Real-Time Estimation of the Risk of Death from Novel Coronavirus (COVID-19) Infection: Inference Using Exported Cases
Abstract
1. Introduction
2. Methods
2.1. Epidemiological Data
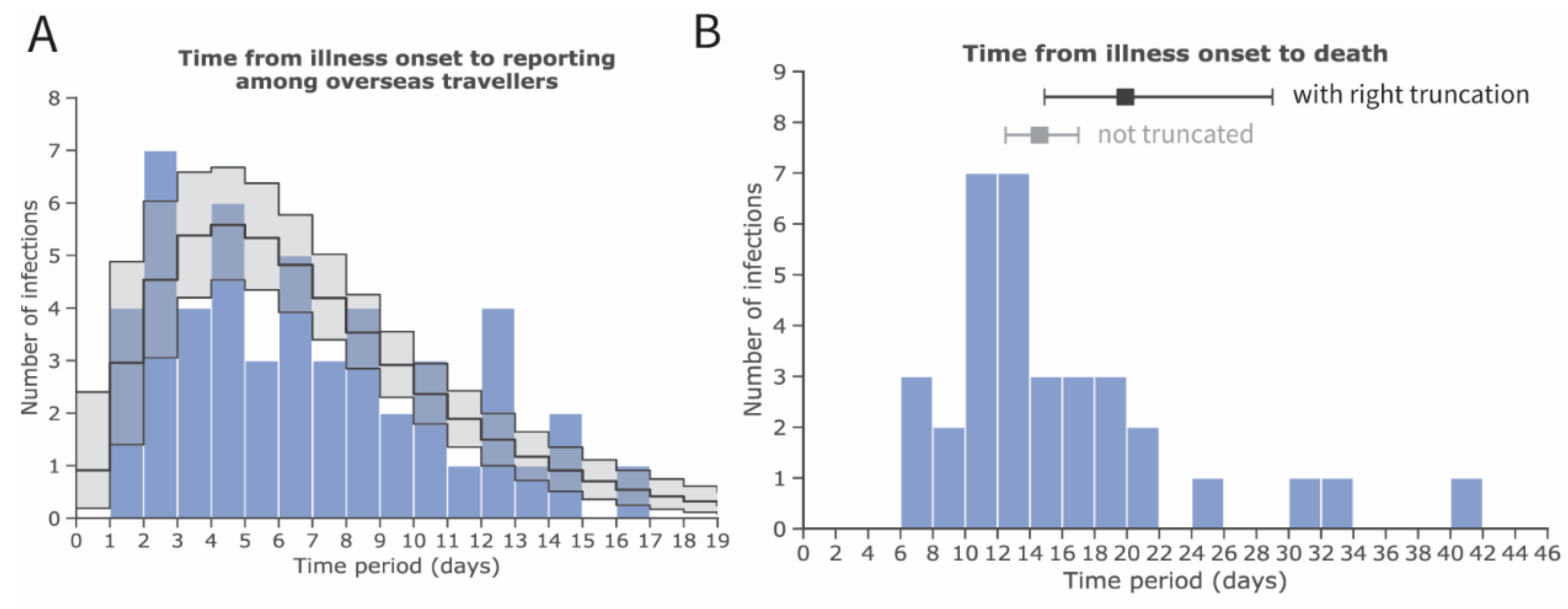
2.2. Estimation of the Delay Distributions
2.3. Statistical Inference
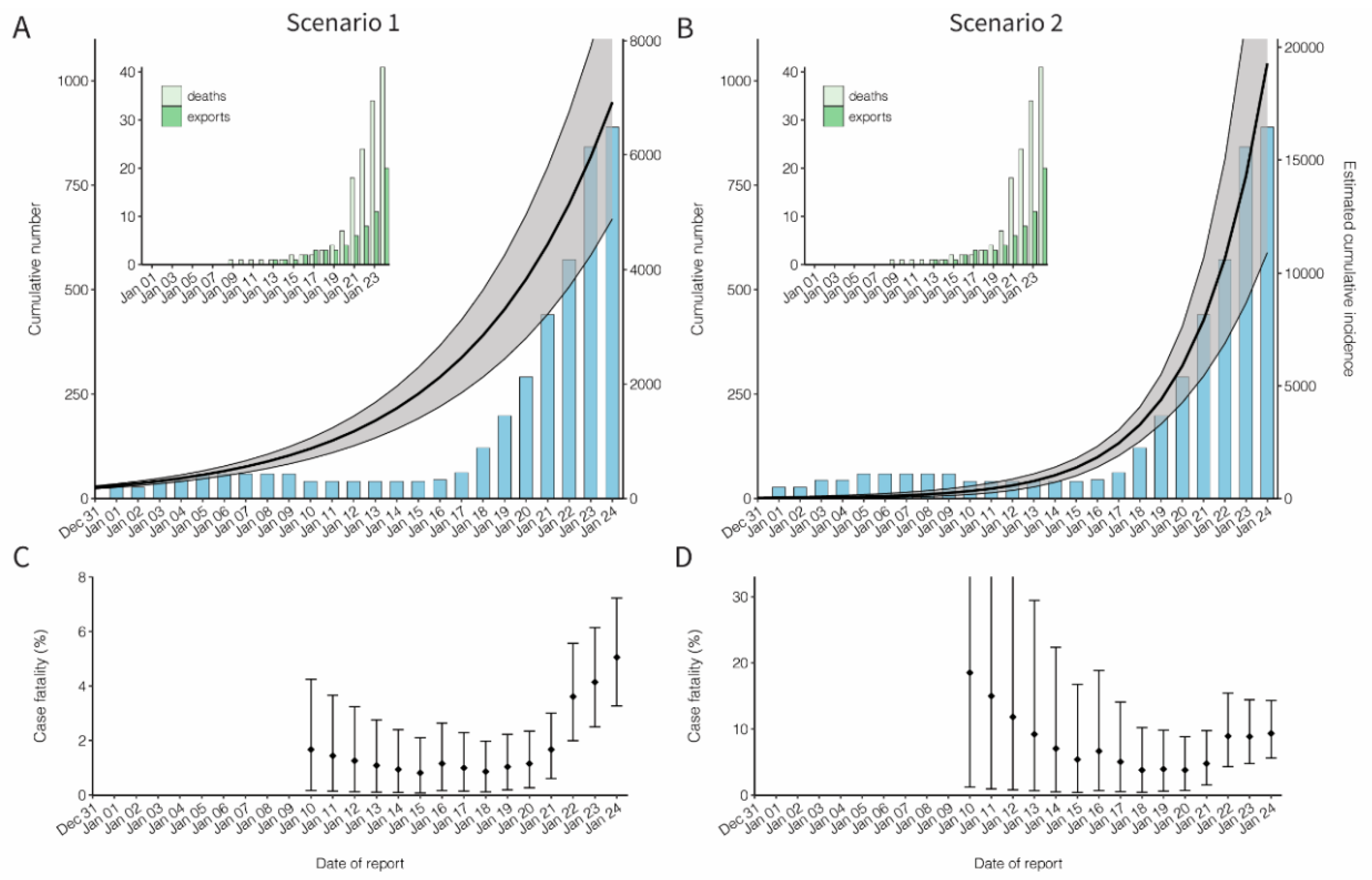
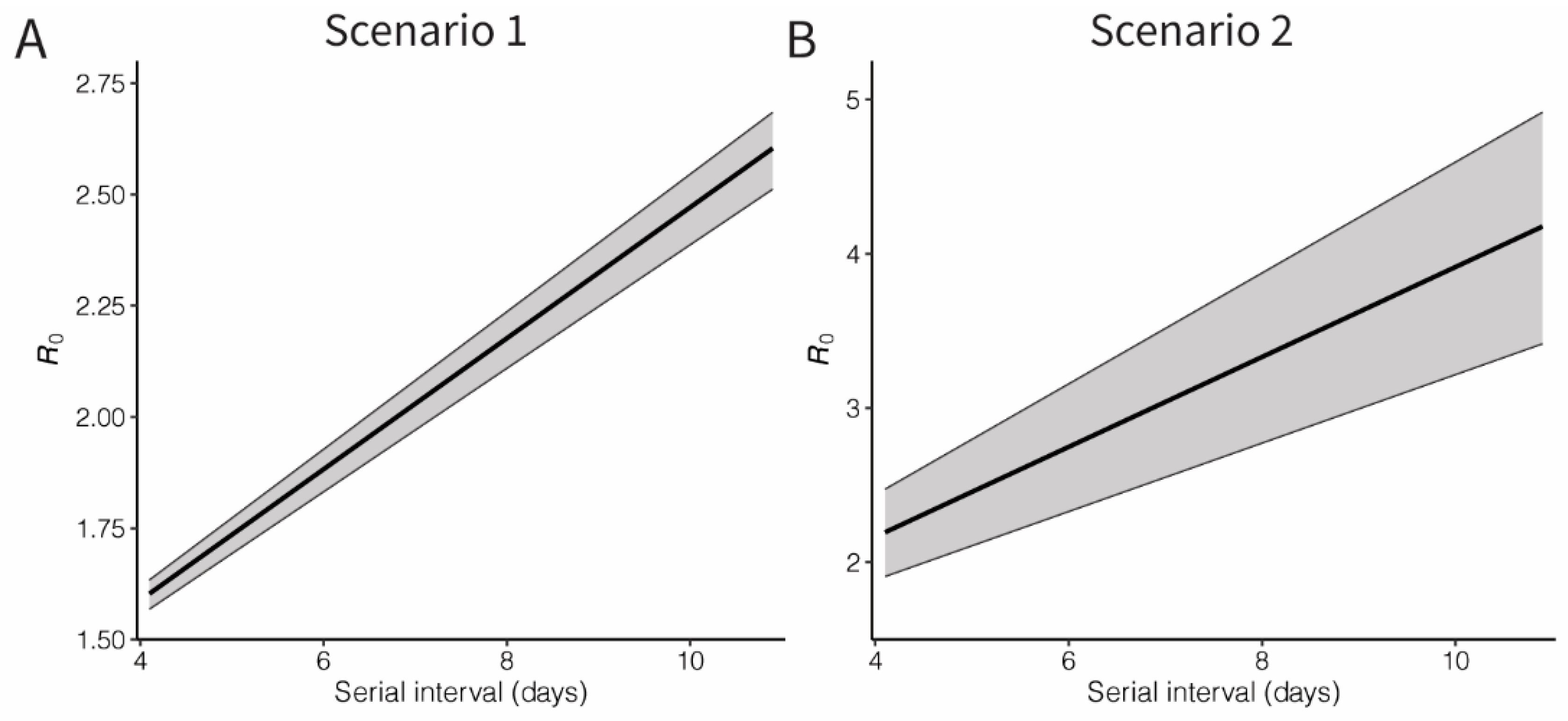
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- World Health Organization. Novel Coronavirus—China Disease Outbreak News. Available online: https://www.who.int/csr/don/12-january-2020-novel-coronavirus-china/en/ (accessed on 9 February 2020).
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Center for Disease Control and Prevention. 2019 Novel Coronavirus, Wuhan, China. Available online: https://www.cdc.gov/coronavirus/COVID-19/about/transmission.html (accessed on 24 January 2020).
