A Humanized Bone Niche Model Reveals Bone Tissue Preservation Upon Targeting Mitochondrial Complex I in Pseudo-Orthotopic Osteosarcoma
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines
2.2. Establishing Pseudo-Orthotopic Osteosarcomas
2.3. Micro Computed Tomography Imaging
2.4. Histology
2.5. Flow Cytometry
2.6. Cytokine Profiling
2.7. Statistical Analyses
3. Results
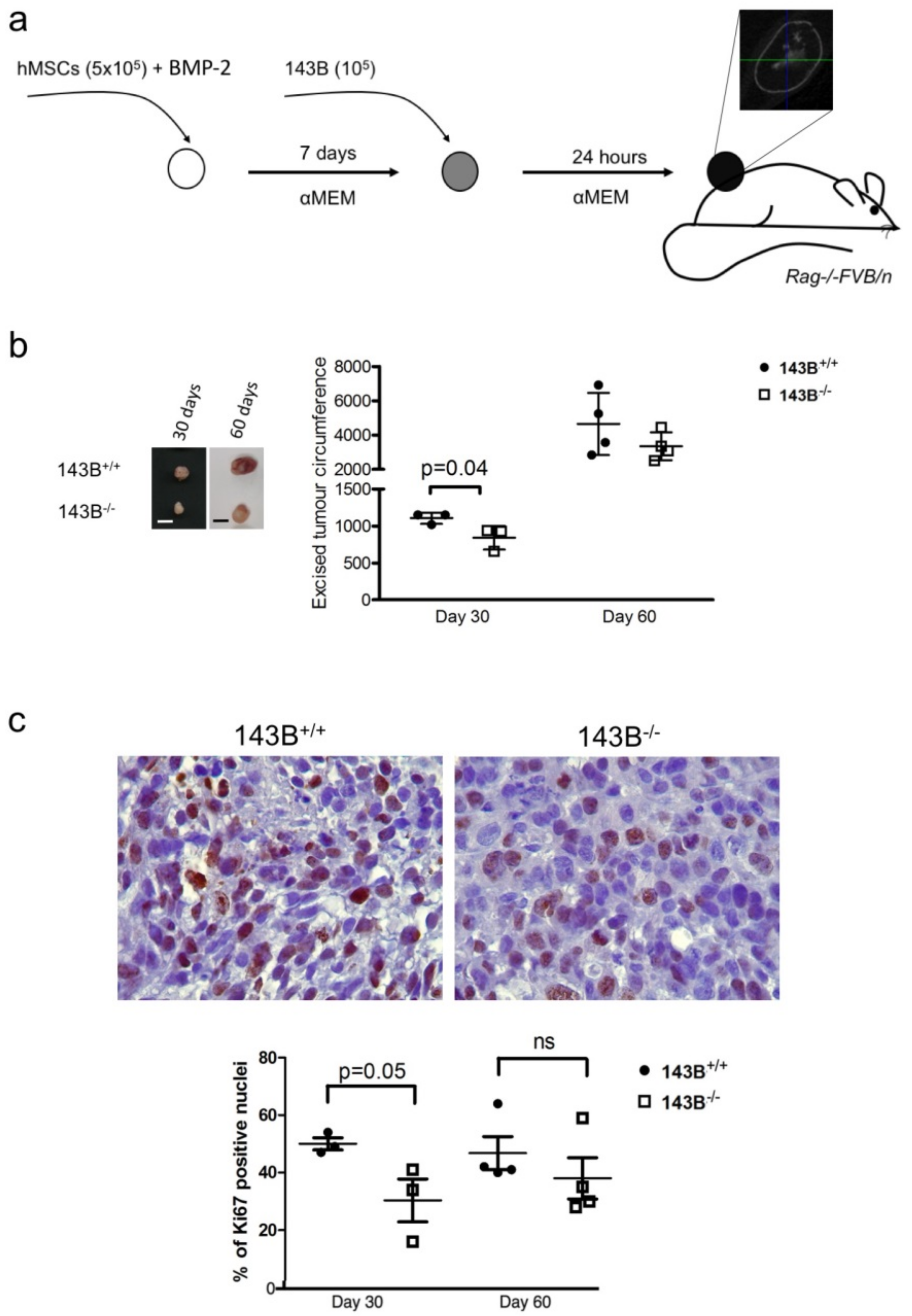
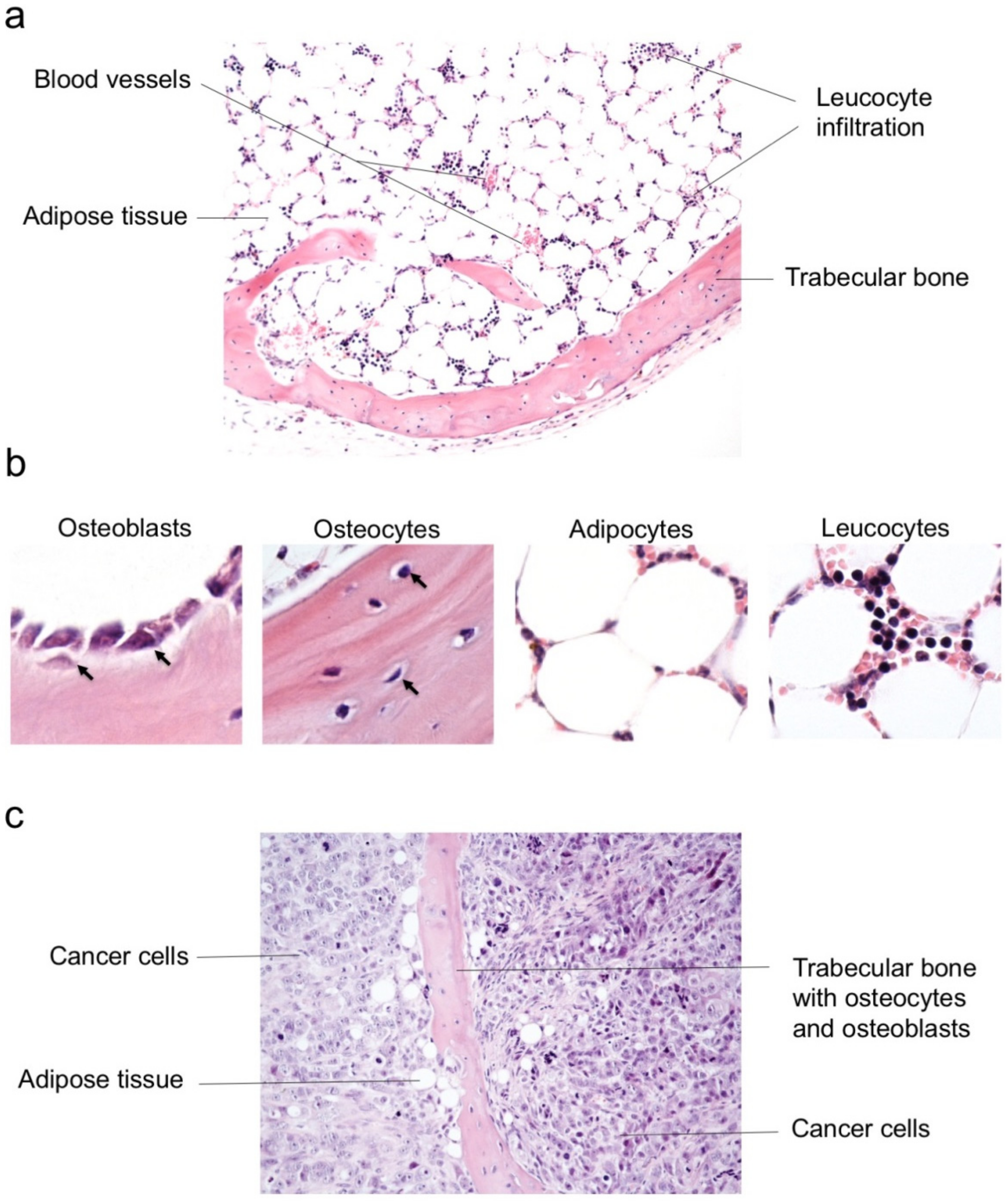
3.1. A Humanized Bone Niche Scaffold Recapitulates Mature Bone Characteristics and Serves as a Pseudo-Orthotopic Osteosarcoma Xenograft Model
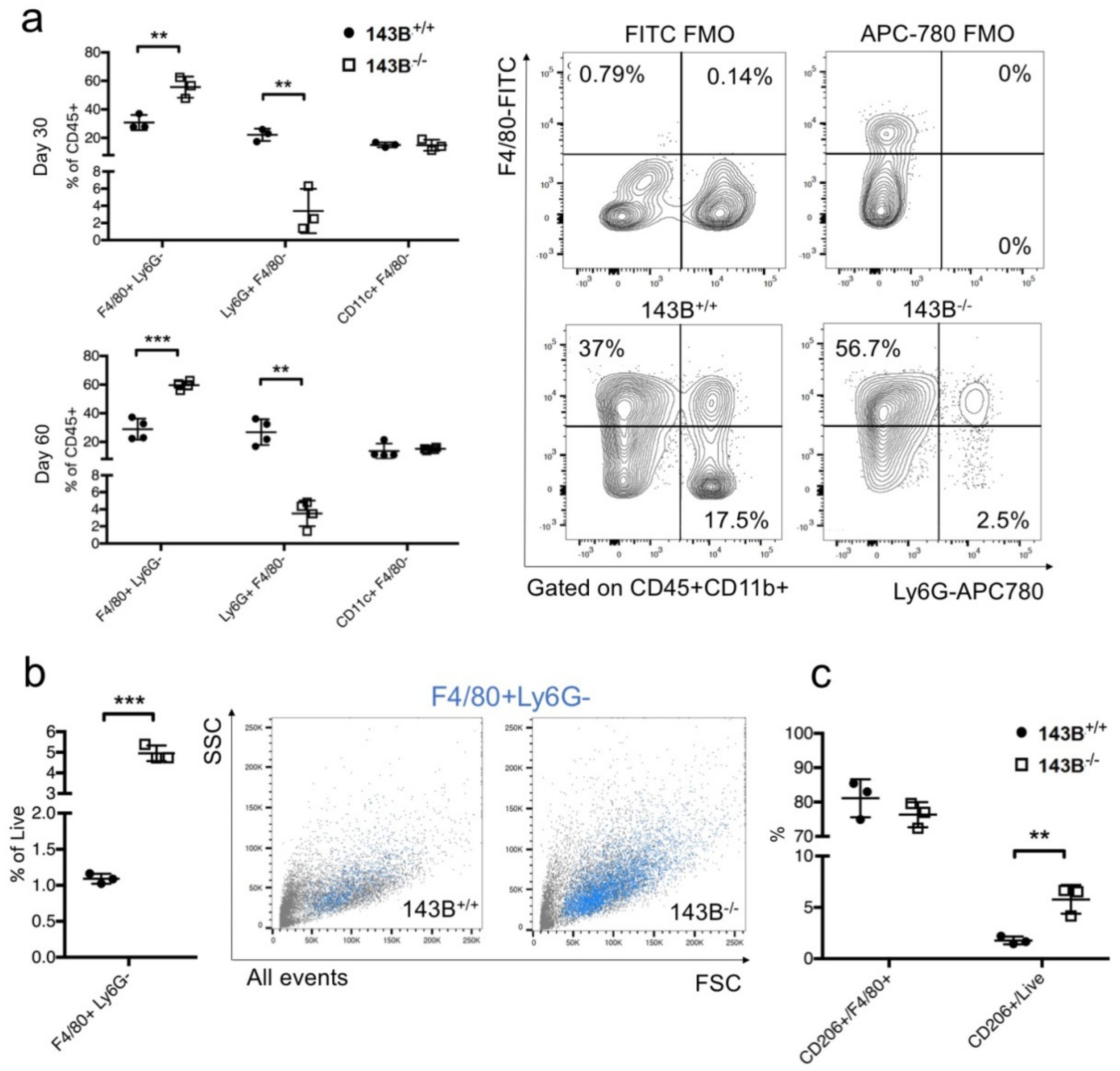
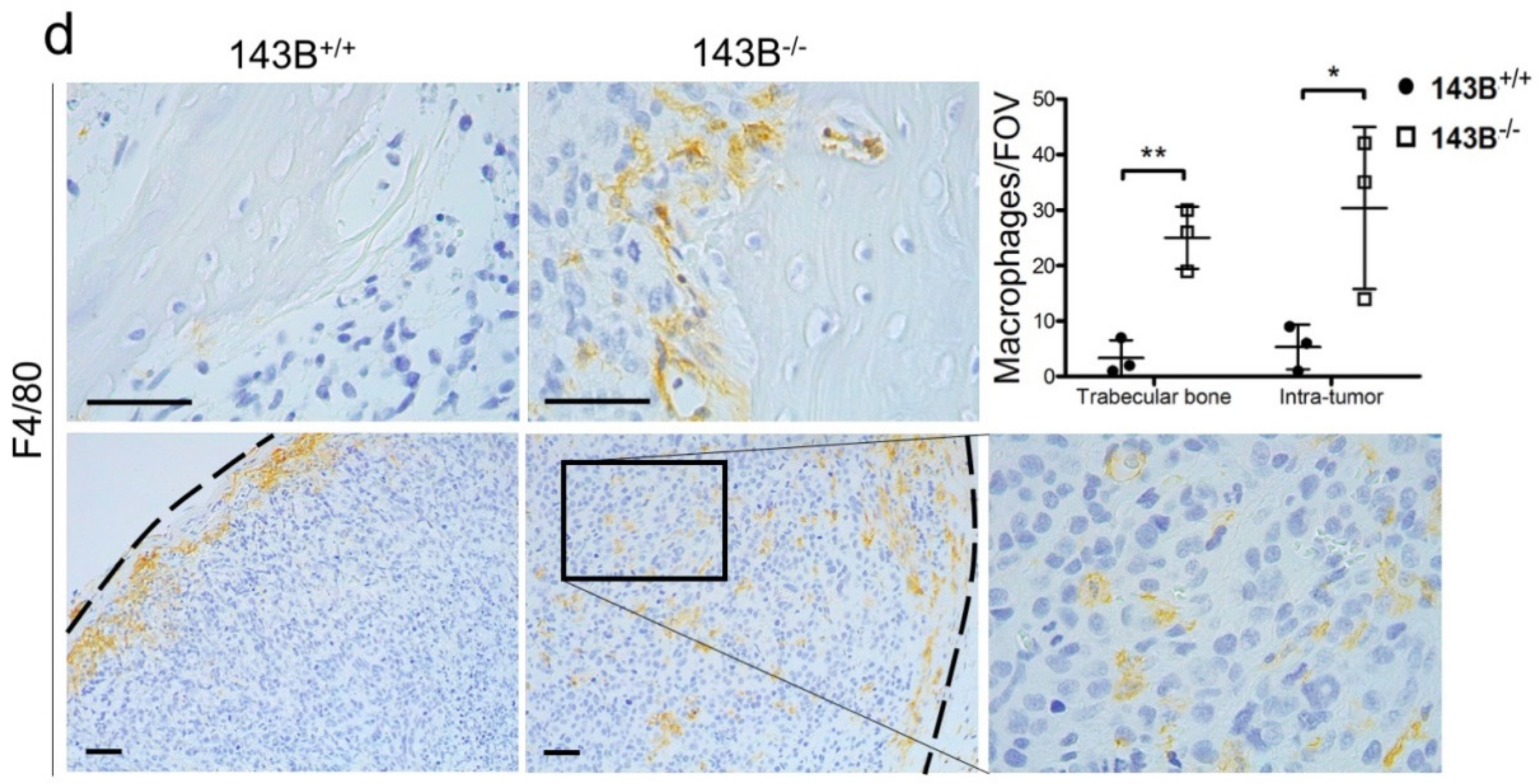
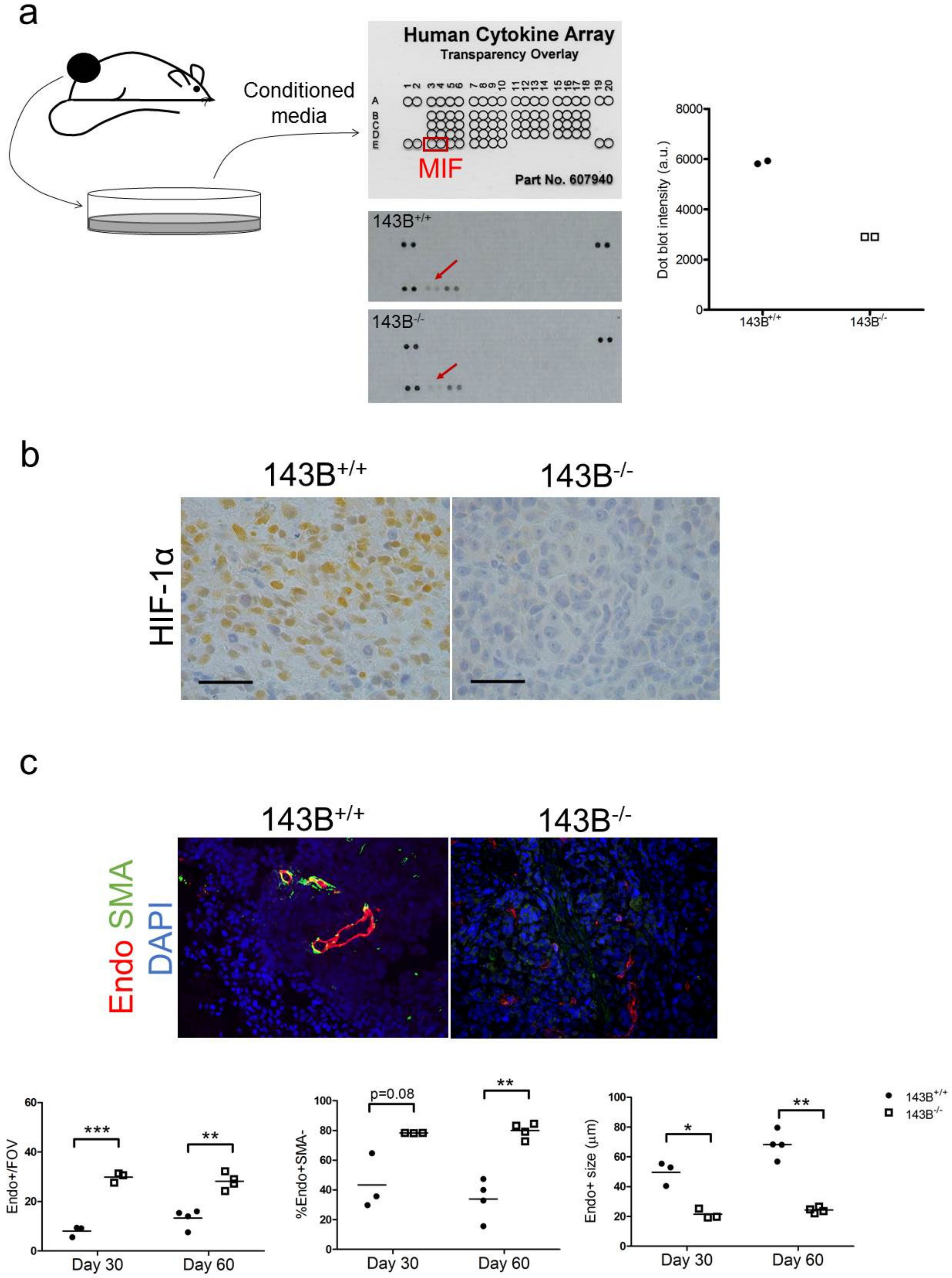
3.2. Tumor Associated Macrophages are the Main Hallmark of the CI-Deficient Osteosarcoma Microenvironment
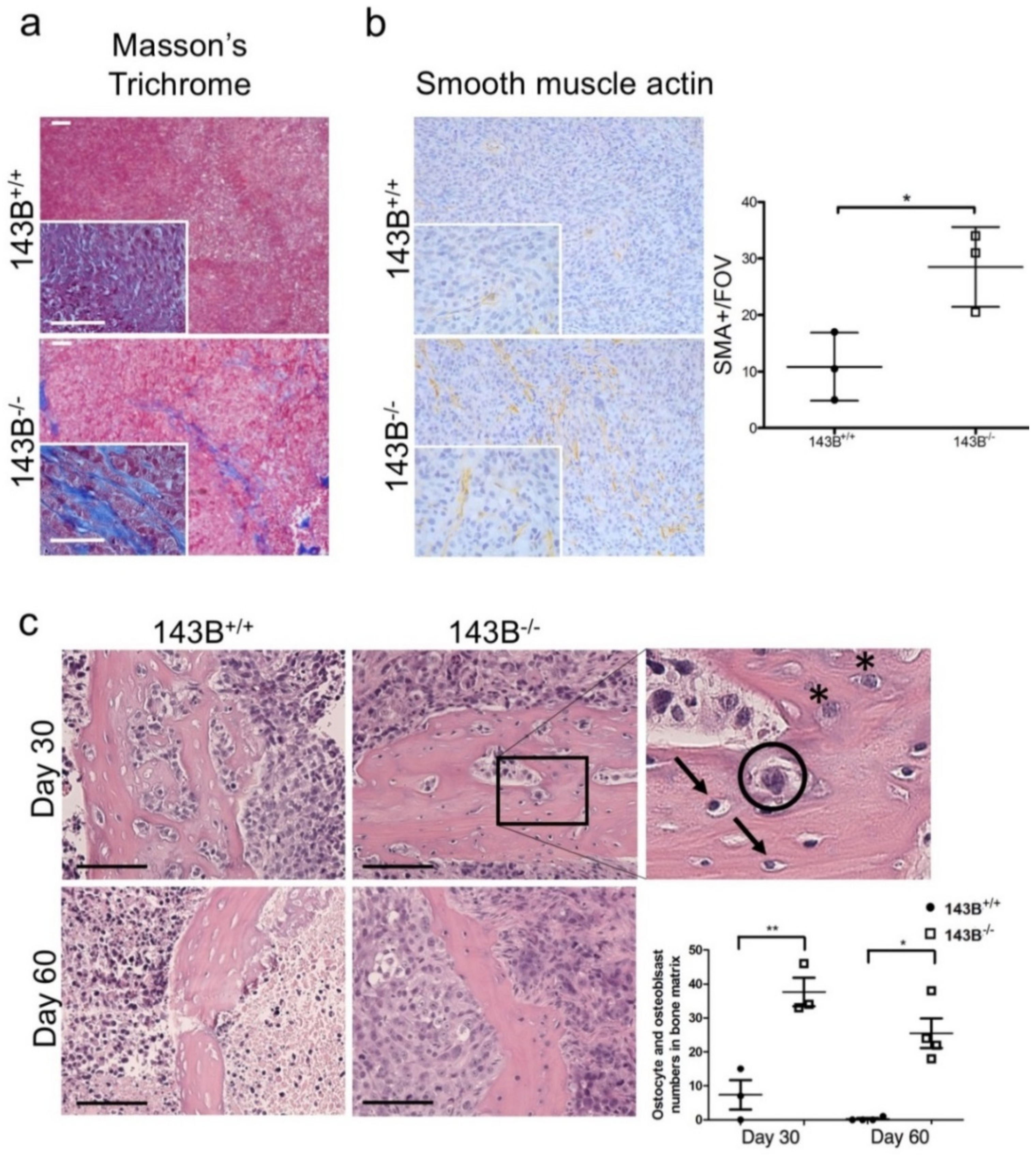
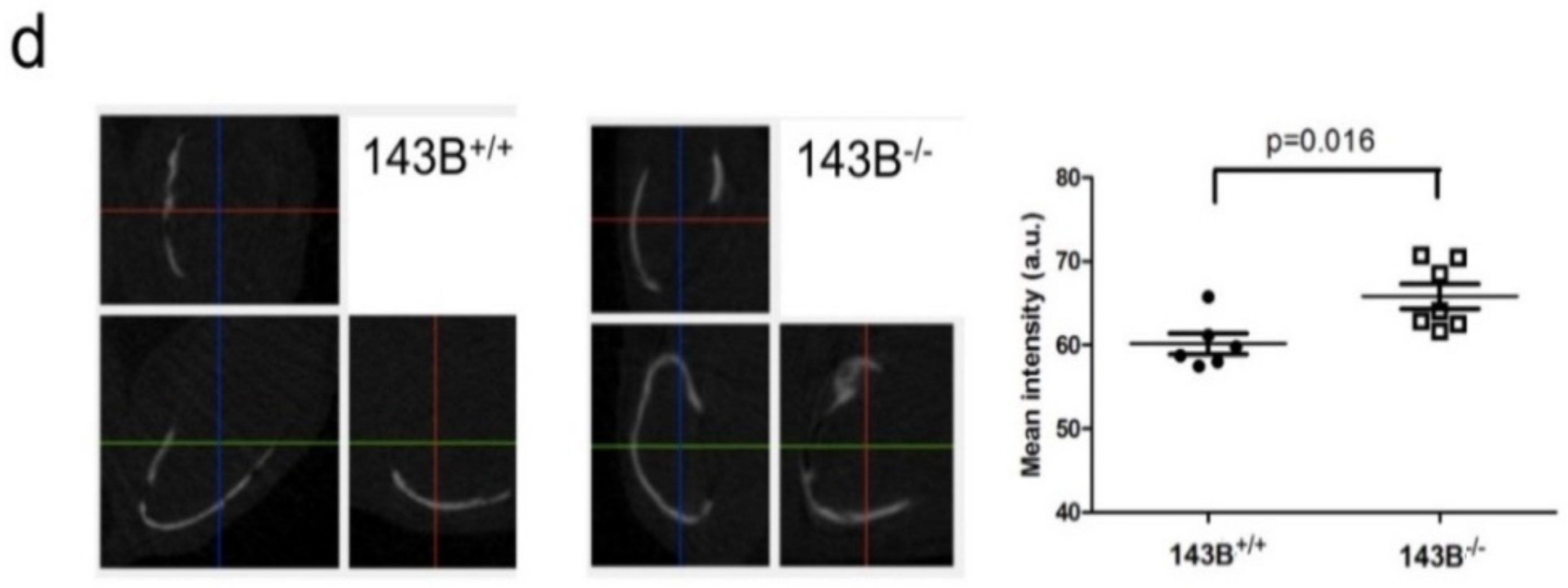
3.3. Osteocytes and Osteoblasts within Trabecular Bone are Preserved in the CI-Deficient Pseudo-Orthotopic Osteosarcoma Xenografts
4. Discussion
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Schulze, A.; Yuneva, M. The big picture: Exploring the metabolic cross-talk in cancer. Dis. Model. Mech. 2018, 11. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.