 1001–1015. [Google Scholar] [CrossRef] [Green Version]
- Zhao, R.; So, M.; Maat, H.; Ray, N.J.; Arisaka, F.; Goto, Y.; Carver, J.A.; Hall, D. Measurement of amyloid formation by turbidity assay—seeing through the cloud. Biophys. Rev. 2016, 8, 445–471. [Google Scholar] [CrossRef] [Green Version]
- Sebastiao, M.; Quittot, N.; Bourgault, S. Thioflavin T fluorescence to analyse amyloid formation kinetics: Measurement frequency as a factor explaining irreproducibility. Anal. Biochem. 2017, 532, 83–86. [Google Scholar] [CrossRef]
- Wolfe, L.S.; Calabrese, M.F.; Nath, A.; Blaho, D.V.; Miranker, A.D.; Xiong, Y. Protein-induced photophysical changes to the amyloid indicator dye thioflavin T. Proc. Natl. Acad. Sci. USA 2010, 107, 16863–16868. [Google Scholar] [CrossRef] [Green Version]
- Mao, X.-B.; Wang, C.-X.; Wu, X.-K.; Ma, X.-J.; Liu, L.; Zhang, L.; Niu, L.; Guo, Y.-Y.; Li, D.-H.; Yang, Y.-L. Beta structure motifs of islet amyloid polypeptides identified through surface-mediated assemblies. Proc. Natl. Acad. Sci. USA 2011, 108, 19605–19610. [Google Scholar] [CrossRef] [Green Version]
- Batista, F.D.; Harwood, N.E. Antigen presentation to B cells. F1000 Biol. Rep. 2010, 2, 87. [Google Scholar] [CrossRef] [Green Version]
- Bennett, K.M.; Gorham, R.D.J.; Gusti, V.; Trinh, L.; Morikis, D.; Lo, D.D. Hybrid flagellin as a T cell independent vaccine scaffold. BMC Biotechnol. 2015, 15, 71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saar, K.L.; Mezzenga, R. Amyloid Fibrils as Building Blocks for Natural and Artificial Functional Materials. Adv. Mater. 2016, 28, 6546–6561. [Google Scholar]
- Chiti, F.; Dobson, C.M. Protein Misfolding, Amyloid Formation, and Human Disease: A Summary of Progress Over the Last Decade. Annu. Rev. Biochem. 2017, 86, 27–68. [Google Scholar] [CrossRef] [PubMed]
- Bourgault, S.; Lim, H.R.; Buxbaum, J.N.; Kelly, J.W.; Price, J.L.; Reixach, N. Mechanisms of transthyretin cardiomyocyte toxicity inhibition by resveratrol analogs. Biochem. Biophys. Res. Commun. 2011, 410, 707–713. [Google Scholar] [CrossRef] [Green Version]
- Reixach, N.; Deechongkit, S.; Jiang, X.; Kelly, J.W.; Buxbaum, J.N. Tissue damage in the amyloidoses: Transthyretin monomers and nonnative oligomers are the major cytotoxic species in tissue culture. Proc. Natl. Acad. Sci. USA 2004, 101, 2817–2822. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Otzen, D.E. Functional amyloid. Prion 2010, 4, 256–264. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Godin, E.; Nguyen, P.T.; Zottig, X.; Bourgault, S. Identification of a hinge residue controlling islet amyloid polypeptide self-assembly and cytotoxicity. J. Biol. Chem. 2019, 294, 8452–8463. [Google Scholar] [CrossRef]
- Wen, Y.; Waltman, A.; Han, H.; Collier, J.H. Switching the Immunogenicity of Peptide Assemblies Using Surface Properties. ACS Nano 2016, 10, 9274–9286. [Google Scholar] [CrossRef]
- Mora-Solano, C.; Wen, Y.; Han, H.; Chen, J.; Chong, A.S.; Miller, M.L.; Pompano, R.R.; Collier, J.H. Active immunotherapy for TNF-mediated inflammation using self-assembled peptide nanofibers. Biomaterials 2017, 149, 1–11. [Google Scholar] [CrossRef]
- Jana, M.; Palencia, C.A.; Pahan, K. Fibrillar amyloid-beta peptides activate microglia via TLR2: Implications for Alzheimer’s disease. J. Immunol. 2008, 181, 7254–7262. [Google Scholar] [CrossRef]
- Westwell-Roper, C.; Denroche, H.C.; Ehses, J.A.; Verchere, C.B. Differential Activation of Innate Immune Pathways by Distinct Islet Amyloid Polypeptide (IAPP) Aggregates. J. Boil. Chem. 2016, 291, 8908–8917. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reed, S.G.; Orr, M.T.; Fox, C.B. Key roles of adjuvants in modern vaccines. Nat. Med. 2013, 19, 1597–1608. [Google Scholar] [CrossRef] [PubMed]
- Eawate, S.; Babiuk, L.A.B.; Emutwiri, G. Mechanisms of Action of Adjuvants. Front. Immunol. 2013, 4, 114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kelley, N.; Jeltema, D.; Duan, Y.; He, Y. The NLRP3 Inflammasome: An Overview of Mechanisms of Activation and Regulation. Int. J. Mol. Sci. 2019, 20, 3328. [Google Scholar] [CrossRef] [Green Version]