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020. [Google Scholar] [CrossRef]
- Chan, J.F.W.; Yuan, S.; Kok, K.H.; To, K.K.W.; Chu, H.; Yang, J.; Xing, J.; Xing, F.; Liu, J.; Yip, C.C.; et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet 2020. [Google Scholar] [CrossRef]
- World Health Organization. Novel Coronavirus (COVID-19) Situation Report-3. Available online: https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200123-sitrep-3-COVID-19.pdf?sfvrsn=d6d23643_4 (accessed on 23 January 2020).
- Nishiura, H.; Jung, S.; Linton, N.M.; Kinoshita, R.; Yang, Y.; Hayashi, K.; Kobayashi, T.; Yuan, B.; Akhmetzhanov, A.R. The extent of transmission of novel coronavirus in Wuhan, China, 2020. J. Clin. Med. 2020, 9, 330. [Google Scholar] [CrossRef]
- Ghani, A.C.; Donnelly, C.A.; Cox, D.R.; Griffin, J.T.; Fraser, C.; Lam, T.H.; Ho, L.M.; Chan, W.S.; Anderson, R.M.; Hedley, A.J.; et al. Methods for estimating the case fatality ratio for a novel, emerging infectious disease. Am. J. Epidemiol. 2005, 16, 479–486. [Google Scholar] [CrossRef]
- Nishiura, H.; Klinkenberg, D.; Roberts, M.; Heesterbeek, J.A. Early epidemiological assessment of the virulence of emerging infectious diseases: A case study of an influenza pandemic. PLoS ONE 2009, 4, e6852. [Google Scholar] [CrossRef]
- Garske, T.; Legrand, J.; Donnelly, C.A.; Ward, H.; Cauchemez, S.; Fraser, C.; Ferguson, N.M.; Ghani, A.C. Assessing the severity of the novel influenza A/H1N1 pandemic. BMJ 2009, 339, b2840. [Google Scholar] [CrossRef]
- Thompson, R.N.; Gilligan, C.A.; Cunniffe, N.J. When Does A Minor Outbreak Become A Major Epidemic? Linking the Risk from Invading Pathogens to Practical Definitions of A Major Epidemic. Available online: https://www.biorxiv.org/content/10.1101/768853v2 (accessed on 9 February 2020).