-H.; Qiu, J.; O’Sullivan, D.; Buck, M.D.; Noguchi, T.; Curtis, J.D.; Chen, Q.; Gindin, M.; Gubin, M.M.; van der Windt, G.J.W.; et al. Metabolic Competition in the Tumor Microenvironment Is a Driver of Cancer Progression. Cell 2015, 162, 1229–1241. [Google Scholar] [CrossRef] [PubMed]
- Kurelac, I.; Iommarini, L.; Vatrinet, R.; Amato, L.B.; De Luise, M.; Leone, G.; Girolimetti, G.; Umesh Ganesh, N.; Bridgeman, V.L.; Ombrato, L.; et al. Inducing cancer indolence by targeting mitochondrial Complex I is potentiated by blocking macrophage-mediated adaptive responses. Nat. Commun. 2019, 10, 903. [Google Scholar] [CrossRef] [PubMed]
- Alfranca, A.; Martinez-Cruzado, L.; Tornin, J.; Abarrategi, A.; Amaral, T.; de Alava, E.; Menendez, P.; Garcia-Castro, J.; Rodriguez, R. Bone microenvironment signals in osteosarcoma development. Cell. Mol. Life Sci. CMLS 2015, 72, 3097–3113. [Google Scholar] [CrossRef] [PubMed]
- Rubio, R.; Abarrategi, A.; Garcia-Castro, J.; Martinez-Cruzado, L.; Suarez, C.; Tornin, J.; Santos, L.; Astudillo, A.; Colmenero, I.; Mulero, F.; et al. Bone environment is essential for osteosarcoma development from transformed mesenchymal stem cells. Stem Cells Dayt. Ohio 2014, 32, 1136–1148. [Google Scholar] [CrossRef] [PubMed]