- Fernandes-Alnemri, T.; Kang, S.; Anderson, C.; Sagara, J.; Fitzgerald, K.A.; Alnemri, E.S. Cutting edge: TLR signaling licenses IRAK1 for rapid activation of the NLRP3 inflammasome. J. Immunol. 2013, 191, 3995–3999. [Google Scholar] [CrossRef]
- Nakanishi, A.; Kaneko, N.; Takeda, H.; Sawasaki, T.; Morikawa, S.; Zhou, W.; Kurata, M.; Yamamoto, T.; Akbar, S.M.F.; Zako, T.; et al. Amyloid beta directly interacts with NLRP3 to initiate inflammasome activation: Identification of an intrinsic NLRP3 ligand in a cell-free system. Inflamm. Regen. 2018, 38, 27. [Google Scholar] [CrossRef]
- Morikawa, S.; Kaneko, N.; Okumura, C.; Taguchi, H.; Kurata, M.; Yamamoto, T.; Osawa, H.; Nakanishi, A.; Zako, T.; Masumoto, J. IAPP/amylin deposition, which is correlated with expressions of ASC and IL-1beta in beta-cells of Langerhans’ islets, directly initiates NLRP3 inflammasome activation. Int. J. Immunopathol. Pharmacol. 2018, 32, 2058738418788749. [Google Scholar] [CrossRef]
- Masters, S.L.; Dunne, A.; Subramanian, S.L.; Hull, R.L.; Tannahill, G.M.; Sharp, F.A.; Becker, C.; Franchi, L.; Yoshihara, E.; Chen, Z.; et al. Activation of the NLRP3 inflammasome by islet amyloid polypeptide provides a mechanism for enhanced IL-1β in type 2 diabetes. Nat. Immunol. 2010, 11, 897–904. [Google Scholar] [CrossRef]
- Stevens, T.L.; Bossie, A.; Sanders, V.M.; Fernandez-Botran, R.; Coffman, R.L.; Mosmann, T.R.; Vitetta, E.S. Regulation of antibody isotype secretion by subsets of antigen-specific helper T cells. Nat. Cell Biol. 1988, 334, 255–258. [Google Scholar] [CrossRef]
- Germann, T.; Bongartz, M.; Dlugonska, H.; Hess, H.; Schmitt, E.; Kolbe, L.; Kölsch, E.; Podlaski, F.J.; Gately, M.K.; Rüde, E. Interleukin-12 profoundly up-regulates the synthesis of antigen-specific complement-fixing IgG2a, IgG2b and IgG3 antibody subclassesin vivo. Eur. J. Immunol. 1995, 25, 823–829. [Google Scholar] [CrossRef]
- Lefeber, D.J.; Benaissa-Trouw, B.; Vliegenthart, J.F.G.; Kamerling, J.P.; Jansen, W.T.M.; Kraaijeveld, K.; Snippe, H. Th1-Directing Adjuvants Increase the Immunogenicity of Oligosaccharide-Protein Conjugate Vaccines Related to Streptococcus pneumoniae Type 3. Infect. Immun. 2003, 71, 6915–6920. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gavin, A.L.; Hoebe, K.; Duong, B.; Ota, T.; Martin, C.; Beutler, B.; Nemazee, D. Adjuvant-Enhanced Antibody Responses in the Absence of Toll-Like Receptor Signaling. Science 2006, 314, 1936–1938. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Denis, J.; Acosta-Ramirez, E.; Zhao, Y.; Hamelin, M.-E.; Koukavica, I.; Baz, M.; Abed, Y.; Savard, C.; Paré, C.; Lopez-Macias, C.I.R.; et al. Development of a universal influenza A vaccine based on the M2e peptide fused to the papaya mosaic virus (PapMV) vaccine platform. Vaccine 2008, 26, 3395–3403. [Google Scholar] [CrossRef] [PubMed]
- Skwarczynski, M.; Toth, I. Recent advances in peptide-based subunit nanovaccines. Nanomedicine 2014, 9, 2657–2669. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Al-Halifa, S.; Zottig, X.; Babych, M.; Côté-Cyr, M.; Bourgault, S.; Archambault, D. Harnessing the Activation of Toll-Like Receptor 2/6 by Self-Assembled Cross-β Fibrils to Design Adjuvanted Nanovaccines. Nanomaterials 2020, 10, 1981. https://doi.org/10.3390/nano10101981
Al-Halifa S, Zottig X, Babych M, Côté-Cyr M, Bourgault S, Archambault D. Harnessing the Activation of Toll-Like Receptor 2/6 by Self-Assembled Cross-β Fibrils to Design Adjuvanted Nanovaccines. Nanomaterials. 2020; 10(10):1981. https://doi.org/10.3390/nano10101981
Chicago/Turabian StyleAl-Halifa, Soultan, Ximena Zottig, Margaryta Babych, Mélanie Côté-Cyr, Steve Bourgault, and Denis Archambault. 2020. "Harnessing the Activation of Toll-Like Receptor 2/6 by Self-Assembled Cross-β Fibrils to Design Adjuvanted Nanovaccines" Nanomaterials 10, no. 10: 1981. https://doi.org/10.3390/nano10101981