- BBC NEWS. Coronavirus: Wuhan Shuts Public Transport over Outbreak. Available online: https://www.bbc.com/news/world-asia-china-51215348 (accessed on 28 January 2020).
- Linton, N.M.; Kobayashi, T.; Yang, Y.; Hayashi, K.; Akhmetzhanov, A.R.; Jung, S.; Yuan, B.; Kinoshita, R.; Nishiura, H. Epidemiological characteristics of novel coronavirus infection: A statistical analysis of publicly available case data. J. Clin. Med. 2020, in press. [Google Scholar]
- Lipsitch, M.; Cohen, T.; Cooper, B.; Robins, J.M.; Ma, S.; James, L.; Gopalakrishna, G.; Chew, S.K.; Tan, C.C.; Samore, M.H.; et al. Transmission dynamics and control of severe acute respiratory syndrome. Science 2003, 300, 1966–1970. [Google Scholar] [CrossRef]
- Li, M.; Dushoff, J.; Bolker, B.M. Fitting mechanistic epidemic models to data: A comparison of simple Markov chain Monte Carlo approaches. Stat. Methods Med. Res. 2018, 27, 1956–1967. [Google Scholar] [CrossRef] [PubMed]
- Kucharski, A.J.; Russell, T.W.; Diamond, C.; CMMID nCoV Working Group; Funk, S.; Eggo, R.M. Early dynamics of transmission and control of COVID-19: A mathematical modelling study. medRxiv 2020. [Google Scholar] [CrossRef]
- Thompson, R.N. 2019–2020 Wuhan coronavirus outbreak: Intense surveillance is vital for preventing sustained transmission in new locations. biorxiv 2020. [Google Scholar] [CrossRef]
- Boldog, P.; Tekeli, T.; Vizi, Z.; Denes, A.; Bartha, F.; Rost, G. Risk assessment of novel coronavirus COVID-19 outbreaks outside China. J. Clin. Med. 2020, 9, 571. [Google Scholar] [CrossRef] [PubMed]
- Donnelly, C.A.; Ghani, A.C.; Leung, G.M.; Hedley, A.J.; Fraser, C.; Riley, S.; Abu-Raddad, L.J.; Ho, L.-M.; Thach, T.-Q.; Chau, P.; et al. Epidemiological determinants of spread of causal agent of severe acute respiratory syndrome in Hong Kong. Lancet 2003, 361, 1761–1766. [Google Scholar] [CrossRef]
- Mizumoto, K.; Saitoh, M.; Chowell, G.; Miyamatsu, Y.; Nishiura, H. Estimating the risk of Middle East respiratory syndrome (MERS) death during the course of the outbreak in the Republic of Korea, 2015. Int. J. Infect. Dis. 2015, 39, 7–9. [Google Scholar] [CrossRef]
- Majumder, M.S.; Kluberg, S.A.; Mekaru, S.R.; Brownstein, J.S. Mortality risk factors for Middle East Respiratory Syndrome outbreak, South Korea, 2015. Emerg. Infect. Dis. 2015, 21, 2088–2090. [Google Scholar] [CrossRef]
- Mizumoto, K.; Endo, A.; Chowell, G.; Miyamatsu, Y.; Saitoh, M.; Nishiura, H. Real-time characterization of risks of death associated with the Middle East respiratory syndrome (MERS) in the Republic of Korea, 2015. BMC Med. 2015, 13, 228. [Google Scholar] [CrossRef]
- Nishiura, H.; Kobayashi, T.; Yang, Y.; Hayashi, K.; Miyama, T.; Kinoshita, R.; Linton, N.M.; Jung, S.-M.; Yuan, B.; Suzuki, A.; et al. The rate of under ascertainment of novel coronavirus (COVID-19) infection: Estimation using Japanese passengers data on evacuation flights. J. Clin. Med. 2020, 9, 419. [Google Scholar] [CrossRef]
- Zhao, S.; Ran, J.; Musa, S.S.; Yang, G.; Lou, Y.; Gao, D.; Yang, L.; He, D. Preliminary Estimation of the Basic Reproduction Number of Novel Coronavirus (COVID-19) in China, from 2019 to 2020: A Data-Driven Analysis in the Early Phase of the Outbreak. Available online: https://www.biorxiv.org/content/10.1101/2020.01.23.916395v1 (accessed on 29 January 2020).