- Baglio, S.R.; Lagerweij, T.; Pérez-Lanzón, M.; Ho, X.D.; Léveillé, N.; Melo, S.A.; Cleton-Jansen, A.-M.; Jordanova, E.S.; Roncuzzi, L.; Greco, M.; et al. Blocking Tumor-Educated MSC Paracrine Activity Halts Osteosarcoma Progression. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2017, 23, 3721–3733. [Google Scholar] [CrossRef]
- Verrecchia, F.; Rédini, F. Transforming Growth Factor-β Signaling Plays a Pivotal Role in the Interplay Between Osteosarcoma Cells and Their Microenvironment. Front. Oncol. 2018, 8, 133. [Google Scholar] [CrossRef]
- Garofalo, C.; Capristo, M.; Manara, M.C.; Mancarella, C.; Landuzzi, L.; Belfiore, A.; Lollini, P.-L.; Picci, P.; Scotlandi, K. Metformin as an adjuvant drug against pediatric sarcomas: Hypoxia limits therapeutic effects of the drug. PLoS ONE 2013, 8, e83832. [Google Scholar] [CrossRef]
- Uehara, T.; Eikawa, S.; Nishida, M.; Kunisada, Y.; Yoshida, A.; Fujiwara, T.; Kunisada, T.; Ozaki, T.; Udono, H. Metformin induces CD11b+-cell-mediated growth inhibition of an osteosarcoma: Implications for metabolic reprogramming of myeloid cells and anti-tumor effects. Int. Immunol. 2019, 31, 187–198. [Google Scholar] [CrossRef]
- Sullivan, L.B.; Luengo, A.; Danai, L.V.; Bush, L.N.; Diehl, F.F.; Hosios, A.M.; Lau, A.N.; Elmiligy, S.; Malstrom, S.; Lewis, C.A.; et al. Aspartate is an endogenous metabolic limitation for tumour growth. Nat. Cell Biol. 2018, 20, 782–788. [Google Scholar] [CrossRef]
- Birsoy, K.; Wang, T.; Chen, W.W.; Freinkman, E.; Abu-Remaileh, M.; Sabatini, D.M. An Essential Role of the Mitochondrial Electron Transport Chain in Cell Proliferation Is to Enable Aspartate Synthesis. Cell 2015, 162, 540–551. [Google Scholar] [CrossRef] [PubMed]
- Griss, T.; Vincent, E.E.; Egnatchik, R.; Chen, J.; Ma, E.H.; Faubert, B.; Viollet, B.; DeBerardinis, R.J.; Jones, R.G. Metformin Antagonizes Cancer Cell Proliferation by Suppressing Mitochondrial-Dependent Biosynthesis. PLoS Biol. 2015, 13, e1002309. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Chen, J.; Yi, G.; Deng, M.; Liu, H.; Liang, M.; Shi, B.; Fu, X.; Chen, Y.; Chen, L.; et al. Metformin suppresses hypoxia-induced stabilization of HIF-1α through reprogramming of oxygen metabolism in hepatocellular carcinoma. Oncotarget 2016, 7, 873–884. [Google Scholar]
- Iommarini, L.; Kurelac, I.; Capristo, M.; Calvaruso, M.A.; Giorgio, V.; Bergamini, C.; Ghelli, A.; Nanni, P.; De Giovanni, C.; Carelli, V.; et al. Different mtDNA mutations modify tumor progression in dependence of the degree of respiratory complex I impairment. Hum. Mol. Genet. 2014, 23, 1453–1466. [Google Scholar] [CrossRef] [PubMed]
- Gasparre, G.; Kurelac, I.; Capristo, M.; Iommarini, L.; Ghelli, A.; Ceccarelli, C.; Nicoletti, G.; Nanni, P.; De Giovanni, C.; Scotlandi, K.; et al. A mutation threshold distinguishes the antitumorigenic effects of the mitochondrial gene MTND1, an oncojanus function. Cancer Res. 2011, 71, 6220–6229. [Google Scholar] [CrossRef]