- Imperial College London-MRC Centre for Global Infectious Disease Analysis. News/Wuhan Coronavirus. Available online: https://www.imperial.ac.uk/mrc-global-infectious-disease-analysis/news--wuhan-coronavirus/ (accessed on 28 January 2020).
- The University of Hong Kong-HKUMed WHO Collaborating Centre for Infectious Disease Epidemiology and Control. Nowcasting and Forecasting the Wuhan COVID-19 Outbreak. Available online: https://www.med.hku.hk/news/press/HKUMed-WHO-Collaborating-Centre-for-Infectious-Disease-Epidemiology-and-Control-releases%20real-time-nowcast-on-the-likely-extent-of-the-Wuhan-coronavirus (accessed on 28 January 2020).
- Read, J.M.; Bridgen, J.R.; Cummings, D.A.; Ho, A.; Jewell, C.P. Novel Coronavirus COVID-19: Early Estimation of Epidemiological Parameters and Epidemic Predictions. medRxiv 2020. [Google Scholar] [CrossRef]
- Fraser, C.; Riley, S.; Anderson, R.M.; Ferguson, N.M. Factors that make an infectious disease outbreak controllable. Proc. Natl. Acad. Sci. USA 2004, 101, 6146–6151. [Google Scholar] [CrossRef] [PubMed]
- Thompson, R.N.; Gilligan, C.A.; Cunniffe, N.J. Detecting presymptomatic infection is necessary to forecast major epidemics in the earliest stages of infectious disease outbreaks. PLoS Comput. Biol. 2016, 12, e1004836. [Google Scholar] [CrossRef] [PubMed]
| Importing Locations | Date of Report (2020) | Cumulative Count | Estimated Incidence in China (95% CI) | |
|---|---|---|---|---|
| Scenario 1 | Scenario 2 | |||
| Thailand | 13 January | 1 | 1828 (1397 2288) | 1369 (1003 1782) |
| Japan | 16 January | 2 | 2120 (1605 2672) | 1829 (1392 2309) |
| Thailand | 17 January | 3 | 2458 (1845 3119) | 2444 (1894 3033) |
| South Korea | 20 January | 4 | 3832 (2802 4962) | 5882 (4252 7629) |
| Taiwan, United States | 21 January | 6 | 4443 (3220 5792) | 7901 (5425, 10,662) |
| Thailand | 22 January | 8 | 5151 (3700 6761) | 10,626 (6897, 15,003) |
| Singapore, Vietnam | 23 January | 11 | 5972 (4252 7892) | 14,308 (8661, 21,250) |
| Japan, Nepal, South Korea, Singapore, Thailand, United States | 24 January | 20 | 6924 (4885 9211) | 19,289 (10,901, 30,158) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jung, S.-m.; Akhmetzhanov, A.R.; Hayashi, K.; Linton, N.M.; Yang, Y.; Yuan, B.; Kobayashi, T.; Kinoshita, R.; Nishiura, H. Real-Time Estimation of the Risk of Death from Novel Coronavirus (COVID-19) Infection: Inference Using Exported Cases. J. Clin. Med. 2020, 9, 523. https://doi.org/10.3390/jcm9020523
Jung S-m, Akhmetzhanov AR, Hayashi K, Linton NM, Yang Y, Yuan B, Kobayashi T, Kinoshita R, Nishiura H. Real-Time Estimation of the Risk of Death from Novel Coronavirus (COVID-19) Infection: Inference Using Exported Cases. Journal of Clinical Medicine. 2020; 9(2):523. https://doi.org/10.3390/jcm9020523
Chicago/Turabian StyleJung, Sung-mok, Andrei R. Akhmetzhanov, Katsuma Hayashi, Natalie M. Linton, Yichi Yang, Baoyin Yuan, Tetsuro Kobayashi, Ryo Kinoshita, and Hiroshi Nishiura. 2020. "Real-Time Estimation of the Risk of Death from Novel Coronavirus (COVID-19) Infection: Inference Using Exported Cases" Journal of Clinical Medicine 9, no. 2: 523. https://doi.org/10.3390/jcm9020523
APA StyleJung, S.-m., Akhmetzhanov, A. R., Hayashi, K., Linton, N. M., Yang, Y., Yuan, B., Kobayashi, T., Kinoshita, R., & Nishiura, H. (2020). Real-Time Estimation of the Risk of Death from Novel Coronavirus (COVID-19) Infection: Inference Using Exported Cases. Journal of Clinical Medicine, 9(2), 523. https://doi.org/10.3390/jcm9020523