- Kurelac, I.; Umesh Ganesh, N.; Iorio, M.; Porcelli, A.M.; Gasparre, G. The multifaceted effects of metformin on tumor microenvironment. Semin. Cell Dev. Biol. 2019. [Google Scholar] [CrossRef]
- Kocatürk, B.; Versteeg, H.H. Orthotopic injection of breast cancer cells into the mammary fat pad of mice to study tumor growth. J. Vis. Exp. JoVE 2015, 96, e51967. [Google Scholar] [CrossRef]
- Reiberger, T.; Chen, Y.; Ramjiawan, R.R.; Hato, T.; Fan, C.; Samuel, R.; Roberge, S.; Huang, P.; Lauwers, G.Y.; Zhu, A.X.; et al. An orthotopic mouse model of hepatocellular carcinoma with underlying liver cirrhosis. Nat. Protoc. 2015, 10, 1264–1274. [Google Scholar] [CrossRef]
- Sasaki, H.; Iyer, S.V.; Sasaki, K.; Tawfik, O.W.; Iwakuma, T. An improved intrafemoral injection with minimized leakage as an orthotopic mouse model of osteosarcoma. Anal. Biochem. 2015, 486, 70–74. [Google Scholar] [CrossRef]
- Geller, D.S.; Singh, M.Y.; Zhang, W.; Gill, J.; Roth, M.E.; Kim, M.Y.; Xie, X.; Singh, C.K.; Dorfman, H.D.; Villanueva-Siles, E.; et al. Development of a Model System to Evaluate Local Recurrence in Osteosarcoma and Assessment of the Effects of Bone Morphogenetic Protein-2. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2015, 21, 3003–3012. [Google Scholar] [CrossRef]
- Abarrategi, A.; Mian, S.A.; Passaro, D.; Rouault-Pierre, K.; Grey, W.; Bonnet, D. Modeling the human bone marrow niche in mice: From host bone marrow engraftment to bioengineering approaches. J. Exp. Med. 2018, 215, 729–743. [Google Scholar] [CrossRef] [PubMed]
- Abarrategi, A.; Perez-Tavarez, R.; Rodriguez-Milla, M.A.; Cubillo, I.; Mulero, F.; Alfranca, A.; Lopez-Lacomba, J.L.; García-Castro, J. In vivo ectopic implantation model to assess human mesenchymal progenitor cell potential. Stem Cell Rev. Rep. 2013, 9, 833–846. [Google Scholar] [CrossRef] [PubMed]
- Abarrategi, A.; Foster, K.; Hamilton, A.; Mian, S.A.; Passaro, D.; Gribben, J.; Mufti, G.; Bonnet, D. Versatile humanized niche model enables study of normal and malignant human hematopoiesis. J. Clin. Investig. 2017, 127, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Passaro, D.; Abarrategi, A.; Foster, K.; Ariza-McNaughton, L.; Bonnet, D. Bioengineering of Humanized Bone Marrow Microenvironments in Mouse and Their Visualization by Live Imaging. J. Vis. Exp. JoVE 2017, 126, e55914. [Google Scholar] [CrossRef]
- Castro, B.A.; Flanigan, P.; Jahangiri, A.; Hoffman, D.; Chen, W.; Kuang, R.; De Lay, M.; Yagnik, G.; Wagner, J.R.; Mascharak, S.; et al. Macrophage migration inhibitory factor downregulation: A novel mechanism of resistance to anti-angiogenic therapy. Oncogene 2017, 36, 3749–3759. [Google Scholar] [CrossRef]
- De Palma, M.; Biziato, D.; Petrova, T.V. Microenvironmental regulation of tumour angiogenesis. Nat. Rev. Cancer 2017, 17, 457–474. [Google Scholar] [CrossRef]
- Ombrato, L.; Nolan, E.; Kurelac, I.; Mavousian, A.; Bridgeman, V.L.; Heinze, I.; Chakravarty, P.; Horswell, S.; Gonzalez-Gualda, E.; Matacchione, G.; et al. Metastatic-niche labelling reveals parenchymal cells with stem features. Nature 2019, 572, 603–608. [Google Scholar] [CrossRef]
- Pavlides, S.; Whitaker-Menezes, D.; Castello-Cros, R.; Flomenberg, N.; Witkiewicz, A.K.; Frank, P.G.; Casimiro, M.C.; Wang, C.; Fortina, P.; Addya, S.; et al. The reverse Warburg effect: Aerobic glycolysis in cancer associated fibroblasts and the tumor stroma. Cell Cycle Georget. Tex. 2009, 8, 3984–4001. [Google Scholar] [CrossRef]
- Yang, L.; Achreja, A.; Yeung, T.-L.; Mangala, L.S.; Jiang, D.; Han, C.; Baddour, J.; Marini, J.C.; Ni, J.; Nakahara, R.; et al. Targeting Stromal Glutamine Synthetase in Tumors Disrupts Tumor Microenvironment-Regulated Cancer Cell Growth. Cell Metab. 2016, 24, 685–700. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kurelac, I.; Abarrategi, A.; Ragazzi, M.; Iommarini, L.; Ganesh, N.U.; Snoeks, T.; Bonnet, D.; Porcelli, A.M.; Malanchi, I.; Gasparre, G. A Humanized Bone Niche Model Reveals Bone Tissue Preservation Upon Targeting Mitochondrial Complex I in Pseudo-Orthotopic Osteosarcoma. J. Clin. Med. 2019, 8, 2184. https://doi.org/10.3390/jcm8122184
Kurelac I, Abarrategi A, Ragazzi M, Iommarini L, Ganesh NU, Snoeks T, Bonnet D, Porcelli AM, Malanchi I, Gasparre G. A Humanized Bone Niche Model Reveals Bone Tissue Preservation Upon Targeting Mitochondrial Complex I in Pseudo-Orthotopic Osteosarcoma. Journal of Clinical Medicine. 2019; 8(12):2184. https://doi.org/10.3390/jcm8122184
Chicago/Turabian StyleKurelac, Ivana, Ander Abarrategi, Moira Ragazzi, Luisa Iommarini, Nikkitha Umesh Ganesh, Thomas Snoeks, Dominique Bonnet, Anna Maria Porcelli, Ilaria Malanchi, and Giuseppe Gasparre. 2019. "A Humanized Bone Niche Model Reveals Bone Tissue Preservation Upon Targeting Mitochondrial Complex I in Pseudo-Orthotopic Osteosarcoma" Journal of Clinical Medicine 8, no. 12: 2184. https://doi.org/10.3390/jcm8122184
APA StyleKurelac, I., Abarrategi, A., Ragazzi, M., Iommarini, L., Ganesh, N. U., Snoeks, T., Bonnet, D., Porcelli, A. M., Malanchi, I., & Gasparre, G. (2019). A Humanized Bone Niche Model Reveals Bone Tissue Preservation Upon Targeting Mitochondrial Complex I in Pseudo-Orthotopic Osteosarcoma. Journal of Clinical Medicine, 8(12), 2184. https://doi.org/10.3390/jcm8122184







